Abstract
The zebrafish (Danio rerio) is a model species that is used to study the circadian clock. It possesses light-entrainable circadian clocks in both central and peripheral tissues, and its core circadian factor cryptochromes (CRYs) have diverged significantly during evolution. In order to elucidate the functional diversity and involvement of CRYs in photoperiodic mechanisms, we investigated the daily expression profiles of six Cry transcripts in central (brain and eye) and peripheral (fin, skin and muscle) tissues. The zCry genes exhibited gene-specific diurnal conserved variations, and were divided into morning and evening groups. Notably, zCry1ab exhibited biphasic expression profiles in the eye, with peaks in the morning and evening. Comparing ocular zCry1ab expression in different photoperiods (18L:6D, 14L:10D, 10L:14D and 6L:18D) revealed that zCry1ab expression duration changed depending on the photoperiod: it increased at midnight and peaked before lights off. zCry1ab expression in constant light or dark after entrainment under long- or short-day conditions suggested that the evening clock and photic input pathway are involved in photoperiod-dependent zCry1ab expression. Laser microdissection followed by qRT-PCR analysis showed that the evening peak of zCry1ab was likely ascribed to visual photoreceptors. These results suggest the presence of an eye-specific photoperiodic time measurement served by zCry1ab.
Similar content being viewed by others
Introduction
The circadian clock consists of three parts: the input pathway(s), the oscillation system and the output pathway(s). Light is the most important signal that activates the input pathway to synchronize the oscillator with daily external cycles. The oscillator generates time signals to control physiological circadian functions through the output pathway. Internal time signals are integrated with the environmental light signal to measure day length, which triggers seasonal photoperiodic responses in many organisms living in temperate zones1.
The zebrafish (Danio rerio) circadian clock is one of the most studied among fish species, and is reported to exist in not only the central tissues but also the embryo, larva, peripheral tissues and cultured cells (refs. 2,3,4,5,6,7,8,9,10,11 in Table 1, refs. 12,13,14,15). These zebrafish circadian clocks are directly light-entrainable and this feature is also seen in the other fish cells16 in contrast to the mammalian circadian clocks that are not light-entrainable except for those in the eye.
Among vertebrate core clock components such as period (PER), CLOCK, BMAL and cryptochrome (CRY) proteins, CRYs have both a wide evolutionary background and divergent molecular functions6. CRYs are structurally classified into several groups, and together form a large protein family with the photo repair enzymes, photolyases (PHRs). Among CRY/PHR family proteins, animal-type CRYs (CRY1 and CRY2) play a central role in the circadian clock oscillator. Animal-type CRY1 and CRY2 function as transcriptional repressors in the core loop of the circadian clock oscillation system17,18, while fruit fly CRY (dCRY) and other non-mammalian CRYs serve as blue light photoreceptor molecules using flavin adenine dinucleotide (FAD) as a chromophore19. In tropical fish, animal-type CRY2 (termed CRY3 in refs. 20,21,22) may function for the lunar timer20,21,22. In addition, CRY4 was found in the chicken pineal gland23 and retina24 and shown to absorb light to change its structure25. Biophysical analyses using recombinant CRY4 proteins indicated their chromophore FAD binding and photoreceptive functions25,26, and hence CRY4 is strongly suggested to be a photoreceptor and/or a light-driven magnetoreceptor27.
Six Cry gene paralogues have been reported in zebrafish: four animal-type Cry1s (zCry1aa, zCry1ab, zCry1ba and zCry1bb), one animal-type Cry2 (zCry2; termed zCry3 in ref. 2) and one Cry4 (zCry4; termed zCry3 in ref. 6)2,6. These genes have evolved through multiple gene duplication events during teleost evolution6, but their functional differences and redundancies are not fully understood. While the animal-type CRY2 (zCRY2) and zCRY4 have only weak or no transcriptional repression ability for the CLOCK:BMAL complex, the four animal-type CRY1s have strong transcriptional repression abilities14, so may not only be involved in the circadian clock but also in clock-associated functions, such as the photoperiodic response. Although daily zCry expression patterns have been investigated both in vivo2,3,4,5,6,8,11 and in vitro7,9,10, we comparatively reinvestigated the spatiotemporal expression profiles and light-responsiveness of zCrys to test the above hypothesis. We found that zCry1ab expression in the eye changes depending on the day length. Based on the zCry1ab expression profiles, we suggest a model of zCry1ab expression regulation in the retinal photoreceptors, in which the light and time information may encode the environmental photoperiod into transcriptional regulatory signals.
Results
Daily variations in zCry mRNA expression levels
We examined the transcript levels of six Cry genes in the brain, eye, pectoral fins, skin and muscle at five time points (ZT1, ZT7, ZT15, ZT19 and ZT1) in a white light, long-day (LD; 14L:10D) cycle by qRT-PCR (Figs. 1 and 2, Supplementary Tables S1–S31), and their acrophases were estimated by cosinor-fitting analyses (Fig. 3, Supplementary Table S32).
Daily profiles of zCry1aa, zCry1ab and zCry2 mRNA expression levels in central and peripheral tissues under white light/dark cycles. Each tissue was collected at ZT1, ZT7, ZT15 or ZT19 from zebrafish (n = 4) entrained to white light/dark cycles (14L:10D). Each mRNA level was estimated using zβ-actin as a reference gene because it was the most stably expressed in the examined tissues among the control genes. Data were analyzed by one-way ANOVA (Supplementary Table S1) and Tukey-Kramer post-hoc tests (Supplementary Tables S2–S31). Error bars represent ± SE. Peak times of cosine-fitting to the data as calculated by CircWave1.4 (red dashed curves; Supplementary Table S32) are shown. A down-pointing triangle in panel g denotes the sustained expression of zCry1ab transcripts at ZT15 in the eye.
