Abstract
Background: People with trisomy 21 (T21) are predisposed to developing hematological tumors, but have significantly lower-than-expected age-adjusted incidence rates of having a solid tumor. Material and methods: To identify novel genetic factors implicated in the lower breast cancer (BC) frequency observed in women with T21 than in the general population, we compared the transcriptome pattern of women with a homogeneous T21, aged more than 30 years, with or without BC, and tumoral BC tissue of control women with a normal karyotype from the study of Varley et al. (2014). Results: Differential analysis of gene expression between the 15 women in the T21 without BC group and BC patients in the other groups (two women with T21 and fifteen control women, respectively) revealed 154 differentially expressed genes, of which 63 were found to have similar expression profile (up- or downregulated). Of those 63 genes, four were in the same family, namely GIMAP4, GIMAP6, GIMAP7 and GIMAP8, and were strongly upregulated in the T21 without BC group compared to the other groups. A significant decrease in mRNA levels of these genes in BC tissues compared to non-tumor breast tissues was also noted. Conclusion: We found that the expression of some GIMAPs is significantly higher in women with T21 without BC than in patients with sporadic BC. Our findings support the hypothesis that GIMAPs may play a tumor-suppressive role against BC, and open the possibility that they may also have the same role for other solid tumors in T21 patients. The search for new prognostic factors and hopefully new therapeutic or preventive strategies against BC are discussed.
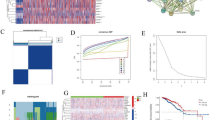
Similar content being viewed by others
Introduction
Trisomy 21 (T21) or Down syndrome is a common genetic disorder that results from the presence of all or part of an extra chromosome 21. It is one of the most frequent and most recognizable form of intellectual disability, appearing in approximately one out of every 700 to 2000 newborns. More than 100 features of people with T21 have been described, encompassing physical, medical, and psychological features1.
Epidemiological studies have shown that people with T21 are more predisposed to developing hematological tumors than the general population, yet their risk of developing solid tumors is at least 12 times lower2,3,4,5,6,7,8. For instance, breast cancer (BC) is almost absent in women with T218,9,10, even though this group shows a higher prevalence of known risk factors for BC, such as nulliparity, higher body mass index and obesity rates, sedentary life, increased chromosomal instability, dysfunctional mitochondria, overexpression of several known oncogenes and attenuation of known tumor suppressors, premature aging, DNA repair anomalies, immune deficiency, and excess of thoracic X-rays for recurrent respiratory infections6,11. Environmental protective factors such as reduced estrogen exposure, early menopause, or no alcohol use10 are not sufficient to explain this paradox.
However, BC incidence in women with intellectual disability that have many risk patterns as women with T21 is similar to that observed in the general population8. This suggests that genetic factors might protect women with T21 against BC12. Although many tumor repressor genes are localized on chromosome 216,12,13, having three copies of chromosome 21 is unlikely to be the only explanation for the protection of women with T21 from BC6.
We hypothesized that other, as yet unknown factors or pathways could be responsible for protecting women with T21 from BC. To identify these genes, we compared the transcriptomes of women with T21 with or without BC to those of women from the general population with or without BC.
Materials and Methods
Patients
More than 5500 clinical files of T21 patients collected at the Jérôme Lejeune Institute (Paris, France), were screened to identify women with a homogeneous T21 with no mosaicism or translocation, aged more than 34 years, and without chronic medication and social problems. Women who did not have BC or any mammary lesion were registered under the subgroup T21-BCF (BCF for Breast Cancer Free); women who had BC were registered under the subgroup T21-BC. No history of BC was noted in the families of these women, and no pathogenic or likely pathogenic variants in genes involved in BC were present in the T21-BC group. Written informed consent from parents or guardians on behalf of the participants was obtained prior to the study.
RNA was extracted from peripheral blood mononuclear cells (PBMCs) using a Qiagen Kit (Qiagen GmBH, Hilden, Germany).
Sequencing and Bioinformatics
Individual differential gene expression profiles were established according to the TruSeq Stranded mRNA protocol (Illumina, CA, USA). After sequencing the RNAseq libraries on the NextSeq (Illumina) platform by Paired-End sequencing (2 × 75 bp), cleaning and trimming of sequences were executed with Trimmomatic software (V0.37), and quality control was checked with the FastQC software. Sequences were then mapped onto the human genome with TopHat2 and aligned reads for each gene were counted with the HTSEQ-count software. Finally, genes were annotated with the GRCh38 human genome version.
