Abstract
Domestic species provides a powerful model for examining genetic mechanisms in the evolution of yield traits. The domestic silkworm (Bombyx mori) is an important livestock species in sericulture. While the mechanisms controlling cocoon yield are largely unknown. Here, using B. mori and its wild relative B. mandarina as intercross parents, 100 BC1 individuals were sequenced by restriction site-associated DNA sequencing (RAD-Seq). The linkage map contained 9,632 markers was constructed. We performed high-resolution quantitative trait locus (QTL) mapping for four cocoon yield traits. A total of 11 QTLs were identified, including one yield-enhancing QTL from wild silkworm. By integrating population genomics and transcriptomic analysis with QTLs, some favourable genes were revealed, including 14 domestication-related genes and 71 differentially expressed genes (DEGs) in the fifth-instar larval silk gland transcriptome between B. mori and B. mandarina. The relationships between the expression of two important candidate genes (KWMTBOMO04917 and KWMTBOMO12906) and cocoon yield were supported by quantitative real-time PCR (qPCR). Our results provide some new insights into the molecular mechanisms of complex yield traits in silkworm. The combined method might be an efficient approach for identifying putative causal genes in domestic livestock and wild relatives.
Similar content being viewed by others
Introduction
Domestication played a central role in formulating Charles Darwin’s theory of evolution1. Since domestication, crops and animals have been under artificial selection, largely directed towards increasing yield, which provides useful model systems to characterize the genetic mechanisms of yield traits. In recent years, domestication-related genes for yield traits have been cloned from several crops2,3. The domestic silkworm (Bombyx mori) is an economically important insect. Compared with its wild relatives, long-term artificial breeding and selection have resulted in a high cocoon yield of domestic silkworm4. However, the molecular mechanisms associated with silk yield remain largely undefined in silkworm5.
QTL mapping is a method for locating the genetic loci within a whole genome that might contain genes of small effect contributing to yield traits. In domestic silkworm, QTL mapping has been conducted using traditional molecular markers, such as amplified fragment length polymorphisms (AFLPs) and simple sequence repeats (SSRs)6,7,8,9,10,11. However, due to the limitation of marker numbers, it is relatively difficult to narrow QTL regions of interest12. With the advance of next-generation sequencing (NGS) technology, restriction-site-associated DNA sequencing (RAD-Seq) has been developed, which scans short sequence regions surrounding all restriction sites for a given restriction endonuclease13. A large number of single nucleotide polymorphism (SNP) markers are available that can be used for constructing a high-density genetic map, QTL mapping and population genomic analyses14,15,16,17. In Plutella xylostella, 2,878 segregating RAD alleles helped construct a linkage map and match linkage groups with homologous chromosomes of Bombyx mori14. In rainbow trout, a three-generation F2 mapping family was genotyped using RAD-Seq to identify 4874 informative SNPs18. Thus, RAD-Seq is a powerful tool for high-density genetic map construction and QTL mapping13.
A clear limitation of QTL mapping is that the confidence intervals are usually large, potentially harbouring up to dozens of genes. Pinpointing candidate gene by means of QTL analysis alone is often extremely challenging19. In crops, one way to finely dissect QTL regions is to breed advanced intercross lines or nearly isogenic lines (NILs)3,19,20,21. However, it is difficult to maintain large mapping populations over multiple generations, and this incurs great cost in terms of time in livestock. Searching for selective signals in the whole genome of domestic species becomes more convenient4,19. Thus, the combination of population genomics and QTL mapping should be an efficient approach for identifying causal genes of domestication-related yield traits22.
Increasing yield is one of principal purposes of animal breeding programs. Furthermore, genetic diversity is a fundamental criterion of high-yield breeding. Generally, the genetic improvement of B. mori has mainly been restricted by the domestic populations23,24. B. mori has been domesticated from the wild mulberry silkworm B. mandarina for at least 5000 years4,25. Utilization of wild silkworm might provide a more powerful way to exploit novel genetic resources23,26,27,28. In this study, we used the domestic silkworm Xiafang (D_XF) and the wild silkworm (W_AK) as intercross parents and employed their backcross progenies (BC1M) as the mapping population. Using RAD-Seq technology, the yield-related loci were mapped in the whole genome of the domestic and wild silkworms. Combining with population genomics and developmental transcriptomic data of the silk gland, integrated analyses were applied to reveal those yield-related loci. It provides a good opportunity for exploring yield-related genes and molecular mechanisms in domestic and wild silkworms, which might be used for high-yield breeding with genome editing, genetic knockdown and overexpression technologies29,30.
Results
Phenotypic variation in the mapping population and correlation between yield traits
In this study, the domestic silkworm strain Xiafang (D_XF), which is widely used in sericulture production in China, and the wild silkworm (W_AK) were used as the parental mapping lines. All four cocoon yield-related traits showed significant differences between D_XF and W_AK (Table 1). In particular, the cocoon shell weight (CSW) and whole cocoon weight (WSW) of the domestic strain Xiafang reached to 4.68 and 2.76 times those of W_AK, respectively. The phenotypic values of the traits measured in the BC1 population were nearer to those of the recurrent parent D_XF (Table 1).
