Abstract
The recognition of and differential responses to salient stimuli are among the main drivers of behavioral plasticity, yet, how animals evolve and modulate functional responses to novel classes of antagonistic stimuli remain poorly understood. We studied free-living male red-winged blackbirds (Agelaius phoeniceus) to test whether gene expression responses in blood are distinct or shared between patterns of aggressive behavioral responses directed at simulated conspecific versus heterospecific intruders. In this species, males defend territories against conspecific males and respond aggressively to female brown-headed cowbirds (Molothrus ater), a brood parasite that commonly lays eggs in blackbird nests. Both conspecific songs and parasitic calls elicited aggressive responses from focal subjects and caused a downregulation in genes associated with immune system response, relative to control calls of a second, harmless heterospecific species. In turn, only the conspecific song treatment elicited an increase in singing behavior and an upregulation of genes associated with metabolic processes relative to the two heterospecific calls. Our results suggest that aspects of antagonistic behaviors to both conspecifics and brood parasites can be mediated by similar physiological responses, suggestive of shared molecular and behavioral pathways involved in the recognition and reaction to both evolutionarily old and new enemies.
Similar content being viewed by others
Introduction
Faced with a suite of relevant and irrelevant stimuli, animals must rapidly perceive, process, and decide whether and how behaviorally to respond to diverse cues1. Upon the recognition of salient stimuli, for example, a cascade of neurophysiological and motor responses can be engaged to facilitate and modulate a reaction2,3. Yet, how recognition systems adapt and enable responses to evolutionarily novel stimuli remain poorly understood4. In particular, the timely and accurate recognition of heterospecific antagonists, such as parasites, predators, or competitors, typically has substantial fitness advantages5. Can evolutionarily established behavioral and physiological responses to conspecific competitors be co-opted in parallel to adaptively respond to heterospecific antagonists?
Hosts of avian brood parasites pay the costs of raising unrelated young, often with the additional expense of losing some or all of their own offspring6. Many hosts have evolved to combat brood parasitism by attacking adult parasites, abandoning parasitized nests, and/or rejecting parasitic offspring7. Anti-parasitic defenses can take categorically different responses from competitive (e.g. territorial defense with song and overt aggression against same-sex conspecific intruders) or anti-predatory behaviors, and can even be evoked by a partial suite of (visual or auditory only) sensory cues8,9.
What constitutes the physiological basis of anti-parasite responses remains largely unknown in avian host-parasite systems10. For example, recent work in wild-caught juvenile male red-winged blackbirds (Agelaius phoeniceus; hereafter: redwings) found no differences in immediate early gene (IEG) expression levels within the auditory forebrain in response to the calls of adult female brood parasites (brown-headed cowbirds Molothrus ater; hereafter: cowbirds) versus a harmless control species (mourning dove Zenaida macroura; hereafter: doves), whereas responses were stronger to conspecific adult female calls11. In the wild, however, juvenile redwings do not hold territories and may not (yet) have been exposed to parasitism by cowbirds. In contrast, breeding adult male redwings are well known for territory defense and anti-brood parasite aggression12 (Fig. 1). The species-typical “conk-a-ree” song is a long-range “keep-out” signal13,14; and is the most common conspecific territory defense display of this species15. Male redwings also respond strongly to parasitic female cowbirds by approaching and attacking the parasites, but they respond little to harmless sympatric species16. Therefore, we set out to test whether antagonistic responses to conspecifics and parasites involve distinct or shared behavioral and physiological responses in freely-behaving adult male redwings on their breeding territories.
Linking evolutionarily relevant and ecologically salient stimuli with their proximate recognition system responses has been difficult in wild animals. This is because, beyond the well-known complexities of field work, lethal collection is required to sample neural tissues during or following the undertaking of a recognition task. Furthermore, such terminal collection prohibits repeated contrasts or ontogenetic comparisons within the same subjects. Alternatively, non-lethal neuroimaging-based techniques, including functional Magnetic Resonance Imaging or micro Positron Emission Tomography, require that subjects become captive during or soon after the recognition task17,18.