To obtain transcriptomic data of women without T21, 30 RNAseq libraries were analyzed. Fifteen were from the tumoral tissue of women with a normal karyotype and with sporadic BC, called C-BC (C for Control), from the study of Varley et al. (RunID: SRR1313090, SRR1313095, SRR1313096, SRR1313098, SRR1313099, SRR1313104, SRR1313105, SRR1313111, SRR1313112, SRR1313115, SRR1313116, SRR1313119, SRR1313120, and SRR1313122, SRR1313128)14. The other 15 RNAseq libraries (RNA extracted from PBMC) were from healthy women without BC (C-BCF) incorporated from the Matrix of RNAseq database that focuses on the RNAseq method, which reflects the expression of the genes in a specific condition, and contains 21,800 human RNAseq profiles. The relative log expression (RLE) strategy15 was chosen as the strategy for normalization.
Differential Expression Analysis
We performed a differential expression analysis between the T21-BCF versus the T21-BC, C-BC, and C-BCF libraries with R software version 3.4.1 and the edgeR package v3.20.1. Selection parameters were an FDR value less than 0.001, a logFC value less than −1 or greater than +1, and a logCPM greater than 5. These stringent parameters were chosen to account for the intrinsic differences between the samples, which were derived from different tissues (PBMCs or breast tumor tissue).
Ethical approval and consent to participate
This study conformed to the tenets of the Declaration of Helsinki and was approved by the Institute Jérôme Lejeune Committee on Clinical Investigation.
Results
Seventeen women with T21, aged 32 to 57 years, were recruited for this study. Of these, 15 did not have BC or any mammary lesion (T21-BCF), while two had BC (T21-BC). From the 17 T21 RNAseq libraries sequenced, 485 million sequences were obtained, of which 93% were validated and mapped to the human genome (version GRCh38) (Supplementary Table 1). In the final count matrix, 57,245 different transcripts were identified. These libraries were compared to existing RNAseq libraries representing women with a normal karyotype and either with BC (C-BC) or without BC (C-BCF). The C-BC and C-BCF groups contain data from 15 samples each. A heat map to overview the similarities and dissimilarities between samples was constructed (Fig. 1). All groups were found to be well structured and homogeneous except the C-BCF group. This latter group was thus eliminated from the study, to avoid introducing bias into the analysis.
A heatmap of the distance matrix showing the similarities and dissimilarities between samples (T21-BCF; T21-BC; C-BCF; and C-BC). The normalized data are used in this figure for sample clustering. Distance between C-BC and T21-BC is the greater indicating the maximum dissimilarities between samples.
Differential analysis of gene expression between T21-BCF versus C-BC revealed a total of 3,261 genes (comparison group A) that were differentially expressed. Among them, 1,581 genes were up regulated in the T21-BCF cohort versus C-BC, and 1,680 were downregulated. Between T21-BCF and T21-BC (comparison group B), a total of 476 genes were differentially expressed, with 127 genes up regulated and 349 down regulated in T21-BCF patients relative to T21-BC patients (Fig. 2).
Differential analysis of gene expression between T21-BCF versus C-BC (group (A) and between T21-BCF versus T21-BC (group B) showing the number of genes that were differentially expressed. At the intersection of the 2 groups, 154 genes were differentially expressed, of which 63 genes had a similar expression profile (15 up- and 48 down-regulated).
In the T21-BCF group, we looked at the expression of genes on chromosome 21, especially those that might predispose to BC or have a positive action against BC occurrence, such as BTG3 (OMIM 605673), TIAM1 (OMIM 600687), ETS2 (OMIM 164740), DYRK1A (OMIM 600855), SIM2 (OMIM 600892), Col18A1 (OMIM 120328), IFNAR1 (OMIM 107450), NRIP1 (OMIM 602490), S100B (OMIM 176990), ERG (OMIM 165080),and RUNX1 (OMIM 151385)6. All these genes were expressed in the T21-BCF group, but no particular or significant differential expression relative to the potential role of each gene was found.
