Abstract
Anthropogenic hybridization is recognized as a major threat to the long-term survival of natural populations. While identifying F1 hybrids might be simple, the detection of older admixed individuals is far from trivial and it is still debated whether they should be targets of management. Examples of anthropogenic hybridization have been described between wolves and domestic dogs, with numerous cases detected in the Italian wolf population. After selecting appropriate wild and domestic reference populations, we used empirical and simulated 39-autosomal microsatellite genotypes, Bayesian assignment and performance analyses to develop a workflow to detect different levels of wolf x dog admixture. Membership proportions to the wild cluster (qiw) and performance indexes identified two q-thresholds which allowed to efficiently classify the analysed genotypes into three assignment classes: pure (with no or negligible domestic ancestry), older admixed (with a marginal domestic ancestry) and recent admixed (with a clearly detectable domestic ancestry) animals. Based on their potential to spread domestic variants, such classes were used to define three corresponding management categories: operational pure, introgressed and operational hybrid individuals. Our multiple-criteria approach can help wildlife managers and decision makers in more efficiently targeting the available resources for the long-term conservation of species threatened by anthropogenic hybridization.
Similar content being viewed by others
Introduction
Over the last decades, thanks to the growing availability of genetic and genomic data, hybridization has been increasingly studied for its evolutionary and conservational implications on the long-term survival of the involved taxa1,2,3,4,5. However, while natural hybridization between closely related taxa is frequently acknowledged as an evolutionary process providing novel adaptive gene assemblages6,7, anthropogenic hybridization (AH), mainly caused by intentional admixture, translocations, habitat modifications and climate changes8,9,10, is globally considered a serious conservation threat to the genetic integrity of local populations, which might be compromised by gene introgression from alien or domesticated species11,12,13,14,15,16. Thus, the consequences of such human-mediated process should be continuously monitored to evaluate their real effects on the viability of natural populations5,17.
However, to date, even in the era of genomics4,18, the concept of hybrid itself is rather fleeting and, consequently, legal status and management of hybrids are often poorly regulated by national and international laws, hampering the conservation of endangered species12,14,19,20,21,22 (for details about AH terms and definitions used through the paper see the Table 1 and the Supplementary Table S1a).
The management of individuals originated from AH is still debated5,16, especially since it is not always clear whether the removal of admixed individuals (Supplementary Table S1a) might represent an appropriate and feasible conservation strategy3,16,21,23,24, or which methods should be applied for their control (e.g. sterilization, captivation, lethal removal). Finally, it is still uncertain whether admixed individuals originated from hybridization events that occurred several generations in the past should be targets of management actions10,16,24,25,26, especially when they retain likely adaptive variants (e.g. the black coat in wolves27 and the domestic goat MHC haplotypes in the Alpine ibex28).
However, solving these doubts is far from trivial. In fact, while identifying first generation (F1) hybrids might be relatively simple, recognizing admixed individuals originated from recurrent or ancient hybridization events, as well as different ranges of generations of admixture, is a demanding or a virtually impossible task5,11. Thus, it would be highly beneficial to introduce standardized criteria to classify admixed individuals based on their actual level of risk to spread alien or domestic variants, allowing to prioritize management actions, more efficiently allocate efforts and resources, and support long-term conservation plans3,5,14,23.
Nevertheless, the detection of hybrids and backcrosses (Supplementary Table S1a) can be further complicated by a number of technical or statistical issues. Many hybridizing taxa are elusive and difficult to detect, therefore hybridization monitoring programs are usually carried out relying on opportunistically-collected and often degraded biological materials23,29,30,31,32,33,34,35,36,37. These approaches limit the use of more diagnostic large panels of markers such as Single Nucleotide Polymorphisms (SNPs) and the employment of more than a few tens of microsatellites (or STRs, Short Tandem Repeats), which generally allow the detection of hybridization events occurred only up to two or three generations in the past5,38,39. Another critical point is represented by the selection of appropriate reference populations to use in the assignment analyses, that should include a sufficient number of pure individuals representative of the genetic variability of the analysed taxa. Furthermore, particular attention should be reserved to the choice of adequate statistical computations for the assignment procedures, in order to minimise the risk that individual assignment probabilities might be affected by the allele frequencies of other samples to be assigned. In addition, the selection of statistically supported assignment q-thresholds based on individual assignment (qi-values) extrapolated from simulated data is necessary to ensure the realistic and reliable identification of admixed individuals5,16,40,41,42.