Transcriptomic analyses have become a critical tool to analyze functionally relevant cell types and tissues both in model species and, increasingly in non-model species collected from the wild19,20. Moreover, non-terminal collection, such as gene expression within peripheral, whole blood of birds, may reveal functional parallels in the recognition of salient stimuli. Peripheral blood is increasingly studied as a potential biomarker for phenotypic changes, such as disease and social recognition of stimuli21,22,23,24,25. Accordingly, in humans gene expression within the peripheral blood can depend on exposure to music26,27. Gene expression detectable by RNAseq within the blood not only represents hematopoietic cells, but also of circulating extracellular mRNA that is contributed from other regions, including the brain28,29. For example, a con- vs. heterospecific acoustic playback paradigm in captive zebra finches (Taeniopygia guttata) found that gene expression patterns were positively correlated for a subset of genes between the auditory forebrain and peripheral blood23.
Here we presented playback stimuli of unfamiliar conspecific (male redwing songs), parasitic heterospecific (female cowbird chatters), and control heterospecific vocalizations (dove coos) to free-living adult male redwings on their breeding territories, recorded their behavioral responses, then caught them to collect blood samples, and assessed peripheral gene expression patterns. Given the well-known behavioral repertoire of territorial male redwings12, we expected them to respond to the playback of conspecific song by increasing their own singing and approaching and remaining in proximity to the playback speaker when compared to playback of a harmless heterospecific song. In turn, we predicted male redwings to approach and maintain proximity to the playback of parasite calls, but not to increase singing, in comparison to the harmless heterospecific. Finally, we tested for distinct and parallel gene activation and behavioral patterns in response to conspecific vs. parasitic stimuli relative to controls.
Results
Behavioral response to playbacks
We performed playbacks to 35 territorial male red-wings. Whether or not a subject was caught within 30 min. had no impact on its response behaviors statistically (all z > −1.9, p > 0.05), nor were there differences in behavioral responses between the 1st and 3rd playback trial segment (see Methods), therefore behavioral data from all subjects and territories in response to the playback paradigm were analyzed statistically (nredwing = 11, ncowbird = 12, ndove = 12). However, transcriptomic data were only available for subjects caught within 30 min of its respective playback set’s delivery (see below for sample sizes). Generalized linear mixed models, with playback type as the independent predictor, revealed that male redwings responses were significantly variable during active playback periods: number of songs (z = −2.82, p = 0.005), number of approaches (z = −2.50, p = 0.01), and nearest distance to the playback speaker (z = −2.10, p = 0.04), but not in the time spent near the playback speaker (z = −0.90, p = 0.17) (Fig. 2). Tukey-corrected post-hoc analyses revealed that subjects sang significantly more in response to redwing than cowbird (p = 0.02) or dove (p = 0.02) playbacks and did not differ significantly in songs in response to cowbird and dove playbacks (p = 0.99). Territorial male redwings approached more frequently to redwing versus dove (p = 0.005) and cowbird versus dove playbacks (p = 0.03), but did not differ between redwing versus cowbird playback (p = 0.79). Territorial male redwings approached more closely to cowbird than dove playbacks (p = 0.003), but did not differ in response to redwing vs. dove (p = 0.10), nor to redwing vs. cowbird playbacks (p = 0.38).
Gene expression
We were able to extract enough RNA from the blood samples of 21 males upon experimental presentations of playback treatments (nredwing = 6, ncowbird = 8, ndove = 7). For these samples, we sequenced an average of 15.9 million reads per sample (range = 11.7–19.1 million reads, Supplementary Table S1). On average, 75.5% of the reads aligned to the proxy reference genome (range = 71.9–78%, Supplementary Table S1).
We first tested for differential gene expression for 20 candidate biomarker genes, previously identified from peripheral (whole blood) of zebra finches23. This analysis yielded no significant differentially expressed genes in response to conspecific songs versus dove coo playback treatments (all Bonferroni corrected p-values > 0.50, Supplementary Table S2).
