Abstract
Long-standing evidence supports the importance of maintaining healthy populations of microbiota for the survival, homeostasis, and complete development of marine mollusks. However, the long-term ecological effects of agricultural runoff on these populations remains largely unknown. Atrazine (6-Chloro-n-ethyl-n′-(1-methylethyl)-triazine-2,4-diamine), a prevalent herbicide in the United States, is often used along tributaries of the Chesapeake Bay where oyster breeding programs are concentrated. To investigate any potential effects atrazine maybe having on mollusk-prokaryote interactions, we used 16S rRNA gene amplicons to evaluate how microbial compositions shift in response to exposure of environmentally relevant concentrations of atrazine previously found within the Chesapeake Bay. The dominant bacterial genera found within all groups included those belonging to Pseudoalteromonas, Burkholderia, Bacteroides, Lactobacillis, Acetobacter, Allobaculum, Ruminococcus, and Nocardia. Our results support previously published findings of a possible core microbial community in Crassostrea virginica. We also report a novel finding: oysters exposed to atrazine concentrations as low as 3 µg/L saw a significant loss of a key mutualistic microbial species and a subsequent colonization of a pathogenic bacteria Nocardia. We conclude that exposure to atrazine in the Chesapeake Bay may be contributing to a significant shift in the microbiomes of juvenile oysters that reduces fitness and impedes natural and artificial repopulation of the oyster species within the Bay.
Similar content being viewed by others
Introduction
Bivalves make up 14% of the 104.3 million tons of annual global marine food production1. The eastern oyster, Crassostrea virginica, is one of the bivalve species that people in North America consume the most. Before the 1970s, wild fisheries produced the majority of the 200 thousand tons of oysters consumed annually; but with the introduction of improved farming technologies, aquaculture has increased production and now accounts for more than 75% of the oysters consumed in North America. Whether they grow naturally along their native shoreline habitats or are reared artificially, exposure to pollution threatens the health and development of oysters. Over the past century, estuaries the world over have witnessed staggering losses to oyster populations: according to one estimate, there are currently 97% fewer oysters than there were about 100 years ago2.
The pollutants of greatest concern include herbicides such as glyphosate and atrazine, general use pesticides such as Metam sodium, and fertilizers such as calcium nitrate. These pollutants are becoming increasingly common in aquatic systems, due to global agricultural intensification practices3,4. The most vulnerable areas of the Chesapeake Bay Estuary are its tributaries, which harbor the highest concentrations of pollutant contamination. Estuarine runoff from agricultural fields occurs when either farmers irrigate their fields excessively or, after a period of drought, intense rain showers saturate the fields. This excessive runoff can transport any chemical that is water-soluble into the surrounding aquatic systems. Once they reach the Chesapeake Bay estuary and its surrounding tributaries, these pollutants mix with estuarine organisms like the eastern oyster, Crassostrea virginica. Although the water in the Chesapeake Bay does respond to the ebb and flow of tidal movements, the elongated shape of the bay tends to trap the polluted water that reaches the estuary. Consequently, the organisms found in the bay suffer long-term exposure to pollutants such as atrazine, which is not the case with those organisms that live along coastal or other shoreline aquatic environments where the water circulates with greater force. Of the various pollutants that get trapped in the estuaries’ water, pesticides pose the greatest threat to aquatic as well as terrestrial ecosystems. Herbicides, which are a kind of pesticide, are of particular concern because they are designed to kill off what are known as producers, or the base of a food web for ecosystems.
Atrazine is the second most widely used herbicide in the world. It has been banned in the European Union, but is still used in the United States, with an estimated 76 million pounds sprayed annually on crops5. Atrazine (6-Chloro-n-ethyl-n′-(1-methylethyl)-triazine-2,4-diamine) is a synthetic herbicide regularly applied to crops like corn and sugar cane during the spring and summer months6. By design, Atrazine inhibits photosynthesis. This inhibition occurs within the plant cell chloroplast thylakoid membrane; it is triggered when atrazine binds to D1 proteins of the photosystem II complex7. The herbicide binding protein blocks CO2 fixation, compromising the targeted plant’s ability to produce the energy that it needs to grow8. The inhibition ultimately leads to the death of the susceptible plant, which generally occurs within 14–21 days after exposure. Atrazine has been known to remain in soils for at least a couple of months and even for up to several years9. The chemical does not break down easily in water either (half-lives > 200 days), which means that once it reaches the estuaries of the Chesapeake Bay, it can remain there for an indefinite period and in ever-increasing concentrations6. A study conducted by the USDA in 2006 found the highest concentration of atrazine in the Chesapeake Bay watershed to be 30 μg/L, 10×, which is at the threshold of the contaminant safety level10. The U.S. Safe Drinking Water Act recognizes the maximum contaminant level of atrazine to be 3 μg/L6,10.
Research on Atrazine’s effects on the reproductive cycles of various species
Independent studies have shown that atrazine can cause chemical castration in frogs11,12. These studies indicate that male frogs that have been induced by atrazine to become females are closer to genetic females than they are to males. It has also been noted that exposure to concentrations “as low as 0.1 parts per billion” of atrazine in surface water have adversely affected frogs by causing the male gonads to produce eggs—effectually turning males into hermaphrodites11,13. Together, the present data suggests that sex-reversal by atrazine is an effect that occurs across nonamniote vertebrate classes11,12,13. This sex-reversal effect may be of consequence to invertebrate animals as well, particularly those that exhibit sequential hermaphroditism (changing one’s sex throughout a lifetime) such as does C. virginica. Further evidence of atrazine’s potential deleterious effects are evidenced in cases of human exposure, where changes in menstrual hormone levels shift follicular and ovarian phases that result in cycle irregularity14. Ultimately, these studies indicate a reproductive reaction to atrazine exposure.