Therefore, using both empirical and simulated data, in this study we developed a simple but effective workflow (Supplementary Fig. S1) to overcome some of the aforementioned issues and to reliably detect different levels of introgression43, maximizing the identification of pure and admixed individuals, either hybrids or backcrosses (Supplementary Table S1a). We then tested this approach on an interesting example of AH16,26, taking place between the wolf (Canis lupus) and its domestic counterpart, the dog (C. l. familiaris). In particular, we investigated samples from the Italian wolf population (C. l. italicus), which experienced a demographic scenario characterized by protracted isolation south of the Alps and recurrent bottlenecks that made Italian wolves sharply genetically differentiated from any other wolf population4,24,26,44,45,46,47. Although the Italian wolf population is now recovering thanks to legal protection and the increased availability of suitable habitats and prey48, it is still threatened by accidental or illegal killings49,50, but also by anthropogenic hybridization, as repeatedly documented by genetic and genomic data showing gene flow from the domestic to the wild subspecies24,25,26,31,33,37,51,52.
Specifically, we used this case-study to:
-
1)
delineate strict criteria for choosing the most appropriate reference parental populations;
-
2)
based on simulated data, determine adequate and reliable assignment q-thresholds to consistently classify individuals into discrete levels of domestic ancestry;
-
3)
apply a standardized and stable Bayesian method to probabilistically assign unknown individuals to one of the ancestry classes;
-
4)
accordingly, define appropriate management categories to prioritize possible mitigation actions.
Results
Selection of the reference populations
Following strict morphological, genetic and genomic criteria for sample selection (see Materials and Methods), we retained from the ISPRA (Italian Institute for Environmental Protection and Research) canid database the genotypes of 190 wolves and 89 dogs typed at a panel of 39 STRs commonly used to reconstruct individual genotypes in some of the most recent studies on wolf x dog hybridization in Europe26,33,47,53. Selected individuals showed no missing data nor the occurrence of allelic dropout and false alleles, thus they were assumed as reference genotypes in the downstream analyses and were used in HybridLab54 to simulate 100 genotypes of wild (PW) and domestic (PD) parentals, first (F1) and second (F2) generation hybrids, and eight backcross generations (BC1W-BC8W) with wild parentals (Supplementary Table S1b).
Evaluation of the relative reliability and replicability of the Bayesian approaches
Results from the four independent Bayesian clustering runs obtained analysing the 1,479 canid individuals (including both reference and simulated genotypes) at K = 2 with the A and I models showed that the “one-by-one” assignment method, implemented in Parallel Structure, confirmed to be highly stable55, with an average variation of only 0.0074 (±0.0085 SD) in individual coefficient values (qi) among runs. This low variation allowed us to present outcomes from the first run, without the need to condense the results from multiple runs as it is usually needed when dealing with larger variations.
Membership proportions and individual coefficients from the assignment tests
We used the assignment results produced by Parallel Structure55 to estimate the average membership proportions (Qi) and individual coefficients (qi) of each predefined group (Fig. 1 and Table 1). We also estimated 90% credibility intervals (CI) for both Qi and qi.
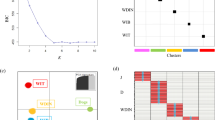
(a) Bar plotting of the individual qi-values obtained assigning the 39-STR genotypes of the simulated wild (PW) and domestic (PD) parentals, first (F1) and second (F2) generation hybrids, and eight backcross generations (BC1W-BC8W) with wild parentals to the wolf (RW) and dog (RD) reference populations. Each individual is represented by a vertical line partitioned into colored segments, whose length is proportional to the individual coefficients of membership (qi) to the wolf and dog clusters inferred by a Bayesian assignment analyses performed by the software Parallel Structure, assuming K = 2 clusters and using the “Admixture” and “Independent allele frequencies” models. (b) Posterior probabilities estimated, for the 39-STR genotypes of the simulated wild (PW) and domestic (PD) parentals, first (F1) and second (F2) generation hybrids, and eight backcross generations (BC1W-BC8W) with wild parentals to the wolf (RW) and dog (RD) reference populations, using the software NewHybrids with the “Jeffreys-like” priors for both mixing proportions and allele frequencies. Each individual is represented by a horizontal bar divided in five segments corresponding to its probability to belong to five classes: wild and domestic parentals (PW and PD), F1, F2, and first backcrosses of F1 with wolves (BC1W).
All the wild reference individuals were probabilistically assigned to the same cluster I with Qw = 0.999 (CI = 0.996–1.000), and with individual qiw ranging from 0.995 (CI = 0.966–1.000) to 1.000 (CI = 0.998–1.000). All the domestic reference individuals were assigned to the same cluster II with Qd = 0.998 (CI = 0.991–1.000), and with domestic qid ranging between 0.993 (CI = 0.949–1.000) and 1.000 (CI = 0.998–1.000) (Fig. 1a and Table 2).
The wild and domestic simulated parental populations showed Qi and qi-values almost completely overlapping with those of the wild and domestic reference populations, and were assigned to their respective clusters with qiw ≥ 0.995 for the wild and qid ≥ 0.989 for the domestic parentals (Fig. 1a).