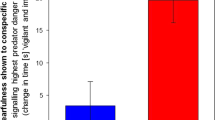
After filtering for lowly expressed genes, we then incorporated the read counts of 7202 genes for co-expression analysis (WGCNA) of the 17249 genes in the proxy reference genome. Two modules were significantly correlated with playback treatments (Figs. 3 and 4, Supplementary Table S3), the “darkturquoise” module included upregulated genes in response to Redwing song treatment (n = 268 genes, r = 0.52, p = 0.01) and the “yellow” module included genes downregulation in response to both Redwing songs and Cowbird chatter relative to Dove coo (n = 475 genes, r = 0.64, p = 0.002).
The top two genes’ expression profiles from each of the (A) the “yellow” module, which includes genes significantly downregulated in redwing and chatter treatments relative to dove coo and (B) the “darkturquoise” module, which includes genes associated with an upregulation in response to the redwing song treatment. Stars denote *p < 0.05 and **p < 0.01 between indicated groups.
Using a rank-ordered approach30, we identified an enrichment for gene ontology (GO) terms for each significant module. Of the 7202 genes, 5916 were considered orthologous in the human genome annotation and 5845 were associated with a GO term (accessed 7 January, 2020). For the “darkturquoise” module, correlated with a response to Redwing song playback, we found significant GO terms associated with the metabolism, regulation of gene expression, and protein ubiquitination (Table 1, Supplementary Table S4). For the “yellow” module, correlated with a response to both Redwing song and Cowbird playback, we find significant GO terms associated with immune system response, such as defense responses to virus and cytokine-mediated pathways (Table 1, Supplementary Table S5).
Discussion
Our expectations that subjects would sing more in response to conspecific playback and approach and perch closer to both redwing and cowbird playback were confirmed. We are therefore confident that we produced meaningful behavioral differences in aggressive responses of our subject free-living male red-winged blackbirds. By studying wild male territorial red-winged blackbirds, we also detected patterns of co-expressed genes in peripheral blood that matched the patterns of variation in aggressive behavioral responses to different classes of playbacks. Accordingly, in one behavioral response metric (approach distance) and in a co-expressed gene module, responses to conspecific songs and brood parasitic heterospecific calls were similar and both were significantly different from responses to harmless heterospecific calls. In turn, in a second behavioral response (number of songs) and another co-expressed gene module, responses to conspecific songs were significantly different from responses to both brood parasitic heterospecific and harmless heterospecific calls. Integrating the discriminability generated by these two patterns of responses provides for unique encoding for each of the three different playback types in both the behavioral and gene expression domains, separating conspecific songs from brood parasitic calls from harmless heterospecific calls.
Our results demonstrate that antagonistic responses to both conspecifics and brood parasites can involve similar physiological responses. The gene module correlated with both conspecific songs and cowbird chatters, relative to dove coos, was enriched for gene ontology immune system terms (Table 1), including defense responses to viruses and regulation of type I interferon production. For example, the top gene from the module, NLRC5 (NOD-like receptor family CARD domain containing 5 gene; Fig. 4), regulates adaptive immune responses against pathogens, including the activation of MHC class I genes31. Unsurprisingly, these genes from the module were downregulated relative to the control playbacks, perhaps as a trade-off between aggressive responses and immune function seen elsewhere in avian and other vertebrate lineages32. Similarly, luteinizing hormone decreases in male red-wing blackbirds in response to simulated territorial intrusion33. Nevertheless, the source of the mRNA expression in our peripheral blood samples, such as erythrocytes, leukocytes or exosomes34, as well as the physiological function of these genes in either immunosuppressive or enhancement of immune response to acute stress remain unclear.
Overall, the behavioral and gene expression data indicate that similar physiological pathways may be involved in the cognitive recognition and motor responses to distinct antagonistic threats, in this case conspecific and heterospecific intruders. Many host species of brood parasites have evolved aggressive anti-parasitic behaviors7, yet the cognitive and physiological mechanisms involved in this behavioral evolution remain undiscovered10. Our study suggests that some of proximate responses to conspecific intruders were co-opted in the evolution of anti-parasitic aggression towards female cowbirds.