Daily profiles of zCry1ba, zCry1bb and zCry4 mRNA expression levels in central and peripheral tissues under white light/dark cycles. The experimental conditions and data analyses were the same as those described in Fig. 1.
Maximum Cry expression in zebrafish tissues. Acrophases of Cry expression as estimated by cosinor analysis (Supplementary Table S32) are shown. Red up-pointing triangles, orange squares, yellow circles, green diamonds, dark-blue down-pointing triangles and light-blue crosses show estimated acrophases for zCry2, zCry1ab, zCry1aa, zCry4, zCry1ba and zCry1bb, respectively.
In all tissues examined, zCry mRNA expression levels exhibited daily changes (p < 0.01, Supplementary Table S1), indicating that their expression was under the control of the circadian clock and/or environmental light. The expression patterns of every gene investigated were similar across the tissues (Figs. 1 and 2), except for a few cases (see below), and the zCry genes were classified into two groups based on their daily expression patterns (Fig. 3, Supplementary Table S32): genes in the ‘morning group’ were zCry1aa, zCry1ab and zCry2 (Fig. 1), the expression patterns of which were well-fitted to curves with peaks in the first half of the light period (ZT0.5-ZT6.9), while genes in the ‘evening group’ were zCry1ba, zCry1bb and zCry4 (Fig. 2), which were well-fitted to curves with peaks at the start of the dark period (ZT11.3-ZT13.4), except for zCry4 in the muscle (ZT8.2) (Figs. 2 and 3).
Interestingly, zCry1ab exhibited sustained expression at ZT15 in the eye (the down-pointing triangle in Fig. 1g; p < 0.01 vs. ZT19, Supplementary Table S8), suggesting possible biphasic expression, which was not observed in any other tissue examined (Fig. 1f,h–j).
Circadian and photic regulation of zCry expression
We next investigated the expression patterns of zCrys under blue light or dark to further characterize their circadian and photic regulations (Fig. 4). Here we focused on the brain and eye, because (i) expression patterns were not substantially different across the tissues (Figs. 1 and 2) and (ii) the expression levels of every Cry were relatively high in these tissues.
zCry mRNA levels under blue light/dark and dark conditions. Zebrafish were entrained to white light/dark cycles (14L:10D) and irradiated with blue (λmax = 462 nm; λ1/2 = 453 nm and 573 nm; blue circles) light or kept in darkness (black squares) at 28 °C for 14 h from ZT0 (Supplementary Fig. S1). Light intensities were set at 5.586 × 1014 photons s−1 m−2 that corresponds to 240 μW cm−2. Tissues were collected from three (brain samples at ZT0 and ZT19, and eye samples at ZT7 and ZT19) or four (other samples) zebrafish at ZT0, ZT1, ZT3, ZT7, ZT14, ZT15, ZT19 or ZT0 (ZT24) (see Methods for details). Each mRNA level was estimated as a relative value to the geometric mean of mRNA levels of zβ-actin, zGapdh and zEf1α (arithmetic means of their CT values were used in the 2−ΔΔCT method because they were stably expressed in the brain and eye). Data were analyzed by two-way ANOVA (Supplementary Table S33) and Tukey-Kramer post-hoc tests (Supplementary Tables S34–S57). Error bars represent ± SE.
Zebrafish were entrained to the 14L:10D cycle and then exposed to blue light (Supplementary Fig. S1) for 14 h from ZT0 or kept in the dark. To evaluate the temporal effects of changing the light conditions, we changed sampling points to include ‘lights on’ and the mid-points of the light and dark periods (ZT0, ZT1, ZT3, ZT7, ZT14, ZT15, ZT19 and ZT0).
In the eye, light effects were observed in all the examined zCry genes (Fig. 4d–f,j–l; interaction p < 0.05, Supplementary Table S33). The zCry1aa expression level under the blue light condition was higher at ZT3 than that under the dark condition (p < 0.01, Supplementary Table S35). Notably, the eye-specific biphasic expression of zCry1ab (Fig. 1g) was reproduced with peaks in both the morning and evening under the blue light condition (Fig. 4e). The zCry1ab expression level at the evening peak was significantly higher than that in the dark (blue light vs dark at ZT14; p < 0.05, Supplementary Table S37). Similarly, all the evening-group genes showed the light-dependent upregulation in the evening (ZT14, Fig. 4j–l; p < 0.01, Supplementary Tables S39, S41, and S45). Under the dark condition, all the morning- and evening-group zCry genes exhibited significant temporal variations with peaks in the morning and evening, respectively (Supplementary Tables S33, S35, S37, S39, S41, S43, and S45). These results suggested that both the morning and evening zCry gene expressions are controlled by the circadian clock and additionally regulated by the light signal in the eye.
In the brain, expressions of the four zCry1 genes were affected by the light (Fig. 4a,b,g,h; interaction p < 0.05, Supplementary Tables S33, S34, S36, S38, and S40), but those of zCry2 and zCry4 were not (Fig. 4c,i; Supplementary Tables S33, S42, and S44). Light-dependent upregulations were observed in zCry1aa (Fig. 4a; p < 0.01, blue light vs dark at ZT3, Supplementary Table S34) and zCry1bb (Fig. 4h; p < 0.01 in the blue light vs dark at ZT7, ZT14, and ZT15, Supplementary Table S40), but not in the other zCry genes. Under the dark condition, no significant temporal variation was observed in zCry1aa, zCry1ba, and zCry1bb, although they showed weak variations (Fig. 4a,g,h; Supplementary Tables S34, S38, and S40).
