Abstract
The function of IM30 (also known as Vipp1) is linked to protection and/or remodeling of the thylakoid membrane system in chloroplasts and cyanobacteria. Recently, it has been revealed that the Arabidopsis IM30 protein exhibits GTP hydrolyzing activity in vitro, which was unexpected, as IM30 does not show any classical GTPase features. In the present study, we addressed the question, whether an apparent GTPase activity is conserved in IM30 proteins and can also be observed for IM30 of the cyanobacterium Synechocystis sp. PCC 6803. We show that Synechocystis IM30 is indeed able to bind and hydrolyze GTP followed by the release of Pi. Yet, the apparent GTPase activity of Synechocystis IM30 does not depend on Mg2+, which, together with the lack of classical GTPase features, renders IM30 an atypical GTPase. To elucidate the impact of this cryptic GTPase activity on the membrane remodeling activity of IM30, we tested whether GTP hydrolysis influences IM30 membrane binding and/or IM30-mediated membrane fusion. We show that membrane remodeling by Synechocystis IM30 is slightly affected by nucleotides. Yet, despite IM30 clearly catalyzing GTP hydrolysis, this does not seem to be vital for its membrane remodeling function.
Similar content being viewed by others
Introduction
IM30, the inner membrane-associated protein of 30 kDa (also known as Vipp1 (vesicle-inducing protein in plastids 1)), is present in cyanobacteria as well as in chloroplasts of higher plants and algae1. In chloroplasts, the protein has a triple localization: It can be found soluble in the stroma as well as bound to the inner chloroplast envelope membrane and the thylakoid membrane (TM)2. Similarly, IM30 is also found in a soluble form as well as bound to the cytoplasmic membrane and TMs of cyanobacteria3.
IM30 is a predominantly α-helical protein predicted to consist of seven α-helices, which are able to form coiled-coil structures3,4,5. In vitro, IM30 monomers assemble to form diverse higher ordered homo-oligomers with molecular masses exceeding 2 MDa4,5,6,7. In vivo, in cyanobacteria and in chloroplasts, GFP-labeled IM30 forms large dynamic assemblies, which are mainly located at TM regions with high membrane curvature8,9,10.
While diverse functions have been attributed to IM30 in the past1,2,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24, most observations clearly indicate that the protein is membrane-active and involved in the biogenesis, dynamics and/or stabilization of internal membranes in chloroplasts and cyanobacteria (recently reviewed in25). Thus, the physiological function of IM30 appears to be linked to dynamic membrane remodeling25,26,27, likely involving membrane fusion/fission events, which have been observed in plant chloroplasts28,29,30,31. In fact, dynamic rearrangement of the TM system is crucial for adaptation of photosynthetic processes to altering environmental conditions, most importantly in response to altering light intensities32,33. While TM dynamics is studied to some extent in chloroplasts, direct observations of TM dynamics in cyanobacteria are still limited, although TMs in cyanobacteria are assumed to be as dynamic as in chloroplasts34,35. Nevertheless, a machinery mediating such membrane remodeling processes has not been unambiguously identified yet, neither in chloroplasts nor in cyanobacteria36, and IM30 is currently the only protein in chloroplast and cyanobacteria recognized to have a membrane fusion activity23. Importantly, light not only triggers rearrangement of the TM system but also rearrangement and redistribution of intracellular IM30 clusters at internal membranes8,26, which again indicates a connection between TM dynamics and IM30 oligomer formation25.
In several aspects, the membrane activity of IM30 resembles other membrane-active proteins, as IM30 is involved in membrane fusion, protection and/or stabilization10,15,16,18,23,37,38 and has a strong tendency to oligomerize and to form higher ordered structures3,4,7. Yet, in most thus far described systems, membrane remodeling, especially membrane fission and fusion, are typically connected to nucleotide hydrolysis. Thus, it was somewhat intriguing to suspect a nucleotide binding/hydrolysis function also for IM30, a protein that can mediate membrane fusion23. Consistent with this idea, Ohnishi et al. recently suggested that recombinant Arabidopsis IM30 (AraIM30) has an intrinsic GTPase activity despite lacking classical features of GTPases, implying that IM30 belongs to a new class of membrane-remodeling GTPases39. Yet, the molecular details of the IM30 membrane activity have been extensively studied in vitro in recent years, and all membrane-related activities were already observed in the absence of nucleotides. Thus, the question arises whether GTP binding and/or hydrolysis have any impact on the structure and/or activity of IM30.
In the present study, we have analyzed the apparent GTPase activity of IM30 of the cyanobacterium Synechocystis sp. PCC 6803. We have tested the influence of nucleotides on IM30-triggered membrane fusion and lipid organization in vitro. Our results suggest that the Synechocystis IM30 also has a low GTP hydrolyzing activity, as observed before with the Arabidopsis homolog, yet nucleotides have only minor effects on the in vitro membrane remodeling activity of IM30, and therefore nucleotide binding/hydrolysis does not appear to critically affect IM30-triggered membrane remodeling.
Results
SynIM30 has an apparent GTPase activity
The Arabidopsis IM30 protein (AraIM30) appears to have a GTPase activity under defined in vitro conditions39. As this has never been described for any IM30 protein before, we first aimed to elucidate whether this apparent activity is conserved in IM30 proteins. Therefore, we analyzed the interaction of a cyanobacterial IM30 homolog (Synechocystis IM30; SynIM30) with GTP.
Initially, we tested whether SynIM30 hydrolyzes GTP, using the malachite green-based assay that has also been used by Ohnishi et al. to determine the apparent GTPase activity of AraIM3039.
Indeed, we observed protein-dependent Pi production, i.e. GTP hydrolysis, in perfect agreement with the findings of Ohnishi et al., reaching about 20 µM Pi release at a protein concentration of 2 µg/100 µL (Fig. 1a). To rule out that Pi release of the protein originated from contamination with a GTPase, we analyzed the purity of the protein via SDS-PAGE, whereby no other protein was identified besides IM30 (Fig. 1g). Although this does not finally exclude any contamination with a highly active GTPase, the missing GTPase activity of an IM30 mutant (Fig. 3a), purified following the exact same protocol, is a strong hint that the observed GTPase activity can indeed be exclusively ascribed to IM30, as further discussed below.
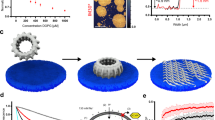
IM30 GTPase activity was characterized by multiple assays. (a) A malachite green-based assay was used to measure the Pi release catalyzed by SynIM30 WT (0.5 mM GTP; 2.5 mM Mg2+; 30 min at 37 °C). (Error bars represent SD, n = 3). (b) The enzymatic properties of the GTP hydrolysis caused by SynIM30 were analyzed with a malachite green-based assay in absence and presence of 0.5 mM GMP-PCP (0.1 µM IM30 WT; 2.5 mM Mg2+; 30 min at 37 °C). Fitting the data with a Hill equation derived model yielded \({V}_{max}\) value of 2.16 ± 0.33 mol GTP per mol IM30 per min, Km = 0.65 ± 0.14 mM; n = 1.29 ± 0.09 and \({V}_{max}\) = 0.83 ± 0.33 min−1; Km = 0.88 ± 0.16 mM; n = 5.19 ± 3.64 in presence of GMP-PCP. (R2 = 0.99835 and R2 = 0.95888; SD, n = 3). (c) A fluorescence-based continuous test for Pi release was used to determine SynIM30 GTPase activity in real-time (0.1 µM IM30 WT; 10 µM GTP; 2.5 mM Mg2+; 37 °C). Fitting the data with a mono-exponential function yielded v0 = 1.79 ± 0.02 min−1 and \(\eta \) = 2.79 ± 0.05 *10−2 (R2 = 0.97898; SD, n = 3). (d) A PK/LDH-coupled assay was used to determine the GTPase activity independent of the Pi release. SynIM30 WT did not show significant activity (SD, n = 3). As a control, a GTPase with known activity was used (SynDLP; n = 1). (e) A luciferase-coupled GTPase assay was used to measure the amount of hydrolyzed GTP over 120 min at 37 °C in the presence of increasing SynIM30 concentrations (4 µM GTP). (Error bars represent SD, n = 3) (f) GTPase activity of IM30 WT was measured by the malachite green-based assay in absence and presence of 30 µM DOPG (2 µg/200 µL protein, 0.5 mM GTP; 2.5 mM Mg2+; 30 min at 37 °C). The GTPase activity is not significantly increased in the presence of the liposomes. (SD, n = 3, two-sample Student´s t-test, significance level p > 0.05). (g) The purity of SynIM30 WT heterologously expressed in E. coli and purified via Ni2+-affinity chromatography was analyzed by SDS-PAGE.