The gene expression responses found here in adult, territorial redwing males in the summer breeding season are different from the IEG patterns detected from juvenile redwing males captured in the fall, which showed that the only conspecific calls generated differential responses in the auditory forebrain relative to cowbird and dove calls11. In turn, our behavioral data from adult red-winged blackbirds confirm previous findings in this and other avian hosts of brood parasitic species, which demonstrated behavioral responses to heterospecific parasitic models and auditory stimuli16,35,36,37.
We did not find differential gene expression in a candidate set of potential biomarker genes, previously identified from peripheral whole blood in a conspecific vs. harmless heterospecific (dove) acoustic playback paradigm in captive female zebra finches23. Given that our experimental treatments in the present study also included conspecific (redwing songs) vs. irrelevant heterospecific (dove) comparisons, we predicted that we would detect some of these same potential biomarker genes to be differentially expressed in free-living redwings. However, the use of different study species (zebra finches vs. red-winged blackbirds) and/or the different sexes of our subjects (female finches vs. male redwings) likely contributed to our inability to identify consistent gene expression differences within whole blood for conspecific recognition in songbirds.
In conclusion, our study demonstrates a parallel in behavioral and gene expression responses to simulated antagonistic threats. In particular, we find support for physiological responses involved in conspecific territorial aggression co-opted in the evolved recognition of heterospecific brood parasites. In addition, our study further demonstrates the utility for peripheral gene expression to study avian recognition and behavioral responses to social stimuli.
Methods
Study species and study area
We studied red-winged blackbirds at Newark Road Prairie in Rock County, Wisconsin, USA (42° 32′ N, 89° 08′ W) during the breeding season of 2018. Newark Road Prairie is a 13-ha wet-mesic remnant prairie and sedge meadow habitat that supports about 35 male redwing territories38. Capture and banding were conducted under United States Geological Survey banding permit #20438 to K.Y. All males were captured in Potter traps placed on platforms and baited with non-viable sunflower seeds. Captured males were banded with USGS numbered aluminum bands and unique color combinations of plastic wraparound bands for individual identification. We did not aim to capture and band females in 2018, although a few females had been captured and individually color banded in previous years.
We used observations of territorial behavior to place an additional seed-baited trap within the breeding territory of each potential subject. We pinned open the doors of each trap, baited it daily, and then observed each baited trap to determine when the male began entering it. The male was then allowed 1–3 weeks of unrestricted access to the bait without capture, at which point the male would usually enter the trap quickly after we baited it. Playbacks were then performed as described below. In our long-term experience working at this site (K.Y.), this method allows most individual males to be reliably recaptured within 30 min. Previous work at this site on redwings’ responses to model cowbirds vs. harmless heterospecifics, coupled with their respective call playbacks, showed strongly graded aggressive responses to the former relative to the latter16.
Playbacks
The playback protocol was approved by the Beloit College IACUC (protocol #18002). We presented one of three playback types at each territory: (1) male conspecific songs (“Redwing”; highly salient), (2) female cowbird chatter calls (“Cowbird”; salient heterospecific vocalization of a brood parasite of redwing nests), or (3) dove coo (“Dove”; non-salient vocalizations of a harmless sympatric heterospecific) for broadcasts. Given that male red-winged blackbirds exhibit greater behavioral responses to female vs. male cowbird models16, the chatter call, a specific call of female cowbirds was chosen to acoustically simulate brood parasite intrusion. To address pseudo-replication39, one out of five different individuals’ song/call exemplars were assigned at random for each playback type. Audacity v 2.2.0 was used to filter playback stimuli above 2000 Hz and below 500 Hz, and to normalize mean amplitude of all stimuli. Acoustic stimuli were matched in peak amplitude and duration. Exemplars were thus similar to each other in peak amplitude with duration.
Playback types and exemplar files were randomly assigned to territorial male redwings using a stratified balanced design to keep sample sizes per type similar. For each playback type we randomly chose one of the exemplars to broadcast from an iPhone 5 or 6 (Apple Inc., Cupertino, California, USA) connected to an Ecoxgear ECOXBT speaker (Grace Digital Audio, Peterborough, Ontario, Canada) via a 30-m auxiliary cable. The speaker was placed on the trap, which was closed and unbaited during playbacks. Playbacks were broadcasted at 80–85 dB SPL at 1 m from the source (as measured by a sound pressure meter: Pyle PSPL01, Pyle Audio Inc., Brooklyn, New York, USA), which approximated natural amplitude (KY personal observations).