Simulated F1 and F2 showed, as expected, intermediate Qi-values (F1: Qw = 0.515, CI = 0.384–0.644; F2: Qw = 0.524, CI = 0.393–0.652), while individual qi were 0.415 (CI = 0.282–0.551) ≤ qiw ≤ 0.661 (CI = 0.525–0.786) for F1 and 0.357 (CI = 0.222–0.496) ≤ qiw ≤ 0.702 (CI = 0.579–0.814) for F2. Backcrossed genotypes showed variable Qi-values that started to completely overlap to those of the parental populations in BC6W (Fig. 1a and Table 3), though a partial overlapping (up to 7%) was observed already from BC2W (Fig. 1a).
Interestingly, as expected, CI values were overall significantly negatively correlated (r = −0.91; P < 0.0001) with qiw-values (Supplementary Fig. S2a), and their mean widths in the simulated admixed individuals were on average significantly larger (t = 82.4, P < 0.0001; t-test) than in parental wolves (Supplementary Fig. S2a).
Bayesian clustering results obtained from Parallel Structure55 analysing a reduced set of 12 STRs, commonly utilized for genotyping low-content DNA samples in non-invasive genetic monitoring projects52,56, showed that, even though all reference parental genotypes were fully assigned to their clusters, the 12 STRs provided a lower resolution in detecting backcrosses. Indeed, 4% of BC1W showed a partial qiw overlap to those of the wild parental population (details are described in the Supplementary Text S1).
The assignment results were robust even when simulating increasingly high levels of allelic dropout (ADO) and missing data, showing less than 2% discrepancy of the individual qi-values even with 30% ADO and missing loci at 39 STRs, and less than 4% at 12 STRs. When we considered 10% ADO and missing data, discrepancies were less than 1% at 39 STRs and less than 3% at 12 STRs.
Selection and performance of the appropriate thresholds
Accuracy, efficiency and performance38 were calculated for different candidate q-thresholds ranging from 0.500 to 0.999 (Fig. 2 and Supplementary Table S2) and each q-threshold was tested between groups of simulated individuals at increasing levels of admixture (e.g. PW vs. BC8W to BCW1, F2 & F1; PW & BC8W vs. BC7W to BCW1, F2 & F1; PW, BC8W & BC7W vs. BC6W to BCW1, F2 & F1; PW, BC8W, BC7W & BC6W vs. BC5W to BCW1, F2 & F1; PW, BC8W, BC7W, BC6W & BC5W vs. BC4W to BCW1, F2 & F1, etc.). A performance higher than 0.90 was obtained only for three category combinations: PW & BCW8 to BCW1 vs. F1 & F2 (best performance = 0.982 at q-threshold = 0.670), PW & BCW8 to BCW2 vs. BC1W, F2 & F1 (best performance = 0.922 at q-threshold = 0.840) and PW & BCW8 to BCW3 vs. BC2W, BC1W, F2 & F1 (best performance = 0.900 at q-thresholds = 0.950–0.955). Therefore, to be as conservative as possible, we decided to retain the highest q-threshold value (0.955) of the latter category combination to efficiently discriminate between recent (F1-BC2W) and older admixed individuals (from BC3W onwards). This q-threshold was able to correctly identify a larger portion (40%) than the other highly performing q-thresholds (39% for a q-threshold = 0.950 for the same categories, 31% for a q-threshold = 0.840 and 20% for q-threshold = 0.670) of the 1000 simulated admixed genotypes, maximizing the recognition of recent admixed individuals (Fig. 3 and Table 3). At this q-threshold, indeed, 100% of the wolf x dog F1, F2, BC1W and 71% of BC2W were coherently classified as recent admixed individuals.
Graphical trends of the average performances (on the y-axis) estimated for increasing values of q-thresholds (on the x-axis). Each performance was computed as the product between the mean efficiency (the ratio of the number of admixed individuals correctly identified on the total number of admixed individuals actually included in the sample) and the accuracy (the number of admixed individuals correctly assigned to a certain admixture class on the total number of individuals actually belonging to that class) obtained considering individual qi-values of the simulated 39-STR genotypes estimated from the Bayesian assignment analyses performed in Parallel Structure, assuming K = 2 clusters and using the “Admixture” and “Independent allele frequencies” models. Each q-threshold was tested considering comparisons between groups (Rn) including simulated individuals for increasing levels of admixture. R1: PW, BC8W to BC1W & F2; R2: PW, BC8W to BC1W; R3: F1 & F2; R4: PW, BC8W to BC2W; R5: BC1W, F2 & F1; R6: PW, BC8W to BC3W; R7: BC2W to BC1W, F2 & F1; R8: PW, BC8W to BC4W; R9: BC3W to BC1W, F2 & F1; R10: PW, BC8W to BC5W; R11: BC4W to BC1W, F2 & F1; R12: PW, BC8W to BC6W; R13: BC5W to BC1W, F2 & F1; R14: PW, BC8W to BC7W; R15: BC6W to BC1W, F2 & F1; R16: PW & BC8W; R17: BC7W to BC1W, F2 & F1; R18: BC8W to BC1W, F2 & F1. The vertical red line identifies the q-threshold (qi = 0.955) selected on the basis of the performance analysis carried out comparing R6 vs. R7 (yellow line).
