Abstract
Triple negative breast cancer (TNBC) is an aggressive breast cancer subtype. Recent studies have shown that MHC class II (MHCII) expression and tumor infiltrating lymphocytes are important prognostic factors in patients with TNBC, although the relative importance of lymphocyte subsets and associated protein expression is incompletely understood. NanoString Digital Spatial Profiling (DSP) allows for spatially resolved, highly multiplexed quantification of proteins in clinical samples. In this study, we sought to determine if DSP could be used to characterize expression of MHCII and other immune related proteins in tumor epithelial versus stromal compartments of patient-derived TNBCs (N = 10) using a panel of 39 markers. We confirmed that a subset of TNBCs have elevated expression of HLA-DR in tumor epithelial cells; HLA-DR expression was also significantly higher in the tumors of patients with long-term disease-free survival when compared to patients that relapsed. HLA-DR expression in the epithelial compartment was correlated with high expression of CD4 and ICOS in the stromal compartment of the same tumors. We also identified candidate protein biomarkers with significant differential expression between patients that relapsed versus those that did not. In conclusion, DSP is a powerful method that allows for quantification of proteins in the immune microenvironment of TNBCs.
Similar content being viewed by others
Introduction
Triple-negative breast cancer (TNBC) is an aggressive subtype of breast cancer that is associated with high rates of local recurrence and a propensity for distant metastases1. Clinically, TNBC is defined by the absence of three biomarkers that are used to inform treatment decisions: ER (estrogen receptor), PR (progesterone receptor), and HER2. While traditional predictive biomarkers are not expressed in TNBCs, there has been considerable interest in quantifying the host anti-tumor immune response in patients with TNBC to determine risk of recurrence and predict response to immunotherapy. Using RNA-seq of patient-derived TNBCs, we have shown that elevated expression of genes in the major histocompatibility complex Class II antigen presentation pathway (MHCII) is a favorable prognostic factor that is associated with long-term DFS2. This genomics discovery led us to develop and validate a clinical prognostic assay that measures expression of MHCII and tumor-infiltrating lymphocyte (TIL) genes in formalin-fixed, paraffin embedded (FFPE) clinical samples using NanoString technology3. We recently confirmed the prognostic significance of this MHCII Immune Activation assay in two independent cohorts of patients with TNBC3. Mechanistic studies support our clinical findings, as ectopic expression of MHCII has been shown to induce Th1-mediated anti-tumor immunity in mouse models of human cancer4,5,6.
We have shown that expression of MHCII mRNA in clinical samples is correlated with the presence of tumor-infiltrating lymphocytes (TILs) and with elevated expression of TIL genes including CD3D, CD4, and CD8A3. Our RNA-seq and NanoString gene expression studies were both performed using macrodissection of clinical TNBC specimens. While macrodissection enriches for carcinoma cells, both tumor and stromal infiltrating immune cells may be present in the macrodissected sample. Therefore, gene expression measured in clinical specimens is often reflective of both carcinoma cells as well as immune and stromal cells within the tumor microenvironment (TME)7,8. A logical question that arises is whether MHCII gene expression in patient tumors is simply a reflection of immune cells within the TME, or whether MHCII is expressed within tumor epithelial cells.
Protein expression and localization in clinical FFPE samples can be assessed using immunohistochemistry (IHC) or immunofluorescence (IF); however, the capacity for multiplexing is generally limited due to spectral overlap9. Furthermore, quantification of IHC is often performed using a semi-quantitative scale that can be subject to interobserver variability, particularly in the research setting10. Laser capture microdissection (LCM) is one method that can be used to isolate cell types and quantify protein and gene expression in distinct populations of cells, though it is technically challenging and difficult to scale8. NanoString Digital Spatial Profiling (DSP) is a highly multiplexed method that allows for protein quantification in spatially resolved, morphologically distinct areas within a single FFPE tissue section9,11,12,13,14,15. This method utilizes UV-photocleavable oligonucleotides that are conjugated to primary antibodies13,14,15. After incubation with a multiplexed cocktail of primary antibodies, oligonucleotides are cleaved in regions of interest (ROIs) using UV LED light and are then hybridized with NanoString optical barcodes for ex situ counting on an nCounter Analysis system13,14,15. In this study, we sought to determine if DSP can be used to characterize expression of MHCII and other immune related proteins in tumor epithelial versus stromal compartments of patient-derived TNBC samples.