Subsequently, we used the malachite green-based assay to further characterize the apparent GTPase activity of IM30. The steady-state activity of IM30 in dependence on the GTP concentration could be described by a slightly sigmoidal curve (Fig. 1b). Fitting the experimental data with the Hill-equation (Eq. 2) resulted in a \({k}_{cat}\) value of 2.16 ± 0.33 mol GTP per mol IM30 per min, a Michaelis-Menten constant of 0.65 ± 0.14 mM and a Hill coefficient of 1.29 ± 0.09. Addition of the non-hydrolyzable GTP analog GMP-PCP, which is expected to act as a competitive inhibitor, strongly increased the cooperativity, as evidenced by the increased Hill coefficient (n = 5.19 ± 3.64) (Fig. 1b).
As the malachite green-based assay is not capable of measuring reaction kinetics, we next applied a continuous assay for Pi release, which is based on a recombinant E. coli phosphate-binding protein labeled with the fluorophore MDCC40. This assay indeed allowed monitoring the kinetics of the apparent IM30 GTPase activity, measured as the Pi release (Fig. 1c). Fitting the data with a model for non-linear, steady-state enzyme kinetics and assuming a first-order process41 yielded a kcat (Eqs. 3 and 4) in presence of 10 µM GTP (1.79 ± 0.02 * 10−2 min−1) (Fig. 1c). However, these measurements could be performed solely at very low GTP concentrations, as the Pi sensor has an extremely high sensitivity for Pi.
Although specifically established for measuring NTPase activities, the malachite green-based assay is prone to artifacts, and e.g. protein binding to the malachite complex or acids can induce NTP hydrolysis, causing false positive results42. Thus, we aimed to apply an established PK/LDH-coupled assay that measured GTP hydrolysis by coupling the regeneration of GTP from GDP to a decline in NADH concentration, which can be monitored by absorbance at 340 nm43. Surprisingly, when using this assay we were not able to measure a significant amount of GTP hydrolysis (Fig. 1d). Even doubling the SynIM30 concentration did not produce a detectable signal (data not shown). Based on the Pi release determined with the malachite green assay (20 µM Pi/30 min) we calculated the expected change in absorption at 340 nm, amounting to ΔOD = 0.124 in 30 min. This should be easily detectable if GDP is released. To ensure that the assay works properly, we also analyzed the GTPase activity of SynDLP, a canonical P-loop GTPase of the cyanobacterium Synechocystis sp. PCC680336, as a positive control (Fig. 1d).
Since we were not able to detect any GTP hydrolysis using the PK/LDH-coupled assay, we wondered whether the observed apparent GTPase activity might be caused by Pi leaking into the reaction mixture, e.g. by release of Pi bound to IM30 rather than by GTP hydrolysis. To test this, we used a luciferase-coupled GTPase assay, which directly depends on the GTP concentration rather than Pi-release. In this assay, non-hydrolyzed GTP is enzymatically converted to ATP and the resulting ATP concentration is subsequently measured in the luciferase reaction. Here we observed a steady decrease of the GTP concentration in the presence of IM30 in a protein concentration-dependent manner, as expected for a GTP hydrolyzing activity (Fig. 1e). Thus, we conclude that SynIM30 can indeed catalyze the hydrolysis of GTP.
The activity of many membrane-active GTPases is modulated when the protein interacts with lipid membranes, and interaction of e.g. dynamins with membrane surfaces can increase the GTPase activity by a factor of 4044,45. As IM30 binds to negatively charged PG liposome surfaces23,24,37,46, we next assayed GTP hydrolysis by SynIM30 in absence vs. in presence of DOPG liposomes. While our measurements were limited to low lipid concentrations, as the liposomes created a strong background signal in the assay, we did not observe a significant increase in GTP hydrolysis in presence of DOPG liposomes (p = 0.066) (Fig. 1f).
In summary, SynIM30 clearly has an apparent GTPase activity in vitro, as observed before with the AraIM30 protein. This activity can be modulated by a classical GTPase inhibitor and does not seem to be affected by membranes.
IM30 binds and hydrolyzes GTP in the presence and absence of Mg2+
Thus far, apparent GTP hydrolysis was always tested in presence of Mg2+, as the activity of NTPases typically requires the presence of Mg2+. Notably, the interplay of IM30, Mg2+ and GTP is of special interest, as also the membrane fusion activity of IM30 depends on Mg2+ 23, and binding of Mg2+ to IM30 even causes rearrangement of the IM30 structure47.
To test the Mg2+ dependence of the IM30 GTPase activity we determined GTP hydrolysis rates in the absence of Mg2+. Surprisingly, the KM value (0.61 ± 0.20 mM) determined for GTP hydrolysis was essentially identical, regardless of the presence of Mg2+. Similarly, the \({v}_{max}\) value (1.65 ± 0.55 min−1) did not significantly differ from the value determined in the presence of Mg2+ (2.16 ± 0.33 min−1) (Fig. 2a). Thus, Mg2+ does obviously not have a strong impact on GTP hydrolysis by SynIM30. Yet, cooperativity was strongly increased in absence of Mg2+, reaching a value of 2.00 ± 0.27, which nicely supports the recently described Mg2+-induced change of the IM30 structure and stability47. In fact, addition of Mg2+ e.g. increased the thermal stability of IM30 as indicated by Tm values of 66.6 ± 0.2 °C and 68.7 ± 0.2 °C in absence and presence of 2.5 mM Mg2+, respectively (Fig. 2b). However, an increased stability of SynIM30 was observed when 0.5 mM GTP and 2.5 mM Mg2+ were present together (Tm 67.6 ± 0.2 °C) compared to SynIM30 in presence of solely 2.5 mM GTP, where the Tm was as low as 64.6 ± 0.2 °C (Fig. 2b). This suggests that GTP binding affects the structure and stability of IM30 to some extent, and Mg2+ and GTP do not operate synergistically but have opposing effects on the SynIM30 stability.
GTP binds to IM30 in presence and absence of Mg2+. (a) GTP hydrolysis caused by IM30 WT in absence of Mg2+ was analyzed using a malachite green-based assay (0.1 µM IM30 WT; 0 mM Mg2+; 30 min at 37 °C). Fitting of the data with a Hill equation derived model yielded \({V}_{max}\) = 1.66 ± 0.55 min−1; Km = 0.61 ± 0.20 mM; n = 2.00 ± 0.27 in absence of Mg2+. (R2 = 0.96951; SD, n = 3). Data in presence of Mg2+ were taken from Fig. 1b (control). (b) Thermal denaturation of IM30 WT in the absence and presence of GTP and Mg2+ was monitored by CD-spectroscopy at 222 nm. The raw signal was converted to the fraction of denatured protein (fD). Fitting the data with an adapted Boltzmann-fit yielded: Tm = 66.6 ± 0.2 °C (0 mM GTP; 0 mM Mg2+) (R2 = 0.99866); Tm = 68.7 ± 0.2 °C (0 mM GTP; 2.5 mM Mg2+) (R2 = 0.99841); Tm = 64.8 ± 0.2 °C (0.5 mM GTP; 0 mM Mg2+) (R2 = 0.99836) and Tm = 67.5 ± 0.2 °C (0.5 mM GTP; 2.5 mM Mg2+) (R2 = 0.99784). Error bars represent errors from the fitting of the data.
Thus, GTP binding to SynIM30 and SynIM30-mediated GTP hydrolysis appear to be independent of Mg2+, which is rather uncommon for GTPases.
GTP hydrolysis depends on the oligomeric state of IM30
Based on studies using truncated AraIM30 it has been suggested that helix 1 is crucial for GTP binding to AraIM30 and for its apparent GTPase activity39. As we assume that the apparent GTPase activity is conserved in IM30 proteins of different origins, we next tested the GTP hydrolyzing activity of the recently described SynIM30 variant H2-7, where helix 1 was deleted46. Although this mutant showed a decreased Pi release rate compared to the wt protein (Fig. 3a), Pi release was decreased only by about 40% (i.e. to roughly 60% of the wt levels), whereas the GTPase activity of the AraIM30 protein was completely abolished when helix 1 was deleted39. However, the AraIM30 protein does not form high molecular mass oligomers anymore when helix 1 is deleted4,39, whereas IM30 of Chlamydomonas rheinhardtii5 and the here analyzed SynIM30 do46, as recently discussed in25. Yet, a fraction of smaller oligomers was also identified in SEC analyses of the SynIM30 H2-7 protein (see Fig. 3b). Thus, the disturbed formation of high molecular mass oligomers might explain the abolished GTPase activity of the AraIM30 H2-7 protein39, and the reduced amount of released Pi by SynIM30 H2-7 (Fig. 3b).
