Abstract
This study sought to evaluate the association of 28 single nucleotide polymorphisms (SNPs) within NFKB and inflammasome pathway genes with the risk of rheumatoid arthritis (RA) and response to TNF inhibitors (TNFi). We conducted a case-control study in a European population of 1194 RA patients and 1328 healthy controls. The association of potentially interesting markers was validated with data from the DANBIO (695 RA patients and 978 healthy controls) and DREAM (882 RA patients) registries. The meta-analysis of our data with those from the DANBIO registry confirmed that anti-citrullinated protein antibodies (ACPA)-positive subjects carrying the NFKB2rs11574851T allele had a significantly increased risk of developing RA (PMeta_ACPA + = 0.0006) whereas no significant effect was found in ACPA-negative individuals (PMeta_ACPA− = 0.35). An ACPA-stratified haplotype analysis including both cohorts (n = 4210) confirmed that ACPA-positive subjects carrying the NFKB2TT haplotype had an increased risk of RA (OR = 1.39, P = 0.0042) whereas no effect was found in ACPA-negative subjects (OR = 1.04, P = 0.82). The meta-analysis of our data with those from the DANBIO and DREAM registries also revealed a suggestive association of the NFKB2rs1056890 SNP with larger changes in DAS28 (OR = 1.18, P = 0.007). Functional experiments showed that peripheral blood mononuclear cells from carriers of the NFKB2rs1005044C allele (in LD with the rs1056890, r2 = 1.00) showed increased production of IL10 after stimulation with LPS (P = 0.0026). These results provide first evidence of a role of the NFKB2 locus in modulating the risk of RA in an ACPA-dependent manner and suggest its implication in determining the response to TNFi. Additional studies are now warranted to further validate these findings.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is a chronic inflammatory disease more frequently diagnosed in females than males, that has a prevalence of about 0.5–1%1. RA perpetuates and amplifies itself through a wide number of molecular mechanisms involving several immune cell types and multiple inflammatory mediators that are released from the damaged tissue2. Although the complexity of inflammatory pathways implicated in RA development and progression remains in part unknown, there are convincing evidences supporting the view that NFKB pathway and its connection with the NLRP3-inflammasome plays a pivotal role in the modulation of the expression of multiple inflammatory genes implicated in RA development3 and drug response or disease progression4.
Activated NFKB has been detected in the synovium of RA patients at both early and late stages of joint inflammation5,6,7,8 and once NFKB is activated (for instance, through the interaction of antigen presenting cells and T cells), it triggers two major signaling pathways in the implicated cells: the canonical and the non-canonical NFKB pathway. Whereas the canonical pathway regulates the activation of NFKB1 p50, RELA and c-REL and leads to rapid but transient NFKB activation, the non-canonical NFKB pathway selectively activates p100-sequestered NFKB members (predominantly NFKB2 p52 and RELB) and produces a long-lasting signaling. Even though a cross-talk between the canonical and non-canonical NFKB pathways has been previously reported, the activation of the canonical NFKB pathway is generally associated with inflammation whereas the induction of the non-canonical NFKB pathway was linked to development processes9. In RA, it is well known that the acute activation of the canonical pathway on antigen presenting cells and T cells quickly leads to the production of a wide range of essential proinflammatory mediators including cytokines (TNFα, IL1α, IL1β, IL1RA, IL2, IL12p40 and IFNγ), chemokines (IL8, CXCL11), immunoreceptors (CD80, CD23, CD48, CD69, IL2R, TNFRs, and CCR5), cell adhesion molecules (ELAM-1, ICAM-1, VCAM-1 and P-selectin) and growth factors (GM-CSF, IGFBP2, and PDGFB) that often facilitate synovial hyperplasia by promoting cell proliferation and apoptosis inhibition of RA fibroblast-like synovial cells10. On the contrary, the activation of the non-canonical pathway involves a slow build-up of long-lasting signals that have been implicated in developmental processes including B-cell development11, secondary lymphoid organ development12,13 and osteoclast differentiation14 but also development of myeloid-related CD4+CD8α− dendritic cells and macrophages15, key players in modulating immune responses in RA.
Besides the role of NFKB in the inflammatory process, recent evidences have shown that the NLRP3-inflammasome is a cytosolic multiprotein complex highly expressed in peripheral blood mononuclear cells of RA patients and in the synovial tissues of osteoarthritis patients. The NLRP3 inflammasome is capable of alerting immune system to the presence of tissue damage and to induce the processing of the IL1β, IL18 and IL33 pro-cytokines into biologically active proinflammatory mediators that drive cartilage destruction16. In addition, it has been reported that the presence of mutations in NLRP3-inflammasome-related proteins (CARD8 and NLRP3) predispose to RA17,18 and that genetic variation in this pathway might also modulate inflammatory activity in early stages of the disease and thereby affect disease progression17,18.
Considering the aspects detailed above, but also previous studies suggesting that the NFKB- and NLRP3-inflammasome pathways are genetically determined19, we decided to conduct a case-control study to investigate whether 28 single nucleotide polymorphisms (SNPs) within the NFKB1, NFKB2, NFKBIB, IKBKB, GBP6, IRF4, NLRP3, REL, RELA, KLRC1, KLRK1 | KLRC4, LOC105376246 (ncRNA), TLR4, TLR5, TLR9, TLR10 and TRAF1 | C5 genes influence the risk of developing RA and the response to TNF inhibitors (TNFi). In addition, we investigated the correlation of selected SNPs with steroid hormone levels and their role in modulating immune responses after stimulation of whole blood, peripheral mononuclear cells (PBMCs) and macrophages with lipopolysaccharide (LPS), phytohemagglutinin (PHA) and Pam3Cys.
Material and Methods
Discovery population
The discovery population consisted of 1194 RA patients and 1328 healthy controls ascertained through the REPAIR consortium (Table 1). RA patients fulfilled the 1987 revised American College of Rheumatology (ACR)20 and the ACR/EULAR 2010 classification criteria21. The study followed the Declaration of Helsinki. Study participants were of European origin and gave their written informed consent to participate in the study, which was approved by the ethical review committee of participant institutions. The Ethics committee of each participant institution approved the study protocol: Virgen de las Nieves University Hospital (2012/89); Santa Maria Hospital-CHLN (CE 877/121.2012); University Clinical Hospital of Santiago de Compostela (2013/156). A detailed description of the discovery population has been reported elsewhere22,23,24.