The correlation between the four cocoon yield traits was evaluated (Table 2). Significantly positive correlations were observed among WCW, CSW, and pupal weight (PW). The highest correlation was shown between WCW and PW (r = 0.99, P < 0.01). The cocoon shell ratio (CSR) was positively correlated with CSW (r = 0.34, P < 0.01) but negatively correlated with WCW (r = −0.61, P < 0.01) and PW (r = −0.73, P < 0.01). In addition, we compared the correlations between cocoon yield traits and sex. The results indicated that WCW and PW were positively correlated with female, i.e., female individuals exhibited higher WCW and PW values than males. The cocoon shell ratio was negatively correlated with female (r = −0.73, P < 0.01). No significant correlation was observed between cocoon shell weight and female/male was observed.
Genetic linkage map and QTL mapping
A total of 100 BC1 individuals were sequenced using RAD-Seq technology (Supplementary Table S1). The number of RAD tags for the 100 BC1 individuals ranged from 720,477 to 4,622,071 with an average of 2,230,620 (Supplementary Table S2). After quality filtration, a total of 49.69 million high-quality SNPs were detected (Supplementary Table S3). The whole genomes of the intercross parents have been resequenced with NGS technology31. The average sequencing depth was 14.42-fold in the female parent (D_XF) and 12.83-fold in the male parent (W_AK). In the mapping parents, 5,748,787 homozygous and polymorphic SNP markers were detected. Through filtration and genotyping in the BC1 mapping population, 51,418 effective markers were identified and used for further linkage map construction. A total of 28 genetic linkage groups were constructed with MSTmap software (Fig. 1 and Supplementary Table S4) and found to exhibit one-to-one correspondence with the silkworm chromosomes32. The linkage map contained 9,632 markers and covered 4,764.96 cM, with a range of 112.31 cM to 254.83 cM.
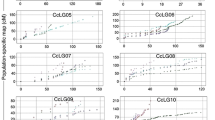
A total of 11 cocoon yield-related QTLs were identified on 7 chromosomes using the composite interval mapping (CIM) algorithm (Fig. 2 and Table 3). The contribution rate (R2) of a single QTL varied from 9.52% to 13.68%, and qtl2-2 had the largest effect. Most of the QTLs showed positive additive effects, implying a higher value for the trait conferred by the allele of the domestic strain Xiafang (Table 3), while qtl2-4 (CSW) showed negative additive effects. Interestingly, the 95% confidence intervals (95% CI) of qtl2-1 (CSW) and qtl4-1 (PW) were 99.90–107.52 cM and 108.52–110.59 cM on chromosome 1, respectively (Table 3). It was suggested that CSW and PW traits showed a certain genetic relatedness.
Identification of putative candidate genes associated with QTLs
Based on the transcriptome of the fifth-instar larval silk gland and the new genome assembly of silkworm33, 194 protein-encoding genes with expression signal (FPKM > 1) were identified in the 11 QTLs (Supplementary Table S5). The number of genes ranged from 8 (qtl1-3 and qtl2-4) to 59 (qtl2-1). Among the expressed genes, binding, organic cyclic compound binding and heterocyclic compound binding were the most enriched GO terms (Supplementary Fig. S1a). Pathway enrichment analysis indicated that metabolic pathway, peroxisome and longevity regulating pathway contained more genes (Supplementary Fig. S1b). To pinpoint candidate genes, the developmental transcriptome of the fifth-instar larval silk gland were compared between the domestic and wild silkworms, including seven stages from the initiation of the fifth instar to wandering (Supplementary Table S5). In total, 71 differentially expressed genes (DEGs) were identified during the seven developmental stages (Fig. 3 and Supplementary Table S5).
Hierarchical clustering of the differentially expressed genes in the QTL regions. In the sample names, D and W indicate domestic and wild silkworms, respectively. 0d: beginning of the fifth-instar larvae; 1d to 5d: day 1, 2, 3, 4, and 5 of the fifth-instar larvae, respectively; w: beginning of the wandering stage.
A combination of selective sweeps and QTL mapping analyses might be an efficient approach for pinpointing the candidate genes associated with domestication34,35. In the silkworm, cocoon yield traits are also under control of domestication. Whole-genome resequencing of eight silkworm strains and seven wild silkworms from ecologically different regions has been conducted in our laboratory36. The selective sweep regions were identified in the whole genome, which showed extremely low heterozygosity and high FST values relative to wild populations (Fig. 4a). As shown in Fig. 4b, selective sweep analysis revealed the genetic complexity underlying some QTLs. The qtl1-1 on chromosome 9 and qtl2-1 on chromosome 1 co-localized with 7 and 6 selective sweep regions, respectively (Supplementary Table S6). Conversely, four QTLs (qtl2-3, qtl3-1, qtl4-1, and qtl4-3) had a simpler genetic profile and contained only one sweep region with one gene, respectively. In total, 14 domestication-related genes were characterized within the QTL regions (Supplementary Table S6), which might provide multiple valuable candidates for understanding molecular mechanisms of cocoon yield.