The oligomeric state of IM30 is crucial for GTP hydrolysis. (a) The malachite green-based assay was used to compare the GTPase activity of IM30 variants with different oligomeric states (0.5 mM GTP; 2.5 mM Mg2+; 30 min at 37 °C). Compared to IM30 WT, H2-7 has a reduced GTPase activity. The tetrameric IM30 FERM_EE has no detectable GTPase activity. (Error bars represent SD, n = 3). (b) The oligomeric state of IM30 variants was determined by SEC (Superose 12 10/300 GL column). The elution volumes determined are 7.4 mL for IM30 WT (>300 kDa); 7.7 mL (>300 kDa) and 10.2 mL (105 kDa) for IM30 H2-7 and 10.3 mL (100 kDa) for IM30 FERM_EE. The asterisk marks the position of the void volume. T marks the position of an IM30 tetramer. (c) The data of the malachite green assay of IM30 WT in absence and presence of Mg2+ and after addition of GMP-PCP were analyzed using a more sophisticated model (described in detail in the Supplemental Data 3), to disentangle the mechanism of GTP-hydrolysis. Error bars represent SD, n = 3. Fit: n = 12, allosteric model (see Supplemental Data 3).
To further test the hypothesis that the decreased oligomeric state, and not deletion of helix 1, has caused the reduced apparent GTPase activity, we next generated a SynIM30 variant that exclusively forms small oligomers (mainly tetramers in buffers with low ionic strength) (SynIM30 FERM_EE) and no high molecular mass oligomers (Fig. 3b). This mutant is based on the SynIM30_FERM variant described recently7,37, but additionally E83 and E84 in the loop region of helix 2 and 3 were mutated to overcome the problem of unspecific aggregation upon prolonged exposure to HEPES buffer. In line with our hypothesis, this variant had no detectable GTPase activity (Fig. 3a). Note that also SynIM30_FERM does not exhibit detectable GTPase activity (Suppl. Fig. 1).
Thus, we conclude that the IM30 GTPase activity depends on the IM30 oligomeric state, and formation of high molecular mass oligomers is a prerequisite of GTP hydrolysis. A lack of high molecular mass oligomers seems to be the major factor for GTP hydrolysis since helix 1 is unchanged in the tetrameric SynIM30 variant. Together, these results clearly suggest that not helix 1 but the formation of high molecular mass oligomers is crucial for GTP hydrolysis. In addition, the absent GTP hydrolyzing activity of the mutant protein confirms that the GTPase activity observed with the IM30 wt (Fig. 1) was not caused by an (unintended) co-purification of an (unknown) GTPase.
Considering that IM30 forms oligomers, the sigmoidal shape of the GTPase activity in presence of GMP-PCP (Fig. 1b) and in absence of Mg2+ (Fig. 2a) might well reflect the interaction of GTP-binding sites within the protein. Indeed, the data describing the GTPase activity of IM30 can be well depicted by a cooperative two-state model (Fig. 3c, see Suppl. Fig. 3 for details). The increase in cooperativity in presence of GMP-PCP can be explained by competitive binding, which would shift the distribution towards the low-affinity state, requiring more GTP to yield measurable activity. Formally, also binding to an allosteric site could explain the data. Both variants of the two-state model describe the data similarly well (see Supplemental Data 3). The number of binding sites interacting, giving rise to the observed cooperativity, is ≥8 (Suppl. Fig. 3d). Thus, these data correspond well with the idea of having multiple subunits interacting with each other during GTP binding/hydrolysis, which explains the necessity of IM30 rings/high molecular weight complexes for GTPase activity.
IM30 binding to membrane surfaces is not influenced by GTP
IM30 binds to negatively charged membrane surfaces and is involved in membrane remodeling16,23,37. Thus, we next studied the impact of GTP on binding of SynIM30 to negatively charged membrane surfaces.
In the presence of Mg2+, SynIM30 exhibits membrane fusion activity, and such membrane fusion events can disturb other assays, involving IM30 and liposomes23. Luckily, GTP binding to SynIM30 and GTP hydrolysis were observed also in absence of Mg2+ (Fig. 2a,b). Thus, we were able to analyze the influence of nucleotide binding to SynIM30 on membrane binding independent of the effect on IM30-mediated membrane fusion.
First, we analyzed whether GTP has an impact on the propensity of SynIM30 to interact with negatively charged DOPG liposomes. It has previously been shown that binding of SynIM30 to DOPG membranes can be monitored via steady-state Laurdan fluorescence spectroscopy23,37. As shown in Fig. 4a,b, the Laurdan fluorescence emission spectrum was already altered upon addition of solely GTP or GDP, even when SynIM30 was not present, albeit only to a minor extent. Thus, the nucleotides appear to interact with the liposomes, resulting in a minor change of the Laurdan fluorescence emission. However, when SynIM30 was added to the liposomes, the Laurdan fluorescence spectrum changed far more significantly, and an altered fluorescence spectrum was observed in absence as well as in presence of GTP or GDP. All spectra recorded in presence of SynIM30 (Fig. 4a) showed an increased fluorescence emission at 440 nm and a decreased emission at 490 nm, leading to an increase of the GP-value. This indicates a decreased polarity of Laurdan’s environment, as occurring when the lipid order increases. The changes in the lipid order, represented by the ΔGP values, showed no significant impact of GTP or GDP on the interaction of SynIM30 with DOPG liposomes (Fig. 4c), indicating that GTP binding and/or hydrolysis by SynIM30 do not affect binding of IM30 to lipid bilayers, at least not in the absence of Mg2+. Unfortunately, it was not possible to analyze SynIM30 binding to DOPG liposomes in the presence of Mg2+ due to direct binding of Mg2+ to DOPG membranes as well as to to SynIM3047 and side-effects caused by SynIM30-mediated membrane fusion23.
The presence of the nucleotides GTP and GDP does not affect the IM30-mediated change of the DOPG lipid order. (a) Laurdan fluorescence emission spectra of 0.1 mM DOPG liposomes were recorded in absence and presence of 2.5 mM GTP or GDP and 1 µM SynIM30 WT. Solid lines show the spectra in absence of IM30. Dashed lines show the spectra in the presence of SynIM30 WT. Binding of IM30 leads to a clear shift of the Laurdan fluorescence maximum. (b) The GP value was calculated for DOPG liposomes in the presence and absence of GTP and GDP without addition of SynIM30. Addition of the nucleotides leads to a small, but still significant decrease of the GP-value. SD, n = 6 two-sample Student´s t-test, *p < 0.05, **p < 0.01, ***p < 0.005. (c) The ΔGP after addition of SynIM30 WT to DOPG liposomes is not significantly changed in the presence of the nucleotides GTP or GDP, respectively. (SD, n = 6 two-sample Student´s t-test).
GTP and GDP do only marginally affect the kinetics of IM30-mediated membrane fusion
SynIM30 has a membrane fusion activity when Mg2+ is present (Hennig et al., 2015). Consequently, we next analyzed the potential impact of GTP and GDP on IM30-induced liposome fusion in the presence of Mg2+.
Membrane fusion was measured using a FRET-based liposome fusion assay, as described in detail previously23. Importantly, in presence of the nucleotides, the concentration of MgCl2 had to be adjusted, as GTP and GDP coordinate Mg2+ via their phosphate groups48 and thereby lower the effective concentration of “free” Mg2+ which clearly influences the membrane fusion process23. The concentration of additionally “needed” MgCl2 was determined experimentally in the presence of GTP and GDP, as described in the “Experimental Procedures” section. Note that controls containing exclusively IM30, Mg2+, the nucleotides or the respective Mg2/nucleotide mixture did not show any membrane fusion activity (data not shown).
In contrast to membrane fusion observed in absence of nucleotides, the fusion assay indicated a slightly increased fusion rate in presence of GTP, while slightly less liposome fusion was observed in presence of GDP (Fig. 5). The initial fusion rate increased from 0.58 ± 0.20%/s in the absence of nucleotides to 0.80 ± 0.22%/s in the presence of GTP. In contrast, addition of GDP lowered the initial fusion rate to 0.33 ± 0.04%/s. However, in all cases the fusion curves appear to level out at approximately the same amount of total fusion.