Response to anti-TNF medications
Six hundred and four RA patients treated with TNFi (adalimumab, etanercept, infliximab, golimumab or certolizumab) were included in the drug response analysis of the discovery population. The change in disease activity score (DAS28) at baseline and at 6 months of treatment with TNFi was calculated for each patient. Linear regression analysis adjusted for age and sex was used to determine the association between selected SNPs and changes in DAS28. Subjects with missing values for DAS28 in any of these time points were not included in the analysis.
SNP selection and genotyping
NFKB- and inflammasome-related polymorphisms were selected on the basis of their potential functionality and linkage disequilibrium (LD) but also because of existing studies reporting their significant association with the risk of developing autoimmune and immune-related diseases or response to TNFi25,26,27,28,29. This strategy resulted in the selection of 28 genetic variants within the GBP6, IKBKB, IRF4, KLRC1, KLRK1, NFKB1, NFKB2, NFKBIB, NLRP3, REL, RELA, RELB, TLR4, TLR5, TLR9, TLR10 and TRAF1/C5 loci that were genotyped in the discovery population (Table 2). Genomic DNA was extracted from peripheral blood using the Qiagen Mini Kit (Qiagen, CA, USA) or from saliva using standard procedures. Genotyping was carried out using KASPar® assays (LGC Genomics, London, UK) in a 384-well plate format (Applied Biosystems, CA, USA) according to manufacturer’s instructions. Five percent of samples were included as duplicates to ensure high-quality genotyping.
Statistical analysis
The Hardy-Weinberg Equilibrium (HWE) test was performed in the control group by a standard observed-expected chi-square (χ2). Logistic and linear regression analyses adjusted for age, sex and country of origin were used to assess the main effects of the selected SNPs on RA risk and the response to TNFi respectively. Statistical power was estimated using Quanto software (http://hydra.usc.edu/gxe/). Correction for multiple testing was performed using the Meff method for SNPs genotyped across all populations30. The threshold used for the risk and drug response analyses was 0.0008 ([0.05/22 independent markersx3 inheritance models).
Linkage disequilibrium (LD) and haplotype analysis
We performed haplotype frequency estimation and haplotype association analysis adjusted for age, sex and country of origin using SNPstats31 and haplo.stats package in STATA. Haplotype frequencies were determined using the Expectation-maximization (EM) algorithm. Haplotypes were reconstructed using SNPtool and Haploview and block structures were determined according to the method of Gabriel et al.32.
Replication populations and meta-analyses for RA risk and drug response
For replication purposes, we genotyped the most promising SNPs associated with RA risk in a cohort of 695 Danish RA patients and 978 healthy controls33. Clinical data from RA patients were collected through the DANBIO registry (The National Danish Registry for Biological Treatment of Rheumatic Diseases)34 and DNA samples were obtained from peripheral blood collected at the Statens Serum Institut (Copenhagen, Denmark), which routinely perform screening for tuberculosis before treatment with biological treatments. Healthy blood donors were recruited in Viborg and Sønderborg (Denmark). In order to replicate the most interesting associations with response to TNFi, we also used genetic data from a genome-wide association study (GWAS) on drug response conducted in 882 Dutch RA patients from the DREAM (Dutch RhEumatoid Arthritis Monitoring) registry. Imputed SNPs reporting potentially interesting overall or ACPA-specific associations with RA risk or drug response were genotyped in a subset of 708 patients. To further validate our results, we also genotyped the most interesting markers associated with drug response in 555 RA patients from the DANBIO registry that were treated with TNFi. A total of 2107 patients were treated with anti-TNF. Demographic and clinical details of the 3 cohorts are included in Supplementary Table 1. The study was approved by the Institutional review board of the Radboud university medical centre and by the Regional Ethics Committee of Central Denmark Region (M-20100153 and S-20120113). All patients provided written informed consent and clinical information was prospectively gathered from the medical records.
To test for genetic association, we conducted a meta-analysis of the discovery data with those from the 2 European registries and the I2 statistic was used to assess statistical heterogeneity between studies. The pooled OR was computed using the random-effect model.
Functional analysis of the NFKB and inflammasome-related variants
Cytokine stimulation experiments were conducted in the 500 Functional Genomics (500FG) cohort from the Human Functional Genomics Project (HFGP; http://www.humanfunctionalgenomics.org/), which was designed to determine the influence of genomic variation on the variability of immune responses. The HFGP study was approved by the Arnhem-Nijmegen Ethical Committee (no. 42561.091.12) and biological specimens (venous blood) were collected after informed consent was obtained. We assessed whether any of the 28 NFKB and inflammasome-related SNPs correlated with cytokine levels (TNFα, IFNγ, IL1β, IL1RA, IL6, IL8, IL10, IL17, and IL22) after the stimulation of whole blood, peripheral blood mononuclear cells (PBMCs) or monocyte-derived macrophages from 408 healthy subjects with LPS (1 or 100 ng/ml), PHA (10 μg/ml), and Pam3Cys (10 μg/ml). After log transformation, linear regression analyses adjusted for age and sex were used to determine the correlation of selected SNPs with cytokine expression quantitative trait loci (cQTLs). All analyses were performed using R software (http://www.r-project.org/). In order to account for multiple comparisons, we used a significant threshold of 0.00025, i.e. 0.05/(22 independent SNPs × 9 cytokines).
Details on PBMCs isolation, macrophage differentiation and stimulation assays have been reported elsewhere35,36,37. Briefly, PBMCs were washed twice in saline and suspended in medium (RPMI 1640) supplemented with gentamicin (10 mg/mL), L-glutamine (10 mM) and pyruvate (10 mM). PBMC stimulations were performed with 5 × 105 cells/well in round-bottom 96-wells plates (Greiner) for 24 hours in the presence of 10% human pool serum at 37 °C and 5% CO2. Supernatants were collected and stored in −20 °C until used for ELISA. LPS (100 ng/ml), PHA (10 μg/ml) and Pam3Cys (10 μg/ml) were used as stimulators for 24 or 48 hours. Whole blood stimulation experiments were conducted using 100 μl of heparin blood that was added to a 48 well plate and subsequently stimulated with 400 μl of LPS and PHA (final volume 500 ul) for 48 hours at 37 °C and 5% CO2. Supernatants were collected and stored in −20 °C until used for ELISA. Concentrations of human TNFα, IFNγ, IL1β, IL1RA, IL6, IL8, IL10, IL17, and IL22 were determined using specific commercial ELISA kits (PeliKine Compact, Amsterdam, or R&D Systems), in accordance with the manufacturer’s instructions.