Genomic regions and candidate genes showing signals of a selective sweep in QTLs. (a) Distribution of ratio (θπ, domestic/θπ, wild) and FST of 500-bp windows with 500-bp steps in the whole genome. At a 5% significance level (corresponding to θπ ratio = 0.1835 and FST = 0.4514), the selected windows were considered as candidate regions with signals of a selective sweep for domestic silkworm (green dots). Red dots represent selected regions in the wild populations. (b) Example of qtl2-1 on chromosome 1 with signals of a selective sweep (grey regions) in domestic silkworms. (c) Example of a gene with a signal of a selective sweep (grey region) in qtl1-1. Gene structure is shown at the bottom. CDS: coding sequence; UTR: untranslated regions. D: domestic silkworm; W: wild silkworm. (d) Neighbour-joining (NJ) tree of KWMTBOMO04917. The NJ tree of the DNA sequences used in (c) was reconstructed with MEGA6.037 using a Kimura 2-parameter model, 500 replicates and pairwise deletion. (e) The miRNA showed sequence diversification between the domestic and wild populations.
KWMTBOMO04917, encoding 5-aminolevulinate synthase (ALAS; E.C.2.3.1.37), was one of the putative genes under artificial selection (Fig. 4c,d) and was differentially expressed between the domestic and wild silkworms at five out of the seven developmental stages of the fifth-instar silk gland (Supplementary Table S5). To understand the relationship of KWMTBOMO04917 with silk yield, its transcriptional level was detected by quantitative real-time PCR (qPCR) in the silk gland of high-yield Xiafang, low-yield Dazao (D_DZ) and the wild silkworm (Fig. 5a,b). Both qPCR and RNA sequencing indicated that the expression of KWMTBOMO04917 was higher in the wild silkworm than in the domestic silkworms (Fig. 5c and Supplementary Table S5). KWMTBOMO12906, encoding for DNA polymerase epsilon (Pol ε) subunit 4, was one of the few DEGs that was highly expressed in the domestic silkworms and showed differential expression at four out of the seven developmental stages (Fig. 3 and Supplementary Table S5). The qPCR result indicated that the expression of KWMTBOMO12906 in the posterior silk gland (PSG) was positively correlated with cocoon yield (Fig. 5d and Supplementary Fig. S2). These results suggested that the genes identified via integrated methods might be important candidates controlling cocoon yield.
qPCR validation of candidate genes in domestic and wild silkworms. (a) Comparison of cocoon of the domestic strain Dazao (D_DZ), Xiafang (D_XF) and wild silkworm. (b) Whole cocoon weight of the wild silkworm and domestic strains used for qPCR analysis. (c,d) qPCR validation of the interesting candidate genes in the silk gland on day 2 of the wild and domestic silkworms. The design of the specific primers and the qPCR strategy were similar to our previous study38. The primer sequences are listed in Supplementary Table S7. One-way analysis of variance (ANOVA) was used to determine significant differences between any two samples. *P < 0.05; **P < 0.01; n.s.: not significant.
Yield-enhancing QTLs from wild silkworm
Based on QTL mapping, it was suggested that qtl2-4 from the wild parent was related to an increase in the trait score (Table 3). The qtl2-4, controlling cocoon shell weight, was located on chromosome 28 (Fig. 2 and Table 3). This QTL contained 8 protein-coding genes showed expression signals (FPKM > 1) in the silk gland of fifth-instar larvae (Supplementary Table S5). KWMTBOMO16286, KWMTBOMO16288 and KWMTBOMO16283 showed differential expression between the domestic and wild silkworms. Especially, KWMTBOMO16283 presented higher expression in silk gland of the domestic silkworm than wild silkworm during all the fifth-instar stages.
MicroRNAs in QTL regions
In the domestic silkworm, miRNAs were characterized in the fifth-instar posterior silk gland using NGS technology39. Based on the previous annotation, 39 miRNAs were found in the QTL regions (Supplementary Table S8). To understand their potential roles, the interaction of the miRNAs and the 3′ untranslated region (3′ UTR) of silk fibroin genes was predicted with miRanda software (Supplementary Table S8). The results indicated that Fib-L (fibroin light chain) is a putative target of bmo-miR-P009-3p, bmo-miR-P1416-3p and bmo-miR-P581-5p. More interestingly, bmo-miR-P100-3p presented some SNPs between the domestic and wild populations (Fig. 4e). In addition, KWMTBOMO04917 is one of the putative targets of bmo-miR-P100-3p in domestic silkworms. However, due to the SNP (A to G) in W_Anyue, W_Beibei and W_Suda, the orthologous bmo-miR-P100-3p exhibited no target site in the 3′ UTR of KWMTBOMO04917 in those wild silkworms.