IM30-mediated membrane fusion is modulated to some extent by GTP and GDP. Liposome fusion was measured in the presence of 1 µM SynIM30 WT, 2.5 mM nucleotide and 6 mM Mg2+. Because of binding of free Mg2+ by the nucleotides, “extra” needed Mg2+, needed to obtain the same membrane destabilization, was also added (2.5 mM for GDP, 4.25 mM for GTP). The presence of GTP leads to a slight increase in the fusion rate, whereas the presence of GDP leads to a slower fusion process. (SD, n = 3).
Clearly, GTP binding/hydrolysis is not unequivocally required for IM30-mediated membrane fusion, and the presence of GTP or GDP does neither abolish nor exclusively trigger membrane fusion. We conclude therefore that the GTP hydrolyzing activity of IM30 is not a key regulator of the membrane fusion process, as it is not switching the IM30 membrane fusion activity from a completely active to an inactive state, or vice versa. Yet, as we did observe small effects on the fusion kinetics in the presence of GTP and GDP, we cannot completely rule out the possibility that nucleotide binding and/or GTP hydrolysis modulate IM30-mediated membrane fusion.
Discussion
SynIM30 has a low GTP binding affinity and hydrolyzes GTP with low rates in the absence of Mg2+
Involvement of the IM30 protein in membrane protection and/or membrane dynamics in chloroplasts and cyanobacteria has been described in the past, and recently, an apparent GTPase activity of AraIM30 has been observed39. Yet, this observation was unexpected, as none of the known IM30 protein sequences does contain predicted domains or amino acid motifs known to be crucial for nucleotide binding and/or hydrolysis (not shown). Thus, the question arose whether the presumed GTPase activity is AraIM30-specific or more generic. Therefore, in the present study, we analyzed the GTPase activity of SynIM30. We here show that SynIM30 can bind GTP, resulting in GTP hydrolysis and release of free Pi. This apparent GTPase activity can be modulated by a classical GTPase inhibitor (GMP-PCP).
In fact, when analyzed with the malachite green assay, SynIM30 has a (rather low) GTPase activity with a kcat of 2.16 ± 0.33 min−1, a value comparable to the activity described for some other membrane remodeling GTPases, such as dynamin in absence of membranes (kcat 2.60 ± 0.98 min−1)49. However, it is worth mentioning that the kcat determined using the malachite green assay (Fig. 1b) is essentially derived from the activity averaged over the entire incubation time, and the determined values do thus not account for any bias on the turnover rates, such as substrate limitation/substrate inhibition, etc. In an attempt to address these limitations, we applied a continuous phosphate release assay (Fig. 1c) that is in principle capable of measuring the actual rate of Pi release and can therefore be used to determine the initial, non-biased turnover rate \({v}_{0}\). Unfortunately, when using this assay we were limited to a very small concentration of the substrate (10 µM GTP), which was significantly lower than the KM value determined for SynIM30 with the malachite green assay (KM 0.65 ± 0.14 mM). Based on the parameters obtained for the Hill-equation, the saturation level of IM30 at this GTP concentration is about 0.5%. Thus, under these conditions, the expected kcat is about 0.01 min−1, which is reasonably well in agreement with the experimentally observed kcat (0.0179 ± 0.02 min−1) (Fig. 1c). However, we were not able to observe GTP hydrolysis when using a PK/LDH-coupled assay (Fig. 1d) that is well-established in the NTPase field. Assuming the same activity as observed with the malachite green assay, we would expect to observe a change of OD 0.124/30 min, which should be easily detectable. This can either be explained by IM30 inhibiting the assay and/or vice versa or by slow koff rates for GDP.
Nevertheless, SynIM30 clearly shows GTP hydrolysis activity, as demonstrated by the luciferase-coupled GTPase assay, which directly depends on the decrease of the GTP concentration. Again, the observed rate is in reasonable agreement with the KM and kcat values obtained from the malachite green-based assay, taking the low GTP concentration of 4 µM into account. However, IM30 binds GTP rather weakly, compared to KD values of typical GTPase. E.g. GTPases of the ras superfamily, which are considered high-affinity GTPases, have KD values in the subnanomolar range50, whereas low-affinity GTPases, such as dynamins or the signal recognition particle (SRP), have KD values in the range of 0.5–5 µM50. Thus, typical low-affinity GTPases still bind GTP with at least 100-fold higher affinities than observed here for SynIM30.
Addition of GMP-PCP, a non-hydrolyzable GTP analog, revealed strong cooperative properties of the apparent GTPase activity, which was essentially not observed in the absence of GMP-PCP. Within the frame of an allosteric two-state model, both allosteric and competitive binding could produce such an increase in cooperativity (see Suppl. Fig. 3). While the data did not allow to identify the exact nature of GMP-PCP inhibition, the observations demonstrate that binding of GTP is a cooperative process, probably involving more than four binding sites (see Suppl. Fig. 3d).
While SynIM30 exhibits a GTPase activity, which can be modulated by a non-hydrolyzable GTP analog, this activity clearly is Mg2+-independent (Fig. 2a), which is rather uncommon for canonical GTPases. Typically, Mg2+ is required for GDP and GTP binding and hydrolysis. Yet, absence of Mg2+ does neither affect GTP binding nor GTP hydrolysis by SynIM30 (Fig. 2), and both KM and \({v}_{max}\) were about identical in presence or absence of Mg2+ (Fig. 2a). While binding of GTP and an apparent GTPase activity in absence of Mg2+ have already been described for other GTPases, involving FtsZ51 or Rho family GTP-binding proteins52, in these cases Mg2+ addition dramatically increased the hydrolysis rates. In fact, here Mg2+ binding was suggested to regulate the kinetics of GTP binding and hydrolysis to achieve high catalytic efficiency and specificity. Thus, a fully Mg2+-independent GTP hydrolyzing activity is rather uncommon. Nevertheless, while \({v}_{max}\) remained unchanged in presence of Mg2+, the cooperativity of GTP hydrolysis by the oligomeric IM30 was significantly increased. Mg2+ is known to directly bind to SynIM30 and to alter the structure of SynIM30 oligomers47, which, vice versa, e.g. results in an increased thermal stability of the protein (Fig. 2b). Thus, a model, where binding of Mg2+ leads to a shift in the conformational distribution between different structural states, is reasonable and explains the observed change in cooperativity well, while \({v}_{max}\) remains constant.
GTP hydrolysis depends on the oligomeric state
While IM30 proteins do not contain any canonical GTPase domain or any motif known to be involved in NTP binding and/or hydrolysis, Ohnishi et al. have identified helix 1 of AraIM30 to be crucial for GTP hydrolysis, as deletion of helix 1 resulted in reduced GTP hydrolysis39. However, deletion of helix 1 also resulted in disassembly of the typical IM30 high molecular mass complexes in case of the AraIM30 protein4,39, and thus the GTP hydrolyzing activity could well be coupled to the oligomeric state of IM30 proteins. In fact, we show here that GTP binding/hydrolysis does depend on the oligomeric state of IM30 but does not crucially involve helix 1. As SynIM30 H2-7 still had considerably high GTP hydrolyzing activity and as we assume a conserved function of the helices in IM30 proteins of different organisms, we rule out the possibility that any amino acid in helix 1 is crucial for GTP binding or hydrolysis. However, the AraIM30 oligomer appears to be less stable than the proteins of Synechocystis or Chlamydomonas reinhardtii, as in the latter cases helix 1-truncated versions reportedly still form high molecular mass complexes5,25,46. Thus, we assumed that oligomer formation, rather than helix 1, is crucial for GTP hydrolysis by IM30 proteins, and we indeed confirmed this assumption by using a SynIM30 full-length mutant protein that forms tetramers but no high molecular mass complexes anymore (Fig. 3b). As this mutant did not hydrolyze GTP anymore, a tetrameric IM30 structure likely is insufficient for GTP hydrolysis and higher oligomeric structures are required, most likely prototypical IM30 rings7. As the analyzed SynIM30 mutant forms tetramers, which are considered to be the basic building blocks of IM30 rings7,37, the putative GTP binding sites are potentially localized within the ring in between adjacent tetrameric building blocks.