Once we examined the correlation of NFKB and inflammasome-related polymorphisms with cytokine levels in our functional experiments, we also used the HaploReg SNP annotation tool (http://www.broadinstitute.org/mammals/haploreg/haploreg.php) to further investigate the functional consequences of each specific variant. We also assessed whether any of the potentially interesting markers correlated with mRNA expression levels of their respective genes using data from GTex portal (www.gtexportal.org/home/).
Correlation between steroid hormone levels and NFKB- and inflammasome-related SNPs
We also measured serum levels of seven steroid hormones (androstenedione, cortisol, 11-deoxy-cortisol, 17-hydroxy progesterone, progesterone, testosterone and 25 hydroxy vitamin D3) in the 500FG cohort, which includes 531 healthy subjects. Steroid hormones were analyzed by Liquid Chromatography Tandem-Mass Spectrometry (LCMSMS) after protein precipitation and solid-phase extraction as described in Ter Horst et al.37 (see also Supplementary Material). Hormone levels and genotyping data were available for a total of 406 subjects.
After log-transform, correlation between steroid hormone levels and NFKB- and inflammasome-related SNPs was evaluated by linear regression analysis adjusted for age and sex. In order to avoid a possible bias, we excluded those subjects that were using oral contraceptives or those subjects in which this information was not known from the analysis. A total of 379 healthy subjects (107 women and 272 men) were finally available for analysis. A Bonferroni significance threshold was set to 0.00033 considering the number of independent SNPs tested (n = 22) and the number of hormones determined (n = 7).
Results
This study was conducted in a discovery population comprised of 1194 RA patients and 1328 healthy controls. RA patients were slightly older than controls (59.22 ± 12.97 vs. 52.67 ± 8.99) and showed an increased female/male ratio compared to healthy controls (4.10 [959/234] vs. 1.39 [773/555]. Sixty percent of the RA patients presented positive values of anti-citrullinated protein antibodies (ACPA) and the median disease duration was of 17.60 years and the disease activity score 28 (DAS28) calculated at patient recruitment was of 5.74 (Table 1).
Association of selected SNPs with RA risk
All SNPs were in Hardy-Weinberg equilibrium in the control group (P > 0.001). Logistic regression analysis adjusted for age, sex and country of origin showed that carriers of the NLRP3rs4612666T allele or the IRF4rs1050975A/A and NFKB2rs12769316T/T genotypes had an increased risk of developing RA at nominal level of P ≤ 0.05 (ORDom = 1.25, 95%CI 1.05–1.49, P = 0.013; ORRec = 1.30, 95%CI 1.04–1.62, P = 0.019; and ORRec = 1.70, 95%CI 1.04–2.78, P = 0.034; Table 3). Interestingly, an ACPA-stratified analysis revealed that ACPA-positive subjects carrying the NFKB1rs4648110A/A genotype or the NFKB2rs11574851T allele had a significantly increased risk of developing RA whereas a non-significant effect was found in ACPA-negative patients (ORRec-ACPA+ = 1.65, 95%CI 1.04–2.63, P = 0.031 vs. ORRec-ACPA− = 0.86, 95%CI 0.39–1.90, P = 0.90 and per-allele ORACPA+ = 1.39, 95%CI 1.06–1.83, P = 0.017 and per-allele ORACPA− = 1.02, 95%CI 0.68–1.52, P = 0.93; Table 3). On the other hand, we found that seronegative subjects carrying the KLRCrs7301582T or KLRK1rs1049174C alleles showed a significantly increased risk of developing RA whereas no effect was detected in ACPA-positive individuals (ORDom-ACPA− = 1.56, 95%CI 1.18–2.09, P = 0.003 vs. ORDom-ACPA+ = 1.05, 95%CI 0.84–1.30, P = 0.67 and ORDom-ACPA− = 1.38, 95%CI 1.03–1.84, P = 0.031 vs. ORDom-ACPA+ = 1.09, 95%CI 0.88–1.35, P = 0.42).
Although none of the above-reported associations survived after correction for multiple testing, we attempted to replicate them through meta-analysis of the discovery data with those from the DANBIO registry. The meta-analysis of these two populations, which included 4194 subjects (1888 RA patients and 2306 healthy controls), confirmed that carriers of the NFKB2rs12769316T/T genotype had an increased risk of developing RA when compared with those carrying the C allele (ORMeta = 1.78, 95%CI 1.21–2.63, P = 0.0037, I2 = 0.0%, PHet = 0.76; Supplementary Table 2). In addition, although the association was only significant at nominal level (P < 0.05), we also found that carriers of the NFKB2rs11574851T allele also had an increased risk of developing RA (ORMeta = 1.29, 95%CI 1.02–1.64, P = 0.035, PHet = 0.27). Given that no population stratification was detected (Supplementary Table 3), these findings suggested that the effect attributed to the NFKB2 locus on the risk of RA was likely true and might depend on a specific haplotype rather than single SNPs. Following this hypothesis, we performed an overall haplotype analysis that revealed that carriers of the NFKB2TC haplotype (including the NFKB2rs11574851T allele) had a significantly increased risk of developing RA (OR = 2.21, 95%CI 1.37–3.56, P = 0.0011). Although this association did not survive multiple testing correction, it pointed to a role of the NFKB2rs11574851 SNP to confer risk to RA development.