Photoperiodic control of the zCry1ab mRNA level
The biphasic expression of zCry1ab in the eye (Fig. 4e) led us to hypothesize that zCry1ab expression is dependent upon day length. To explore this possibility, we entrained zebrafish to blue light/dark cycles of the different photoperiods and investigated the expression patterns of ocular zCry1ab at 2-h intervals (Fig. 5). Under all of the photoperiodic conditions examined, the evening peaks were more pronounced than the morning (shoulder) peaks. Notably, the expression profiles of zCry1ab changed depending on the photoperiod (interaction p < 0.01, Supplementary Table S58): both the morning and evening peaks advanced with the shortening of the light period.
Daily profiles of zCry1ab mRNA expression levels in the eye under different photoperiods. Zebrafish were entrained to 18L:6D extra-long-day (ELD; panel a), 14L:10D long-day (LD; panel b), 10L:14D short-day (SD; panel c) or 6L:18D extra-short-day (ESD; panel d) cycles. Eyes (n = 5, no mark; n = 4, +; n = 3, *) were collected to measure zCry1ab mRNA levels (see Methods for details) at the indicated time points from ZT1. Each mRNA level was estimated as a relative value to the geometric mean of mRNA levels of zβ-actin, zGapdh and zEf1α. Error bars represent ± SD. Light and dark periods are indicated in blue and grey, respectively. Data were analyzed by two-way ANOVA (Supplementary Table S58) and Tukey-Kramer post-hoc tests (Supplementary Tables S59–S63).
In order to identify the underlying regulatory mechanisms, we compared zCry1ab profiles (relative mRNA levels) under the four photoperiodic conditions using different reference points (Fig. 6). When the profiles were plotted with ‘lights off’ as the reference point (Fig. 6b), the evening peaks were superimposed 1 h before lights off, except under ELD conditions (“E” in panel b), while the morning (shoulder) peaks diverged. Under ELD conditions (Figs. 5a and 6b), the evening peak occurred 3 h before lights off (ZT15), and the zCry1ab level decreased 1 h before lights off (ZT17). The fact that zCry1ab mRNA decreased during the light period refutes the hypothesis of dark-triggered mRNA downregulation around the end of the light period. In contrast to the evening peaks, the morning (shoulder) peaks seemed to be closer together when the profiles were plotted with midnight (broken lines in Fig. 6c) or noon as the reference points. In this plot, troughs and increasing phases coincided at midnight (“M” in Fig. 6c). These comparisons highlight the photoperiod-dependent changes in the duration of zCry1ab expression.
Double plots of 24-h profiles of relative zCry1ab expression levels in the eye under different photoperiods. Relative values of zCry1ab expression levels (Fig. 5; normalized by the peak) were double-plotted using ‘lights on’ (panel a), ‘lights off’ (panel b) or the middle of the night (panel c) as reference points. Data for two points at ZT1 (the start and end of sampling) were averaged. Evening peaks and increasing phases to morning (shoulder) peaks are indicated by E and M, respectively. Light conditions are indicated by bars with colored lines at the top of each panel.
Contributions of the light input pathway and circadian clock
zCry1ab mRNA level was maintained in the evening under constant dark condition (black square at ZT14 in Fig. 4e), implying that zCry1ab expression was regulated by not only the light signal but also the circadian clock in the evening. In order to validate this hypothesis, we investigated the evening levels of zCry1ab mRNA at 1-h intervals under blue light or in the dark after entrainment to the LD or SD conditions (Fig. 7). When the fish were kept under constant blue light conditions after entrainment to a LD condition (LD-LL, orange up-pointing triangles in Fig. 7a), the zCry1ab expression level increased from ZT11 and peaked at ZT13, before decreasing to a very low level at ZT16. This decrease in constant light confirmed that the decrease after the evening peak was not triggered by lights off, and instead suggested that downregulation was probably dependent on the circadian regulatory mechanism. In fact, a small peak was observed in the evening (12–14 h after ZT0) when the fish were kept under constant dark conditions after entrainment to a LD condition (LD-DD, black up-pointing triangles in Fig. 7a). The level of the peak at ZT13 in LD-LL was significantly higher than that in LD-DD (2.45-fold, p < 0.01, Light13 vs Dark13 in Supplementary Table S65), suggesting combinatorial regulation by both the circadian clock and photic upregulation to form the evening peak. We then investigated levels of zCry1ab transcripts after entrainment to short-day conditions (SD-LL and SD-DD, Fig. 7b, Supplementary Tables S68–S71). The evening peak was observed at ZT10-11 in SD-LL (blue down-pointing triangles in Fig. 7b), and a lower peak was observed at ZT11 under SD-DD conditions (black down-pointing triangles in Fig. 7b).
zCry1ab mRNA levels in the eye under constant light (LL) or dark (DD) conditions after entrainment to different photoperiods. (a) Zebrafish were entrained to a 14L:10D long-day cycle and kept in constant blue light (LL; yellow up-pointing triangles) or in the dark (DD; black up-pointing triangles). (b) Zebrafish were entrained to a 10L:14D short-day cycle and kept in constant blue light (LL; blue down-pointing triangles) or in the dark (DD; black down-pointing triangles). Eyes (n = 4, no mark; n = 3, +; n = 2, *) were collected to measure zCry1ab mRNA levels (see Methods for details) at the indicated time points from ZT0 on the last day. Each mRNA level was estimated as a relative value to the geometric mean of mRNA levels of zβ-actin, zGapdh and zEf1α. Data were analyzed by two-way ANOVA (Supplementary Tables S64 and S68) and Tukey-Kramer post-hoc tests (Supplementary Tables S65–S67 and S69–S71). Error bars represent ± SD.