We then examined the expression of genes in pathways that play an important role in BC development, in particular those involved in reducing apoptosis allowing proliferation, or those that have been linked to diagnosis, prognosis and hereditary susceptibility (KEGG database; Kyoto Encyclopedia of Genes and Genomes: www.kegg.jp/kegg/kegg1.html). Specifically, we studied the PI3K/AKT/mTOR, EGFR, ovarian steroidogenesis, tyrosine kinase inhibitor resistance, RAP1 thyroid hormone signaling, parathyroid hormone synthesis, GNRH signaling, prolactin signaling pathways, and immune pathways. In all of them, different transcript levels were noted in some of the genes, but nothing obvious to conclude to a particular alteration of a signaling pathway.
We thus compared transcript expression between 2 different groups: T21-BCF versus C-BC (group A) and T21-BCF versus T21-BC (group B). At the intersection of comparison groups A and B, 154 differentially expressed genes were identified. Among them, 63 genes were found to have the same regulation profile (15 up- and 48 down-regulated) (Table 1). Among the identified genes, a group of four genes in the GTPases of the immunity-associated proteins family (GIMAP) – GIMAP4, GIMAP6, GIMAP7 and GIMAP8 – were strongly upregulated (fold ration >10) in the T21-BCF group compared to the T21-BC or C-BC patients (Fig. 3).
Discussion
To determine why BC occurs at a lower frequency in women with T21 compared to the general population, we studied the transcriptome profile of 17 women with T21: 15 without BC (T21-BCF) and 2 with BC (T21-BC), in addition to 15 control women with a normal karyotype and BC (C-BC). The comparison of transcript expression between groups identified four genes, namely, GIMAP4, GIMAP6, GIMAP7 and GIMAP8, that were strongly upregulated in the T21-BCF group compared to the T21-BC or C-BC groups.
The GIMAP genes are located within a region on chromosome 7q35–7q36.1. They code for a family of proteins mainly expressed in the immune system. GIMAP genes promote immunological functions such as thymocyte development, apoptosis of peripheral lymphocytes and T helper cell differentiation. When altered these genes are linked to immunological diseases such as T cell lymphopenia, acute myeloid leukemia and autoimmune diseases16,17,18,19.
Interestingly, lymphocytes, including T cells, T regulatory cells, and natural killer cells, and their cytokine release patterns, are involved in both primary prevention and recurrence of BC20. Furthermore, it has been hypothesized that chronic inflammation involving T lymphocytes is a possible pathophysiological pathway to breast adenocarcinoma20,21. Latter observations lead to a possible role of GIMAPs in cancer. In fact, in addition to their involvement in the regulation of the immune system, GIMAPs might act as tumor suppressor genes, as was speculated by Krucken et al. who showed that all GIMAPs are expressed at very low levels in diverse cancer tissues and cell lines22. This hypothesis was further confirmed by other studies that proved that GIMAP4 accelerates the execution of programmed cell death15, while GIMAP6 plays a role in the autophagic process23. On the other hand, a study on non-small cell lung cancer (NSCLC) showed that the expression of GIMAP4, GIMAP6, and GIMAP8 were lower in tumor tissues than in adjacent non-tumor tissues24. Interestingly, GIMAP8 mRNA level was abnormally elevated in the adjacent non-tumor tissues compared to that in the control lung tissues24. Furthermore, on a recent study performed on hepatocellular carcinoma (HCC), Huang et al. found that the mRNA expression of GIMAP5 and GIMAP6 were significantly downregulated in the HCC tumor samples and in the blood samples from HCC patients in comparison to matched non-tumor tissue samples, and blood from healthy subjects25. GIMAP5 and GIMAP6 proteins followed the same scheme of expression. Results suggesting the involvement of GIMAP5 and GIMAP6 in the pathogenesis of HCC24.
Despite the low number of patients with T21 and BC included in our study (2 patients), our findings suggest that the overexpression of different GIMAPs might have a tumor repression function in women with T21. In order to further support these results and check if GIMAPs play also a role in BC in women in general and not only in T21 patients, we evaluated the expression of GIMAP family members in 62 RNAseq libraries of 18 women without BC, 16 with DCIS (Ductal Carcinoma In Situ), 13 with triple-negative BC, and 15 with HER2 (human epidermal growth factor receptor 2) positive BC from the study of Varley et al. (Supplementary Table 2)14. Interestingly, our data show a significant decrease in mRNA levels of GIMAP4, GIMAP6, GIMAP7 and GIMAP8 in BC tissues, compared to DCIS tissues, and non-tumor breast tissues from either women with or without BC (supplementary 2). Furthermore, the mRNA levels of those 4 GIMAPs were significantly downregulated in the blood samples from patients with HER2+ or Triple-negative BC. These findings align with previous research related to the association between GIMAP genes and cancer and suggest that these genes act as breast tumor suppressing genes, perhaps by inhibiting cell proliferation, enhancing apoptosis or controlling the cell cycle. On the other hand, the observed overexpression of the GIMAP genes in T21 patients might explain the paradox that although people with T21 have an increased risk of leukemia and often show immune biological abnormalities and clinical immunodeficiency, they seem to be protected against solid tumors26.