Most importantly, an ACPA-stratified meta-analysis of our data with those from the DANBIO registry also revealed that each copy of the NFKB2rs11574851T allele conferred an additive risk of developing RA in ACPA-positive subjects (ORMeta = 1.48, 95%CI 1.18–1.86, P = 0.0006) that was not detected in ACPA-negative individuals (Table 4 and Fig. 1). Of note, the association of the NFKB2rs11574851 SNP with an increased risk of RA remained significant after correction for multiple testing and the direction of the effect was consistent with no significant heterogeneity between cohorts (PHet = 0.40; Fig. 1). The ACPA-stratified meta-analysis of both populations also showed an increased risk of RA in ACPA-positive and ACPA-negative subjects carrying the NFKB2rs12769316T/T genotype (P = 0.013 and P = 0.004; Table 4 and Supplementary Table 4). Even though none of the associations of the NFKB2rs12769316T/T genotype with RA remained significant after correction for multiple testing, these findings supported the notion of a relevant role of the NFKB2 locus in modulating the RA risk. In order to further confirm this hypothesis, we decided to evaluate whether there was an ACPA-specific haplotype that could influence the risk of developing RA. Interestingly, the ACPA-stratified haplotype analysis including both the discovery and DANBIO cohorts also confirmed that ACPA-positive subjects carrying the NFKB2TT haplotype (including the NFKB2rs11574851T risk allele) had a significantly increased risk of RA (ORHaplotype-ACPA+ = 1.39, 95%CI 1.11–1.74, P = 0.0042) whereas no effect was detected in ACPA-negative individuals (ORHaplotype-ACPA− = 1.04, 95%CI 0.75–1.44, P = 0.82; Table 5). These results again pointed to an ACPA-specific effect of the NFKB2 locus to modulate the risk of RA. No additional overall or ACPA-specific associations were confirmed in the meta-analysis of both cohorts.
On the basis of the effect found for the NFKB2rs11574851 or NFKB2rs12769316 SNPs on the risk of developing RA, we decided to analyse whether these SNPs might exert their biological function directly through the modulation of NFKB2-mediated immune responses or indirectly through the regulation of steroid hormone levels. To do that we evaluated if there were any correlation between the NFKB2rs11574851 and NFKB2rs12769316 SNPs and levels of 9 cytokines (TNFα, IFNγ, IL1β, IL1RA, IL6, IL8, IL10, IL17, and IL22) after stimulation of whole blood, PBMCs or macrophages with LPS, PHA or Pam3Cys in a cohort of 408 healthy subjects. Although our functional experiments were well powered, we did not find any significant correlation between the NFKB2rs11574851 and NFKB2rs12769316 SNPs and cytokine or steroid hormone levels (data not shown). Although these results might suggest no impact of the NFKB2 variants in modulating immune responses, it is important to mention that we could not evaluate whether the effect of the NFKB2rs11574851 and NFKB2rs12769316 SNPs on the modulation of immune responses could be dependent on ACPA status as the genetic analyses indicate.
Association of selected SNPs with the response to anti-TNF drugs
When we evaluated the effect of any of the selected SNPs on the response to TNFi (defined as a change in DAS28 after 6 months of treatment), we found a significant effect of the NFKB2rs1056890 SNP to modulate the response to TNFi at nominal level (P < 0.05). Thus, each copy of the NFkB2rs1056890T allele additively increased the drop in DAS28 by 22% after the treatment with TNFi (per-allele OR = 1.22, 95%CI 1.03–1.44, P = 0.025; Table 6). Importantly, when we attempted to replicate this association through a well-powered meta-analysis of our data from the discovery population with those from the DREAM and DANBIO registries (n = 2107), we could confirm that carriers of the NFKB2rs1056890T allele showed a significantly higher improvement in DAS28 after treatment with TNFi (ORMeta = 1.18, 95%CI 1.05–1.33, P = 0.0077, I2 = 51.7%, PHet = 0.13; Fig. 2A). Although this association did not remain significant after correction for multiple testing and therefore need to be further validated, this finding suggested that the NFKB2rs1056890 SNP might modulate the response to anti-TNF drugs through the regulation of the NFKB2-related immune responses.
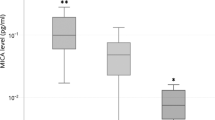
Meta-analysis of the association of the NFKB2rs1056890 SNP with response to TNFi [A] and correlation with higher levels of IL10 after stimulation of PBMCs (n = 377) with LPS [B]. [A] Association estimates according to a random effect model. PMeta = 0.0077. [B] Correlation with IL10 was analysed using genotype data of the NFKB2rs1005044 SNP, a marker in strong LD with the rs1056890 (r2 = 1.00).
In order to test this hypothesis, we assessed whether the NFKB2rs1056890 SNP was associated with cytokine and steroid hormone levels in the HFGP cohort. Although this SNP was not included in the genome-wide association data available from the HFGP cohort, we could evaluate the association of this marker with cytokine and steroid hormone levels through the analysis of neighbouring SNPs in strong LD with it. Our stimulation experiments showed that PBMCs from carriers of the NFKB2rs1005044C allele (in complete LD with the rs1056890T allele, r2 = 1.00) showed an increased production of IL10 after stimulation of PBMCs with LPS for 24 h (P = 0.0025; Fig. 2B). The analysis of additional neighbouring SNPs belonging to the same LD block allowed us to confirm the association of the rs1056890T allele with increased levels of IL10 (Supplementary Table 5). Although the association of the NFKB2rs1056890 SNP with a better response to TNFi and its correlation with higher levels of IL10 did not remain statistically significant after correction for multiple testing, altogether these findings point to a role of this marker in determining the response to TNFi likely through the modulation of IL10-mediated immune responses. No significant association of the NFKB2rs1056890 SNP with response to TNFi was observed when association analysis was stratified by ACPA, which dismissed the implication of ACPA in the functional effect attributed to this polymorphism. We did not find correlation of any of the NFKB2 SNPs with steroid hormone levels (data not shown), which also ruled out the implication of steroid hormones in the modulation of the IL10-mediated immune responses.