Research on Atrazine’s effects on infectious diseases that harm bivalves
There are very few studies examining microbiome communities in bivalves and even fewer that focus on C. virginica. These microbes can have positive, negative, or neutral effects on the organism. Their interactions may also change based on the composition of other microbial species, the population sizes of the other microbes, and the genetic expression of the host. The majority of oyster-prokaryote research has focused on the “protective and preventative role” of certain species of bacteria against a potentially fatal pathogen, Vibrio15. In one study, Alteromonas haloplanktis, isolated from scallops, “inhibited the growth of both Vibrio anguillarum and Vibrio alginolyticus in vitro”16. Similarly, a study concluded that Aeromonas media A199 protects scallops against Vibrio tubiashii via interactions with an aromatic heterocyclic organic compound known as indole (C8H7N)17,18.
According to one study, 20% of bacterial isolates from C. virginica are capable of producing indole (“an intracellular signal molecule which regulates various aspects of bacterial physiology, including spore formation, plasmid stability, resistance to drugs, biofilm formation, and virulence”)19. Another study has found that adding isolated bacteria from C. virginica samples to larvae that have been infected with Vibrio corallilyticus significantly increases the oyster’s chances of survival20. Finally, it has been discovered that Phaeobacter sp. S4, which is a bacterial isolate from the inner shell of C. virginica, also protects C. virginica larvae and juveniles challenged with R. crassostreae and V. tubiashii21. However much these beneficial bacterial help to defend their host, the protection they provide may be short-lived20,21. Additional work is required to fully comprehend the protective and possibly opportunistic role of the microbial community of C. virginica.
As filter-feeders, oysters are exposed to many different microbes. Since they also have unique physical structures, there are many surfaces and micro-ecosystems in which microbes can reside. It is thus unsurprising that oysters harbor an order of magnitude greater concentration of bacteria than the surrounding water in which they live22,23. Through next-generation microbiome analysis, we characterized the oyster’s microbial community composition and recorded a snapshot of these integral dynamics24,25,26,27,28. As we aim to demonstrate, this experiment offers a unique opportunity to improve our understanding of how oyster–prokaryotic interactions respond to ecologically relevant chemical contaminants.
In this study, we used 16s rRNA gene amplicon sequencing to characterize the microbiome of hatchery raised C. virginica juveniles. We sought to determine the extent to which atrazine may be affecting the bacterial composition of juvenile oysters, after having been exposed to environmentally relevant concentrations. As a herbicide commonly detected in the Chesapeake Bay Watershed10, atrazine was chosen to be the focus of this study investigating the effects of herbicide-induced microbiome shifts in C. virginica juveniles.
Methods
Examined concentrations
This study used a range of atrazine concentrations based upon those commonly detected in the Chesapeake Bay10,29,30,31,32,33. The following outlines the chosen concentrations and the reasoning for using that concentration in the experiment:
-
1.
30 µg/L Atrazine: The peak concentration of atrazine detected in any sample within the Chesapeake Bay was 30 µg/L (station L0002488), which is located in a tributary on the Eastern Shore10. The highest concentration recorded is of interest, as it represents an extreme concentration.
-
2.
20 µg/L Atrazine: A half-way point in between 30 µg/L Atrazine and 10 µg/L Atrazine for data exploration purposes.
-
3.
10 µg/L Atrazine: During June of 2011, atrazine concentrations from the upper Choptank river were logged. The average of all recorded concentrations was 0.29 µg/L. The highest recorded concentration was 10 µg/L10,13,29. The general trend in the data indicates that the main detections of atrazine are in the tributaries while significantly lower concentrations have been found in the Bay itself. Oyster restoration grounds are generally found within tributaries along the upper, middle and lower sections of the Choptank river30.
-
4.
3 µg/L Atrazine: The EPA’s Maximum Residue Limit10,13 for atrazine is relevant because it is the limit which the EPA has set forth for regulating our own drinking water, further investigation into the MRL of atrazine is merited based on health concerns cited in many scholarly papers throughout the last decade, the last official measurement was carried out in 201730,31,32, 34,35,36,37.
-
5.
30 µg/L Acetone: Atrazine stock solution was dissolved in a 100% acetone solution. In order to assume continuity throughout the experiment this treatment was added to assure that any noticed effect on development was due solely to atrazine exposure.
-
6.
0 µg/L Atrazine/Acetone: Control group.
Oyster acquisition and stabilization
300 juvenile Crassostrea virginica oysters of similar size and weight were purchased from Horn-Point Laboratory. The oysters were separated into six groups of 50 specimens and randomly assigned to an exposure group (30 µg/L Atrazine, 20 µg/L Atrazine, 10 µg/L Atrazine, 3 µg/L Atrazine, 30 µg/L Acetone and a atrazine and acetone-free Control of instant ocean seawater). 3.0 mm square mesh sieves were used to separate each group. No oyster was smaller than 5.0 mm long × 4.0 mm wide when placed in the mesh sieves. Oysters underwent a 2-week long acclimation period inside of a large holding tank filled with 300 L of pressure filtered instant ocean seawater at a salinity of 25 parts per thousand (ppt). Frequent water changes (25% twice weekly) were used in order to minimize buildup of both ammonium and nitrate within the closed water system. Each oyster group was fed 6 L of a concentrated phytoplankton mixture of (Tetraselmis Chuii, isochrysis galbana, and Nannochloropsis oculata) approximately ~ 400,000 cells/mL) every day.