Discussion
In this study, the domestic silkworm and its wild relative B. mandarina were used for mapping parents. A linkage map of 9,632 SNP markers was constructed, and 11 QTLs related to WCW, CSW, CSR, and PW traits were identified (Figs. 1 and 2). In the previous study, an integrated consensus map contained 692 unique SSR sites was developed7. Based on the linkage map, at least six QTLs involved in WCW, CSW, CSR, and PW were detected, which located in chromosomes 1, 18, 19, 21, 22, and 23. In addition, using the pooling sequencing-based methodology, 8 SNPs linked with CSW trait were also identified, which located on chromosomes 11, 22 and 2312. These results suggested that the silk yield traits were mainly located on some specific chromosomes. However, different studies might find diverse QTLs on the same chromosome. For instance, SNP marker193648 on chromosome 2312 was not included in qtl2-2/ qtl2-3 regions related to CSW in present study. It was indicated that silk yield is controled by multiple QTLs, and different mapping parents might obtain some novel loci.
Silk gland is responsible for silk protein synthesis and excretion in silkworm40. In this study, the developmental transcriptome of the fifth-instar silk glands was compared between the domestic and wild silkworms, in which KWMTBOMO04917 and KWMTBOMO12906 were DEGs in multiple stages (Supplementary Table S5). At the same time, KWMTBOMO04917 was one of the domestication-related genes that showed higher expression in the middle and posterior silk gland of the wild silkworm compared to those of the domestic strains (Fig. 5c). However, no significant difference was found in domestic strains Xiafang and Dazao, which might be affected by the direction of artificial selection, which has resulted in interspecies differences in gene expression and low within-species variance41. KWMTBOMO04917 encodes 5-aminolevulinate synthase (BmALAS), which may function as a homodimer and converts glycine into 5-aminolevulinate in a single step42. A previous study revealed that glycine was one of the major amino acid residues in the silk fibroin heavy chain43. Due to the lower expression of BmALAS, more glycines might be used for the synthesis of silk proteins in domestic silkworm. In addition, the expression of KWMTBOMO12906, encoding DNA polymerase epsilon (Pol ε) subunit 4 (POLE4), was positively related to cocoon yield (Fig. 5d and Supplementary Fig. S2). The protein sequences of POLE4 were identical among Dazao, D_XF, and W_AK (Supplementary Fig. S3). Previous studies suggested that the synthesis of silk proteins is proportional to the DNA dosage per unit silk gland cell44,45. We assumed that the higher expression of POLE4 in the high-yield strain might increase the DNA content in silk gland cells and result in an increased cocoon yield.
Except for B. mori, wild silkworm might also possess genes potentially involved in silk improvement. In this study, one yield-enhancing QTLs (qtl2-4) was identified from wild silkworm (Table 3), which provides an opportunity to mine favourable genes from wild relatives. Only 8 protein-encoding genes were identified in qtl2-4, in which 3 genes showed differential expression in developmental transcriptomes of the fifth-instar silk glands between the domestic and wild silkworms (Supplementary Table S5). KWMTBOMO16283 has higher expression in the domestic silkworm than wild silkworm during all the fifth-instar stages. While its function is unknown through homologous search in GenBank (Supplementary Table S5). It is worth exploring whether KWMTBOMO16283 plays roles in mediating the biosynthesis and secretion of silk proteins.
MicroRNAs (miRNAs) recognized as important regulators of gene expression in higher eukaryotes46. In recent years, several miRNAs have been reported to control rice grain yield47,48. In the domestic silkworm, miRNAs were characterized in silk gland39. Based on the annotation, 39 miRNAs were found in the QTL regions (Supplementary Table S8), which may potentially regulate the genes related to protein synthesis and processing39. It was found that no miRNA within QTLs was located in the candidate selective sweep regions (Supplementary Table S6). While, three SNPs was detected in bmo-miR-P100-3p between the domestic and wild populations (Fig. 4e). Especially, the SNP (G to A) can affect formation and stability of miRNA-target duplex, which may down-regulate the expression of KWMTBOMO04917 in the domestic silkworm (Fig. 5c). In addition, some differentially expressed miRNAs were found in the silk gland among three silkworm strains with differential silk production49. It was suggested that differential expression of miRNAs may also affect transcriptional and translational level of the target genes. Further studies are needed to determine whether those miRNAs within the QTLs would influence expression of the target genes and cocoon yield traits in silkworm.