Discussion
Our data provided, for the first time, evidence that NFKB2 locus might modulate the risk of RA. The meta-analysis of the data obtained in the discovery population with those from the DANBIO cohort showed a potentially interesting overall association of the NFKB2rs11574851 SNP with the risk of RA that was further confirmed in an overall haplotype analysis. Most importantly, we found that the effect attributed to the NFKB2 locus on RA risk depended on the ACPA status. An ACPA-stratified meta-analysis of the discovery and DANBIO populations including 3712 subjects revealed that ACPA-positive subjects carrying the NFKB2rs11574851T allele had a significantly increased risk of developing RA whereas no effect was detected in ACPA-negative individuals. Of note, the association of the NFKB2rs11574851T allele with an increased risk of RA in ACPA-positive subjects remained significant even after correction for multiple testing and was further confirmed in an ACPA-stratified haplotype analysis that showed that the presence of the NFKB2rs11574851T allele was driving the effect of the NFKB2TA haplotype on the risk of RA in ACPA positive subjects but not in ACPA-negative individuals.
The NFKB2 gene is located on chromosome 10q24 and it encodes for a subunit of the NFKB complex (p100/p52) that is expressed in multiple immune cells and modulates the inflammation. Other important processes involved in the RA pathology such as Th1 immune responses, activation, abnormal apoptosis and osteoclast differentiation and proliferation10 are also impacted. It is broadly known that RA arises as a consequence of the interaction between genetic and environmental factors and that the NFKB pathway plays a central role in determining the onset of the disease and its progression. In addition, it has been reported that the genetic and environmental factors that predispose to RA development are substantially different between ACPA-positive and ACPA-negative subjects. Recent studies have demonstrated, for instance, that the effect attributed to the two major genetic risk factors for RA (shared epitope of the HLADRB1 and a SNP on the PTPN22 gene) is clearly dependent on the ACPA status having a more evident effect in ACPA-positive subjects than in those lacking of these antibodies38. Furthermore, recent GWAS studies have reported the existence of a completely different genetic component or even a gene-smoking interaction pattern between ACPA-positive and ACPA-negative patients, again suggesting a relevant role of ACPA in determining the onset of the disease39,40. However, up to now, little is known about the effect of ACPA on the control of the NFKB pathway. Interestingly, recent investigations have demonstrated that the treatment of PBMCs-derived macrophages with ACPA induced the activation of the NFKB pathway and subsequently the induction of the NLRP3-inflammasome and the production of pro-inflammatory cytokines41. Mechanistically, it was demonstrated that ACPA induces the activation of the NFKB pathway through the induction of the interaction between CD147 and integrin β1 or ATGB1, which in turn activates the downstream Akt/NFKB signalling pathway, resulting in the upregulation of NLRP3 and pro-IL-1β expression and further NLRP3 inflammasome activation41. Considering these interesting findings, we decided to assess in the HFGP cohort if there was any correlation between the NFKB2 SNPs and pro- and anti-inflammatory cytokine production after stimulation of whole blood, PBMCs or monocyte-derived macrophages with LPS, PHA or Pam3Cys. We also analysed whether NFKB2 variants could indirectly affect immune responses through the modulation of steroid hormone levels. Despite the use of a large cohort of healthy subjects from the HFGP cohort, we could not find any significant correlation between the NFKB2rs11574851 and NFKB2rs12769316 SNPs and cytokine or steroid hormone levels. Although these results suggested that these variants might not exert their effect on RA risk through the modulation of NFKB2- or steroid hormone-mediated immune responses, we could not rule out the possibility of a true effect of these variants on the immune response as their effect might depend on the presence of ACPA (as suggested by our genetic data) or even specific haplotypes. In line with this hypothesis, in silico analysis using Haploreg data showed that the NFKB2rs11574851 and NFKB2rs12769316 SNPs mapped among histone marks in multiple primary T helper naïve and memory cells and primary B cells from peripheral blood and they were predicted to act as enhancers in T helper memory cells and to change motifs for Po6fu1, AP-4, CEBPB, Mef2 and RP58. Even though these data supported the idea of a role of NFKB2 variants in modulating immune responses, we think that additional experiments are still needed to determine whether ACPA or specific haplotypes are factors involved in modulating the effect of the NFKB2 locus on the risk of RA.
Besides the role of the NFKB2 locus in determining the risk of RA, this study also showed a noticeable impact of the NFKB2 gene in the modulation of the response to TNFi. In particular, the meta-analysis of the discovery population with data from the DREAM and DANBIO registries, including 2107 RA patients, showed that carriers of the NFKB2rs1056890T allele had an improvement in DAS28 after treatment with TNFi. We found that the direction of the effect of the NFKB2rs1056890 SNP on drug response was consistent across populations and that the effect was statistically significant in 2 of the 3 populations analysed. Although at this point it tempting to speculate that this SNP constitutes a biomarker for good response to TNFi in RA patients that might help to design more individualized treatment strategies, the association did not remain significant after correction for multiple testing and, therefore, need to be confirmed in independent populations. Mechanistically, we found that the presence of neighbouring genetic markers in strong LD with the NFKB2rs1056890 SNP were associated with increased levels of IL10, suggesting that the NFKB2 locus might be implicated in modulating IL10-mediated immune responses. Although the association of the NFKB2rs1056890 SNP with IL10 levels neither survive correction for multiple testing, our results were in agreement with previous studies demonstrating that NFKB2 unlikely NFKB1 is implicated in the control of antigen presenting cell function and not in the activation of T and B cells. Likewise, recent studies have also identified genetic polymorphisms within the NFKB pathway as genetic biomarkers for response to TNFi in RA42 but also other autoimmune diseases42, which further supported our hypothesis suggesting a key role of the NFKB2 gene in modulating the response to TNFi. In addition, in silico tools such as Regulome showed that the rs1056890 SNP has a score of 4, which means that this polymorphism could affect transcription factor affinity and DNase peak43. Using haploreg it was also suggested that the NFKB2rs1056890 SNP might play a role in modulating immune responses as it mapped among histone marks in primary T helper naïve and T helper memory cells, T regulatory and primary NK cells and it was predicted to alter binding motifs for NRSF, Sin3Ak-20 and PLAG1. These transcription factors have been implicated in bone-related diseases44 and their activation results in up-regulation of multiple target genes including immune-related genes such as macrophage colony stimulator factor (MCSF) and insulin growth factor (IGF)-2.