Relevant tank water parameters were monitored every other day and water changes were completed when required. The tank was consistently maintained to fit the following water parameters:
Salinity (ppt) | Ammonium (ppb) | Nitrate (ppb) | Plankton concentration |
|---|---|---|---|
25 | 0 | 0 | > 9,000 cells/ml |
Water quality parameters were monitored using: Salinity Refractometer—Salinity, API Testing Kit—Ammonium levels, API Testing Kit—Nitrate levels, Mass Spectrophotometer at 654 nm—Plankton Concentration, Hemocytometer count—Plankton Concentration.
Assigning experimental groups
Individual oysters were randomly separated into six groups containing 50 oysters each. To prevent potential overcrowding of the organisms and ensure that each oyster in the study was filtering the same amount of water, each of these groups was subsequently divided into a sub-group of 10 oysters and placed into 3.0 mm square mesh sieves. These sub-groups of 10 oysters were kept together throughout the entirety of the experiment. The acetone group was used because the atrazine we used for the experiment was diluted in a solution of HPLC-grade acetone. Each group of 50 oysters was assigned to one of six treatments: 0 μg/L Atrazine/Acetone, 3 μg/L Atrazine, 10 μg/L Atrazine, 20 μg/L Atrazine, 30 μg/L Atrazine, and 30 μg/L Acetone.
Exposures
Following the 2-week long stabilization period, the experimental groups were exposed to their first atrazine trial. All oyster groups were kept in separate tanks to avoid any potential cross contamination. In order to expose the oyster groups to atrazine and acetone, each group was removed from its holding tank and placed into a separate 4-L glass tank containing 2 L of newly made seawater (salinty = 25 ppt). Atrazine was added to each 4-L glass tank according to the corresponding treatment concentration. In order to mimic the heavy rainfall pattern surrounding the Chesapeake Bay38, treated groups spent a total of 3 h submerged in the treated 2 L of saltwater three times per week, every other day (Monday, Wednesday, Friday). Aeration was provided in both holding and treatment tanks.
In order to minimize any potential contamination of each group’s respective holding tanks after the 3-h exposure period, each group was washed using a constant stream of pressure-filtered water for three one-minute rinses. Oyster groups were then relocated to their respective 40-L glass bio-cube tanks. The bio-cube tanks were set to have the same parameters as the large holding tank.
Prokaryotic DNA extraction
Each oyster was carefully opened using sterilized forceps. Whole tissue extraction was done by scraping and removing all existing tissue from each of the two shells. The whole oyster tissue was then placed in a labeled 1.5 mL microcentrifuge tube with the treatment and group to which it belonged. In order to extract only the prokaryotic DNA a QIAGEN QIAamp DNA Microbiome Kit protocol was used.
Oyster weights
Oysters were weighed on a weekly basis using a Mettler Toledo PB303-S Digital Balance Scale MonoBloc Weighing Technology. During each weighing session, every oyster group within each of the six treatments was carefully removed from the holding bags and placed into a dish (previously weighed separately in order to set scale to zero). After placing the dish with the oysters on the scale, oyster weights were recorded to the nearest ten thousandth place. Oysters were then removed from the dish and laid out on a surface for pictures. All previous steps were repeated for each of the groups within the six treatments. Pictures of each group were taken independently to make sure that the correct number of specimens were there. All weights were recorded in grams. The last set of weights (g) were recorded on September 25, 2018, the same date on which five out of the ten oysters belonging to each of the six treatment groups were removed for DNA isolation. Following the removal of half of the oysters in each treatment group, the remaining five specimens in each group were kept under the same environmental conditions, except that they were no longer exposed to atrazine. All the dates under the post-atrazine treatment section of the graph therefore belong only to the remaining five oysters from each group; in order to account for this loss, group averages were taken and divided by the total number of individuals per group. September 30, 2018 marks the last day on which oyster weights (g) were recorded.
16S sequencing
The following protocol was produced and completed with CD Genomics.
Amplicons were sequenced on a paired-end Illumina HiSeq2500 platform to generate 250 bp paired-end raw reads, and then pretreated. The forward and reverse primers used were 338F:5′ ACTCCTACGGGAGGCAGCA-3′ and 806R:5′- GGACTACHVGGGTWTCTAAT-3′ respectively. Specific processing steps for both sequencing events were completed as follows: (1) Paired-end reads were assigned to a sample by their unique barcode, and the barcode and primer sequence were then truncated. (2) Paired-end reads were merged using FLASH (V1.2.7, https://ccb.jhu.edu/software/FLASH/), a very fast and accurate analysis tool to merge pairs of reads when the original DNA fragments are shorter than twice of the reads length. (3) Quality filtering was then performed on the raw tags under specific filtering conditions of Trimmomatic v0.33 (https://www.usadellab.org/cms/?page=trimmomatic) quality control process. After filtering, high-quality clean tags were obtained. (4) The resulting high quality tags were compared with the reference database (Gold database, https://drive5.com/uchime/uchime_download.html) using UCHIME algorithm (UCHIME Algorithm (https://www.drive5.com/usearch/manual/uchime_algo.html) to detect chimeric sequences and then the chimeric sequences were removed.
Statistical analysis
The following protocol was produced and completed with CD Genomics.