GTP binding and hydrolysis do not critically affect IM30-mediated membrane remodeling
While we could clearly establish that SynIM30 high molecular mass oligomers do hydrolyze GTP (also in absence of Mg2+), the important question arose whether this cryptic GTPase activity has any impact on the physiological function of the protein. Unfortunately, the exact function of IM30 is still not finally resolved. Yet, in vitro analyses clearly show that the protein binds to negatively charged membrane surfaces and is able to mediate membrane fusion16,23,24,37,46. In fact, in many cases a membrane remodeling activity of NTPases is nucleotide-dependent and typically requires nucleotide hydrolysis (reviewed in53). Nevertheless, membrane interaction and membrane remodeling by IM30 clearly do not require nucleotide binding and/or hydrolysis, as NTPs were never present in any assay reported thus far4,16,17,23,24,37,46. As a stimulating effect of a lipid surface on a protein’s GTPase activity has been shown for many other proteins, such as dynamin-like proteins49, it was well possible that GTP binding and/or hydrolysis modulate the membrane interaction and/or the fusion activity of SynIM30. However, our results suggest that the presence of GTP or GDP does not affect membrane binding of SynIM30, at least in absence of Mg2+ (Fig. 4a,b), and also the GTPase activity of SynIM30 is not significantly enhanced in presence of DOPG liposomes (Fig. 1f). However, a minor effect on GTPase activity might have been unnoticed due to the rather low affinity of IM30 to lipid membranes37, leaving a considerable fraction of the protein unaffected upon presence of the membranes.
Analysis of the fusion events was challenging, as direct binding of Mg2+ is necessary for IM30 activation23,47 as well as for membrane destabilization, when Mg2+ is interacting with negatively charged liposomes54,55,56,57,58,59. Thus, while the exact same concentration of “free” Mg2+ is necessary for comparing the fusion data, the amount of free Mg2+ is clearly altered in the presence of GTP or GDP due to the nucleotide-Mg2+ interaction48. We minimized this effect via experimentally adjusting the concentration of Mg2+ specifically when GTP or GDP was present (as described in the “Experimental Procedures” section). Nevertheless, we cannot rule out that the observed differences of the fusion kinetics (Fig. 5) were biased by our approach to experimentally determine the required Mg2+ concentrations in presence of GTP or GDP to gain similar fusion curves in absence of IM30 (Suppl. Fig. 4).
Thus, the results of the membrane fusion assay have to be treated with caution. Yet, the kinetics of IM30-mediated membrane fusion were somewhat altered in the presence of GDP or GTP. While GTP enhanced the fusion rate, the membrane fusion rate was somewhat decreased in the presence of GDP (Fig. 5).
However, any influence of potential direct interaction of the nucleotides with the lipid bilayer, as possibly observed in the present study (Fig. 4a and b), is not considered in our analysis. In fact, in presence of Mg2+, GTP or GDP could well interact with the negatively charged DOPG head group, as Mg2+ can “bridge” the phosphate groups of the nucleotides and the PG head group. This has e.g. been discussed for binding of DNA to anionic and zwitterionic membrane surfaces60,61. Consequently, nucleotides binding to the membrane surface might directly affect fusion-related membrane properties, which eventually alters fusion kinetics. Furthermore, membrane fusion is affected by multiple, complex equilibria involving IM30, Mg2+, liposomes and the nucleotides that are furthermore constantly changing during membrane fusion and GTP hydrolysis. The small changes in the fusion rate observed here are consequently no decisive hint for any influence on the membrane fusion. Importantly, major impacts on the IM30 (in vitro) fusion activity can be detected via the fusion assay, as e.g. observed analyzing IM30 mutants7.
We conclude that GTP binding and/or hydrolysis are no key regulators of IM30 membrane binding or membrane fusion. If at all, GTP binding/hydrolysis only slightly modulates the kinetics of the membrane fusion process. Hence, membrane remodeling by SynIM30 is vastly nucleotide-independent, despite the cryptic GTP hydrolyzing activity.
IM30 is an atypical GTPase
Besides apparently hydrolyzing GTP, at least in vitro, SynIM30 does not exhibit features expected for a canonical GTPase, involving (i) a G-domain and (ii) Mg2+ dependent substrate binding.
Although the G-domain or the P-loop motif, respectively, are fingerprints of GTPases, a few examples of G-proteins lacking the P-loop motif are described62. The most extensively studied members of such untypical GTPases are Tubulin and FtsZ, which have a highly conserved nucleotide-binding site that clearly differs from typical GTPases63. While IM30 proteins do not contain any sequences involved in nucleotide binding to any thus far studied protein, we cannot per se exclude that GTP binds to a new, currently not described GTP-binding pocket to IM30 proteins. Furthermore, as our results suggest that GTP hydrolysis requires oligomeric structures larger than tetramers, it is well possible that amino acids of different IM30 monomers together form the GTPase site, which would not be easily predictable and could only be visualized in a high-resolution structure of oligomeric IM30. Unfortunately, neither a high-resolution structure of an oligomer nor of the monomer is available for IM30 yet. Thus, structural comparisons are limited to homology modeling based on the structure of the PspA coiled-coil fragment that was solved recently64. However, no NTP-binding site was identified in such a model7.
Furthermore, as outlined above, typically nucleotide binding and hydrolysis depend on Mg2+ and in cases where nucleotides are hydrolyzed even in the absence of Mg2+, the hydrolysis rates increased in the presence of Mg2+. In contrast, here we show that the cooperativity of the apparent IM30 GTPase activity is increased when Mg2+ is absent, which could well be ascribed to an allosteric action of Mg2+ (see Suppl. Fig. 3). Binding of Mg2+ to IM30 has been demonstrated recently, and Mg2+ binding clearly induces structural rearrangements of oligomeric IM3047, in line with the allosteric function of Mg2+. These structural rearrangements appear to affect the GTP hydrolysis activity of oligomeric SynIM30, as discussed above. The GTP hydrolyzing activity of SynIM30 was altered in the presence of GMP-PCP, a non-hydrolyzable GTP analog, typically used as a competitive inhibitor. This lead to a further increase in cooperativity, due to an increased population of a low-affinity state.
Taken together, IM30 exhibits many characteristics that clearly separates it from canonical GTPases, such as the missing G-domain and Mg2+-independent GTP hydrolysis. Thus, IM30 might found a new class of GTPases39. Yet, GTP binding and hydrolysis clearly have no major effect on the membrane remodeling activity of IM30, at least in vitro, and it remains to be shown whether GTP hydrolysis by IM30 proteins is relevant in vivo.
Experimental Procedures
Cloning, expression and purification of IM30
Construction of the plasmid used for expression of N-terminally His-tagged Synechocystis IM30 (pRSET IM30 WT) was described recently13. The plasmid used for expression of helix 1 truncated SynIM30 is described in46. The plasmid used for expression of the IM30 FERM_EE mutant was created via introducing six mutations (E83A, E84A, F168A, E169A, R170A, M171A) into IM30 WT using the Quick-Change method65,66.
All proteins were expressed in E.coli BL21 (DE3) at 37 °C for 3–4 h after induction via the addition of 0.5 mM Isopropyl-β-D-thiogalactopyranoside (IPTG).
Cells were resuspended in lysis buffer (50 mM HEPES pH 7.6, 300 mM NaCl, 20 mM imidazole), lysed via sonification (4 °C) and cell debris was removed by centrifugation (8200 g, 10 min, 4 °C). His-tagged IM30 was isolated via Ni2+-affinity-chromatography (Ni-NTA agarose matrix, 3 wash steps with 20, 50 and 100 mM imidazole, elution with 1000 mM imidazole). After isolation of the protein, the buffer was exchanged to 20 mM HEPES pH 7.6 by gel filtration (Sephadex G25). Where necessary, the protein solution was concentrated using a Centriprep filter unit (MWCO 10 kDa, regenerated cellulose membrane, 3000 g, 30 min, 4 °C). The protein concentration was estimated using a Bradford assay with a BSA standard curve. For storage, the protein solution was mixed 1:1 (v/v) with glycerol and stored at −20 °C. The purification was completed within 8 hours (time from cell lysis to storage at −20 °C).
Malachite green assay
SynIM30 (0.1 µM) was mixed with 0.5 mM (if not stated otherwise) of GTP and Mg2+ (2.5 mM if not stated otherwise) in 20 mM HEPES and incubated for 30 min at 37 °C. 200 µL of this sample was transferred to a 96-well plate and mixed with 50 µL of the malachite green reaction mixture (“Gold mix”, PiColorLock Gold Phosphate Detection Kit by Innova Biosciences) and incubated for 5 min at RT. Finally, 20 µL of “stabilizer” (PiColorLock Gold Phosphate Detection Kit by Innova Biosciences) was added. Absorbance at 635 nm was measured with an OMEGA FLUOstar Platereader (BMG Labtech). Buffer blank (including the respective GTP concentration) absorbance was subtracted, and the concentration of released phosphate was determined by linear regression from a phosphate standard curve.