Localization of the cell layer expressing zCry1ab mRNA in the evening by laser microdissection (LMD) and qRT-PCR
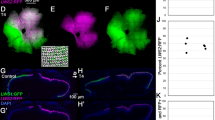
The photoperiodic and biphasic expression of zCry1ab in the retina raised questions about its physiological importance and its transcriptional regulation mechanisms. To approach these questions, we next tried to identify the retinal cell layer, in which the retina-specific biphasic expression of zCry1ab occurs, by LMD followed by qRT-PCR (Fig. 8). We collected two retinal regions, photoreceptor layer (PRL) and inner retinal layer (IRL)(Fig. 8a,b); PRL dominantly contains visual photoreceptors and IRL contains inner nuclear layer, inner plexiform layer, and ganglion cell layer. The zCry1ab mRNA level in PRL at ZT13 was significantly higher than those in IRL at ZT13 and PRL at ZT17 (Fig. 8c; p < 0.05, Supplementary Tables S72 and S73). The zCry1ab mRNA levels in PRL well agreed with the profile obtained in the analyses of the eye (Fig. 5b), while those in IRL were rather consistent with zCry1ab mRNA profile in the brain (Fig. 4b).
qRT-PCR analysis of zCry1ab mRNA levels in the zebrafish retinal cells captured by laser microdissection (LMD). Retinas were collected at ZT1 or ZT13 or ZT17 from zebrafish entrained to long-day cycles (14L10D). Photoreceptor cell layer (PRL) mainly containing outer nuclear layer (visual photoreceptors) and inner retinal layer (IRL) containing inner nuclear layer, inner plexiform layer and retinal ganglion cells were obtained from the retinal sections. (a) A retinal section (ZT1) showing PRL and IRL. (b) An example of retinal sections after the collection of PRL and IRL samples by LMD. (c) Levels of zCry1ab mRNA in PRL and IRL. The levels of zCry1ab mRNA and 18S rRNA were measured by qRT-PCR, and the zCry1ab mRNA levels relative to 18S rRNA levels were shown. Error bars represent ± SE. Data were analyzed by two-way ANOVA (Supplementary Table S72) and Tukey-Kramer post-hoc tests (Supplementary Table S73). Asterisks represent significant difference (p < 0.05).
Discussion
The zCry genes studied were classified into two groups based on their mRNA expression profiles in five zebrafish tissues (Figs. 1 and 2): the morning group (zCry1aa, zCry1ab and zCry2) and the evening group (zCry1ba, zCry1bb and zCry4), which exhibited expression peaks in the morning and evening, respectively (Fig. 3). The daily expression peaks of all six Cry genes seemed to be conserved in most of the tissues examined, except for ocular zCry1ab and muscle zCry4 (Fig. 3, Table 1), suggesting possible tissue-specific regulation mechanisms of zCry1ab and zCry4 in the eye and muscle, respectively. The further examination of photic and circadian regulations of the zCry genes between the brain and eye suggested that not only zCry1ab and zCry4 but also zCry1ba is likely regulated in a tissue-specific manner (Fig. 4g,j). The diurnal variation and/or light responsiveness of zCry mRNA expression in zebrafish cell lines and living animals have been reported by several groups so far (Table 1). Although some previous reports had different experimental conditions to the present study, such as sampling regime, tissues and light conditions, our results are mainly consistent with those obtained in previous studies. Together, they indicate that each zebrafish organ has light input and oscillation systems.
The divergent, or almost anti-phase, profiles of the morning and evening groups (Figs. 1–3) suggest that group-specific expression control occurred and there was functional divergence between the two groups. In fact, zCRYs exhibit varying repressive activity against CLOCK-BMAL complexes2,6,14, which could be explained by combinations of three CLOCKs and BMALs forming varying complexes. In mammals, E-box/E′-box, D-box and RRE are morning-time, day-time and night-time elements, respectively18 and CREB and AP-1 sites possibly function as light-responsive elements28,29. On the other hand, E-box and D-box are involved in light-dependent gene expression as well as circadian regulation in zebrafish30,31,32. Thus, zCry genes in the morning and evening groups may be differentially regulated by these elements. This hypothesis led us to searche for putative core clock and light-responsive elements (E-box/E′-box, D-box, RRE, CREB, and AP-1) in zCry genes (Supplementary Fig. S2, Supplementary Tables S74–S79). There is no clear difference between numbers of the putative elements in the morning- and evening-group genes, but clusters of E-box/E′-box and D-box elements were found in or near Exon 1 of the morning-group genes (Supplementary Fig. S2, red bars). On the other hand, the E-box/E′-box and D-box clusters were accompanied by RREs in the evening-group genes (Supplementary Fig. S2, red bars). Combinatorial regulations depending on the context of promoter/enhancer may occur in those clusters, and hence temporal and tissue-dependent changes in their regulatory activities should be further examined by genome-wide analyses such as ChIP-seq.
We found that the previously unreported unique expression properties of zCry1ab may be governed by both photoperiod- and tissue-specific regulation: zCry1ab expression peaks in the morning and evening, and the evening peak probably depends on phase-specific photic induction (Fig. 7). Weak rhythmic changes were observed around the projected time points, which corresponded with the end of the light period in the dark (LD-DD and SD-DD; Fig. 7a,b), indicating that the timing of the evening peak was synchronized with the internal clock, the phase of which was defined by the end of the light period in the previous cycle(s) (Fig. 9). The evening signal might trigger the phase-specific photoinduction of the zCry1ab promoter/enhancer region (Supplementary Fig. S2), in which light and circadian signals combinatorially regulate zCry1ab transcription. Such regulation may fine-tune the evening peak to occur just before lights off in a wide range of photoperiods (Figs. 5 and 6). In addition to the evening peak, zCry1ab transcription was probably tuned to start at midnight (Fig. 6c), which resulted in morning zCry1ab expression (between midnight and noon). It is unclear what mechanism underlies how the ocular cells anticipate midnight, so future studies should conduct transcriptome analyses of zebrafish eyes under different photoperiods in order to identify the mechanism involved.