Box plots of individual qiw-values (on the y-axis) observed in parental and simulated 39-STR genotypes (on the x-axis) estimated from the Bayesian assignment analyses performed in Parallel Structure, assuming K = 2 clusters and using the “Admixture” and “Independent allele frequencies” models. White solid boxes include 90% of the observed data values. Dashed gray boxes contain the 5th percentile and the 95th percentile of the observed data values. Black dots indicate mean data values. Middle transversal lines inside boxes show median data values (the 50th percentile). Box plot wiskers include the ranges of the confidence intervals. The dashed line represents the q-threshold of 0.955, selected on the basis of the performance analysis (see Supplementary Table S2), which, minimizing the risk of both type I and type II errors, is able to efficiently discriminate between recent admixed (F1-BC2W) and older admixed (BC3W-BC8W) individuals. The red line indicates the q-threshold identified at 0.995, value corresponding to the minimum individual qiw of both simulated and real wolf parentals (see Table 2), which, minimizing the risk of both type I and type II errors, is able to efficiently discriminate between older admixed (BC3W-BC8W) and pure individuals.
However, since none of the highly-performing q-thresholds was able to reliably discriminate between older admixed and pure individuals, we introduced a second q-threshold at 0.995, representing the minimum individual qiw assignment value of the simulated and real wild parentals. Therefore, we assumed that the assignment interval in the range 0.955-0.995 could include older admixed individuals, showing only a marginal dog ancestry (<5%). In this way, 22% of BC2W were classified as older admixed individuals, together with 40% of BC3W, 21% of BC4W, 7% of BC5W, 1% of BC6W and 1% of BC7W (Table 3). Above this second q-threshold type I errors confirmed to be absent but type II errors were further minimized since we found 7% of BC2W, 40% BC3W, 76% BC4W, and more than 90% of BC5W-BC8W clustering together with reference and simulated wolf parentals (Table 3).
When we considered their 90% confidence intervals, their mean widths in the older admixed individuals were significantly larger than in pure wolves (t = 61.1, P < 0.0001; t-test) as well as they were significantly larger in recent admixed than in older admixed (t = 38.5, P < 0.0001; t-test) individuals (Supplementary Fig. S2b). Additionally, all individuals from BC3W to BC8W showed CI values higher than 0.955, thus representing an additional criterion to identify recent admixed individuals5,57.
Also when we considered the reduced 12-marker panel, we correspondingly retained two q-thresholds: the first value of 0.975 was chosen since it efficiently discriminated between recently admixed individuals (F1-BC2W) and all the other simulated classes, including older admixed (from BC3W onwards) plus pure (PW) individuals. A second q-threshold of 0.990, representing the minimum individual qiw assignment value of the simulated and real wild parentals, was selected to reliably discriminate between older admixed and pure individuals (for details see the Supplementary Text S1).
The four replicated runs in NewHybrids58 showed almost identical outcomes using both “Jeffreys-like” (t ≥ 0.006, P ≥ 0.91; t-tests for all pairwise combinations) and “Uniform” (t ≥ 0.032, P ≥ 0.97; t-tests) priors, with no significant differences even between the two models (t = 0.45, P = 0.65; t-test between average values of the four runs), therefore we decided to present only the results from the first NewHybrids run obtained with the “Jeffreys-like” priors (Table 2). Overall, the Bayesian assignment performed in NewHybrids58 also proved to be efficient (Fig. 1b), showing proportions of real and simulated samples correctly assigned to their own categories up to BC1W, not significantly (\({\chi }_{6}^{2}=1.74\), P = 0.94; χ2-test) different from the proportions achieved using Parallel Structure (Table 3), despite the very different assumptions they rely on55,58. All wild and domestic references and all wild and domestic simulated parentals had the best posterior probabilities (P ≥ 0.999) to be purebred animals (Table 2). Most F1 (98.3%), F2 (92.2%) and BC1W (98.4%) were clearly assigned to their own categories (P ≥ 0.900) showing posterior probabilities to belong to pure wolves or dogs always <0.001 (Fig. 1b). Interestingly and coherently with the detection power of the software58, looking at BC2W 60% of them were misclassified as BC1W and 40% as pure wild parentals (Table 2 and Table 3). Additionally, 88% BC3W, 99% BC4W and all the other BCW showed significant posterior probabilities (P ≥ 0.900) to be pure wolves (Table 2 and Table 3).