Conclusions
In this study, 11 QTLs related to cocoon yield traits were mapped to 7 chromosomes in silkworm. Some of those traits shared the adjacent genomic regions, which shows a certain genetic relatedness. Integrated QTL mapping, population genomics, and transcriptional profiling analyses allowed us to implicate some potential causative genes for cocoon yield traits. Simultaneously, one trait-improving QTL allele (qtl2-4) from the wild parent was identified, which will help to explore yield-enhancing genes in wild silkworms. The top candidate genes will be particularly interesting targets for experimental verification.
Materials and Methods
Silkworm rearing and sample preparation
The domestic silkworm was reared on fresh mulberry leaves at 25 ± 1 °C and 75% ± 3% relative humidity (14 hours light: 10 hours dark). The wild silkworm was collected from Ankang City, Shaanxi Provence in China. An F1 intercross was performed from a female of the domestic silkworm Xiafang (D_XF) and a male of the Ankang wild silkworm (W_AK). We generated backcross (BC1M) progeny through single-pair matings between an F1 male with a Xiafang female. Unhealthy individuals were discarded during rearing. On the eighth day after wandering, the cocoon yield traits of each individual were investigated, including whole cocoon weight (WCW), cocoon shell weight (CSW), cocoon shell ratio (CSR), and pupal weight (PW). Thereafter, the pupae were stored at −80 °C for genomic DNA isolation.
RAD sequencing
Genomic DNA was extracted from 100 BC1 individuals using the classical phenol/chloroform method50. The quality of DNA was checked with a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA), and through 1% agarose gel electrophoresis. DNA from each individual was digested with the restriction endonuclease EcoR I. Solexa P1 adapters were added to the digested fragments. The ligated DNA samples were pooled and sheared using a Covaris S-Series ultrasonicator (Covaris) to an average size of 500 bp. The sheared samples were separated in a 1.5% agarose gel, and DNA fragments of 300 to 700 bp were isolated and enzymatically blunted. The Klenow fragment (exo-; New England Biolabs) was then used to add an adenine to the 3′ end. An adapter with divergent ends (P2 adapter) was ligated to conduct selective PCR. The libraries were run for sequencing of paired-end reads (100 bp) on the Illumina HiSeq. 2000 Genome Analyzer platform (Novogene, Beijing, China).
Reads mapping to the reference genome
The genomes of the mapping parents D_XF and W_AK were resequenced in a previous study by our group and deposited in the ENA database (The European Bioinformatics Institute) under accession number PRJEB545831. The raw reads of the mapping population and parents were filtered by removing adaptor sequences and low-quality sequences containing >10% poly-N or >50% bases with Phred quality scores ≤5. The genome version 3.0 of the silkworm was downloaded from SilkBase (http://silkbase.ab.a.u-tokyo.ac.jp). The clean reads were mapped to the reference genome using Burrows-Wheeler Alignment (BWA) tool51, with the following parameters: aln -o 1 -m 100000 -t 4 -l 32 -i 15 -q 10. Reads that were not uniquely mapped were discarded and not considered in further analysis.
SNP detection and genetic marker development
Alignments were piped with SAMtools v0.1.1852 and reformatted into BAM and pileup files for SNP identification, with the following parameters: -m 2 -F 0.002 -d 1000. Among the homozygous SNPs in the intercross parents, polymorphic SNP markers were developed. To be considered for genotyping design, an SNP had to exhibit minimum sequencing coverage of 4× in the intercross parents and 2× in BC1 individuals. SNPs that were consistently identified in parents and the progenies were retained. Some abnormal bases existed in offspring rather than parents, which were treated as a deficiency in the offspring. To maintain the integrity of markers, the degree of genotype coverage in offspring was over 85%. If the distribution of markers in offspring showed segregation distortion (chi-square test, P < 0.001)53, it was discarded. After filtration based on abnormal bases, integrity and segregation distortion, the high-quality markers were used for genotyping.
Linkage group construction and QTL analysis
The genetic linkage map was constructed using MSTmap software54 and visualized with Map Draw V2.155. The composite interval mapping (CIM) algorithm implemented in WinQTLCart v2.5 software (http://statgen.ncsu.edu/qtlcart/WQTLCart.htm) was employed to scan QTLs. The CIM analysis was based on Model 6, with a 1 cM step size, forward and backward stepwise regression, and five control markers. The LOD threshold was determined based on 1,000 permutations with a genome-wide level of significance of 5%. Sex as a categorical trait was used as a factor that can be “regressed out” in regression analysis.