Results
TNBC tumor specimens have variable numbers of immune cells within epithelial and stromal compartments (Fig. 1). Additionally, stromal compartments within individual tumors can vary from lymphocyte-rich (Fig. 1A) to lymphocyte-poor (Fig. 1B). Stromal TIL density positively correlates with improved prognosis in patients with TNBC, although the relative importance of lymphocyte subsets and associated protein expression is incompletely understood16. The first main goal of this study was to determine whether DSP could be used to quantify proteins in morphologically distinct compartments within patient-derived TNBCs (N = 10). We selected distinct tumor cell epithelial and stromal ROIs and performed protein expression profiling using DSP; three epithelial and three stromal ROIs were selected from each tumor (Fig. 2A). We found that epithelial and stromal ROIs had distinct patterns of protein expression that were consistent with the predicted cell types in each region (Fig. 2B). Pan-cytokeratin expression was higher in epithelial ROIs, whereas the expression of CD3 and other immune cell proteins were significantly higher in stromal ROIs (Fig. 2B,C).
Stromal lymphocyte density in TNBCs. (A) Example of lymphocyte-rich TNBC containing sheets of syncytial epithelial cells and broad bands of stromal lymphocytes. (B) Lymphocyte-poor TNBC with hyalinized collagenous stroma and rare stromal-infiltrating lymphocytes. Right panel highlights epithelial compartments in green. Magnification: 10x(left-most panels); 20x(middle and right panels).
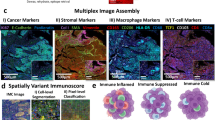
Region of interest selection and protein quantification in TNBC tumors. (A) Multi-label immunofluorescence was used to visualize tissue morphology and select regions of interest (ROIs). We selected six ROIs from each tumor, including three epithelial and three stromal ROIs. Four representative ROIs are shown from a single tumor. Magnification: 20x for all panels. (B) Epithelial and stromal ROIs had distinct patterns of protein expression that were consistent with the predicted cell types in each region. (C) CD3 expression was elevated in stromal ROIs whereas Pan-cytokeratin expression was elevated in epithelial ROIs.
Traditionally, expression of MHCII was thought to be limited to antigen presenting cells (APCs) such as dendritic cells, B cells, and macrophages, although previous studies demonstrate that a subset of patient-derived TNBCs and TNBC cell lines can also express genes in the MHCII pathway2. To confirm that breast cancer epithelial cells can express MHCII proteins, we used DSP to evaluate HLA-DR expression in epithelial ROIs. We detected elevated HLA-DR protein expression in 18 TNBC epithelial ROIs from 6 tumors and low HLA-DR protein expression in 12 epithelial ROIs from 4 tumors (Fig. 3A). Pancytokeratin expression was likewise elevated within these HLA-DR positive ROIs, thus confirming epithelial origin of the signal. We then evaluated HLA-DR expression using IHC in these tumors, and results were concordant with DSP (Fig. 3A). Importantly, histomorphologic features were consistent between ROIs that had been assessed using DSP, H&E, and IHC (Fig. 3B). We then compared semi-quantitative IHC scores to normalized DSP protein expression counts, where we found that the measurements were highly correlated between the two methods (Spearman correlation coefficient R2 = 0.71; Fig. 3C). Additionally, HLA-DR protein expression was relatively homogenous across the 3 epithelial ROIs from each tumor (Fig. 3E). These findings validate the DSP detection of HLA-DR expression in epithelial tumor ROIs and confirm that a subset of patient-derived breast cancer epithelial cells can express MHCII proteins such as HLA-DR, which is consistent with our previous observations2,3, as well as with those of other investigators 4,17,18,19,20,21.