Model of a possible zCry1ab regulatory circuit in the zebrafish eye. The zCry1ab mRNA level was regulated by morning, evening, light and day length signals. Based on their mRNA expression patterns (Figs. 1–3), Cry1aa/Cry2 and Cry1ba/Cry1bb were considered morning and evening oscillators, respectively. The morning signal induced the expression of zCry1ab at midnight, regardless of light and day length conditions. In addition to resetting the morning and evening clocks, the external light signal induced zCry1ab expression in the evening to form the evening peak, which may have been based on an integrated signal of light, day length and evening information. The morning and evening peaks were independent and additive. Shown on the right are schematic drawings of zCry1ab expression profiles under long-day and short-day cycles (modified from data shown in Figs. 4 and 7). Biological signal integration and regulation are represented using logic gates, with representative images of daily regulation patterns.
Another important issue is whether both the morning and evening peaks originated from multiple oscillatory loops present in a single cell or different subsets of ocular cells. The present analysis of zCry1ab mRNA levels in the retinal photoreceptor and the other layers (Fig. 8) implied that the biphasic expression occurs exclusively in the photoreceptor cells. Although it is difficult to show the presence of dual loops in a single photoreceptor cell, analyses of cultured photoreceptor cells or a photoreceptor-derived cell line would resolve the issue.
zCry1ab activity around dusk may be particularly important, because photoperiodic responses are generally triggered by a light stimulus at dusk called the “photo-inducible phase”1. In Japanese quail (Coturnix japonica), which is a photoperiodic species, the photo-inducible phase occurs around ZT13-ZT14, and downstream photoperiodic responses, such as the induction of Dio2 and thyroid hormones in the hypothalamic area, are well-characterised1. In the quail pars tuberalis, which is a key site for its seasonal responses, Cry1 is induced by light in a photoinducible-phase-specific manner33. The upstream molecular mechanism that opens the temporal window of the photoinducible phase has not yet been identified in quail or any other animal, so the zebrafish eye is a suitable model for investigating photoperiodic gene expression mechanisms.
When compared with a strongly photoperiodic fish, medaka (Oryzias latipes)34, photoperiodic responses had been less characterised in zebrafish. Zebrafish are naturally found in northern and north-eastern India (~26.5°N, ~89.5°E), and their breeding season is reportedly between April and August35. The day length change in their natural habitat is approximately 10–14 h that is not so large as those in temperate zones but may be enough to trigger the photoperiodic responses, because some tropical or neotropical animals respond to changes in the day length less than 1 h36,37.
Based on the result of LMD followed by qRT-PCR (Fig. 8), we consider that the light signal received in the visual photoreceptors likely controls the photoperiodic expression of zCry1ab. Possibly, it regulates the photoperiod-responsive genes in the photoreceptor cells, thereby triggering the seasonal change in the retinal photoreceptor physiology relevant to vision. In the retina, visual sensitivity, retinomotor movement and melatonin synthesis may change with light and circadian time38. In the medaka, the mRNA expression of opsins in the retina changes with the environmental temperature as a seasonal response39. To the best of our knowledge, there have been few reports of the eye controlling photoperiodic responses, either locally or systemically, but our results suggest that the eye might play a role in photoperiodic responses, at least in the eye itself. In this regard, a recent paper reported that the neural network and intercellular synchronization in the mouse suprachiasmatic nucleus (SCN) are dependent upon day length40, which suggests that tissue-specific photoperiodic measurement occurs in neural tissues. Because zCry1ab is a potent inhibitor of E-box/E′-box-mediated circadian gene expression, it is possible that it plays a role in photoreceptor-specific physiological phenomena. In fact, mRNA levels of retinal Opn4 decrease when zebrafish are transferred from 14L:10D to 18L:6D41. Future studies should analyze the downstream molecules of zCry1ab and the upstream regulatory mechanism of Opn4. The in vitro and in vivo functional characterization of zCry1ab and its promoter/enhancer are also required in order to ascertain how and why two peaks occur in the morning and evening.
Methods
Ethics statement
All experiments were conducted in accordance with the guidelines of Waseda University. All protocols were approved by the Committee for the Management of Biological Experiment at Waseda University, and experimental animal care was conducted with permission from the Committee for Animal Experimentation of the School of Science and Engineering, Waseda University (permission # WD15-060; 2015-A028; #2018-A104; #2019-A039).
Animals
Wild-type zebrafish (EkkWill or commercial fish obtained from a local supplier) were maintained in stock tanks at 26–30 °C under 14L:10D long days and white light/dark cycles with lights on at 9:00 am. Fluorescent lamps (FHF32EX-N-HX-S, three-wavelength type, daylight-white, 32 W; Supplementary Fig. S1, black curve) were used as a light source, and the intensity was 5–20 μW cm−2 at water level.
Collection of tissues during the white light, long-day cycle
Zebrafish (EkkWill line) were transferred from the stock tanks to another tank and fasted under the same cycles (14L:10D, lights on at 9:00 am) at 26 °C for 2 days before sampling. The light source was fluorescent lamps (FL20SS-EX-N/18-F, three-wavelength type, daylight-white, 18 W; Supplementary Fig. S1, black dotted curve) with an intensity of 210–240 μW cm−2 at water level. The fish were chilled on ice before sampling to avoid possible RNA synthesis or degradation. The brain, eye, pectoral fin, skin and skeletal muscle were taken at ZT1, ZT7, ZT15, ZT19 and ZT1 (n = 4 at each time point). During the light (ZT1 and ZT7) and dark (ZT15 and ZT19) periods, sampling was conducted under a white fluorescent lamp (240 μW cm−2; Supplementary Fig. S1, black curve) and a dim red light (>650 nm and <80 μW cm−2), respectively. The samples were kept in RNAlater® (Ambion) for 24 h at 4 °C and stored at −80 °C until RNA extraction.