We used a 30-min paradigm to induce (differential) gene expression (e.g.23), but to avoid habituation in the field, each playback consisted of three 10-min segments for a total period of 30 min. The first and third segments were active sound broadcast periods and the second segment was a 10-min silent period. Each active period consisted of 10 1-min sub-segments in which a single exemplar (song, chatter, or coo) played at 0, 10, 20, 30, and 40 s, followed by 20 s of silence. As soon as the 30-min playback was completed, we removed the playback equipment and baited and set the trap; we aimed to capture each subject within 30 min of the end of the playback and to band all previously unmarked subjects as described above. We recorded the time to capture for all subjects obtained within the 30-min maximum. Each captured bird was removed from its trap within 1 minute (usually less than 30 s), processed, and released within 5 min of capture. A blood sample of about 50 μl was collected from the brachial vein of each hand-held, unanesthesized subject using a 1.27-cm 27-g needle and heparinized caraway tube (Fisher Scientific, Pittsburgh, Pennsylvania, USA). The birds remained calm, did not react to the insertion of the needle, and resumed normal behavior within minutes of release. Each blood sample was placed in 500 µl of RNAlater and then stored in a −80 °C freezer for subsequent RNA extraction and sequencing.
Prior to playback, we placed two markers each 10 m to the right and left of from the speaker to facilitate measuring proximity time spent within 10 m of the speaker. During each 10-min segment we recorded (1) number of songs (songs), (2) number of flights towards the speaker (approaches), (3) time spent within 10 m of the speaker (time in proximity), and (4) the distance (m) of the closest approach (closest approach). These behavioral variables are well known to indicate a male redwing’s aggressiveness40. We analyzed behavioral data collected during the 1st and 3rd 10-min periods, which preceded the capture timepoint by up to 30 min and, thus, is representative of the time sampled for the gene-expression patterns, too. Number of songs and number of approaches were the totals for periods 1 and 3 (active playback). Time (min) within 10 m was the total time for the active playback periods. Closest approach was the shortest distance between the subject and the speaker during the two active playback periods.
We used generalized linear mixed models with a negative binomial response and individual male identity as a random effect with glmmTMB in R (version 3.5.1) to analyze the responses of male redwings to the three playbacks. Each model included playback treatment, playback trail segment, and whether the male was captured as explanatory variables. A significant result was further examined with a Tukey post-hoc analysis to identify significantly different pairs of treatment. The alpha level was set at the p < 0.05.
RNA extraction and sequencing
RNA was extracted with RiboPure blood kits (Life Technologies, Carlsbad, California, USA) and treated with DNAse for purification. We assessed the quality of purified RNA on a Bioanalyzer (Agilent, Wilmington, Delaware, USA) (RIN > 7.0).
All library preparations and sequencing were performed at the University of Illinois at Urbana-Champaign Roy J. Carver Biotechnology Center, Urbana, IL, USA. A library for each sample was prepared with an Illumina TruSeq Stranded RNA sample prep kit. All libraries were pooled, quantitated by qPCR, and sequenced on one lane of an Illumina HiSeq 4000 with a HiSeq 4000 sequencing kit version 1, producing single-end 100 bp reads. Fastq files were demultiplexed with bcl2fastq v 2.17.1.14 (Illumina).
Preparation of reference genome
Lacking an annotated reference genome for the red-winged blackbird, we created a proxy reference following the pseudo-it pipeline (https://github.com/bricesarver/pseudo-it). Briefly, we extracted DNA from liver and muscle tissue of a male red-winged blackbird (cataloged at the Museum of Southwestern Biology MSB:Bird:40979) and performed paired-end (200 bp) whole-genome sequencing on one lane of HiSeq (2500) at the Duke Genome Center. After removing the Illumina adapters with Trim Galore! v 0.3.7 (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/), which incorporates Cutadapt v 1.7.141, we aligned the DNA reads to the closest related species publicly available, the white-throated sparrow (Zonotrichia alibicolis) genome (assembly Zonotrichia_albicollis-1.0.1), with BWA-mem42. Using GATK43 we then identified and inserted red-winged blackbird SNPs into the white-throated sparrow reference genome. To improve the proxy reference genome, we performed an additional iteration of the pseudo-it pipeline.