The phosphate release value was used to calculate the average NTPase activity ANTPase [min−1] per mol protein:
[Pi] refers to the concentration of released phosphate and t to the incubation time of the hydrolysis reaction.
The Michaelis-Menten constant KM was determined by using a Hill equation-derived fitting model and the resulting \({V}_{max}\) was used to determine \({k}_{cat}\):
[S] denotes for the substrate concentration and n for the Hill coefficient.
For a more detailed analysis of the dependence of GTPase activity on GTP, Mg2+ and GMP-PCP, a two-state model was fitted to the data. This is described in more detail in the supplement.
Continuous phosphate release assay
SynIM30 was mixed with a phosphate sensor (recombinant E. coli phosphate-binding protein labeled with the fluorophore MDCC; Sigma Aldrich)40 and Mg2+ in a 96-well plate. The mixture was preincubated at 37 °C and prewarmed GTP was added directly via an automatic titrator prior to the measurement (OMEGA FLUOstar Platereader (BMG Labtech)). The final concentration in the wells were 0.1 µM protein, 2.5 mM Mg2+ and 10 µM GTP. Fluorescence of the samples was measured at 37 °C with a 420/10 nm excitation filter and a 460/10 nm emission filter over the indicated timescale.
From the change in the fluorescence signal over time (after subtraction of the fluorescence signal generated by GTP auto-hydrolysis in the buffer), the concentration of released phosphate was determined via linear regression using a phosphate standard curve. The initial, non-biased turnover rate \({v}_{0}\) [min−1] was determined by a monoexponential fit for non-linear, steady-state enzyme kinetics, assuming a first-order process41:
[Pi] is the measured concentration of released phosphate and \(\eta \) accounts for a term describing the bias on \({v}_{0}\) that causes non-linearity, e.g. reduced velocity due to substrate limitation or product inhibition41. \({v}_{0}\) was used to determine the \({k}_{cat}\):
PK/LDH-coupled GTPase assay
The reaction buffer containing 5 mM MgCl2, 150 mM NaCl, 7.5 mM KCl, 0.6 mM NADH, 1 mM PEP (phosphoenolpyruvate), PK(final concentration 12–20 units/mL)/LDH(final concentration 18–28 units/mL) mix (Pyruvate Kinase /Lactic Dehydrogenase mix from Sigma-Aldrich) (and 0–5 mM GTP in 20 mM HEPES pH 7.6 was preincubated at room temperature for 15 min to convert already present GDP to GTP. SynIM30 WT or SynDLP was mixed with the reaction buffer to a final concentration of 0.5 µM. The GTP to GDP turnover rate was determined indirectly via the decrease of NADH absorbance at 340 nm43 on a timescale of 1–2 h at 37 °C with an OMEGA FLUOstar Platereader (BMG Labtech). The decline in NADH concentration is proportional to the concentration of GDP produced. The slope of the measured NADH absorbance was determined via fitting with a linear function. The GTPase activity \({A}_{NTPase}\) [min−1] was calculated as follows:
with \(\frac{\Delta {A}_{340}}{\Delta t}\) being the linear slope of the absorbance at 340 nm, l the filling height of the well and \({\varepsilon }_{NADH}\) the extinction coefficient of NADH (ε = 6220 M−1 cm−1). Data were corrected for autohydrolysis and the 0 mM GTP value was subtracted from all values to account for protein caused perturbations.
The maximum GTP turnover rate Vmax was determined via Eq. 2. The GTP turnover rate was used to determine kcat.
Luciferase-coupled GTPase assay
To analyze the GTPase activity of IM30 by directly measuring changes in the GTP concentration, we used a luciferase-coupled GTPase assay (GTPase-Glo Assay Kit by Promega). The assay was performed according to the instructions of the manufacturer. In short, 4 µM GTP and IM30 (0–6.5 µM IM30) were incubated for 120 min at 37 °C. After the reaction, the remaining GTP was converted to ATP by addition of GTPase-Glo Reagent (including ADP and a luciferase) and further incubation for 30 min at 37 °C. After 10 min incubation with the detection reagent at 37 °C, the luminescence was measured in a 384-wellplate using an OMEGA FLUOstar Platereader (BMG Labtech). GTP concentrations were calculated from the luminescence intensities using GTP standards that were treated as the samples.
Size exclusion chromatography (SEC)
Proteins were analyzed on an ÄKTA basic system (GE Healthcare) with a Superose12 10/300 GL column (GE Healthcare) equilibrated with 20 mM HEPES pH 7.6 at 8 °C. Protein elution was monitored via absorbance at 280 nm. The column was calibrated against standards of known size (blue dextran >2000 kDa, β-amylase (200 kDa), alcohol dehydrogenase (150 kDa), albumin (66 kDa), ribonuclease A (13.7 kDa).
Thermal denaturation
Thermal stability of IM30 (3.2 µM) was determined via CD spectroscopy (JASCO-815 CD spectrometer with an MPTC-490S temperature-controlled cell holder) in presence and absence of 0.5 mM GTP and/or 2.5 mM Mg2+. CD spectra were collected from 250 to 200 nm (cell length 0.1 cm, 1 nm data pitch, 5 nm bandwidth, 200 nm/min, 1 s data integration time, averaged over 3 accumulations of the same sample) over a temperature ramp from 15 to 95 °C (2 °C steps, overall heating rate 0.27 °C/min). Spectra were smoothened with a Savitzky-Golay filter. The ellipticity at 222 nm was used as a measure for the folding state of the protein. To estimate the transition temperature Tm of the protein, an indicator for the thermal stability, the resulting melting curve was fitted with an adapted Boltzmann-Fit, which allows linear slopes in the plateau regions of the curve, assuming a simple two-state unfolding mechanism:
\({\theta }_{meas}\) refers to the measured ellipticity. T is the temperature, θN and θD are the ellipticities at the plateaus of native and denatured protein, mN and mD are the slopes of the respective plateaus.
Liposome preparation
The lipids DOPG (1,2-dioleoyl-sn-glycero-3-phosphoglycerol), MGDG (monogalactosyldiacylglycerol) and the fluorescently labeled lipids NBD-PE (1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-(7-nitro-2-1,3-benzoxadiazol-4-yl) and LissRhod-PE (Lissamine Rhodamine PE; 1,2-Dioleoyl-sn-glycero-3-phosphoethanolamine-N-(lissamine rhodamine B sulfonyl) (ammonium salt)) were purchased from Avanti Polar Lipids, Inc. (Birmingham, AL, USA). For liposome preparation, lipids were dissolved in chloroform/methanol (2:1, v/v). The organic solvent was evaporated under a gentle stream of nitrogen gas followed by overnight vacuum desiccation to remove any traces of solvent. Unilamellar liposomes were prepared by hydration of the dried lipid film with 20 mM HEPES buffer (pH 7.6) and five cycles of freeze-thawing. For the liposome fusion assay, the liposomes were subsequently extruded 15 times through a 100-nm filter, using an extruder from Avanti Polar Lipids, Inc. (Alabaster, AL, USA).
Laurdan fluorescence measurements
Laurdan (6-dodecanoyl-N,N-dimethyl-2-naphthylamine) is a fluorescent dye that incorporates into lipid bilayers. Its fluorescence is sensitive to changes in the polarity of the environment and is therefore used to report changes of the membrane fluidity. In order to quantify the spectral changes, the Generalized Polarization (GP) value defined by67 is calculated for each spectrum.
here, I440 and I490 are the fluorescence emission intensities at 440 and 490 nm, respectively.
Laurdan (Sigma, Taufkirchen, Germany) was added to the dissolved lipid DOPG in a molar ratio of 1:500. Unilamellar liposomes were prepared as described before. To analyze the effect of nucleotides on the binding of IM30 to DOPG, 1 µM IM30 WT, 0.1 mM liposomes and 2.5 mM GTP (or GDP) were mixed and incubated for 2 h at 25 °C. As the protein was stored in 50% glycerol, the final concentration of glycerol was 15%. For samples without IM30, the corresponding amount of 50% glycerol (20 mM HEPES, pH 7.6) was added.
The fluorescence emission spectra were recorded on a FluoroMax-4 spectrometer (Horiba Scientific, Kyoto, Japan) from 400 to 550 nm with excitation at 350 nm at 25 °C. The slit width was set at 4 nm for excitation and emission of Laurdan.