Collection of eyes and brain under blue light conditions
Zebrafish obtained from a local supplier were entrained to a 14L:10D white light/dark cycle (lights on at 9:00 am) in stock tanks over 3 months. Subsequently, they were transferred to another tank the day before sampling. At ZT0 of the sampling day, the fish were exposed to blue light (λmax = 462 nm, λ1/2 = 20 nm; Supplementary Fig. S1, blue curve) or kept in the dark for 14 h. The light intensities were set at 5.586 × 1014 photons cm−2 s−1 which was equivalent to 240 μW cm−2. We took three or four samples at ZT0 (start of the light treatment), ZT1 (1-h light treatment), ZT3 (3-h light treatment), ZT7 (midpoint of the light period in the previous LD cycle), ZT14 (end of the light treatment), ZT15 (1 h after the light treatment), ZT19 (midpoint of the dark period in the previous LD cycle) and ZT24 (start of the light period in the previous LD cycle). At each sample collection, the fish were chilled as described above and their heads were collected under white light (light conditions) or a dim red light (>650 nm) (dark conditions; ZT15, ZT19 and ZT24 for the light-treated fish and all time points for the control fish kept in the dark) and incubated in RNAlater® (Ambion) for 24 h at 4 °C. The eyes and brain were taken from the head and homogenized in TRIzol reagent (Invitrogen).
Collection of eyes under different photoperiods
Zebrafish obtained from a local supplier were entrained to a 14L:10D white light/dark cycle (lights on at 9:00 am) in stock tanks. Subsequently, they were entrained to 18L:6D extra-long-day (ELD), 14L:10D long-day (LD), 10L:14D short-day (SD) or 6L:18D extra-short-day (ESD) cycles under white light/dark cycles for 7 days. They were then transferred to another tank and kept under blue light (Supplementary Fig. S1, blue curve)/dark cycles for each photoperiod (lights on at 9:00 am) at 28 °C for 4 days until sampling. On the day of sampling, we took three to five fish for 13 points from ZT1 at 2-h intervals. At each sampling, the fish were chilled as described above. The eyes were collected and immediately homogenized in TRIzol reagent under white light (light conditions) or a dim red light (>650 nm) (dark conditions).
Collection of eyes after entrainment to LD or SD photoperiod
For the long-day sampling, zebrafish were entrained to a 14L:10D white light/dark cycle (lights on at 9:00 am; for the light conditions, see Animals) for more than 10 days and then transferred to another tank and kept under 14L:10D blue light (240 μW cm−2; Supplementary Fig. S1, blue curve)/dark cycles at 28 °C for 4 days until sampling. For the short-day sampling, zebrafish were entrained to the 10L:14D white light/dark cycle (lights on at 9:00 am; for the light conditions, see Animals) for 10 days and then transferred to another tank and kept under 10L:14D blue light (240 μW cm−2; Supplementary Fig. S1, blue curve)/dark cycles at 28 °C for 4 days until sampling. At ZT0 on the sampling day, the fish were exposed to blue light (light conditions) or kept in the dark (dark conditions). We took three or four samples at each time point. For each sampling, the fish were chilled as described in above and the eyes were collected and treated as described immediately homogenized in TRIzol reagent under white light (light conditions) or a dim red light (>650 nm) (dark conditions).
RNA extraction and cDNA synthesis
Total RNA was extracted using TRIzol reagent (Invitrogen) following the manufacturer’s instructions, then treated with RNase-free Recombinant DNase I (TakaraBio). We synthesized cDNA from 0.4 or 0.5 μg (experiments shown in Figs. 1, 2 and 4) or 1 μg (experiments shown in Figs. 5 and 7) of total RNA using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems).
Quantitative reverse transcription-polymerase chain reaction
For each primer pair (Table 2), we verified the amplification efficiency and accuracy of the target sequence by quantitative reverse transcription-polymerase chain reaction (qRT-PCR) using a dilution series of head cDNA (a mixture of ZT7 and ZT19). qRT-PCR except for the case using LMD samples (see below) was conducted using 2 × Fast SYBR® Green Master Mix (Applied Biosystems). A StepOnePlus Real-Time PCR System (Applied Biosystems) was used according to the manufacturer’s instructions. The relative level of each mRNA was calculated using the 2−ΔΔCT method. Each CT value was the mean of three to five biological replicates.
LMD followed by qRT-PCR
Heads of zebrafish (ZT1, ZT13, and ZT17 under 14L:10D blue light/dark cycle entrainment) were embedded in a 1:2 (vol/vol) mixture of optimal cutting temperature (OCT) embedding compound (Tissue-Tek, Sakura) and 0.1 M sodium phosphate buffer (pH 7.4) without fixation, and frozen using isopentane and liquid nitrogen. Tissues were cut at 20 μm onto membrane slides (PEN-Membrane, 2.0 μm, Leica) using a cryostat (CM1850, Leica). Sections were fixed with 70% ethanol for 30 sec and washed with diethyl pyrocarbonate (DEPC) water for 30 sec. Morphology was exposed with 0.05% toluidine blue (pH 4.1, Wako) for 30 sec and washed with DEPC water for 30 sec for two times and air-dried for 30 minutes. Then the sections were immediately subjected to LMD system (LMD6000, Leica) to collect two regions of the retina; photoreceptor layer (PRL) and inner retinal layer (IRL) including inner nuclear layer, inner plexiform layer and ganglion cell layer. Tissues of each region from three fish were pooled together (45,000 μm2–70,000 μm2).
Total RNA was extracted with the RNeasy Mini kit (Qiagen) and the On-column DNase digestion (RNase-free DNase set, Qiagen). cDNAs were synthesized with High Capacity cDNA Reverse Transcription Kit (Applied Biosystems) according to the manufacturer’s instructions. The total RNA was divided into four aliquots for triplicate RT reactions and one negative control reaction without RTase to confirm no amplification from possibly contaminating residual gDNA.