However, when we tried to extend NewHybrids assignment to classes older than BC1W (from BC2W to BC8W), results were highly different from expected since both real and simulated genotypes were never clearly attributed to their own genotypic categories, with the only exceptions of F1 and F2 individuals (Supplementary Table S3).
Application to real data
The application of the selected q-thresholds and Bayesian methodology to the 39-STR real genotypes (in which neither missing data nor allelic dropout were ever detected) belonging to 569 putative wolves collected from 1987 to 2019 throughout the whole wolf distribution range in Italy26,33,47,52 highlighted that 12.7% of them were diagnosable as recent admixed individuals (qiw < 0.955), that we thus assigned to the management class of operational hybrids. Another 13.5% were diagnosable as older admixed individuals (0.955 ≤ qiw < 0.995), thus operationally classified as introgressed individuals (Table 1 and Table 4). Conversely, the remaining 73.8% of the analysed real genotypes were identified as assignment-pure wolves (qiw ≥ 0.995), thus falling in the management class of the operational pure individuals (Table 1 and Table 4).
When real data assignments were completed with uniparental and coding markers (mtDNA, Y-STRs and K-locus), 68.0% of the analysed real genotypes did not show any traces of dog ancestry (Table 4). In particular, none of the individuals identified as operational pure animals showed dog mtDNA haplotypes, whereas 6% of them (corresponding to 10% of the pure males) showed dog Y-STR haplotypes and 3% had the melanistic 3-bp deletion (Table 4). These animals could represent additional older admixed individuals retaining domestic alleles in other genetic markers not included in the nuclear STR-based workflow. Interestingly, another 19 operational pure individuals (4.5%) showed dog-like phenotypic traits (white claws and/or spur on the hind legs26), which were not genetically detected.
When we applied the selected q-thresholds of 0.975 and 0.990 to the 12-STR genotypes of the real 569 putative wolves, the percentages of operational pure animals and of operational hybrids respectively increased to 77.% and 15.5%, whereas the percentage of introgressed individuals decreased to 7.1% since a part of them was misclassified as pure and another part was misclassified as recent admixed individuals (for details see the Supplementary Text S1).
Discussion
While natural hybridization has been widely acknowledged as a powerful evolutionary force6,7, during last decades anthropogenic hybridization considerably contributed to threat the genomic integrity and survival of a number of taxa through the introgression of alien or domestic alleles in the gene pool of natural populations3,11,12,15,41,42,59. In particular, though some studies documented cases of beneficial introgression of domestic mutations in wild populations of North American wolves27 and Alpine ibexes28, introgressive hybridization with domestic forms is globally recognized as a significant risk factor for the conservation of several wild taxa14,24,28,60,61,62. However, though being essential to understand the real impact of the phenomenon and to design sound conservation strategies16,23, the identification of hybrids and their backcrosses remains far from trivial even in the genomic era3,4,5,10,13. In the common practice, the domestic ancestry of biological samples is usually assessed typing their DNA at presumably neutral molecular markers and probabilistically assigning the obtained genotypes to reference parental populations by Bayesian statistics57,63. Consequently, Bayesian assignment values (qi-values) are considered key parameters for management initiatives5,16 and well relate to genomic proportions of parental ancestry estimated by genomic approaches such as the PCA-based admixture deconvolution methods26,64,65.
Nonetheless, detecting admixture signals between subspecies sharing a very recent common ancestry is often hampered by the difficulty to a priori identify pure individuals24,26 and a number of pitfalls may sway the analyses, thus strict criteria should be applied for a reliable identification of admixed individuals: (1) reference parental populations should be composed by the genetic profiles of a sufficient number of individuals (e.g. at least 40 for each reference population5), obtained through the genotyping of high-quality samples at a large number of markers, and lacking any genetic - and possibly morphological - signature of hybrid ancestry; (2) qi-values of unclassified individuals should be estimated by assigning them to parental populations through a repeatable and standardized Bayesian statistical approach; (3) the a posteriori classification of individuals should be based on q-thresholds previously established from the distribution of qi-values observed in simulated genotypes5,33,38.
In this study, we implemented a rapid and efficient standardized workflow (Supplementary Fig. S1) to molecularly detect and classify different levels of admixture in individuals belonging to the Italian wolf population (C. l. italicus), a taxon in which wild x domestic hybridization has been repeatedly documented24,25,26,31,33,37,51,66,67.
The selection of a sufficient number of non-admixed parental individuals to use as reference populations in the assignment analyses was made possible by testing a large national database that includes hundreds of individuals sampled from the entire subspecies distribution range, which had been all formerly morphologically described and molecularly characterized at different sets of genome-wide (STRs and SNPs) markers26,33,52. Therefore, initiatives aiming at systematically collecting population-wide samples of target species should be strongly sustained by national or local authorities, possibly including also samples from nearby populations in order to take into account possible gene flows22,68 and, whenever achievable, detailed information on possible phenotypical anomalies5,24,26.