Liposome fusion assay
The influence of nucleotides on IM30-triggered liposome fusion was measured using a FRET-based assay, as described earlier23,68. Unlabeled liposomes were mixed in 10-fold excess with labeled liposomes containing two fluorescent dyes that form a FRET-pair. Upon fusion of labeled with unlabeled liposomes, the FRET dyes redistribute over the membrane and consequently FRET decreases, resulting in increasing donor emission intensity. To simulate complete membrane fusion, liposomes containing only a 10th of the fluorescently labeled lipids were used as a positive control.
For the fusion assay, 1 µM IM30 WT and 0.1 mM liposomes (MGDG/DOPG, 40:60, w/w) were used. As described for the Laurdan measurements, the final concentration of glycerol was 15% for all samples. The concentration of MgCl2 was 6 mM for samples without nucleotides.
The measurements were performed using a Fluoromax-4 spectrometer (Horiba Scientific, Kyoto, Japan). The IM30-containing solutions were preincubated with Mg2+ for 15 min, whereas the liposomes were premixed with the nucleotides for about 1 min. After fast mixing of all compounds, the measurement was started immediately. Upon excitation of the FRET-donor NBD-PE at 460 nm, the donor emission was monitored at 525 nm over 1000 s at 25 °C. The slit widths for excitation and emission were set at 5 nm.
The raw fluorescence data was converted to a fusion rate in percentage by Eq. 8 using the intensities of the negative control (INC), positive control (IPC) and the measured sample (I) at every point in time t.
The initial fusion rate was determined by the first derivative of the fusion curve after 50 s.
Since GTP and GDP bind Mg2+ via their phosphate groups, higher concentrations of MgCl2 had to be used to obtain the same amount of free Mg2+ in samples containing GTP or GDP as in the control. To determine the additionally needed amount of Mg2+, we made use of the fact that Mg2+ alone can induce fusion at a concentration of 10 mM. Increasing amounts of MgCl2 were added to the assay containing 10 mM Mg2+ in presence of 2.5 mM GTP or GDP, until the observed fusion rates matched the one obtained in absence of the nucleotides. In the case of GTP, 4.25 mM MgCl2 was additionally needed, in the presence of GDP 2.5 mM MgCl2 (Suppl. Fig. 2).
References
Vothknecht, U. C., Otters, S., Hennig, R. & Schneider, D. Vipp1: A very important protein in plastids?! Journal of Experimental Botany 63, 1699–1712 (2012).
Li, H. M., Kaneko, Y. & Keegstra, K. Molecular cloning of a chloroplastic protein associated with both the envelope and thylakoid membranes. Plant Mol. Biol. 25, 619–632 (1994).
Fuhrmann, E. et al. The vesicle-inducing protein 1 from Synechocystis sp. PCC 6803 organizes into diverse higher-ordered ring structures. Mol. Biol. Cell 20, 4620–4628 (2009).
Otters, S. et al. The first α-helical domain of the vesicle-inducing protein in plastids 1 promotes oligomerization and lipid binding. Planta 237, 529–540 (2013).
Gao, F., Wang, W., Zhang, W. & Liu, C. α-Helical Domains Affecting the Oligomerization of Vipp1 and Its Interaction with Hsp70/DnaK in Chlamydomonas. Biochemistry 54, 4877–4889 (2015).
Aseeva, E. et al. Complex formation of Vipp1 depends on its α-helical PspA-like domain. J. Biol. Chem. 279, 35535–35541 (2004).
Saur, M. et al. A Janus-Faced IM30 Ring Involved in Thylakoid Membrane Fusion Is Assembled from IM30 Tetramers. Structure 25(1380–1390), e5 (2017).
Gutu, A., Chang, F. & O’Shea, E. K. Dynamical localization of a thylakoid membrane binding protein is required for acquisition of photosynthetic competency. Mol. Microbiol. 108, 16–31 (2018).
Bryan, S. J. et al. Localisation and interactions of the Vipp1 protein in cyanobacteria. Mol. Microbiol. 94, 1179–1195 (2014).
Zhang, L., Kato, Y., Otters, S., Vothknecht, U. C. & Sakamoto, W. Essential role of VIPP1 in chloroplast envelope maintenance in Arabidopsis. Plant Cell 24, 3695–3707 (2012).
Kroll, D. et al. VIPP1, a nuclear gene of Arabidopsis thaliana essential for thylakoid membrane formation. Proc. Natl. Acad. Sci. U. S. A. 98, 4238–4242 (2001).
Westphal, S., Heins, L., Soll, J. & Vothknecht, U. C. Vipp1 deletion mutant of synechocystis: A connection between bacterial phage shock and thylakoid biogenesis? Proc. Natl. Acad. Sci. U. S. A. 98, 4243–4248 (2001).
Fuhrmann, E., Gathmann, S., Rupprecht, E., Golecki, J. & Schneider, D. Thylakoid membrane reduction affects the photosystem stoichiometry in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 149, 735–744 (2009).
Lo, S. M. & Theg, S. M. Role of vesicle-inducing protein in plastids 1 in cpTat transport at the thylakoid. Plant J 71, 656–668 (2012).
Zhang, L. & Sakamoto, W. Possible function of VIPP1 in thylakoids: Protection but not formation? Plant Signal. Behav. 8 (2013).
McDonald, C., Jovanovic, G., Ces, O. & Buck, M. Membrane stored curvature elastic stress modulates recruitment of maintenance proteins pspa and vipp1. MBio 6 (2015).
McDonald, C., Jovanovic, G., Wallace, B. A., Ces, O. & Buck, M. Structure and function of PspA and Vipp1 N-terminal peptides: Insights into the membrane stress sensing and mitigation. Biochim. Biophys. Acta - Biomembr 1859, 28–39 (2017).
Thurotte, A., Brüser, T., Mascher, T. & Schneider, D. Membrane chaperoning by members of the PspA/IM30 protein family. Commun. Integr. Biol 10, e1264546 (2017).
Benning, C., Xu, C. & Awai, K. Non-vesicular and vesicular lipid trafficking involving plastids. Current Opinion in Plant Biology 9, 241–247 (2006).
Benning, C. A role for lipid trafficking in chloroplast biogenesis. Progress in Lipid Research 47, 381–389 (2008).
Nordhues, A. et al. Evidence for a role of VIPP1 in the structural organization of the photosynthetic apparatus in Chlamydomonas. Plant Cell 24, 637–659 (2012).
Zhang, S., Shen, G., Li, Z., Golbeck, J. H. & Bryant, D. A. Vipp1 is essential for the biogenesis of Photosystem I but not thylakoid membranes in Synechococcus sp. PCC 7002. J. Biol. Chem. 289, 15904–15914 (2014).
Hennig, R. et al. IM30 triggers membrane fusion in cyanobacteria and chloroplasts. Nat. Commun. 6 (2015).
Hennig, R. et al. The IM30/Vipp1 C-terminus associates with the lipid bilayer and modulates membrane fusion. Biochim. Biophys. Acta - Bioenerg 1858, 126–136 (2017).
Siebenaller, C., Junglas, B. & Schneider, D. Functional Implications of Multiple IM30 Oligomeric States. Front. Plant Sci. 10 (2019).
Junglas, B. & Schneider, D. What is Vipp1 good for? Molecular Microbiology 108, 1–5 (2018).
Heidrich, J., Thurotte, A. & Schneider, D. Specific interaction of IM30/Vipp1 with cyanobacterial and chloroplast membranes results in membrane remodeling and eventually in membrane fusion. Biochimica et Biophysica Acta - Biomembranes 1859, 537–549 (2017).
Westphal, S., Soll, J. & Vothknecht, U. C. A vesicle transport system inside chloroplasts. FEBS Lett 506, 257–261 (2001).
Morré, D. J., Selldén, G., Sundqvist, C. & Sandelius, A. S. Stromal low temperature compartment derived from the inner membrane of the chloroplast envelope. Plant Physiol. 97, 1558–1564 (1991).
Lindquist, E. & Aronsson, H. Chloroplast vesicle transport. Photosynth. Res. 138, 361–371 (2018).
Khan, N. Z., Lindquist, E. & Aronsson, H. New Putative Chloroplast Vesicle Transport Components and Cargo Proteins Revealed Using a Bioinformatics Approach: An Arabidopsis Model. PLoS One 8, e59898 (2013).