In our cohort of women with T21, no cases of BC were reported in the families. In contrast, in some reported women with T21 and BC, the occurrence of BC in other family members was observed. It was linked to the presence of a pathogenic mutation predisposing to BC and associated with a risk of developing BC equals to that of the general population11. Whether the protective action of GIMAPs works only in the absence of mutations in known BC-promoting genes remains to be investigated.
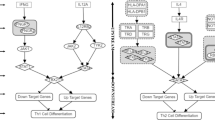
Recently, different studies have shown interest in IL-12 as a potential agent for anti-tumor immunotherapy27. This immunocytokine, that is essential for the differentiation of the Th1 lineage, was found to upregulate the GIMAP1 and the GIMAP4 genes, in addition to Th1 cytokines (IL-2, TNF-alpha, IFN-gamma, etc.)17,18,28,29. The contribution of IL-12 to cell cycle control, cell proliferation inhibition and/or apoptosis induction, might thus be mediated by the GIMAP genes and/or the cytokines of the Th1 lineage that are interestingly overexpressed in T2130.
Conclusions
This is the first time that a transcriptomic approach was used in a cohort of women with T21 with the aim of finding BC-related genetic factors. Our study identified a higher GIMAP expression in women with T21 without BC compared to BC patients either with T21 or a normal karyotype. It also showed that GIMAP genes are downregulated in BC tissues compared to non-tumor tissues. The characterization of the molecular mechanisms leading to GIMAP overexpression might help to better understand the decreased incidence of BC in women with T21. A similar approach could be used to identify if GIMAPs and/or other factors are involved in other solid tumors and secondary cancers. The ultimate goal would be to search for new prognostic factors, and hopefully new therapeutic or preventive approaches against cancer.
Data availability
The data and the material are available at the CRB BioJel, Jerome Lejeune Institute, Paris, France.
References
Mégarbané, A. et al. The 50th anniversary of the discovery of trisomy 21: the past, present, and future of research and treatment of Down syndrome. Genet Med. 11, 611–616 (2009).
Satgé, D. et al. A tumor profile in Down syndrome. Am. J. Med. Genet. 78, 207–216 (1998).
Hasle, H., Clemmensen, I. H. & Mikkelsen, M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet. 355, 165–169 (2000).
Patja, K., Pukkala, E., Sund, R., Iivanainen, M. & Kaski, M. Cancer incidence of persons with down syndrome in Finland: A population-based study. Int. J. Cancer. 118, 1769–1772 (2006).
Sullivan, S. G., Hussain, R., Glasson, E. J. & Bittles, A. H. The profile and incidence of cancer in Down syndrome. J. Intel. Disab. Res. 51, 228–231 (2007).
Nižetić, D. & Groet, J. Tumorigenesis in Down’s syndrome: big lessons from a small chromosome. Nat. Rev. Cancer. 12, 721–732 (2012).
Hasle, H., Friedman, J. M., Olsen, J. H. & Rasmussen, S. A. Low risk of solid tumors in persons with Down syndrome. Genet. Med. 18, 1151–1157 (2016).
Rethoré, M. O., Rouëssé, J. & Satgé, D. Cancer screening in adults with down syndrome, a proposal. Eur. J. Med. Genet. 9, 103783, https://doi.org/10.1016/j.ejmg.2019.103783 (2019).
Scholl, T., Stein, Z. & Hansen, H. Leukemia and other cancers, anomalies and infections as causes of death in Down’s syndrome in the United States during 1976. Develop. Med. Child. Neurol. 24, 817–829 (1982).
Satgé, D., Sasco, A. J., Pujol, H. & Rethoré, M. O. Breast cancer in women with trisomy 21. Bull. Acad. Natl. Med. 185, 1239–1252 (2001).