The simulation of hybrid and backcrossed genotypes, as well as a sufficient number of ancestry-informative markers able to discriminate even closely-related species or subspecies, is then required in order to establish reliable q-thresholds discriminating between different levels of admixture classes5,18,38.
In addition, stable statistical Bayesian approaches, such as that implemented in Parallel Structure55, are strongly recommended to minimize the risk of biased assignment probabilities to an a priori assumed number of populations40, which might occur when sample sizes vary among analyses or when unknown samples with variable levels of admixture (namely including both pure and admixed individuals) are analysed simultaneously instead of one by one40,69,70, conversely to other fully (Structure, NewHybrids, Baps) or partially (GeneClass) Bayesian assignment methods commonly applied for admixture identifications33,41,42,71,72,73,74,75,76. As expected, the “one-by-one” approach with Parallel Structure55 performed reliably, with very limited fluctuations of both Qi and qi among different replicates of the same runs. Up to BC1W, results were also highly concordant with the results obtained from the assignment method implemented in NewHybrids58, despite the very different assumptions and algorithms the two approaches rely on55,58.
Though anthropogenic hybridization has been deeply investigated for a number of animal species, only a few studies applied reliable statistical criteria to define adequate assignment q-thresholds to correctly identify non-admixed individuals and distinguish different admixture classes[1,41,42,73,77. Conversely, most genetic investigations about hybridization in canids were mainly based on q-thresholds selected arbitrarily or chosen among those widely used in the literature (e.g. Malde et al.41) and rarely using simulated data to estimate error rates associated to the choice of a certain threshold31,33,37,66,68,74. A third challenge is thus represented by the adoption of objective criteria based on a Performance Analysis38 for setting the most appropriate q-thresholds to classify individuals into different admixture classes (e.g. pure vs. older admixed vs. recent admixed individuals) that could result into different management categories (e.g. operational pure, introgressed and operational hybrid individuals), minimizing the risk of both type I (pure individuals erroneously identified as admixed animals) and type II (admixed individuals falsely identified as pure animals) errors5,12,16,33,38.
Analysing the 39-STR marker panel, our assignment values appeared strongly robust even when introducing increasingly high levels of allelic dropout and missing data, nonetheless we remind that stringent filters on the quality and reliability of multilocus genotypes are essential to avoid significant biases in all downstream analyses. Our first selected q-threshold allowed us to correctly classify as admixed 100% of F1, F2, BC1W and 71% of BC2W, without any type I error. The remaining 29% of BC2W were classified as pure individuals likely due to a combination of: (i) higher mean qiw, closer to the identified q-threshold (0.955) compared to earlier generations of backcrossing (F1, F2, and BC1W), and (ii) wider CI compared to further generations of backcrossing (BC3W, BC4W, etc.).
Further backcrossing categories showed increasing percentages of assignment as pure individuals (40% in BC3W and 76% in BC4W), clearly showing the limits of the method in our study system when dealing with older backcrossing generations.
Nonetheless, the second empirical q-threshold allowed us to reliably discriminate also between real pure wolves and older admixed individuals, that only show a marginal dog ancestry and possibly deserve additional investigations.
Our results agree with other hybridization studies based on a comparable number of microsatellites, which highlighted the difficulty to reliably detect individuals with a domestic ancestry tracing back to more than two-three generations in the past5,31,33,42,72,78.
When the selected q-thresholds obtained with the 39-STR panel were applied to a large sample (c. 600 genotypes) of putative free-living wolves collected in Italy during the last 20 years, 73.8% of the analysed genotypes resulted operational pure animals (i.e. without relevant signs of domestic ancestor), while 13.5% were classifiable as introgressed individuals and 12.7% as operational hybrids, compatible with multiple and recurrent admixture events that might have occurred trough time, mostly during the phase of population re-expansion26,31,33. However, as shown by simulated data and confirmed by the genetic information derived from the analysis of the uniparental and coding markers, the operational pure category might include a proportion (in our case, 5.8%) of older admixed individuals not reliably detectable using the applied set of molecular markers.
Nonetheless, these percentages of admixed individuals cannot be intended as estimates of prevalence of admixed individuals in the Italian wolf population because the analysed samples had not been randomly collected, but mostly derived from specific monitoring projects focused on hybrid detection and from heterogeneously monitored areas26,31,33,52,56. Conversely, reliable estimates of hybridization prevalence could be assessed through statistical multi-event models applied to capture-recapture data obtained from well-planned long-term genetic and camera-trapping monitoring projects carried out through the entire Italian wolf distribution range79,80,81.
Despite 39 STRs represent a very limited portion of the genetic makeup of the analysed individuals that could be routinely applied to wide monitoring programs, the assignment values of recently-admixed individuals well correlate with those obtained from thousands of genome-wide markers26.