Comparison of HLA-DR expression in epithelial ROIs using DSP and IHC. (A) Examples of DSP measurements along with IHC staining are shown for a case with negative HLA-DR staining (Left panels) and for a case with elevated HLA-DR expression (Right panels). (B) Histomorphologic features were consistent between methods. (C) HLA-DR DSP counts and immunohistochemical scoring were positively correlated (Spearman R2 = 0.71). (D) HLA-DR protein expression in both epithelial and stromal ROIs was significantly higher in patients that did not relapse (P < 0.001; P = 0.006; Mann-Whitney tests). (E) HLA-DR DSP counts were relatively homogenous across the 3 epithelial ROIs from each tumor. Magnification for all images is 20x.
We have previously demonstrated that elevated expression of HLA-DRB1 mRNA in patient-derived TNBC tumors is significantly associated with long-term disease-free survival (DFS)3, though it has been unclear whether stromal or epithelial expression is more predictive of patient outcomes. Using DSP, we found that HLA-DR protein expression in both epithelial and stromal ROIs was significantly higher in patients with long-term DFS when compared to patients that relapsed (P < 0.001; Fig. 3D). Notably, the magnitude of differential HLA-DR expression between patient groups was more pronounced in the epithelial compartment (Fig. 3D). This finding is consistent with the hypothesis that aberrant expression of MHCII expression by TNBC epithelial cells results in the presentation of tumor-specific neoantigens to CD4+ T cells, thus contributing to the host anti-tumor immune response and improving patient outcomes2. In agreement with this hypothesis, we found that epithelial expression of HLA-DR was highly correlated with stromal expression of CD4 ((Pearson correlation coefficient R2 = 0.67; Fig. 4A), likely representing recruitment of CD4+ T lymphocytes to the tumor microenvironment. Correspondingly, CD4 protein expression in stromal ROIs was significantly higher in patients with long-term DFS when compared to patients that relapsed (P < 0.0001; Fig. 4B). In addition, we found that epithelial HLA-DR expression was highly correlated with stromal expression of ICOS (CD248; Pearson correlation coefficient R2 = 0.48; Fig. 4C), and that ICOS expression is significantly higher in patients that did not experience relapse (P = 0.0001; Fig. 4D). ICOS is a T-cell co-stimulator that enhances T-cell responses including proliferation and lymphokine proliferation; thus, it may mediate host anti-tumor immunity.
Epithelial HLA-DR expression is correlated with stromal CD4 and ICOS expression. (A) By linear regression, epithelial HLA-DR expression was positively correlated with expression of CD4 in the stroma (P = 0.0040; Pearson R2 = 0.67). (B) Stromal CD4 expression was significantly higher in patients with long-term DFS (P < 0.0001). (C) Epithelial HLA-DR expression was positively correlated with expression of ICOS (CD278) in the stroma (P = 0.0273; Pearson R2 = 0.48). (D) Stromal ICOS (CD278) expression was significantly higher in patients with long-term DFS (P < 0.0001).
The multiplexed protein quantification provided by DSP allowed us to identify other proteins, besides HLA-DR, that may be involved in host immune response and that that may ultimately influence patient outcome. In order to identify proteins that were differentially expressed between tumors with disparate clinical behavior, we analyzed DSP data using DESEQ2. We observed striking differences in protein expression between tumors from patients that relapsed versus those that did not, and notably, we discovered that there were differences between the significant proteins within epithelial versus stromal ROIs (Fig. 5A,B). Among patients that did not relapse, we found that elevated expression of HLA-DR, IDO-1, and Beta-2 microglobulin were significant only within the epithelial compartment, whereas CD45RO, CD4, PD-1, and MS4A1 were significantly elevated in only the stromal compartment (Fig. 5A–C; Supplementary Table 1). We also identified 5 proteins that were significant across both epithelial and stromal ROIs (ICOS, CD45, CD11c, CD3, and CD8A; Fig. 5C). Future studies are warranted to determine if these candidate prognostic protein markers are significant in larger cohorts, and to investigate their role in anti-tumor immunity.