Screening the genes under artificial selection within QTL regions
The putative candidate genes were explored in the genomic region located within the boundary markers of QTLs and its 200 kb flanking regions. Gene Ontology (GO) enrichment analysis was performed with BLAST2GO56. Pathway analysis of the candidate genes was performed with the KOBAS 2.0 online tool57. Whole-genome resequencing data of eight domestic and seven wild individuals could be archived from National Genomics Data Center (NGDC, http://bigd.big.ac.cn/) under accession number PRJCA002125. These deeply resequenced genomes (approximately 40-fold coverage) were aligned to the silkworm reference genome using BWA-MEM v0.7.7 with the default parameters58. We conducted a series of procedures before SNP calling, including removing PCR duplicates with Picards v1.41, realignment of indels using GATK v3.459. SNPs and insertion/deletion polymorphisms (INDELs) were identified with the GATK, SAMtools, and FreeBayes tools52,60,61. The shared variants supported by at least five reads were used in the subsquent analysis of population genetics. To identify the genes that may have been subjected to selection during domestication, we calculated the ratio of genetic diversity (θπ, domestic/θπ, wild) and the population differentiation statistic (FST)62,63. We used an empirical procedure and selected windows with a 5% significance level of the θπ ratio and a 5% significance level of FST as candidate regions with strong signals of a selective sweep for the domestic silkworm17,64. In the QTL regions, genes showing evidence of a selective sweep were considered as important candidates35,64.
Developmental transcriptome analysis of the fifth-instar silk gland
The domestic strain Xiafang and the wild silkworm from Chongqing in China were used for comparative transcriptome analysis. The rearing method of the wild silkworm was similar to that applied in our previous study38. Seven time points of fifth-instar larvae were selected in the wild silkworm, including beginning (0d), day 1 (1d), day 2 (2d), day 3 (3d), day 4 (4d), and day 5 (5d) of the fifth-instar and beginning of wandering (w). Due to the longer feeding period of the domestic silkworm, the corresponding time points were averagely partitioned in relation to those of the wild silkworm. The intact silk glands of three individuals were dissected at each time point. Each sample was repeated two times for RNA sequencing. Transcriptome sequencing and analysis were performed similarly to our previous study38. The analysis of differential expression was conducted at the same time point between the domestic and wild silkworms using the edgeR package65. The P-value was adjusted by the method of Benjamini & Hochberg66. Differentially expressed genes (DEGs) were identified according to threshold values of an adjusted P-value <0.05 and a|log2 fold-change|>1.
Prediction of microRNA-target duplexes
Based on miRNA annotation in silkworm39, miRNAs located in QTL regions were identified. The putative targets of the miRNAs were mainly focused on silk fibroin heavy chain (Fib-H), light chain (Fib-L) and P25, and the QTL-harbored DEGs in developmental transcriptomes of the fifth-instar silk gland between the domestic and wild silkworms. The 3′ untranslated regions (3′-UTRs) of Fib-H, Fib-L and P25 were retrieved from BB988047.1, NM_001044023.1, and X04226.1, respectively, in GenBank. The target site of 3′-UTR region was predicted with miRanda v3.3a software67 based on the following parameters: score cutoff = 140, energy cutoff ≤ −7.0, gap opening: −9.0, and gap extension −4.0.
Ethics approval and consent to participate
Experiments were conducted in accordance with the protocol approved by the Institutional Animal Care and Use Committee of the Chongqing University (permit number CBE-A201405010).
Data availability
RAD sequencing datasets of 100 BC1 individuals were available at NCBI Sequence Read Archive (SRA) database under accession number PRJNA453838. Whole-genome resequencing data and the developmental transcriptome of the fifth-instar larval silk gland could be retrived from NGDC (http://bigd.big.ac.cn/) under the accession numbers PRJCA002125 and PRJCA001835, respectively.
References
Darwin, C. The variation of animals and plants under domestication. London: John Murrary (1868).
Li, M., Zhong, W., Yang, F. & Zhang, Z. Genetic and molecular mechanisms of quantitative trait loci controlling maize inflorescence architecture. Plant Cell Physiol. 59, 448–457 (2018).
Zuo, J. & Li, J. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu. Rev. Genet. 48, 99–118 (2014).
Xia, Q. et al. Complete resequencing of 40 genomes reveals domestication events and genes in silkworm (Bombyx). Science 326, 433–436 (2009).
Hemmatabadi, R. N., Seidavi, A. & Gharahveysi, S. A review on correlation, heritability and selection in silkworm breeding. J. Appl. Anim. Res. 44, 9–23 (2016).
Lu, C., Li, B., Zhao, A. & Xiang, Z. QTL mapping of economically important traits in silkworm (Bombyx mori). Sci. China C Life Sci. 47, 477–484 (2004).
Zhan, S. et al. An integrated genetic linkage map for silkworms with three parental combinations and its application to the mapping of single genes and QTL. BMC Genomics 10, 389, https://doi.org/10.1186/1471-2164-10-389 (2009).
Mirhoseini, S. Z., Rabiei, B., Potki, P. & Dalirsefat, S. B. Amplified fragment length polymorphism mapping of quantitative trait loci for economically important traits in the silkworm, Bombyx mori. J. Insect Sci. 10, 153 (2010).
Zhang, L., Lu, C., Dai, F. Y. & Fang, S. M. Mapping of major quantitative trait loci for economic traits of silkworm cocoon. Genet. Mol. Res. 9, 78–88 (2010).