From a management perspective, known limits and efficiency in identifying different admixture classes allow to conceive corresponding management categories as robust as possible. However, a complication in the management of hybrids and backcrosses arises from the use of ambiguous or imprecise terminologies for defining different classes of admixed individuals. Therefore, in this study, we propose to categorize admixed individuals on the basis of empirically-defined q-thresholds, where “operational hybrids” correspond to recent admixed individuals (that include F1-F2 hybrids and most of the first two generations of backcrosses), while “operational pure individuals” correspond either to pure wolves or to older admixed individuals that could not be reliably distinguished from pure ones with the applied panel of molecular markers, but may retain marginal dog ancestry. Between them, we proposed an intermediate assignment class which mostly includes older admixed individuals that cannot be considered as operational pure animals, but do not require priority management actions given their limited domestic ancestry.
Given that hybridization should be primarily counteracted by (i) preventive measures aimed at reducing the number of free-ranging dogs, and (ii) proactive strategies to preserve prey availability, social cohesion, structure and connectivity of wolf packs, since habitat loss, rapid pack turnovers and recent population expansions are known to favor hybridization82, the proposed categorization would permit to avoid management interventions on pure animals erroneously classified as admixed individuals and their negative effects on the genetic and demographic viability of small or threatened wild populations26,47,49,50. Moreover, this categorization would allow to better focus efforts and resources toward “operational hybrids”, which carry significant portions of domestic genome ancestry and likely belong to the first generations of admixture, more efficiently than without any prioritization (e.g. genetically speaking, the removal of one hybrid with 50% dog ancestry would equal to the biological removal of 10 admixed individuals with 5% dog ancestry).
However, in those cases where an active management on operational hybrids is needed, the social acceptability of the applicable methods should be carefully considered, possibly avoiding controversial interventions such as lethal removal3,16,82. Indeed, among other more acceptable management methods, life-long captivation in welfare-respectful structures or sterilization and release of admixed individuals might represent feasible mitigation strategies16,23.
On the other side, the active management of introgressed individuals might become a necessary option where they locally occur at a high prevalence (that can be sometimes much higher than region- or population-wide estimates), thus increasing the probability of interbreeding between hybrids and retaining domestic variants on the long term81,82.
Conversely, dog-derived phenotypic traits, though validated by robust phenotype-genotype association tests26, when found in operational pure individuals should not be considered sufficient reasons for any intervention, since they might reflect old introgression events. Nonetheless they could represent useful clues for identifying potential hybrids with preliminary field surveying methods, such as camera trapping79,80,83, to be followed by further careful genetic investigations.
These classes appear to be more suitable for practical and management purposes compared to categories based on the supposed hybrid generations that, unless they are formally estimated based on genome-wide data26, are largely hazardous since a virtually infinite number of hybrid classes exists, with individual membership proportions widely overlapping.
These findings, together with the results derived from the analyses performed with our 12-STR marker panel, suggest that reduced molecular marker sets and empirical assignment q-thresholds can represent an effective first approach to orientate the most appropriate management actions.
Moreover, the recent possibility to access genome-wide SNP data to investigate anthropogenic hybridisation in a number of taxa7,41,61, including canids24,26,44,77, allows to gain a better resolution on the domestic ancestry proportions and to infer the real generations since the hybridization events26,64,84, that could be needed for the discrimination between real pure and older admixed individuals. Subsequently, the selection of reduced panels of ancestry-informative SNPs, including both neutral and coding mutations26, diagnosable by quantitative or microfluidic PCR techniques77,85,86,87, could be particularly suitable for cost-effective future monitoring projects based on the genotyping of invasive and non-invasive samples to be collected with a standardized design in hybridization hot-spots.
Our workflow, though designed on the case-study of the Italian wolf population, could be easily adapted to monitor the status of other populations and species potentially threatened by anthropogenic hybridization, although each study should adopt ad-hoc q-thresholds, based on the genetic distance between wild and domestic reference populations, their genetic diversity and possible substructure, but also on the number and type of analysed molecular markers. Moreover, when gene flow is known to occur between multiple wild populations (e.g. in Northeastern Alps and Carpathian Mountains88,89,90), the number of reference populations and the optimal number of genetic clusters K should be modified accordingly, in order to avoid the identification of false wild x domestic hybrids (type I errors). Nonetheless, we also remind that such complex systems also require large parental populations to be used as reference. Of course, such an effort is worth using only when dealing with complex levels of admixture, whereas for simpler systems (e.g. when a few individuals could be assigned to recent crosses (F1, F2) or backcrosses (BC1)) standard approaches are sufficient.