Gene expression
We removed Illumina adapters from RNA reads with Trim Galore! v 0.3.7. We then aligned the reads to the proxy reference genome with Hisat244 and quantified read abundance with HTSeq-count45. With DeSeq 246, we then included the playback treatments and time to capture (minutes) to analyze the gene expression patterns and also to compare candidate genes identified from the parallel expression patterns of peripheral (whole blood) in a previous RNA-sequencing study on zebra finches in response conspecific vs. heterospecific songs (n = 20 differentially expressed genes; Table 1 in23).
Next, we sought to identify networks of genes specifically responsive to Redwing song, Cowbird chatter or both Redwing and Cowbird relative to Dove coo. We performed weighted gene co-expression network analysis (WGCNA), which is used for finding clusters (modules) of highly correlated genes and determine the relationship of modules to treatments. We used the WGCNA package in R47 to identify modules of co-expressed genes in our dataset. To remove genes with low read abundance, we filtered for genes with <1 count per million in at least 10 samples. We then normalized for read-depth and extracted variance stabilizing transformed (vst) read counts from DEseq. 2 into WGCNA. To build the co-expression matrix, we chose a soft thresholding power (β) value of 12, at which at which we observed a plateau in Mean Connectivity, thus representing a scale-free topology47. We generated a signed network with minimum module size of 30 genes and merged highly correlated modules (dissimilarity threshold = 0.25). We then correlated the eigengene, which is the first principal component of a module, of these merged modules with playback treatments (redwing, cowbird, dove). Modules with p < 0.05 were considered significantly correlated with a given trait.
Finally, we tested for functional enrichment of gene ontology (GO) categories with GOrilla30. For each module, genes were ranked based on their module membership score determined in the WGCNA analysis. We preferred this rank-order based approach (as opposed to strict module assignment) as it reflects the correlation among modules, and because some genes could be assigned to multiple modules48. GOrilla performs ranked-order analyses with human gene IDs, so we identified orthologous genes from the annotated red-winged blackbird proxy-genome. Statistical significance of GO categories was determined with p-values corrected for multiple hypothesis testing (FDR < 0.05). We used REVIGO to remove redundant and overlapping GO categories, with an allowed semantic similarity measure of 0.549.
Data availability
The RNAseq data that support the findings of this study are available at NCBI, BioProject PRJNA608291.
References
Mendelson, T. C. et al. Cognitive Phenotypes and the Evolution of Animal Decisions. Trends Ecol. Evol. 31, 850–859 (2016).
Patrick, C. J. Psychophysiological correlates of aggression and violence: an integrative review. Phil. Trans. R. Soc. B 363, 2543–2555 (2008).
Louder, M. I. M. et al. An acoustic password enhances auditory learning in juvenile brood parasitic cowbirds. Curr. Biol. 29, 4045–4051 (2019).
Louder, M. I. M., Lawson, S., Lynch, K. S., Balakrishnan, C. N. & Hauber, M. E. Neural mechanisms of auditory species recognition in birds. Biol. Rev. 94, 1619–1635 (2019).
Peiman, K. S. & Robinson, B. W. Ecology and evolution of resource-related heterospecific aggression. Q. Rev. Biol. 85, 133–158 (2010).
Davies, N. B. Cuckoos, Cowbirds and Other Cheats. (T & AD Poyser Ltd, 2000).
Feeney, W. E., Welbergen, J. A. & Langmore, N. E. The frontline of avian brood parasite–host coevolution. Anim. Behav. 84, 3–12 (2012).
Burgham, M. C. J. & Picman, J. Effect of brown-headed cowbirds on the evolution of yellow warbler anti-parasite strategies. Anim. Behav. 38, 298–308 (1989).