Xu, H. M. et al. A new mapping method for quantitative trait loci of silkworm. BMC Genet. 12, 19, https://doi.org/10.1186/1471-2156-12-19 (2011).
Li, B., Wang, X. Y., Hou, C. X., Xu, A. Y. & Li, M. W. Genetic analysis of quantitative trait loci for cocoon and silk production quantity in Bombyx mori (Lepidoptera: Bombycidae). Eur. J. Entomol. 110, 205–213 (2013).
Li, C. et al. QTL analysis of cocoon shell weight identifies BmRPL18 associated with silk protein synthesis in silkworm by pooling sequencing. Sci. Rep. 7, 17985, https://doi.org/10.1038/s41598-017-18277-y (2017).
Davey, J. W. et al. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 12, 499–510 (2011).
Baxter, S. W. et al. Linkage mapping and comparative genomics using next-generation RAD sequencing of a non-model organism. PLoS One 6, e19315, https://doi.org/10.1371/journal.pone.0019315 (2011).
Wu, K. et al. High-density genetic map construction and QTLs analysis of grain yield-related traits in Sesame (Sesamum indicum L.) based on RAD-Seq techonology. BMC Plant Biol. 14, 274, https://doi.org/10.1186/s12870-014-0274-7 (2014).
Nadeau, N. J. et al. Population genomics of parallel hybrid zones in the mimetic butterflies, H. melpomene and H. erato. Genome Res. 24, 1316–1333 (2014).
Li, M. et al. Genomic analyses identify distinct patterns of selection in domesticated pigs and Tibetan wild boars. Nat. Genet. 45, 1431–1438 (2013).
Liu, S. X. et al. Identification of single-nucleotide polymorphism markers associated with cortisol response to crowding in rainbow trout. Marine Biotech. 17, 328–337 (2015).
Jensen, P. Behavior genetics and the domestication of animals. Annu. Rev. Anim. Biosci. 2, 85–104 (2014).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Tan, L. et al. Quantitative trait loci underlying domestication- and yield-related traits in an Oryza sativa × Oryza rufipogon advanced backcross population. Genome 51, 692–704 (2008).
Price, N. et al. Combining population genomics and fitness QTLs to identify the genetics of local adaptation in Arabidopsis thaliana. Proc. Natl. Acad Sci. USA 115, 5028–5033 (2018).
Ruiz, X. & Almanza, M. Implications of genetic diversity in the improvement of silkworm Bombyx mori L. Chil. J. Agr. Res. 78, 569–579 (2018).
Zamani, P., Ghanipoor, M., Mirhosseini, S. Z., Abdoli, R. & Seidavi, A. Comparison of different selection strategies for mulberry silkworm, Bombyx mori. Int. J. Trop. Insect Sci. 39, 139–145 (2019).
Xiang, H. et al. The evolutionary road from wild moth to domestic silkworm. Nat. Ecol. Evol. 2, 1268–1279 (2018).
Xiao, J. et al. Genes from wild rice improve yield. Nature 384, 223–224 (1996).
Wu, Y.C., P., Q. & S.M., H. Breeding of a new silkworm variety “Yesanyuan” with healthiness and hypersilk. J. Anhui Agric. Sci. 36, 14621–14622 (In Chinese) (2008).
Xiao, J. et al. Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genet. 150, 899–909 (1998).
Ma, S. Y., Smagghe, G. & Xia, Q. Y. Genome editing in Bombyx mori: new opportunities for silkworm functional genomics and the sericulture industry. Insect Sci. 26, 964–972 (2019).
Ali, A. et al. Knockdown of broad-complex gene expression of Bombyx mori by oligopyrrole carboxamides enhances silk production. Sci. Rep. 7, 805, https://doi.org/10.1038/s41598-017-00653-3 (2017).
Zhao, Q., Han, M. J., Sun, W. & Zhang, Z. Copy number variations among silkworms. BMC Genomics 15, 251, https://doi.org/10.1186/1471-2164-15-251 (2014).
International Silkworm Genome Consortium. The genome of a lepidopteran model insect, the silkworm Bombyx mori. Insect Biochem. Mol. Biol. 38, 1036–1045 (2008).
Kawamoto, M. et al. High-quality genome assembly of the silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 107, 53–62 (2019).
Johnsson, M. et al. Feralisation targets different genomic loci to domestication in the chicken. Nat. Commun. 7, 12950, https://doi.org/10.1038/ncomms12950 (2016).
Lim, J. H., Yang, H. J., Jung, K. H., Yoo, S. C. & Paek, N. C. Quantitative trait locus mapping and candidate gene analysis for plant architecture traits using whole genome re-sequencing in rice. Mol. Cells 37, 149–160 (2014).