Differential protein expression and association with relapse status. DESEQ2 was used to identify proteins with significant differential expression between patients that relapsed versus those with that did not (adjusted p-values < 0.05). Heatmaps depict the quantification of proteins that were identified as significant in epithelial (A) and stromal (B) ROIs. (C) Overview of significantly elevated proteins in patients with long-term DFS. A subset of these proteins was found to be significantly elevated across both epithelial and stromal ROIs.
Discussion
NanoString DSP is a unique method that allows for multiplexed protein quantification in clinical patient samples11,12,13,14,15. The method is gaining traction in the immune oncology environment and has been used to develop signatures that are predictive of patient response to immune checkpoint inhibitors22,23. We applied DSP to patient-derived TNBC tumors in order to interrogate differential protein expression in tumor epithelial and stromal compartments between patients who relapsed and those who did not. DSP provided quantitative protein expression measurements that allowed us to gain new insight into the complexities of the breast cancer immune microenvironment. The ability of DSP to simultaneously quantify multiple protein biomarkers in clinical samples is promising; however, extensive validation studies are required before this technology can be applied to guide patient care.
We applied DSP to patient-derived TNBCs, where we confirmed that a subset of breast tumors has elevated expression of HLA-DR in epithelial tumor cells. We also established that HLA-DR protein expression is significantly higher in the tumors of patients with long-term DFS compared to patients who relapsed. This finding was significant in both epithelial and stromal ROIs, although HLA-DR protein expression in the epithelial compartment of breast tumors was a better discriminator of patient outcome than it was in the stromal compartment. We also found that HLA-DR expression in the epithelial compartment is correlated with high expression of CD4 and ICOS in the stromal compartment of the same tumors. This finding is consistent with our hypothesis that MHCII positive TNBC epithelial cells present antigen and activate CD4+ T cells2. Although antigen presentation is typically thought to be limited to APCs, there is evidence in the literature to suggest that unique epithelial subsets may be capable of antigen presentation. For example, MHCII+ thyroid follicular epithelial cells have been shown to present viral peptide antigens to human T cells24. More recently, corneal epithelial cells were shown to induce MHCII upregulation in response to HSV-1 infection; these epithelial cells were found to have amateur antigen-presenting capabilities and could induce CD4+ T cell proliferation in vitro25. Our findings suggest that MHCII expression in epithelial TNBC tumor cells is an important factor in host immune response.
In addition to confirming expression of HLA-DR in tumor epithelial cells, the multiplexed protein quantification provided by DSP allowed us to identify novel candidate prognostic biomarkers that could enhance the power of existing MHCII biomarker assays, and inform future functional studies of the anti-tumor immune response. We found that tumors with elevated expression of HLA-DR expression in epithelial ROIs had elevated expression of CD4, ICOS, CD45RO, PD-1, CD11c, CD8 and MSA4A1 in the stroma. This result suggests that, in addition to CD4+ T helper cells, multiple types of immune cells may be present in the tumor microenvironment including memory T cells (CD45RO), dendritic cells (CD11c), cytotoxic T-cells (CD8A), and B cells (MSA4A1/CD20). In summary, DSP provided a novel and powerful method examining the immune microenvironment of TNBCs and identifying proteins associated with prognosis.
Methods
Patient-derived tissue sections
This project was performed under an approved University of Utah IRB protocol (#24487). Since the tissue specimens were leftover, de-identified archival FFPE blocks, the IRB waived the requirement for informed consent. Patient-derived TNBC specimens were collected as previously described3. All specimens were de-identified by the Huntsman Cancer Institute Biorepository and Molecular Pathology Shared Resource facility. For this study, a subset of tumors (N = 10) with adequate pathologic material in the archives were selected for protein expression profiling using NanoString DSP. In order to interrogate the immune microenvironment of TNBCs with disparate clinical behavior, we selected samples from patients that experienced disease recurrence (N = 5) and samples from patients that remained disease-free (N = 5) after a median follow-up time of 7.4 years. Clinical characteristics of these patients are provided in Supplementary Table 2. Only primary stage I-III breast cancers were included in the study. Detailed clinicopathologic and outcome data were obtained through review of the medical record.