Campobello, D. & Sealy, S. G. Enemy recognition of reed warblers (Acrocephalus scirpaceus): threats and reproductive value act independently in nest defence modulation. Ethology 116, 498–508 (2010).
Abolins-Abols, M. & Hauber, M. E. Host defences against avian brood parasitism: an endocrine perspective. Proc Biol Sci 285, (2018).
Lynch, K. S., Louder, M. I. M. & Hauber, M. E. Species-specific auditory forebrain responses to non-learned vocalizations in juvenile blackbirds. Brain Behav. Evol. 91, 193–200 (2018).
Searcy, W. A. & Yasukawa, K. Red-winged Blackbird (Agelaius phoeniceus), version 2.0. In The Birds of North America (Cornell Lab of Ornithology, 2019).
Peek, F. W. An experimental study of the territorial function of vocal and visual display in the male red-winged blackbird (Agelaius phoeniceus). Anim. Behav. 20, 112–118 (1972).
Yasukawa, K. Song repertoires in the red-winged blackbird (Agelaius phoeniceus): A test of the Beau Geste hypothesis. Anim. Behav. 29, 114–125 (1981).
Peek, F. W. Seasonal change in the breeding behavior of the male red-winged blackbird. Wilson Bull. 83, 383–395 (1971).
Yasukawa, K., Lindsey-Robbins, J., Henger, C. S. & Hauber, M. E. Antiparasitic behaviors of red-winged blackbirds (Agelaius phoeniceus) in response to simulated brown-headed cowbirds (Molothrus ater): further tests of the frontloaded parasite-defense hypothesis. Wilson J. Ornithol. 128, 475–486 (2016).
Louder, M. I. M. et al. Shared neural substrates for song discrimination in parental and parasitic songbirds. Neurosci. Lett. 622, 49–54 (2016).
Tokarev, K. et al. Sexual dimorphism in striatal dopaminergic responses promotes monogamy in social songbirds. eLife 6, e25819 (2017).
Ekblom, R. & Galindo, J. Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 107, 1–15 (2011).
Jax, E., Wink, M. & Kraus, R. H. S. Avian transcriptomics: opportunities and challenges. J Ornithol. 159, 599–629 (2018).
Mohr, S. & Liew, C.-C. The peripheral-blood transcriptome: new insights into disease and risk assessment. Trends Mol Med. 13, 422–432 (2007).
Liew, C.-C., Ma, J., Tang, H.-C., Zheng, R. & Dempsey, A. A. The peripheral blood transcriptome dynamically reflects system wide biology: a potential diagnostic tool. J. Lab. Clin. Med. 147, 126–132 (2006).
Louder, M. I. M., Hauber, M. E. & Balakrishnan, C. N. Early social experience alters transcriptomic responses to species-specific song stimuli in female songbirds. Behav. Brain Res. 347, 69–76 (2018).
Guelfi, G. et al. A cross-talk between blood-cell neuroplasticity-related genes and environmental enrichment in working dogs. Sci. Rep. 9, 1–11 (2019).
Newhouse, D. J., Barcelo−Serra, M., Tuttle, E. M., Gonser, R. A. & Balakrishnan, C. N. Parent and offspring genotypes influence gene expression in early life. Mol. Ecol. 28, 4166–4180 (2019).
Kanduri, C. et al. The effect of music performance on the transcriptome of professional musicians. Sci. Rep. 5, 1–7 (2015).
Kanduri, C. et al. The effect of listening to music on human transcriptome. PeerJ 3, e830 (2015).
Tylee, D. S., Kawaguchi, D. M. & Glatt, S. J. On the outside, looking in: a review and evaluation of the comparability of blood and brain “-omes”. Am. J. Med. Genet. B 162, 595–603 (2013).
Sullivan, P. F., Fan, C. & Perou, C. M. Evaluating the comparability of gene expression in blood and brain. Am. J. Med. Genet. B 141B, 261–268 (2006).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Kobayashi, K. S. & van den Elsen, P. J. NLRC5: a key regulator of MHC class I-dependent immune responses. Nat. Rev. Immunol. 12, 813–820 (2012).