To characterize microbial communities and to perform functional analyses, the following wrappers were employed in R statistical software: summarize_taxa_through_plots (to produce the taxonomical files and charts), alpha_rarefaction and beta_diversity_through_plots (to assess respectively the alpha- and beta-rarefaction diversity indices), and principal_coordinates.py (to compare groups of samples based on phylogenetic distance metrics). To compare treatment groups (Control versus all treatment concentrations), AMOVA (analysis of molecular variance) analyses was performed at genus level (P-value < 0.005; FDR < 0.01) using Excel version 16.13. In order to further test for significant variation between groups in the metagenomic data, METASTATS (a software for detecting differentially abundant features in metagenomic case/control studies) was used. It is the first tool developed in the context of metagenomic data comprising multiple samples and relies on a non-parametric t-test39. Finally, a heatmap analysis was also created via clustering of high abundance and low abundance genera. The heatmap was aggregated by color gradient and similarity to reflect the similarity and difference of the bacterial composition of multiple samples. “According to the taxonomic composition and relative abundance of each sample, the genera heatmap analysis was carried out to extract the species at each taxonomic level and plotted using the R language tools, the heatmap clustering analysis was conducted at the level of phylum, class, order, family, genus and species respectively” (CD Genomics).
Results
Diversity of microbial communities and taxonomic richness
As seen in Fig. 1, the relative abundance of the most plentiful genera found between all samples was significantly decreased in specimens exposed to concentrations of atrazine higher than 10 µg/L. According to the AMOVA test scores, this decrease was most significant in the Control vs 30 µg/L (p < 0.001). The same overall decrease in abundance holds true for bacterial composition on the species and phyla levels (supplementary material S1, S3). All groups included dominant bacteria from the orders Rickettsiales, Vibrionales, and Alteromonadales.
As seen in Fig. 2A,B, both Acetobacter and Bacteroides abundance dropped to negligible levels if the animal was treated with 30 µg/L of atrazine. In Fig. 2C,D we find similar reductions in abundance within Ruminococcus and Allobaculum, respectively. Allobaculum, Oscillospira, Escherichia, and Bacteroides predominate amongst the control samples. We detected a significant dose-dependent relationship to relative abundance for both Bacteroides and Allobaculum: as oysters were exposed to higher concentrations of atrazine, a decrease in their relative abundance was observed. Acetobacter and Ruminococcus experienced a particular loss of relative abundance when exposed to concentrations of atrazine higher than 10 µg/L.
The box graph visually displays the results of the METASTATs, t-tests with sample permutation for detecting differentially abundant features in a metagenome. A non-parametric T-test determines whether there are any taxa that are differentially represented between samples; significance is depicted by the asterisks. Plots were made in R, the first name listed refers to the bacterial family, the second the genera. The vertical axis denotes relative abundance.
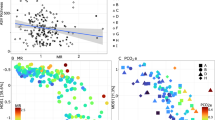
A principal coordinate analysis (PCoA) was performed using unweighted UniFrac distances (Fig. 3). We used these analyses to test for significant differences in microbial genera composition between all groups. Figure 3 shows a clear grouping along both the x-axis (PC1 = 29.36% of variability) and the y-axis (PC2 = 12.24%) between the control, acetone and 3 µg/L atrazine samples. The remaining treatment groups (atrazine 10 µg/L, 20 µg/L and 30 µg/L, respectively) clustered farther away with increasing dosages of atrazine.
Similarly, the acetone and control groups were found to contain more similar microbial compositions than any of the atrazine groups did (Fig. 3). However, 30 µg/L atrazine is the only group found in the bottom left quadrant of the plot, indicating this treatment results in the most dissimilar microbial composition. According to the heatmap in Fig. 4, Allobaculum, Akkermansia, Ruminococcus, and Bacterioides all severely decreased in the atrazine 30 µg/L group. However, Allobaculum, Akkermansia, Ruminococcus, Bacteroides, Brevibacterium, Aceobacter, and Halomicronema genera all showed decreased levels of abundance in the oyster groups treated with atrazine (Fig. 4).
In the heatmap analysis the vertical clustering indicates the similarity of the abundance between different genera. The shorter the distance between the two genera, the more similar abundance between the samples. In the horizontal clustering, the closer and shorter of the branch length between the samples, the more similarity of the abundance.
An analysis of molecular varience (AMOVA) test was performed on all treatment groups for the most abundant genera (Table 1).
Psuedoalternamonas, Fusobacterium, Bacterioides and Clostridium were found to be present within both Atrazine 20 and Atrazine 30 groups, but at significantly lower levels when compared to the control. Additionally, a significant difference was detected between the Atrazine 20 group when compared to Atrazine 3 group when comparing between the most abundant genera.
Finally, bacteria in the genus Nocardia were detected only in groups exposed to atrazine (S2). These differences in microbial composition may shed some light on the differences between each groups’ average growth over the experiment.
Oyster growth data
The general trend when comparing slope values was that as atrazine concentrations increased, slope values tended to decrease (Fig. 5). Both the control (0.0371) and acetone (0.0318) treatments displayed the highest slope values (Fig. 5). The highest slope values correlate to the treatment groups with the highest average weights over time, which in this case is approximately over the course of 3 months. The Atrazine group with 10 µg/L experienced the fastest growth when compared to the other atrazine treatment groups. The atrazine 30 µg/L group was expected to have undergone the least growth as it was the group that was exposed to the most elevated concentration; however, the Atrazine 20 µg/L treatment group had the lowest slope value (0.0128). On October 16, 2018, the Atrazine 30 µg/L average growth line drops, which could be attributed to the death of an oyster that was not detected at the time (Fig. 5).
Discussion
Interpreting the results
The present study investigates the effects of atrazine exposure on the microbiome of C. virginica. Today, we understand that the microbiome of marine mollusks is vitally important to their survival, homeostasis, and overall development40. The microbiome of filter-feeding bivalves such as oysters is comprised of both continuously occupant and temporary transitory microbiota. Eastern oysters are suspension-feeders and “filter large quantities of water (e.g., 3–9 L/h/g dry mass for oysters”41,42. As such, they are exposed to both free-living and particle-associated microbes. Therefore, the oysters’ microbiome may be particularly important for physiological and biochemical enantiostasis (open system maintenance).