Satgé, D., Mircher, C., Tretarre, B. & Rethore, M. O. Additional data for breast cancer surveillance in women with Down syndrome. World Down Syndrome Congress. Cape Town. South Africa. August 14–17 (2012).
Chatterjee, A. et al. Potential contribution of SIM2 and ETS2 functional polymorphisms in Down syndrome associated malignancies. BMC. Med. Genet. 14, 12 (2013).
Ohgaki, K. et al. Mapping of a new target region of allelic loss to a 6-cM interval at 21q21 in primary breast cancers. Genes. Chromosomes. Cancer. 23, 244–247 (1998).
Varley, K. E. et al. Recurrent read-through fusion transcripts in breast cancer. Breast. Cancer. Res. Treat. 146, 287–297 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome. Biol. 11, R106 (2010).
Schnell, S., Démollière, C., van den Berk, P. & Jacobs, H. Gimap4 accelerates T-cell death. Blood. 108, 591–599 (2006).
Filén, J. J. et al. Quantitative proteomics reveals GIMAP family proteins 1 and 4 to be differentially regulated during human T helper cell differentiation. Mol. Cell. Proteomics. 8, 32–44 (2009).
Filén, S. & Lahesmaa, R. GIMAP Proteins in T-Lymphocytes. J. Signal. Transduct. 2010, 268589 (2010).
Döhner, K. et al. Molecular cytogenetic characterization of a critical region in bands 7q35-q36 commonly deleted in malignant myeloid disorders. Blood. 92, 4031–4035 (1998).
Standish, L. J. et al. Breast cancer and the immune system. J. Soc. Integr. Oncol. 6, 158–168 (2008).
Morrow, E. S., Roseweir, A. & Edwards, J. The role of gamma delta T lymphocytes in breast cancer: a review. Transl. Res. 203, 88–96 (2019).
Krücken, J. et al. Comparative analysis of the human gimap gene cluster encoding a novel GTPase family. Gene. 341, 291–304 (2004).
Pascall, J. et al. GIMAP6 is required for T cell maintenance and efficient autophagy in mice. PLoS. One. 13, e0196504 (2018).
Shiao, Y. M. et al. Dysregulation of GIMAP genes in non-small cell lung cancer. Lung. Cancer. 62, 287–294 (2008).
Huang, Z. et al. Dysregulation of GTPase IMAP family members in hepatocellular cancer. Mol. Med. Rep. 14, 4119–4123 (2016).
Satgé, D. & Seidel, M. G. The Pattern of Malignancies in Down Syndrome and Its Potential Context With the Immune System. Front. Immunol. 9, 3058 (2018).
Fallon, J. et al. The immunocytokine NHS-IL12 as a potential cancer therapeutic. Oncotarget. 5, 1869–1884 (2014).
Heinonen, M. T., Kanduri, K., Lähdesmäki, H. J., Lahesmaa, R. & Henttinen, T. A. Tubulin- and actin-associating GIMAP4 is required for IFN- γ secretion during Th cell differentiation. Immunol. Cell. Biol. 93, 158–166 (2015).
Heinonen, M. T. et al. GIMAP GTPase family genes: potential modifiers in autoimmune diabetes, asthma, and allergy. J. Immunol. 194, 5885–5894 (2015).
Murphy, M., Friend, D. S., Pike-Nobile, L. & Epstein, L. B. Tumor necrosis factor-alpha and IFN-gamma expression in human thymus. Localization and overexpression in Down syndrome (trisomy 21). J. Immunol. 149, 2506–2512 (1992).
Acknowledgements
We are thankful to the patients for their cooperation. This work was funded by Jerome Lejeune Foundation.
Author information
Authors and Affiliations
Contributions
A.M. and D.P. designed research. A.M., D.P., S.S., F.P., R.B., F.N. performed main experiments. D.P., F.P., R.B., F.N. did the bioinformatical analysis. A.M., D.P., G.L. wrote the paper. A.M., D.P., A.S.R., S.S., F.P., R.B., F.N., C.M., A.R., M.V.M., S.D., G.L. critically analyzed the results. All authors discussed and interpreted data, and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mégarbané, A., Piquemal, D., Rebillat, AS. et al. Transcriptomic study in women with trisomy 21 identifies a possible role of the GTPases of the immunity-associated proteins (GIMAP) in the protection of breast cancer. Sci Rep 10, 9447 (2020). https://doi.org/10.1038/s41598-020-66469-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66469-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.