Conclusions
In conclusion, this study reports, for the first time, a consistent association of the NFKB2rs11574851 polymorphism and NFKB2TT haplotype with an increased risk of developing RA in ACPA-positive subjects. In addition, this study suggests a possible role of the NFKB2 locus in the modulation of the response to TNFi. Mechanistically, the functional experiments in the 500FG cohort suggested that the effect attributed to the NFKB2 gene in the modulation of the response to TNFi might be mediated by IL10-mediated immune responses. However, additional studies are still warranted to shed light into the biological processes that link NFKB2 SNPs and RA risk and drug response.
Data availability
All data used in this project have been meticulously cataloged and archived in the BBMRI-NL data infrastructure (https://hfgp.bbmri.nl/) using the MOLGENIS open source platform for scientific data45. This allows flexible data querying and download, including sufficiently rich metadata and interfaces for machine processing (R statistics, REST API) and using FAIR principles to optimize Findability, Accessibility, Interoperability and Reusability46. Genetic data from the discovery and DANBIO populations can be accessed at ftp.genyo.es and data from the DREAM registry are available at https://www.synapse.org/#!Synapse:syn3280809/wiki/194735 and https://www.synapse.org/#!Synapse:syn3280809/wiki/194736.
References
Silman, A. J. & Pearson, J. E. Epidemiology and genetics of rheumatoid arthritis. Arthritis Res. 4(Suppl 3), S265–272 (2002).
McInnes, I. B. & Schett, G. The pathogenesis of rheumatoid arthritis. N. Engl. J. Med. 365, 2205–2219 (2011).
Barnes, P. J. & Karin, M. Nuclear factor-kappaB: a pivotal transcription factor in chronic inflammatory diseases. N. Engl. J. Med. 336, 1066–1071 (1997).
Han, Z., Boyle, D. L., Manning, A. M. & Firestein, G. S. AP-1 and NF-kappaB regulation in rheumatoid arthritis and murine collagen-induced arthritis. Autoimmunity 28, 197–208 (1998).
Asahara, H., Asanuma, M., Ogawa, N., Nishibayashi, S. & Inoue, H. High DNA-binding activity of transcription factor NF-kappa B in synovial membranes of patients with rheumatoid arthritis. Biochem. Mol. Biol. Int. 37, 827–832 (1995).
Marok, R. et al. Activation of the transcription factor nuclear factor-kappaB in human inflamed synovial tissue. Arthritis rheumatism 39, 583–591 (1996).
Gilston, V. et al. NF-kappa B activation in human knee-joint synovial tissue during the early stage of joint inflammation. Biochemical Soc. Trans. 25, 518S (1997).
Miyazawa, K., Mori, A., Yamamoto, K. & Okudaira, H. Constitutive transcription of the human interleukin-6 gene by rheumatoid synoviocytes: spontaneous activation of NF-kappaB and CBF1. Am. J. Pathol. 152, 793–803 (1998).
Pomerantz, J. L. & Baltimore, D. Two pathways to NF-kappaB. Mol. Cell 10, 693–695 (2002).
Makarov, S. S. NF-kappa B in rheumatoid arthritis: a pivotal regulator of inflammation, hyperplasia, and tissue destruction. Arthritis Res. 3, 200–206 (2001).
Weih, D. S., Yilmaz, Z. B. & Weih, F. Essential role of RelB in germinal center and marginal zone formation and proper expression of homing chemokines. J. immunology 167, 1909–1919 (2001).
Dejardin, E. et al. The lymphotoxin-beta receptor induces different patterns of gene expression via two NF-kappaB pathways. Immun. 17, 525–535 (2002).
Basak, S., Shih, V. F. & Hoffmann, A. Generation and activation of multiple dimeric transcription factors within the NF-kappaB signaling system. Mol. Cell. Biol. 28, 3139–3150 (2008).
Vaira, S. et al. RelB is the NF-kappaB subunit downstream of NIK responsible for osteoclast differentiation. Proc. Natl Acad. Sci. U S Am. 105, 3897–3902 (2008).
Wu, L. et al. RelB is essential for the development of myeloid-related CD8alpha- dendritic cells but not of lymphoid-related CD8alpha+ dendritic cells. Immun. 9, 839–847 (1998).
Guo, C. et al. NLRP3 inflammasome activation contributes to the pathogenesis of rheumatoid arthritis. Clin. Exp. immunology 194, 231–243 (2018).
Kastbom, A., Johansson, M., Verma, D., Soderkvist, P. & Rantapaa-Dahlqvist, S. CARD8 p.C10X polymorphism is associated with inflammatory activity in early rheumatoid arthritis. Ann. rheumatic Dis. 69, 723–726 (2010).
Kastbom, A. et al. Genetic variation in proteins of the cryopyrin inflammasome influences susceptibility and severity of rheumatoid arthritis (the Swedish TIRA project). Rheumatol. 47, 415–417 (2008).
Verma, D. et al. The Q705K polymorphism in NLRP3 is a gain-of-function alteration leading to excessive interleukin-1beta and IL-18 production. PLoS one 7, e34977 (2012).
Arnett, F. C. et al. The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum. 31, 315–324 (1988).
van Gestel, A. M. et al. Development and validation of the European League Against Rheumatism response criteria for rheumatoid arthritis. Comparison with the preliminary American College of Rheumatology and the World Health Organization/International League Against Rheumatism Criteria. Arthritis Rheum. 39, 34–40 (1996).
Canet, L. M. et al. Genetic variants within the TNFRSF1B gene and susceptibility to rheumatoid arthritis and response to anti-TNF drugs: a multicenter study. Pharmacogenet Genomics 25, 432–443 (2015).
Canet, L. M. et al. Genetic variants within immune-modulating genes influence the risk of developing rheumatoid arthritis and anti-TNF drug response: a two-stage case-control study. Pharmacogenet Genomics 25, 432–443 (2015).
Canet, L. M. et al. Genetic variants within the TNFRSF1B gene and susceptibility to rheumatoid arthritis and response to anti-TNF drugs: a multicenter study. Pharmacogenet Genomics 25, 323–333 (2015).
Sode, J. et al. Confirmation of an IRAK3 polymorphism as a genetic marker predicting response to anti-TNF treatment in rheumatoid arthritis. Pharmacogenomics J. 18, 81–86 (2018).