NanoString DSP and gene expression analysis
The DSP assay was performed on FFPE TNBC tumor sections through the NanoString Technology Access Program service. The specimens were sent to NanoString and ROI selection was performed through a web-based interface by a study team pathologist with the assistance of NanoString scientists. The DSP workflow can be divided into four main steps: (1) Tissue staining and imaging (2) Region of interest selection; (3) Cleavage and aspiration of oligonucleotide tags; (4) Hybridization and counting using the NanoString nCounter system. For this study, tissue sections (4 µm) were stained using a multiplexed cocktail of primary antibodies; each antibody was conjugated to a unique, UV-photocleavable indexing oligonucleotide tag. For this experiment we included all DSP validated antibodies-oligo conjugates available at the time of the study. A complete list of protein targets is provided in Supplementary Table 3. Tissue sections were imaged using three-color immunofluorescence in order to guide ROI selection. Morphology markers included CD3, pan-cytokeratin, and DAPI. A board-certified pathologist selected three epithelial ROIs and three stromal ROIs from each TNBC tumor section; each ROI measured 300 µm. UV LED light was used to release oligonucleotide tags from discrete ROIs. These tags were then aspirated, transferred to a 96-well plate and were then hybridized to target-specific barcode-labeled probes, as has been previously described26. Using the nCounter Prep Station, hybridized probes were immobilized in a cartridge, excess probe was washed, and the cartridge was transferred to the nCounter Digital Analyzer for analysis. All DSP values for the 39 markers are included in Supplementary Table 4.
Immunohistochemistry (IHC)
IHC staining for HLA-DR was performed as previously described3. Briefly, 4 µm FFPE sections were stained using an HLA-DR antibody (Santa Cruz Biotechnology (sc-53319) in a Ventana BenchMark™ ULTRA autostainer. Sections were de-paraffinized and subject to antigen retrieval using Cell Conditioning 1 (CC1, pH 8.5) for 64 minutes at 95 °C. The primary antibody was applied at a concentration of 1:1000. Interpretation was performed as previously described3.
Statistical analysis
Statistical analyses were performed using GraphPad version 7.0 C and R version 3.5.0. Raw counts for each of the photo-cleavage fluorescent probes corresponding to each of the proteins measured in the DSP assay were normalized to engineered RNA sequences (ERCC RNA controls) to correct for background signal in the assay. DESEQ2 version 1.20.027 was used to identify proteins with significantly different normalized counts (adjusted p-value < 0.05) between stromal and epithelial ROIs as well as tumor from patients who recurred and those who did not. Heatmaps of log normalized protein counts were created using the R package ‘pheatmap’ version 1.0.10. Differences between groups were analyzed using Fisher exact test, 2-tailed t test with Welch correction, or 1-way analysis of variance with post hoc Tukey test.
References
Newman, L. A., Reis-Filho, J. S., Morrow, M., Carey, L. A. & King, T. A. The 2014 Society of Surgical Oncology Susan G. Komen for the Cure Symposium: triple-negative breast cancer. Ann Surg Oncol 22, 874–882, https://doi.org/10.1245/s10434-014-4279-0 (2015).
Forero, A. et al. Expression of the MHC Class II Pathway in Triple-Negative Breast Cancer Tumor Cells Is Associated with a Good Prognosis and Infiltrating Lymphocytes. Cancer Immunol Res 4, 390–399, https://doi.org/10.1158/2326-6066.CIR-15-0243 (2016).