Our results indicate that oysters can act as a host for a specific set of core microbiota. Although the oysters in the study were comprised of hatchery-reared juveniles, many of the bacteria sequenced in this study were also found residing within oysters raised and sampled in the wild43,44,45,46. Our results also indicate that atrazine exposure may cause C. virginica juveniles to develop at a slower rate than they would without exposure, possibly due to a loss of beneficial bacteria (Fig. 5).
The microbial composition of the control group’s four samples bear a greater resemblance among themselves than when they are compared to those other groups that had been exposed atrazine (Fig. 3). This may be a sign of a healthy core microbiome existing within the control groups but not in the atrazine exposed experimental groups. It may be that as the core microbiome’s relative abundance decreases, it gives way to new, potentially pathogenic bacteria which take hold within the animal. According to the PCOA, this was particularly evident in those groups that were exposed to at least 3 μg/L.
A significant increase of Nocardia (a gram-positive, partially acid-fast actinomycete, pathogenic bacteria) was observed in the 3, 10, 20, and 30 μg/L atrazine treatment groups. A subsequent decrease in Nocardia was detected in the 30 μg/L treatment group, which may account for this group’s increased growth rate between 10/16/18 and 10/23/18. Nonetheless, Nocardia was found only within the atrazine treated groups (Supplementary material S2), which suggests the presence of atrazine may have artificially selected for the colonization and subsequent survival of pathogenic Nocardia spp.
Nocardia is often found in aquatic and terrestrial environments, existing as soil saprophytes, but can also appear in fresh and saltwater, dust, decaying vegetation and fecal deposits from animals. Nocardia was found to be lethal to Ostrea edulis, flat oysters45,46,47,48 and may also be lethal to C. virginica in advanced stages of colonization49. In a study by Carella et al., Nocardia crassostreae was observed in Mytilus galloprovincialis and Ostrea edulis sampled from raft-culture farms in the Gulf of Naples49. Carella et al. found that the inflammatory lesions observed were characteristics also found in other Nocardia-related cases in different animals, both in invertebrates and vertebrates alike. However, there have been no other studies that compare and confirm these findings.
The microbial communities of oysters include both resident (or core) as well as transient microbiota. Oysters are thus highly susceptible to pathogen introduction and atrazine may be affecting the ability of beneficial oyster microbes to fight off both known and unknown bacterial pathogens. One of the beneficial roles of the oyster microbiome is to defend the host oyster against pathogen inoculation and environmental stress43. However, in compromised hosts, the helpful microbes may well become opportunistic pathogens50,51,52.
Psuedoalternamonas, Fusobacterium and Clostridium species, which all form part of Crassostrea virginica’s core microbiome, were found to a lesser degree within the 20 μg/L atrazine treated sample when compared to the control (Table 1). This finding indicates that atrazine exposure may play a role in crafting an environment which does not permit Psuedoalternamonas, Clostridium and/or Fusobacterium species to effectively colonize the oysters during their juvenile life stage, a stage of rapid organismal development. There have been little to no studies done investigating the effects these bacteria can have on oysters, which means that the reduction of these bacteria could have a positive, negative, or neutral effect on the oysters. However, these groups have often been found to be part of the microbiome of bivalves worldwide53,54,55.
Known bacterial interactions
Studies have found that Proteobacteria and Cyanobacteria are integral members of bivalve microbiomes, irrespective of their host56,57. Bacteroides species normally form mutualistic relationships within the gut of bivalve species, where they process complex molecules into simpler ones within the host intestine, and at times by produce indole58,59,60. Our results indicate that atrazine has decoupled this interaction by decreasing Bacteroides relative abundance in particular (Figs. 1, 4). It may be possible that Bacteroides indole production helps to keep pathogenic bacteria from colonizing the oyster microbiome. However, these genera are not the most plentiful within the microbiome of oysters. In multiple studies, and in particular one conducted by Wexler, H. M, it was found that Proteobacteria can account for a majority of the total abundance33,57,61,62,63,64. Other dominant phyla that have been identified include Acteroidetes, Actinobacteria, Firmicutes, Chlamydiae, Fusobacteria, Spirochates, Chroloflexi, Plantomycetes, and Verrucomicrobia12,48,56, 65,66. Our study corroborates these findings (S3).
Achromobacter, Aeromonas, Flavobacterium, Micrococcus, and Vibrio are all bacterial genera commonly isolated from bivalves53,60, 67,68,69,70, however, more recent culture-independent work has shown that while these genera are commonly present, they do not necessarily represent most of the community54,55,71. We detected all of the above genera within our samples except Aeromonas. Because these genera do not represent the majority of the community, it is unsurprising that we did not detect all of the bacterial genera known to occur in oyster microbiomes.
Conclusion
Because the microbiome of C. virginica is vital for its homeostasis and survival—especially during its developmental stages—the persistence of atrazine in the Bay may increase the fatality levels of juvenile oysters by selecting for the colonization of pathogenic bacteria and the subsequent decrease in the relative abundance of common beneficial microbes. This interaction may also be affecting the growth rate of C. virginica juveniles, causing them to grow at a slower rate. If so, atrazine contamination can significantly harm the Chesapeake Bay’s oyster restoration efforts, which target the reseeding of baby oysters and not adults. More research must be done on the long-term effects of this herbicide, but our findings indicate that growing atrazine concentrations in the Chesapeake Bay may be accelerating the drastic decline of this keystone species. This study’s sequence data could be used to compare oyster populations along the eastern seaboard as well as the Gulf Coast where the eastern oyster grows naturally and is reared in aquaculture farms. These microbiome shifts may not be isolated to just herbicide exposures; thus, further research into individual and combined effects of other contaminants would pinpoint which ones are the most detrimental to healthy microbial communities.