Liu, C. et al. Genome-wide association scan identifies candidate polymorphisms associated with differential response to anti-TNF treatment in rheumatoid arthritis. Mol. Med. 14, 575–581 (2008).
Sode, J. et al. Genetic Variations in Pattern Recognition Receptor Loci Are Associated with Anti-TNF Response in Patients with Rheumatoid Arthritis. PLoS one 10, e0139781 (2015).
Bek, S. et al. Systematic review and meta-analysis: pharmacogenetics of anti-TNF treatment response in rheumatoid arthritis. Pharmacogenomics J. 17, 403–411 (2017).
Gregersen, P. K. et al. REL, encoding a member of the NF-kappaB family of transcription factors, is a newly defined risk locus for rheumatoid arthritis. Nat. Genet. 41, 820–823 (2009).
Nyholt, D. R. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am. J. Hum. Genet. 74, 765–769 (2004).
Sole, X., Guino, E., Valls, J., Iniesta, R. & Moreno, V. SNPStats: a web tool for the analysis of association studies. Bioinforma. 22, 1928–1929 (2006).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Sci. 296, 2225–2229 (2002).
Canet, L. M. et al. Polymorphisms at phase I-metabolizing enzyme and hormone receptor loci influence the response to anti-TNF therapy in rheumatoid arthritis patients. Pharmacogenomics J. (2018).
Ibfelt, E. H., Jensen, D. V. & Hetland, M. L. The Danish nationwide clinical register for patients with rheumatoid arthritis: DANBIO. Clin. Epidemiol. 8, 737–742 (2016).
Li, Y. et al. A Functional Genomics Approach to Understand Variation in Cytokine Production in Humans. Cell 167, 1099–1110 e1014 (2016).
Schirmer, M. et al. Linking the Human Gut Microbiome to Inflammatory Cytokine Production Capacity. Cell 167, 1897 (2016).
Ter Horst, R. et al. Host and Environmental Factors Influencing Individual Human Cytokine Responses. Cell 167, 1111–1124 e1113 (2016).
Viatte, S. et al. Genetic markers of rheumatoid arthritis susceptibility in anti-citrullinated peptide antibody negative patients. Ann. rheumatic Dis. 71, 1984–1990 (2012).
Padyukov, L. et al. A genome-wide association study suggests contrasting associations in ACPA-positive versus ACPA-negative rheumatoid arthritis. Ann. rheumatic Dis. 70, 259–265 (2011).
Jiang, X. et al. An Immunochip-based interaction study of contrasting interaction effects with smoking in ACPA-positive versus ACPA-negative rheumatoid arthritis. Rheumatol. 55, 149–155 (2016).
Dong, X. et al. ACPAs promote IL-1beta production in rheumatoid arthritis by activating the NLRP3 inflammasome. Cellular & molecular immunology, (2019).
Bank, S. et al. Polymorphisms in the NFkB, TNF-alpha, IL-1beta, and IL-18 pathways are associated with response to anti-TNF therapy in Danish patients with inflammatory bowel disease. Alimentary pharmacology therapeutics 49, 890–903 (2019).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Zhang, Y. et al. Endochondral ossification pathway genes and postmenopausal osteoporosis: Association and specific allele related serum bone sialoprotein levels in Han Chinese. Sci. Rep. 5, 16783 (2015).
Swertz, M. A. et al. The MOLGENIS toolkit: rapid prototyping of biosoftware at the push of a button. BMC Bioinforma. 11(Suppl 12), S12 (2010).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Ovejero-Benito, M. C. et al. Polymorphisms associated with etanercept response in moderate-to-severe plaque psoriasis. Pharmacogenomics 18, 631–638 (2017).
Kichaev, G. et al. Leveraging Polygenic Functional Enrichment to Improve GWAS Power. Am. J. Hum. Genet. 104, 65–75 (2019).
Iwaszko, M. et al. Influence of CD94 and NKG2A variants on susceptibility to rheumatoid arthritis and efficacy of anti-TNF treatment. Jt. Bone Spine 83, 75–79 (2016).
Iwaszko, M. et al. Influence of NKG2D Genetic Variants on Response to Anti-TNF Agents in Patients with Rheumatoid Arthritis. Genes (Basel) 9 (2018).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
Piotrowski, P., Lianeri, M., Olesinska, M. & Jagodzinski, P. P. Prevalence of the NKG2D Thr72Ala polymorphism in patients with systemic lupus erythematosus. Mol. Biol. Rep. 39, 1343–1347 (2012).
Kabalak, G. et al. Association of an NKG2D gene variant with systemic lupus erythematosus in two populations. Hum. Immunol. 71, 74–78 (2010).
Stahl, E. A. et al. Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat. Genet. 42, 508–514 (2010).
Eyre, S. et al. High-density genetic mapping identifies new susceptibility loci for rheumatoid arthritis. Nat. Genet. 44, 1336–1340 (2012).
Ali, F. R. et al. An investigation of rheumatoid arthritis loci in patients with early-onset psoriasis validates association of the REL gene. Br. J. Dermatol. 168, 864–866 (2013).
Varade, J. et al. Analysis of the REL polymorphism rs13031237 in autoimmune diseases. Ann. rheumatic Dis. 70, 711–712 (2011).
Chen, F. et al. Genetic Variation in the REL Gene Increases Risk of Behcet’s Disease in a Chinese Han Population but That of PRKCQ Does Not. PLoS one 11, e0147350 (2016).
Ellinghaus, E. et al. Genome-wide meta-analysis of psoriatic arthritis identifies susceptibility locus at REL. J. Invest. Dermatol. 132, 1133–1140 (2012).
Bowes, J. et al. Comprehensive assessment of rheumatoid arthritis susceptibility loci in a large psoriatic arthritis cohort. Ann. rheumatic Dis. 71, 1350–1354 (2012).
Tang, H. et al. TLR10 and NFKBIA contributed to the risk of hip osteoarthritis: systematic evaluation based on Han Chinese population. Sci. Rep. 8, 10243 (2018).
Vrgoc, G. et al. Interleukin-17 and Toll-like Receptor 10 genetic polymorphisms and susceptibility to large joint osteoarthritis. J. Orthop. Res. 36, 1684–1693 (2018).