Stewart, R. L. et al. A Multigene Assay Determines Risk of Recurrence in Patients with Triple-Negative Breast Cancer. Cancer Res 79, 3466–3478, https://doi.org/10.1158/0008-5472.CAN-18-3014 (2019).
Axelrod, M. L., Cook, R. S., Johnson, D. B. & Balko, J. M. Biological Consequences of MHC-II Expression by Tumor Cells in Cancer. Clin Cancer Res 25, 2392–2402, https://doi.org/10.1158/1078-0432.CCR-18-3200 (2019).
Meazza, R., Comes, A., Orengo, A. M., Ferrini, S. & Accolla, R. S. Tumor rejection by gene transfer of the MHC class II transactivator in murine mammary adenocarcinoma cells. Eur J Immunol 33, 1183–1192, https://doi.org/10.1002/eji.200323712 (2003).
Mortara, L. et al. CIITA-induced MHC class II expression in mammary adenocarcinoma leads to a Th1 polarization of the tumor microenvironment, tumor rejection, and specific antitumor memory. Clin Cancer Res 12, 3435–3443, https://doi.org/10.1158/1078-0432.CCR-06-0165 (2006).
Cleator, S. J. et al. The effect of the stromal component of breast tumours on prediction of clinical outcome using gene expression microarray analysis. Breast Cancer Res 8, R32, https://doi.org/10.1186/bcr1506 (2006).
Onuchic, V. et al. Epigenomic Deconvolution of Breast Tumors Reveals Metabolic Coupling between Constituent Cell Types. Cell Rep 17, 2075–2086, https://doi.org/10.1016/j.celrep.2016.10.057 (2016).
Decalf, J., Albert, M. L. & Ziai, J. New tools for pathology: a user’s review of a highly multiplexed method for in situ analysis of protein and RNA expression in tissue. J Pathol 247, 650–661, https://doi.org/10.1002/path.5223 (2019).
Lawrie, C. H. et al. Inter- and intra-observational variability in immunohistochemistry: a multicentre analysis of diffuse large B-cell lymphoma staining. Histopathology 61, 18–25, https://doi.org/10.1111/j.1365-2559.2012.04179.x (2012).
Toki, M. I. et al. High-Plex Predictive Marker Discovery for Melanoma Immunotherapy-Treated Patients Using Digital Spatial Profiling. Clin Cancer Res 25, 5503–5512, https://doi.org/10.1158/1078-0432.CCR-19-0104 (2019).
Prokop, S. et al. Impact of TREM2 risk variants on brain region-specific immune activation and plaque microenvironment in Alzheimer’s disease patient brain samples. Acta Neuropathol 138, 613–630, https://doi.org/10.1007/s00401-019-02048-2 (2019).
Beechem, J. M. High-Plex Spatially Resolved RNA and Protein Detection Using Digital Spatial Profiling: A Technology Designed for Immuno-oncology Biomarker Discovery and Translational Research. Methods Mol Biol 2055, 563–583, https://doi.org/10.1007/978-1-4939-9773-2_25 (2020).
Hinerfeld, D., Barker, K., Merritt, C. & Beechem, J. Validation of Antibody Panels for High-Plex Immunohistochemistry Applications. J Biomol Tech 30, S40–S41 (2019).
Ihle, C. L. et al. Distinct tumor microenvironments of lytic and blastic bone metastases in prostate cancer patients. J Immunother Cancer 7, 293, https://doi.org/10.1186/s40425-019-0753-3 (2019).
Salgado, R. et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann Oncol 26, 259–271, https://doi.org/10.1093/annonc/mdu450 (2015).
Johnson, D. B. et al. Melanoma-specific MHC-II expression represents a tumour-autonomous phenotype and predicts response to anti-PD-1/PD-L1 therapy. Nat Commun 7, 10582, https://doi.org/10.1038/ncomms10582 (2016).
Oldford, S. A. et al. Tumor cell expression of HLA-DM associates with a Th1 profile and predicts improved survival in breast carcinoma patients. Int Immunol 18, 1591–1602, https://doi.org/10.1093/intimm/dxl092 (2006).