Qiu, C. Z. et al. Evidence of peripheral olfactory impairment in the domestic silkworms: insight from the comparative transcriptome and population genetics. BMC Genomics. 19, 788, https://doi.org/10.1186/s12864-018-5172-1 (2018).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Fang, S. M., Hu, B. L., Zhou, Q. Z., Yu, Q. Y. & Zhang, Z. Comparative analysis of the silk gland transcriptomes between the domestic and wild silkworms. BMC Genomics 16, 60, https://doi.org/10.1186/s12864-015-1287-9 (2015).
Li, J. et al. MicroRNA expression profiling of the fifth-instar posterior silk gland of Bombyx mori. BMC Genomics 15, 410, https://doi.org/10.1186/1471-2164-15-410 (2014).
Xia, Q., Li, S. & Feng, Q. Advances in silkworm studies accelerated by the genome sequencing of Bombyx mori. Annu. Rev. Entomol. 59, 513–536 (2014).
Romero, I. G., Ruvinsky, I. & Gilad, Y. Comparative studies of gene expression and the evolution of gene regulation. Nat. Rev. Genet. 13, 505–516 (2012).
Hunter, G. A. & Ferreira, G. C. 5-aminolevulinate synthase: catalysis of the first step of heme biosynthesis. Cell Mol. Biol. (Noisy-le-grand). 55, 102–110 (2010).
Zhou, C. Z. et al. Silk fibroin: structural implications of a remarkable amino acid sequence. Proteins 44, 119–122 (2001).
Patel, C. V. & Gopinathan, K. P. Development stage-specific expression of fibroin in the silk worm Bombyx mori is regulated translationally. Indian J. Biochem. Biophys. 28, 521–530 (1991).
Gage, L. P. Polyploidization of the silk gland of Bombyx mori. J. Mol. Biol. 86, 97–108 (1974).
Yates, L. A., Norbury, C. J. & Gilbert, R. J. The long and short of microRNA. Cell 153, 516–519 (2013).
Liu, Q. et al. Integrating small RNA sequencing with QTL mapping for identification of miRNAs and their target genes associated with heat tolerance at the flowering stage in rice. Front. Plant Sci. 8, 43, https://doi.org/10.3389/fpls.2017.00043 (2017).
Zhang, J. P. et al. miRNA miR408 regulates grain yield and photosynthesis via a phytocyanin protein. Plant Physiol. 175, 1175–1185 (2017).
Qin, S. et al. MicroRNA profile of silk gland reveals different silk yields of three silkworm strains. Gene. 653, 1–9 (2018).
Yasukochi, Y. A dense genetic map of the silkworm, Bombyx mori, covering all chromosomes based on 1018 molecular markers. Genet. 150, 1513–1525 (1998).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kakioka, R., Kokita, T., Kumada, H., Watanabe, K. & Okuda, N. A RAD-based linkage map and comparative genomics in the gudgeons (genus Gnathopogon, Cyprinidae). BMC Genomics 14, 32, https://doi.org/10.1186/1471-2164-14-32 (2013).
Wu, Y., Bhat, P. R., Close, T. J. & Lonardi, S. Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet. 4, e1000212, https://doi.org/10.1371/journal.pgen.1000212 (2008).
Liu, R. H. & Meng, J. L. MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas (Beijing) 25, 317–321 (In Chinese) (2003).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Xie, C. et al. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic. Acids Res. 39, W316–22 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. ArXiv e-prints 1207, 3907 (2012).
Wang, M. S. et al. Positive selection rather than relaxation of functional constraint drives the evolution of vision during chicken domestication. Cell Res. 26, 556–573 (2016).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Qiu, Q. et al. Yak whole-genome resequencing reveals domestication signatures and prehistoric population expansions. Nat Commun. 6, 10283, https://doi.org/10.1038/ncomms10283 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Statist. Soc. B. 57, 289–300 (1995).
Enright, A. J. et al. MicroRNA targets in. Drosophila. Genome Biol. 5, R1, https://doi.org/10.1186/gb-2003-5-1-r1 (2003).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31772524), the Venture & Innovation Support Progam for Chongqing Overseas Returnees (cx2018066), and Initiation Fund of China West Normal University (No. 15E022). We thank Hong-Bo Zhang for preparing the genomic resequencing data. The authors sincerely thank the anonymous peer reviewers for their helpful comments and suggestions.
Author information
Authors and Affiliations
Contributions
Y.Q.Y. and Z.Z. conceived and designed the study. S.M.F. and Y.Q.Y. performed the analysis of QTL mapping and related experiments. Q.Z.Z. performed the developmental transcriptome analysis. S.M.F and Y.Q.Y. drafted the manuscript. Y.Q.Y. and Z.Z. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, SM., Zhou, QZ., Yu, QY. et al. Genetic and genomic analysis for cocoon yield traits in silkworm. Sci Rep 10, 5682 (2020). https://doi.org/10.1038/s41598-020-62507-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-62507-9
This article is cited by
-
Application of biotechnology in sericulture: Progress, scope and prospect
The Nucleus (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.