This research should expand to sample more time points, repeat herbicidal challenges with the inclusion of pathogens, and sequence molecular data to determine the oyster’s response to both challenge treatments and shifts in microbial community composition. This is a vital baseline for future research aimed at understanding the role of the microbiome in oyster development and physiology. As such, future research efforts should also focus on more robust sampling in natural habitats, working with oyster farmers, hatcheries, and restoration labs to help revitalize the oyster population.
References
Home. https://www.fao.org/home/en/.
Delorenzo, M. E. Impacts of climate change on the ecotoxicology of chemical contaminants in estuarine organisms. Curr. Zool. 61(4), 641–652 (2015).
Banerjee, M. R. & Chapman, S. J. The significance of microbial biomass sulphur in soil. Biol. Fertil. Soils 22(1–2), 116–125 (1996).
Khan, A. et al. Scrutinizing impact of atrazine on stress biomarker: Plasma glucose concentration of fresh water fish, grass carp, (Ctenopharyngodon idella). Pure Appl. Biol. PAB 6(4), 1319–1327 (2017).
Toxic Substances Portal—Atrazine. https://www.atsdr.cdc.gov/phs.asp?id=336&tid-59 (2015).
Toxic Substances Portal—Atrazine. https://www.atsdr.cdc.gov/phs.asp?id=336&tid-59 (2003).
Univeristy of California, D. Photosystem II Inhibitors. https://herbicidesymptoms.ipm.ucanr.edu/MOA/Photosystem_II_Inhibitors.
U.S. Environmental Protection Agency, National pesticide survey—Glossary: Washington, D.C., U.S. Government Printing Office, (7) (2011).
U.S. Environmental Protection Agency, National pesticide survey—Glossary: Washington, D.C., U.S. Government Printing Office, (7) (2006).
USDA dataset 2006. https://usda.custhelp.com/.
Hayes, T. B. et al. Atrazine-induced hermaphroditism at 0.1 ppb in American leopard frogs (Rana pipiens): Laboratory and field evidence. Environ. Health Perspect. 111(4), 568–575 (2003).
de Voogd, N. J., Cleary, D. F., Polónia, A. R. & Gomes, N. C. Bacterial community composition and predicted functional ecology of sponges, sediment and seawater from the thousand islands reef complex, West Java, Indonesia. FEMS Microbiol. Ecol. 91(4), fiv019 (2015).
DeLorenzo, M. E., Scott, G. & Ross, P. E. Toxicity of pesticides to aquatic microorganisms: A review. Environ. Toxicol. Chem. SETAC. 20, 84–98. https://doi.org/10.1897/1551-5028(2001)020<0084:TOPTAM>2.0.CO;2 (2001).
Cragin, L. A. et al. Menstrual cycle characteristics and reproductive hormone levels in women exposed to atrazine in drinking water. Environ. Res. 111(8), 1293–1301 (2011).
Riquelme, C. A. R. L. O. S. et al. Isolation of a native bacterial strain from the scallop Argopecten purpuratus with inhibitory effects against pathogenic vibrios. J. Shellfish Res. 15(2), 369–374 (1996).
Ma, Y. X. et al. The use of Pseudoalteromonas sp. F15 in larviculture of the Yesso scallop, Patinopecten yessoensis. Aquac. Res. 50(7), 1844–1850 (2019).
Gibson, L. F., Woodworth, J. & George, A. M. Probiotic activity of Aeromonas media on the Pacific oyster, Crassostrea gigas, when challenged with Vibrio tubiashii. Aquaculture 169(1–2), 111–120 (1998).
Lategan, M. J., Booth, W., Shimmon, R. & Gibson, L. F. An inhibitory substance produced by Aeromonas media A199, an aquatic probiotic. Aquaculture 254(1–4), 115–124 (2006).
Brown, C. The effects of some selected bacteria on embryos and larvae of the American oyster, Crassostrea virginica. J. Invertebr. Pathol. 21(3), 215–223 (1973).
Lim, H. J., Kapareiko, D., Schott, E. J., Hanif, A. & Wikfors, G. H. Isolation and evaluation of new probiotic bacteria for use in shellfish hatcheries: I. Isolation and screening for bioactivity. J. Shellfish Res. 30(3), 609–616 (2011).
Karim, M. et al. Probiotic strains for shellfish aquaculture: Protection of eastern oyster, Crassostrea virginica, larvae and juveniles againsl bacterial challenge. J. Shellfish Res. 32(2), 401–409 (2013).
Colwell, R. R. & Liston, J. Microbiology of shellfish. Bacteriological study of the natural flora of Pacific oysters (Crassostrea gigas). Appl. Microbiol. 8(2), 104 (1960).
Cavallo, R. A., Acquaviva, M. I. & Stabili, L. Culturable heterotrophic bacteria in seawater and Mytilus galloprovincialis from a Mediterranean area (Northern Ionian Sea, Italy). Environ. Monit. Assess. 149(1–4), 465–475 (2009).
Lazarevic, V. et al. Metagenomic study of the oral microbiota by Illumina high-throughput sequencing. J. Microbiol. Methods 79(3), 266–271 (2009).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6(8), 1621 (2012).