Oldford, S. A., Robb, J. D., Watson, P. H. & Drover, S. HLA-DRB alleles are differentially expressed by tumor cells in breast carcinoma. Int J Cancer 112, 399–406, https://doi.org/10.1002/ijc.20441 (2004).
Lazzaro, B., Anderson, A. E., Kajdacsy-Balla, A. & Hessner, M. J. Antigenic characterization of medullary carcinoma of the breast: HLA-DR expression in lymph node positive cases. Appl Immunohistochem Mol Morphol 9, 234–241, https://doi.org/10.1097/00129039-200109000-00007 (2001).
Tamiolakisl, D. et al. Gains and losses of HLA class II (DR) and CD4 in atypical hyperplasia, carcinoma in situ and infiltrating ductal carcinoma of the breast. Acta Medica (Hradec Kralove) 47, 257–262 (2004).
Amaria, R. N. et al. Neoadjuvant immune checkpoint blockade in high-risk resectable melanoma. Nat Med 24, 1649–1654, https://doi.org/10.1038/s41591-018-0197-1 (2018).
Blank, C. U. et al. Neoadjuvant versus adjuvant ipilimumab plus nivolumab in macroscopic stage III melanoma. Nat Med 24, 1655–1661, https://doi.org/10.1038/s41591-018-0198-0 (2018).
Londei, M., Lamb, J. R., Bottazzo, G. F. & Feldmann, M. Epithelial cells expressing aberrant MHC class II determinants can present antigen to cloned human T cells. Nature 312, 639–641, https://doi.org/10.1038/312639a0 (1984).
Royer, D. J., Elliott, M. H., Le, Y. Z. & Carr, D. J. J. Corneal Epithelial Cells Exhibit Myeloid Characteristics and Present Antigen via MHC Class II. Invest Ophthalmol Vis Sci 59, 1512–1522, https://doi.org/10.1167/iovs.17-23279 (2018).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol 26, 317–325, https://doi.org/10.1038/nbt1385 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550, https://doi.org/10.1186/s13059-014-0550-8 (2014).
Acknowledgements
We acknowledge the direct financial support for the research reported in this publication that was provided by the William L. Roberts Memorial Fund Award at ARUP Laboratories and the University of Utah Department of Pathology. The project described was also supported by the NIH National Center for Advancing Translational Sciences through grant number KL2TR001996 (RL Stewart). Research reported in this publication utilized the University of Utah Huntsman Cancer Institute Biorepository and Molecular Pathology Shared Resource which is supported by the National Cancer Institute of the National Institutes of Health under Award Number P30CA042014. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
R.L.S. analyzed the data, generated figures, and wrote the manuscript. R.E.F. reviewed clinical specimens and edited the manuscript. A.P.M. analyzed data and generated figures. K.E.V. analyzed data, revised the manuscript, and supervised the project. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
K.E.V. and R.L.S. are inventors on US Provisional Patent Application No. 62/829,440 that was filed on April 4, 2019 (Title: Multigene assay to assess risk of recurrence of cancer; Inventors: Katherine E. Varley, Rachel L. Stewart, Philip S. Bernard, and Katherine Updike). The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stewart, R.L., Matynia, A.P., Factor, R.E. et al. Spatially-resolved quantification of proteins in triple negative breast cancers reveals differences in the immune microenvironment associated with prognosis. Sci Rep 10, 6598 (2020). https://doi.org/10.1038/s41598-020-63539-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-63539-x
This article is cited by
-
Distinct spatial immune microlandscapes are independently associated with outcomes in triple-negative breast cancer
Nature Communications (2023)
-
A framework for multiplex imaging optimization and reproducible analysis
Communications Biology (2022)
-
Protein-based immune profiles of basal-like vs. luminal breast cancers
Laboratory Investigation (2021)
-
Multiplexed imaging analysis of the tumor-immune microenvironment reveals predictors of outcome in triple-negative breast cancer
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.