The PCR reaction mixture was prepared with cDNA and 2x TaqMan Fast Advanced Master Mix (Applied Biosystems), 200 nM TaqMan probe ([FAM] TCTGT TTGTG AGCTT GTCCA ACCAT [TAMRA]) and 300 nM primers for zCry1ab (Table 2). PCR cycling conditions were 50 °C for 2 min, 95 °C for 20 sec and 40 cycles of 95 °C for 1 sec and 60 °C for 20 sec. 18S rRNA was used as internal control (18S_F1, TCGCT AGTTG GCATC GTTTA TG; 18S_R1, CGGAG GTTCG AAGAC GATCA). A StepOnePlus Real-Time PCR System (Applied Biosystems) was used according to the manufacturer’s instructions.
Statistical analyses
Statistical analyses of the qRT-PCR data were performed using R v. 3.3.2 and 3.5.2 (https://cran.r-project.org). We investigated differences in gene expression using one-way and two-way analyses of variance (ANOVA) and Tukey-Kramer post-hoc tests. Results were considered statistically significant if p < 0.05. The acrophases of the daily expression patterns were analyzed by CircWave (v.1.4) (http://www.euclock.org/results/item/circ-wave.html).
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information Files.
References
Ikegami, K. & Yoshimura, T. Seasonal time measurement during reproduction. J. Reprod. Dev. 59, 327–333, https://doi.org/10.1210/en.2014-1741 (2013).
Kobayashi, Y. et al. Molecular analysis of zebrafish photolyase/cryptochrome family: two types of cryptochromes present in zebrafish. Genes Cells 5, 725–738 (2000).
Haug, M. F., Gesemann, M., Lazović, V. & Neuhauss, S. C. F. Eumetazoan cryptochrome phylogeny and evolution. Genome Biol. Evol. 7, 601–619 (2015).
Doria, H. B. et al. Time does matter! Acute copper exposure abolishes rhythmicity of clock gene in Danio rerio. Ecotoxicol. Environ. Saf. 155, 26–36 (2018).
Hamilton, N., Diaz-De-Cerio, N. & Whitmore, D. Impaired light detection of the circadian clock in a zebrafish melanoma model. Cell Cycle 14, 1232–1241 (2015).
Liu, C. et al. Molecular evolution and functional divergence of zebrafish (Danio rerio) cryptochrome genes. Sci. Rep. 5, 8113, https://doi.org/10.1038/srep08113 (2015).
Mracek, P. et al. ERK Signaling regulates light-induced gene expression via D-box enhancers in a differential, wavelength-dependent manner. Plos One 8, 1–10 (2013).
Li, Y., Li, G., Wang, H., Du, J. & Yan, J. Analysis of a gene regulatory cascade mediating circadian rhythm in zebrafish. Plos Comput. Biol. 9, e1002940, https://doi.org/10.1371/journal.pcbi.1002940 (2013).
Farhat, F. P., Martins, C. B., Ribeiro Graciani De Lima, L. H., Isoldi, M. C. & Castrucci, A. M. D. L. Melanopsin and clock genes: Regulation by light and endothelin in the zebrafish Zem-2S cell line. Chronobiol. Int. 26, 1090–1119 (2009).
Sousa, J. C., Magalhães-Marques, K. K., Da Silveira Cruz-Machado, S., Moraes, M. N. & Castrucci, A. M. D. L. Dexamethasone modulates nonvisual opsins, glucocorticoid receptor, and clock genes in Danio rerio ZEM-2S cells. Biomed Res. Int. 2017, 8459385, https://doi.org/10.1155/2017/8459385 (2017).
Weger, B. D. et al. The light responsive transcriptome of the zebrafish: Function and regulation. Plos One 6, e17080, https://doi.org/10.1371/journal.pone.0017080 (2011).
Whitmore, D., Foulkes, N. S. & Sassone-Corsi, P. Light acts directly on organs and cells in culture to set the vertebrate circadian clock. Nature 404, 87–91, https://doi.org/10.1038/35003589 (2000).
Pando, M. P., Pinchak, A. B., Cermakian, N. & Sassone-Corsi, P. A cell-based system that recapitulates the dynamic light-dependent regulation of the vertebrate clock. Proc. Natl. Acad. Sci. USA 98, 10178–10183, https://doi.org/10.1073/pnas.181228598 (2001).
Ishikawa, T., Hirayama, J., Kobayashi, Y. & Todo, T. Zebrafish CRY represses transcription mediated by CLOCK-BMAL heterodimer without inhibiting its binding to DNA. Genes Cells 7, 1073–1086 (2002).
Hirayama, J., Cardone, L., Doi, M. & Sassone-Corsi, P. Common pathways in circadian and cell cycle clocks: light-dependent activation of Fos/AP-1 in zebrafish controls CRY-1a and WEE-1. Proc. Natl. Acad. Sci. USA 102, 10194–10199, https://doi.org/10.1073/pnas.0502610102 (2005).
Okano, K. et al. Light- and circadian-controlled genes respond to a broad light spectrum in Puffer Fish-derived Fugu eye cells. Sci. Rep. 7, 46150, https://doi.org/10.1038/srep46150 (2017).
Kume, K. et al. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98, 193–205 (1999).
Ueda, H. R. et al. System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nat. Genet. 37, 187–192, https://doi.org/10.1038/ng1504 (2005).
Michael, A. K., Fribourgh, J. L., Van Gelder, R. N. & Partch, C. L. Animal cryptochromes: Divergent roles in light perception, circadian timekeeping and beyond. Photochem. Photobiol. 93, 128–140, https://doi.org/10.1111/php.12677 (2017).
Fukushiro, M. et al. Lunar phase-dependent expression of cryptochrome and a photoperiodic mechanism for lunar phase-recognition in a reef fish, goldlined spinefoot. Plos One 6, e28643, https://doi.org/10.1371/journal.pone.0028643 (2011).