Takahashi, A., Flanigan, M. E., McEwen, B. S. & Russo, S. J. Aggression, Social Stress, and the Immune System in Humans and Animal Models. Front. Behav. Neurosci. 12 (2018).
Harding, C. & Follett, B. Hormone changes triggered by aggression in a natural population of blackbirds. Science 203, 918 (1979).
Li, M. et al. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Phil. Trans. R. Soc. B 369, 20130502 (2014).
Capper, C.-L., Guigueno, M. F. & Sealy, S. G. Acceptance of Simulated Cowbird Parasitism in a Northern Population of Red-Winged Blackbirds. Am. Midl. Nat 167, 127–135 (2012).
Gill, S. A., Neudorf, D. L. H. & Sealy, S. G. Do hosts Discriminate between Sexually Dichromatic Male and Female Brown-headed Cowbirds? Ethology 114, 548–556 (2008).
Henger, C. S. & Hauber, M. E. Variation in antiparasitic behaviors of red-winged blackbirds in response to simulated brown-headed cowbirds. Wilson J. Ornithol. 126, 488–499 (2014).
Yasukawa, K. The costs and benefits of a vocal signal: the nest-associated ‘Chit’ of the female red-winged blackbird, Agelaius phoeniceus. Anim. Behav. 38, 866–874 (1989).
Kroodsma, D. E., Byers, B. E., Goodale, E., Johnson, S. & Liu, W.-C. Pseudoreplication in playback experiments, revisited a decade later. Anim. Behav. 61, 1029–1033 (2001).
Yasukawa, K. Aggressive tendencies and levels of a graded display: Factor analysis of response to song playback in the redwinged blackbird (Agelaius phoeniceus). Behav. Biol. 23, 446–459 (1978).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Van der Auwera, G. A. et al. From fastq data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nature Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq 2. Genome Biol. 15, 550 (2014).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Horton, B. M., Ryder, T. B., Moore, I. T. & Balakrishnan, C. N. Gene expression in the social behavior network of the wire-tailed manakin (Pipra filicauda) brain. Genes Brain Behav. e12560 (2019).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLOS One 6, e21800 (2011).
Acknowledgements
For financial support we thank the National Science Foundation IOS Program in Behavioral Systems (nos. 1456524 and 1456612 to M.E.H. and C.B.), the National Academies Keck Futures Initiative Grant (to F.M.K.U. and M.E.H). Additional funding was provided by the National Geographic Society (NGS-60453R-19) and the Harley Jones Van Cleave Professorship at the University of Illinois and (to M.E.H.) and the Mead-Witter Foundation (to K.Y.). For discussions, we are grateful to Kathleen Lynch, Mikus Āboliņš-Ābols, Shelby Lawson, and Sarah London.
Author information
Authors and Affiliations
Contributions
M.I.M.L., K.Y., F.M.K.U. and M.E.H. designed the project; M.L., K.Y. collected field data and samples; A.A.L. generated the laboratory data, M.I.M.L., C.B. and M.E.H. analyzed the data; M.I.M.L., and M.E.H. wrote the first draft, and all authors provided critical feedback, reviewed, and edited the manuscript. All authors approve the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Louder, M.I.M., Lafayette, M., Louder, A.A. et al. Shared transcriptional responses to con- and heterospecific behavioral antagonists in a wild songbird. Sci Rep 10, 4092 (2020). https://doi.org/10.1038/s41598-020-60231-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60231-y
This article is cited by
-
Functional neurogenomic responses to acoustic threats, including a heterospecific referential alarm call and its referent, in the auditory forebrain of red-winged blackbirds
Scientific Reports (2024)
-
Evolution and function of neurocognitive systems in non-human animals
Scientific Reports (2021)
-
Parasitic begging calls of nestmate-evictor common cuckoos stimulate more parental provisions by red-winged blackbirds than calls of nest-sharing brown-headed cowbirds
Behavioral Ecology and Sociobiology (2021)
-
Heterospecific eavesdropping on an anti-parasitic referential alarm call
Communications Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.