Kirchhoff, H. et al. Dynamic control of protein diffusion within the granal thylakoid lumen. Proc. Natl. Acad. Sci. U. S. A. 108, 20248–53 (2011).
Chuartzman, S. G. et al. Thylakoid membrane remodeling during state transitions in Arabidopsis. Plant Cell 20, 1029–39 (2008).
Stingaciu, L. R. et al. Revealing the Dynamics of Thylakoid Membranes in Living Cyanobacterial Cells. Sci. Rep. 6 (2016).
Nagy, G. et al. Reversible membrane reorganizations during photosynthesis in vivo: Revealed by small-angle neutron scattering. Biochem. J. 436, 225–230 (2011).
Jilly, R., Khan, N. Z., Aronsson, H. & Schneider, D. Dynamin-like proteins are potentially involved in membrane dynamics within chloroplasts and cyanobacteria. Front. Plant Sci. 9, 206 (2018).
Heidrich, J. et al. Organization into higher ordered ring structures counteracts membrane binding of IM30, a protein associated with inner membranes in chloroplasts and cyanobacteria. J. Biol. Chem. 291, 14954–14962 (2016).
Zhang, L., Kondo, H., Kamikubo, H., Kataoka, M. & Sakamoto, W. VIPP1 has a disordered C-terminal tail necessary for protecting photosynthetic membranes against stress. Plant Physiol. 171, 1983–1995 (2016).
Ohnishi, N., Zhang, L. & Sakamoto, W. VIPP1 Involved in Chloroplast Membrane Integrity Has GTPase Activity in Vitro. Plant Physiol. 177, 328–338 (2018).
Brune, M., Hunter, J. L., Corrie, J. E. T. & Webb, M. R. Direct, Real-Time Measurement of Rapid Inorganic Phosphate Release Using a Novel Fluorescent Probe and Its Application to Actomyosin Subfragment 1 ATPase. Biochemistry 33, 8262–8271 (1994).
Cao, W. & De La Cruz, E. M. Quantitative full time course analysis of nonlinear enzyme cycling kinetics. Sci. Rep 3, 2658 (2013).
Zhang, Y.-Z. et al. Interaction of malachite green with bovine serum albumin: Determination of the binding mechanism and binding site by spectroscopic methods. J. Hazard. Mater. 163, 1345–1352 (2009).
Ingerman, E. & Nunnari, J. A continuous, regenerative coupled GTPase assay for dynamin-related proteins. in Methods in Enzymology 404, 611–619 (Academic Press Inc., 2005).
Song, B. D., Leonard, M. & Schmid, S. L. Dynamin GTPase domain mutants that differentially affect GTP binding, GTP hydrolysis, and clathrin-mediated endocytosis. J. Biol. Chem. 279, 40431–40436 (2004).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Thurotte, A. & Schneider, D. The fusion activity of IM30 rings involves controlled unmasking of the fusogenic core. Front. Plant Sci. 10, 108 (2019).
Heidrich, J. et al. Mg2 binding triggers rearrangement of the IM30 ring structure, resulting in augmented exposure of hydrophobic surfaces competent for membrane binding. J. Biol. Chem. 293, 8230–8241 (2018).
Rudack, T., Xia, F., Schlitter, J., Kötting, C. & Gerwert, K. The role of magnesium for geometry and charge in GTP hydrolysis, revealed by quantum mechanics/Molecular mechanics simulations. Biophys. J. 103, 293–302 (2012).
Song, B. D., Leonard, M. & Schmid, S. L. Dynamin GTPase domain mutants that differentially affect GTP binding, GTP hydrolysis, and clathrin-mediated endocytosis. J. Biol. Chem. 279, 40431–40436 (2004).
Shan, S. O., Schmid, S. L. & Zhang, X. Signal recognition particle (SRP) and SRP receptor: A new paradigm for multistate regulatory GTPases. Biochemistry 48, 6696–6704 (2009).
De Boer, P., Crossley, R. & Rothfield, L. The essential bacterial cell-division protein FtsZ is a GTPase. Nature 359, 254–256 (1992).
Zhang, B., Zhang, Y., Wang, Z. X. & Zheng, Y. The role of Mg2+ cofactor in the guanine nucleotide exchange and GTP hydrolysis reactions of Rho family GTP-binding proteins. J. Biol. Chem. 275, 25299–25307 (2000).
McNew, J. A., Sondermann, H., Lee, T., Stern, M. & Brandizzi, F. GTP-Dependent Membrane Fusion. Annu. Rev. Cell Dev. Biol. 29, 529–550 (2013).
Newton, C., Pangborn, W., Nir, S. & Papahadjopoulos, D. Specificity of Ca2+ and Mg2+ binding to phosphatidylserine vesicles and resultant phase changes of bilayer membrane structure. BBA - Biomembr. 506, 281–287 (1978).
Nir, S., Newton, C. & Papahadjopoulos, D. 201 - Binding of Cations to Phosphatidylserine Vesicles. Bioelectrochemistry Bioenerg 5, 116–133 (1978).
Düzgüne, N. et al. Calcium- and magnesium-induced fusion of mixed phosphatidylserine/phosphatidylcholine vesicles: Effect of ion binding. J. Membr. Biol. 59, 115–125 (1981).
Wilschut, J., Düzgüne, N. & Papahadjopoulos, D. Calcium/Magnesium Specificity in Membrane Fusion: Kinetics of Aggregation and Fusion of Phosphatidylserine Vesicles and the Role of Bilayer Curvature. Biochemistry 20, 3126–3133 (1981).
Ohki, S. A mechanism of divalent ion-induced phosphatidylserine membrane fusion. BBA - Biomembr 689, 1–11 (1982).
Mondal Roy, S. & Sarkar, M. Membrane Fusion Induced by Small Molecules and Ions. J. Lipids 2011, 1–14 (2011).
Liang, H., Harries, D. & Wong, G. C. L. Polymorphism of DNA-anionic liposome complexes reveals hierarchy of ion-mediated interactions. Proc. Natl. Acad. Sci. U. S. A. 102, 11173–11178 (2005).
Antipina, A. Y. & Gurtovenko, A. A. Molecular-level insight into the interactions of DNA with phospholipid bilayers: Barriers and triggers. RSC Adv 6, 36425–36432 (2016).
Wittinghofer, A. & Vetter, I. R. Structure-Function Relationships of the G Domain, a Canonical Switch Motif. Annu. Rev. Biochem. 80, 943–971 (2011).
Nogales, E., Downing, K. H., Amos, L. A. & Löwe, J. Tubulin and FtsZ form a distinct family of GTPases. Nat. Struct. Biol 5, 451–8 (1998).
Osadnik, H. et al. PspF-binding domain PspA1-144 and the PspA·F complex: New insights into the coiled-coil-dependent regulation of AAA+ proteins. Mol. Microbiol. 98, 743–759 (2015).
Shenoy, A. R. & Visweswariah, S. S. Site-directed mutagenesis using a single mutagenic oligonucleotide and DpnI digestion of template DNA. Anal. Biochem. 319, 335–6 (2003).
Xia, Y., Chu, W., Qi, Q. & Xun, L. New insights into the QuikChangeTM process guide the use of Phusion DNA polymerase for site-directed mutagenesis. Nucleic Acids Res 43, e12 (2015).
Parasassi, T., Krasnowska, E. K., Bagatolli, L. & Gratton, E. Laurdan and Prodan as Polarity-Sensitive Fluorescent Membrane Probes. J. Fluoresc 8, 365–373 (1998).
Meers, P., Ali, S., Erukulla, R. & Janoff, A. S. Novel inner monolayer fusion assays reveal differential monolayer mixing associated with cation-dependent membrane fusion. Biochim. Biophys. Acta - Biomembr 1467, 227–243 (2000).
Acknowledgements
This work was funded by the Max-Planck Graduate Center at the Max Planck Institutes and the University of Mainz. We thank Dr. Ruven Jilly for preparing a sample of SynDLP and assisting with the PK/LDH-coupled GTPase assay. We thank Renate Genswein for help with the protein purification and for excellent technical support.
Author information
Authors and Affiliations
Contributions
D.S., B.J., C.S. and N.H. designed the experiments and analyzed data. B.J., C.S. and L.S. performed the experiments. D.S., B.J., C.S. and N.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Junglas, B., Siebenaller, C., Schlösser, L. et al. GTP hydrolysis by Synechocystis IM30 does not decisively affect its membrane remodeling activity. Sci Rep 10, 9793 (2020). https://doi.org/10.1038/s41598-020-66818-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66818-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.