Toda, R. et al. Hypothalamic expression and moonlight-independent changes of Cry3 and Per4 implicate their roles in lunar clock oscillators of the lunar-responsive Goldlined spinefoot. Plos One 9, e109119, https://doi.org/10.1371/journal.pone.0109119 (2014).
Takeuchi, T. et al. Moonlight controls lunar-phase-dependency and regular oscillation of clock gene expressions in a lunar-synchronized spawner fish, Goldlined spinefoot. Sci. Rep. 18, 6208, https://doi.org/10.1038/s41598-018-24538-1 (2018).
Kubo, Y., Akiyama, M., Fukada, Y. & Okano., T. Molecular cloning, mRNA expression, and immunocytochemical localization of a putative blue-light photoreceptor CRY4 in the chicken pineal gland. J. Neurochem. 97, 1155–1165, https://doi.org/10.1111/j.1471-4159.2006.03826.x (2006).
Watari, R. et al. Light-dependent structural change of chicken retinal Cryptochrome4. J. Biol. Chem. 287, 42634–42641, https://doi.org/10.1074/jbc.M112.395731 (2012).
Mitsui, H. et al. Overexpression in yeast, photocycle, and in vitro structural change of an avian putative magnetoreceptor cryptochrome4. Biochemistry 54, 1908–1917, https://doi.org/10.1021/bi501441u (2015).
Ozturk, N. et al. Comparative photochemistry of animal type 1 and type 4 cryptochromes. Biochemistry 48, 8585–8593, https://doi.org/10.1021/bi901043s (2009).
Wiltschko, R. & Wiltschko, W. Magnetoreception in birds. J. R. Soc. Interface 16, 20190295, https://doi.org/10.1098/rsif.2019.0295 (2019).
Gau, D. et al. Phosphorylation of CREB Ser142 regulates light-induced phase shifts of the circadian clock. Neuron 34, 245–253 (2002).
Kornhauser, J. M., Nelson, D. E., Mayo, K. E. & Takahashi, J. S. Regulation of jun-B messenger RNA and AP-1 activity by light and a circadian clock. Science 255, 1581–1584 (1992).
Vallone, D., Gondi, S. B., Whitmore, D. & Foulkes, N. S. E-box function in a period gene repressed by light. Proc. Natl. Acad. Sci. USA 101, 4106–4011 (2004).
Vatine, G. et al. Light directs zebrafish period2 expression via conserved D and E boxes. Plos Biol. 7, e1000223, https://doi.org/10.1371/journal.pbio.1000223. (2009).
Mracek, P. et al. Regulation of per and cry genes reveals a central role for the D-box enhancer in light-dependent gene expression. Plos One 7, e51278, https://doi.org/10.1371/journal.pone.0051278 (2012).
Yasuo, S. et al. Photoinducible phase-specific light induction of Cry1 gene in the pars tuberalis of Japanese quail. Endocrinology 145, 1612–1616, https://doi.org/10.1210/en.2003-1285 (2004).
Koger, C. S., Teh, S. J. & Hinton, D. E. Variations of light and temperature regimes and resulting effects on reproductive parameters in Medaka (Oryzias latipes). Bio. Reprod. 61, 1287–1293 (1999).
Engeszer, R. E., Pattersion, A. B., Rao, A. A. & Parichy, D. M. Zebrafish in the wild: A review of natural history and new notes from the field. Zebrafish 4, 20–41, https://doi.org/10.1089/zeb.2006.9997 (2007).
Wayne, N. L. & Rissman, E. F. Tropical photoperiods affect reproductive development in the musk shrew, Suncus murinus. Physiol. Behav. 50, 549–553 (1991).
Hau, M., Wikelski, M. & Wingfield, J. C. A neotropical forest bird can measure the slight changes in tropical photoperiod. Proc. R. Soc. B 265, 89–95 (1998).
Besharse, J. C. & McMahon, D. G. The retina and other light-sensitive ocular clocks. J. Biol. Rhythms. 31, 223–243, https://doi.org/10.1177/0748730416642657 (2016).
Shimmura, T. et al. Dynamic plasticity in phototransduction regulates seasonal changes in color perception. Nat. Commun. 8, 412, https://doi.org/10.1038/s41467-017-00432-8 (2017).
Myung, J. et al. GABA-mediated repulsive coupling between circadian clock neurons in the SCN encodes seasonal time. Proc. Natl. Acad. Sci. USA 112, E3920–3929, https://doi.org/10.1073/pnas.1421200112 (2015).
Matos-Cruz, V. et al. Unexpected diversity and photoperiod dependence of the zebrafish melanopsin system. Plos One 6, e25111, https://doi.org/10.1371/journal.pone.0025111 (2011).
McCurley, A. T. & Callard, G. V. Characterization of housekeeping genes in zebrafish: male-female differences and effects of tissue type, developmental stage and chemical treatment. B.M.C. Mol. Biol. 9, 102, https://doi.org/10.1186/1471-2199-9-102 (2008).
Acknowledgements
This work was supported by the Japanese Society for the Promotion of Science (JSPS, Nos. 23248033, 24657109, 26650024, 17H03710) of Japan awarded to T.O.
Author information
Authors and Affiliations
Contributions
K.O., Y. Saratani, A.T., Y. Shoji and R.T. treated the zebrafish. K.O., Y. Saratani, A.T. and R.T. performed the qRT-PCRs. K.O., Y. Saratani, Y. Shoji and T.O. performed the data analysis. K.O. and Y. Saratani performed laser microdissection and K.O., A.T. and T.O. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okano, K., Saratani, Y., Tamasawa, A. et al. A photoperiodic time measurement served by the biphasic expression of Cryptochrome1ab in the zebrafish eye. Sci Rep 10, 5056 (2020). https://doi.org/10.1038/s41598-020-61877-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61877-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.