Pedrós-Alió, C. The rare bacterial biosphere. Annu. Rev. Mar. Sci. 4, 449–466 (2012).
Widerström, M., Wiström, J., Sjöstedt, A. & Monsen, T. Coagulase-negative staphylococci: Update on the molecular epidemiology and clinical presentation, with a focus on Staphylococcus epidermidis and Staphylococcus saprophyticus. Eur. J. Clin. Microbiol. Infect. Dis. 31(1), 7–20 (2012).
Volety, A. K. Effects of salinity, heavy metals and pesticides on health and physiology of oysters in the Caloosahatchee Estuary, Florida. Ecotoxicology 17(7), 579–590 (2008).
Powell, K. et al. A retrospective analysis of agricultural herbicides in surface water reveals risk plausibility for declines in submerged aquatic vegetation. Toxics 5(3), 21 (2017).
Hively, W. D. et al. Relating nutrient and herbicide fate with landscape features and characteristics of 15 subwatersheds in the Choptank River watershed. Sci. Total Environ. 409(19), 3866–3878 (2011).
U.S. Environmental Protection Agency, National pesticide survey—Glossary: Washington, D.C., U.S. Government Printing Office, (7). (2017)
Christopher G., et al. Choptank ecological assessment: digital atlas: Baseline status report. NOAA technical memorandum NOS NCCOS; 213 115–132 (2016)
Arfken, A. et al. Denitrification potential of the eastern oyster microbiome using a 16S rRNA gene based metabolic inference approach. PLoS ONE 12(9), e0185071 (2017).
Hussein, S. Y., El-Nasser, M. A. & Ahmed, S. M. Comparative studies on the effects of herbicide atrazine on freshwater fish Oreochromis niloticus and Chrysichthyes auratus at Assiut, Egypt. Bull. Environ. Contam. Toxicol. 57(3), 503–510 (1996).
Trentacoste, S. V. et al. Atrazine effects on testosterone levels and androgen-dependent reproductive organs in peripubertal male rats. J. Androl. 22(1), 142–148 (2001).
Pogrmic-Majkic, K. et al. Atrazine suppresses FSH-induced steroidogenesis and LH-dependent expression of ovulatory genes through PDE-cAMP signaling pathway in human cumulus granulosa cells. Mol. Cell. Endocrinol. 461, 79–88 (2018).
Wirbisky, S. E., Weber, G. J., Schlotman, K. E., Sepúlveda, M. S. & Freeman, J. L. Embryonic atrazine exposure alters zebrafish and human miRNAs associated with angiogenesis, cancer, and neurodevelopment. Food Chem. Toxicol. 98, 25–33 (2016).
USGS Climate/Precipitation Data. https://www.usgs.gov/centers/cba.
Yabuuchi, E. et al. Proposal of Burkholderia gen. nov. and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. Nov.. Microbiol. Immunol. 36(12), 1251–1275 (1992).
Altura, M. A. et al. The first engagement of partners in the Euprymna scolopes–Vibrio fischeri symbiosis is a two-step process initiated by a few environmental symbiont cells. Environ. Microbiol. 15(11), 2937–2950 (2013).
Kemp, W. M. et al. Eutrophication of Chesapeake Bay: Historical trends and ecological interactions. Mar. Ecol. Prog. Ser. 303, 1–29 (2005).
Cranford, P. J., Ward, J. E., & Shumway, S. E. Bivalve filter feeding: Variability and limits of the aquaculture biofilter. Shellfish Aquac. Environ. 81–124. (2011).
Garnier, M., Labreuche, Y., Garcia, C., Robert, M. & Nicolas, J. L. Evidence for the involvement of pathogenic bacteria in summer mortalities of the Pacific oyster Crassostrea gigas. Microb. Ecol. 53(2), 187–196 (2007).
Estrada-De Los Santos, P., Bustillos-Cristales, R. & Caballero-Mellado, J. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Appl. Environ. Microbiol. 67(6), 2790–2798 (2001).
Trabal Fernández, N. et al. Changes in the composition and diversity of the bacterial microbiota associated with oysters (Crassostrea corteziensis, Crassostrea gigas and Crassostrea sikamea) during commercial production. FEMS Microbiol. Ecol. 88(1), 69–83 (2014).
Friedman, C. S. et al. Nocardia crassostreae sp. Nov., the causal agent of nocardiosis in Pacific oysters. Int. J. Syst. Evol. Microbiol. 48(1), 237–246 (1998).
Meyer, G. R., Lowe, G. J., Gilmore, S. R. & Bower, S. M. Disease and mortality among Yesso scallops Patinopecten yessoensis putatively caused by infection with Francisella halioticida. Dis. Aquat. Org. 125(1), 79–84 (2017).
Yee-Duarte, J. A., Ceballos-Vázquez, B. P., Shumilin, E., Kidd, K. A. & Arellano-Martínez, M. Parasitic castration of chocolate clam Megapitaria squalida (Sowerby, 1835) caused by trematode larvae. J. Shellfish Res. 36(3), 593–599 (2017).
Carella, F. et al. Nocardiosis in Mediterranean bivalves: First detection of Nocardia crassostreae in a new host Mytilus galloprovincialis and in Ostrea edulis from the Gulf of Naples (Italy). J. Invertebr. Pathol. 114(3), 324–328 (2013).
Cerf-Bensussan, N. & Gaboriau-Routhiau, V. The immune system and the gut microbiota: Friends or foes?. Nat. Rev. Immunol. 10(10), 735 (2010).
Chestnut, T. et al. Heterogeneous occupancy and density estimates of the pathogenic fungus Batrachochytrium dendrobatidis in waters of North America. PLoS ONE 9(9), e106790 (2014).