Nishikawa, R. et al. Genetic prediction of the effectiveness of biologics for psoriasis treatment. J. Dermatol. 43, 1273–1277 (2016).
Gebura, K. et al. Polymorphisms within Genes Involved in Regulation of the NF-kappaB Pathway in Patients with Rheumatoid Arthritis. Int. J. Mol. Sci. 18 (2017).
Wang, H., Zhou, S., Zhang, J., Lei, S. & Zhou, J. Correlations between TLR polymorphisms and inflammatory bowel disease: a meta-analysis of 49 case-control studies. Immunol. Res. 67, 142–150 (2019).
Bank, S. et al. Polymorphisms in the Toll-Like Receptor and the IL-23/IL-17 Pathways Were Associated with Susceptibility to Inflammatory Bowel Disease in a Danish Cohort. PLoS one 10, e0145302 (2015).
Bank, S. et al. Genetically determined high activity of IL-12 and IL-18 in ulcerative colitis and TLR5 in Crohns disease were associated with non-response to anti-TNF therapy. Pharmacogenomics J. 18, 87–97 (2018).
Loft, N. D. et al. Associations between functional polymorphisms and response to biological treatment in Danish patients with psoriasis. Pharmacogenomics J. 18, 494–500 (2018).
Oliveira-Tore, C. F. et al. Genetic Polymorphisms of Toll-like receptors 2 and 9 as Susceptibility Factors for the Development of Ankylosing Spondylitis and Psoriatic Arthritis. J. Immunol. Res. 2019, 1492092 (2019).
Yi, X., Xu, E., Xiao, Y. & Cai, X. Evaluation of the Relationship Between Common Variants in the TLR-9 Gene and Hip Osteoarthritis Susceptibility. Genet. Test. Mol. Biomarkers 23, 373–379 (2019).
Zheng, M. et al. Association between TLR-9 gene rs187084 polymorphism and knee osteoarthritis in a Chinese population. Bioscience reports 37 (2017).
Huang, C. M. et al. Association of toll-like receptor 9 gene polymorphism in Chinese patients with systemic lupus erythematosus in Taiwan. Rheumatol. Int. 32, 2105–2109 (2012).
Bank, S. et al. Polymorphisms in the inflammatory pathway genes TLR2, TLR4, TLR9, LY96, NFKBIA, NFKB1, TNFA, TNFRSF1A, IL6R, IL10, IL23R, PTPN22, and PPARG are associated with susceptibility of inflammatory bowel disease in a Danish cohort. PLoS one 9, e98815 (2014).
Cho, W. K. et al. Association of Polymorphisms in Toll-Like Receptors 4 and 9 with Autoimmune Thyroid Disease in Korean Pediatric Patients. Int. J. Endocrinol. 2017, 2304218 (2017).
Bank, S. et al. Associations between functional polymorphisms in the NFkappaB signaling pathway and response to anti-TNF treatment in Danish patients with inflammatory bowel disease. Pharmacogenomics J. 14, 526–534 (2014).
Plenge, R. M. et al. TRAF1-C5 as a risk locus for rheumatoid arthritis–a genomewide study. N. Engl. J. Med. 357, 1199–1209 (2007).
Acknowledgements
We thank all participants who have agreed to participate in this study. Authors also thank María Dolores Casares, Ángeles Molina, Carmen Oloriz for the collection of Spanish samples and Hans Jurgen Hoffmann, Marianne Thomsen, Vibeke Østergaard Thomsen, Malene Rohr Andersen, Lise Lotte B. Laursen, Helle Jørgensen, Ram Benny Christian Dessau, Niels Steen Krogh, Ulla Vogel, Paal Skytt Andersen, Ivan Brandslund, Steffen Bank, Frederik Trier Møller, Nikolai Toft and Niels Møller Andersen for the participation in collection and purification of Danish samples. We also thank the Danish Departments of Rheumatology for their implication in the collection of clinical data from RA patients included in the DANBIO cohort and the Danish Rheumatologic Biobank. Likewise, we would like to thank Teun van Herwaarden for steroid hormone measurements in serum samples from subjects ascertained through the HFGP initiative. This work was partially supported by intramural funds of GENYO and FIBAO foundation (Granada, Spain); Novo Nordisk Fonden (NNF15OC0016932, VA); and Knud og Edith Eriksens Mindefond (VA) and Gigtforeningen (A2037, A3570, VA). MGN was supported by a Spinoza grant from the Netherlands Organization for Scientific Research.
Author information
Authors and Affiliations
Contributions
R. Caliz and J. Sainz designed the study and drafted the manuscript. J. Sanchez-Maldonado, A. Moñiz-Díez, S. Muñoz-Peña, and A. Rodríguez-Ramos were responsible for genotyping. H. Canhão, A. Escudero, S.B. Sorensen, M.L. Hetland, M.A. Ferrer, B. Glintborg, I. Filipescu, E. Pérez-Pampin, P. Conesa-Zamora, A. García, A. den Broeder, S. de Vita, S.E.H. Jacobsen, E. Collantes, L. Quartuccio, J.E. Fonseca, M. Jurado, M.A. López-Nevot, M.J.H. Coenen, V. Andersen, R. Cáliz and J. Sainz coordinated the sample collection and H. Canhão, I. Filipescu, A. García, and M.A Ferrer were involved in the records review and data acquisition. M. Martínez-Bueno and J. Sainz performed data quality control and statistical analysis. M. Netea, R. Ter Horst, Y. Li provided the functional raw data and J. Sainz performed the analysis of functional data. All authors contributed to and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
V.A. has received compensation for consultancy and for being a member of an advisory board from MSD (Merck) and Janssen. B.G. received funding for research from AbbVie, Biogen, and Pfizer. M.L.H. received funding for research from Abbvie, Biogen, BMS, CellTrion, MSD, Novartis, Orion, Pfizer, Samsung and UCB. The rest of authors declare that they have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Manuel Sánchez-Maldonado, J., Martínez-Bueno, M., Canhão, H. et al. NFKB2 polymorphisms associate with the risk of developing rheumatoid arthritis and response to TNF inhibitors: Results from the REPAIR consortium. Sci Rep 10, 4316 (2020). https://doi.org/10.1038/s41598-020-61331-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61331-5
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.