Altizer, S. et al. Social organization and parasite risk in mammals: Integrating theory and empirical studies. Annu. Rev. Ecol. Evol. Syst. 34(1), 517–547 (2003).
Colwell, R. R. & Liston, J. Microbiology of shellfish Bacteriological study of the natural flora of Pacific oysters (Crassostrea gigas). Appl. Microbiol. 8(2), 104 (1960).
Romero, J. et al. Bacterial 16S rRNA gene analysis revealed that bacteria related to Arcobacter spp. constitute an abundant and common component of the oyster microbiota (Tiostrea chilensis). Microb. Ecol. 44(4), 365–371 (2002).
Lokmer, A., Kuenzel, S., Baines, J. F. & Wegner, K. M. The role of tissue-specific microbiota in initial establishment success of Pacific oysters. Environ. Microbiol. 18(3), 970–987 (2016).
King, G. M., Judd, C., Kuske, C. R. & Smith, C. Analysis of stomach and gut microbiomes of the eastern oyster (Crassostrea virginica) from coastal Louisiana, USA. PLoS ONE 7(12), e51475 (2012).
Wexler, H. M. Pump it up: Occurrence and regulation of multi-drug efflux pumps in Bacteroides fragilis. Anaerobe 18(2), 200–208 (2012).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285), 59 (2010).
Wu, X. et al. Molecular characterisation of the faecal microbiota in patients with type II diabetes. Curr. Microbiol. 61(1), 69–78 (2010).
Pujalte, M. J., Ortigosa, M., Macián, M. C. & Garay, E. Aerobic and facultative anaerobic heterotrophic bacteria associated to Mediterranean oysters and seawater. Int. Microbiol. 2(4), 259–266 (1999).
Hernández-Zárate, G. & Olmos-Soto, J. Identification of bacterial diversity in the oyster Crassostrea gigas by fluorescent in situ hybridization and polymerase chain reaction. J. Appl. Microbiol. 100(4), 664–672 (2006).
Green, T. J. & Barnes, A. C. Bacterial diversity of the digestive gland of Sydney rock oysters, Saccostrea glomerata infected with the paramyxean parasite, Marteilia sydneyi. J. Appl. Microbiol. 109(2), 613–622 (2010).
Fernandez-Piquer, J. et al. Molecular analysis of the bacterial communities in the live Pacific oyster (Crassostrea gigas) and the influence of postharvest temperature on its structure. J. Appl. Microbiol. 112(6), 1134–1143 (2012).
Jovel, J. et al. Characterization of the gut microbiome using 16S or shotgun metagenomics. Front. Microbiol. 7, 459 (2016).
Pierce, M. L. & Ward, J. E. Microbial ecology of the Bivalvia, with an emphasis on the family Ostreidae. J. Shellfish Res. 37(4), 793–806 (2018).
Trabal, N. et al. Molecular analysis of bacterial microbiota associated with oysters (Crassostrea gigas and Crassostrea corteziensis) in different growth phases at two cultivation sites. Microb. Ecol. 64(2), 555–569 (2012).
Vasconcelos, G. J. & Lee, J. S. Microbial flora of Pacific oysters (Crassostrea gigas) subjected to ultraviolet-irradiated seawater. Appl. Environ. Microbiol. 23(1), 11–16 (1972).
Pillai, C. T. Microbial flora of mussels in the natural beds and farms. CMFRI Bull. 29, 41–43 (1980).
Olafsen, J. A. et al. Indigenous bacteria in hemolymph and tissues of marine bivalves at low temperatures. Appl. Environ. Microbiol. 59(6), 1848–1854 (1993).
Kueh, C. S. & Chan, K. Y. Bacteria in bivalve shellfish with special reference to the oyster. J. Appl. Bacteriol. 59(1), 41–47 (1985).
Wrange, A. L. et al. Massive settlements of the Pacific oyster, Crassostrea gigas, in Scandinavia. Biol. Invasions 12(5), 1145–1152 (2010).
Acknowledgements
I would like to thank Horn-point laboratory for providing the spat used throughout the experiment; Dr. Tara Scully for acting as the lead investigator for the project and encouraging me throughout the process to perfect the required techniques, Karen Gedan for her continued support reviewing written drafts, Courtney Smith for advice and instrument use, and Sony Dalapati, Benjamen McSweeney, and Sophia Duchin for help in conducting extractions. And finally, The George Washington University, for both housing the project and providing the funds necessary for it to be carried out.
Author information
Authors and Affiliations
Contributions
M.B. and K.O. helped to write the conclusion. B.M., S.D., S.D., K.N.T.P. and K.C. helped to prepare the protocol for bacterial DNA acquisition. N.S., S.N., M.R. and G.D. helped to care for animals used in study and conducted background research into bacteria found within obtained samples. T.S. assisted with the discussion and introduction. A.B. conducted the protocols for DNA extraction and edited the entire paper. DNA Sequence Accession number: PRJNA575277.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Britt, A., Bernini, M., McSweeney, B. et al. The effects of atrazine on the microbiome of the eastern oyster: Crassostrea virginica. Sci Rep 10, 11088 (2020). https://doi.org/10.1038/s41598-020-67851-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-67851-4
This article is cited by
-
Toxicity of Atrazine to Marine Invertebrates Under Flow-Through Conditions—Eastern Oyster (Crassostrea virginica) and Mysid Shrimp (Americamysis bahia)
Water, Air, & Soil Pollution (2021)
-
Endocrine Disrupting Chemicals in Aquatic Ecosystem: An Emerging Threat to Wildlife and Human Health
Proceedings of the Zoological Society (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.