In conclusion, the identification of operational categories based on admixture classes outlined through simulations can support scientists, practitioners and decision-makers in the implementation of more efficient conservation strategies mostly focusing on recent hybrids, whose diffusion and consequent spread of domestic alleles could be limited by active management actions to be defined upon local context and acceptance levels toward the presence of free-ranging admixed individuals, but taking into account that nonlethal actions such as captivation or sterilization are often considered by scientists and the public opinion as more feasible and ethically acceptable conservation tools16.
Materials and Methods
Ethical statements
No ethics permit was required for this study, and no animal research ethics committee prospectively was needed to approve this research or grant a formal waiver of ethics approval since the collection of wolf samples involved dead animals. Fieldwork procedures were specifically approved by ISPRA as a part of national wolf monitoring activities.
Dog blood samples were collected by veterinarians during health examinations with a not-written (verbal) consent of their owners (students/National park volunteers/or specialised technician personnel of the Italian Forestry Authority (CFS)), since they were interested on wolf conservation studies and monitoring projects in Italy. Moreover there is not a relevant local law/legislation that exempts our study from this requirement.
Additionally no anesthesia, euthanasia, or any kind of animal sacrifice was applied for this study and all blood samples were obtained aiming at minimizing the animal suffering.
Selection of the reference populations
Reference wild parentals were selected from found-dead wolves collected across the Italian peninsular distribution range that showed the typical wild coat colour pattern and no other apparent dog-like traits such as white claws or spurs on the hind legs26,31,33,91.
Reference domestic parentals were selected from free-ranging mongrels and village dogs sampled in the same areas of the reference wolves, plus one male and one female randomly chosen from 14 wolf-sized dog breeds. Given the high between-breed variation92 these samples could represent a good proxy of the diversity in dogs while avoiding significant sub-structuring during clustering analyses26,33,47,93.
As wild and domestic reference individuals, all available in the ISPRA canid database26,33,52,56, we only retained those whose genotypes showed no missing data and proportions of membership qi > 0.990 to the respective wild or domestic clusters estimated in previous Bayesian assignment procedures performed, using the software Structure v.2.3.457,94, on 39 canine STRs commonly used to reconstruct individual genotypes in some of the most recent studies on wolf x dog hybridization in Europe26,33,47,53. This conservative q-threshold was selected to avoid the inclusion of older admixed individuals among the wild reference population, thus reducing the power to correctly identify admixed individuals in the tested dataset. Furthermore, 90 of the selected reference wolves and 30 of the selected reference dogs were also tested in Maximum-Likelihood assignment procedures performed analysing 156 K genome-wide canine SNPs in the software Admixture v.1.2395 and confirmed their pure status showing qi > 0.99026.
Simulation of pure and admixed populations
Reference samples were used in HybridLab54 to simulate 100 genotypes (a sufficient number to well represent the parental allele frequencies40) for each of the following pure and admixed classes: wild (PW) and domestic (PD) parentals, first (F1) and second (F2) generation hybrids, and eight backcross generations (BC1W-BC8W) with wild parentals (Supplementary Table S1b). In a selectively neutral perspective, BC8W individuals should theoretically retain less than 0.2% of the domestic parental ancestry (Supplementary Table S1b). Simulations were performed both with the complete set of 39 STRs33,47 and with a reduced set of 12 STRs commonly utilized for genotyping low-content DNA samples through a multiple-tube approach in non-invasive genetic monitoring projects52,56.
Bayesian assignment tests
To perform admixture analyses and assign individuals to their reference populations, empirical and simulated multilocus microsatellite genotypes were run using the R package Parallel Structure55, which uses the back-end executable of Structure57,94 parallelizing the Markov Chain Monte Carlo (MCMC) algorithm to: (i) distribute computation jobs among multiple processors, thus speeding up analysis times, and: (ii) automatically subdivide a dataset of genotypes to be assigned to predefined reference populations into multiple single projects (each project is composed by the reference populations and one of the genotypes to be assigned) which are independently run, preventing that sample sizes or the simultaneous analysis of samples with different levels of admixture might affect results69,70. Custom bash and excel macro scripts were designed to assembly output files, that are equal to the number of the analysed samples, and to create a single summary result file (See the Supplementary Fig. S1, the Supplementary Text S2 and the Supplementary Table S7 for the detailed pipeline).
We ran four independent replicates of Parallel Structure with 5 × 105 iterations following a burn-in period of 5 × 104 iterations, using the Admixture (A) and Independent allele frequencies (I) models, which are the most suitable ones to investigate gene flow between populations with reasonably different allele frequencies and independently evolving66,94, and assuming K = 2 a priori clusters (corresponding to the optimal number of genetic clusters in which reference populations are split to identify the proportion of admixture33,52). For each group, we assessed the average proportion of membership (Qi) to the two clusters and individual assignments were based on proportions of membership (qi) estimated for every single individual. We also estimated 90% credibility intervals (CI) for both Qi and qi in order to evaluate their overlap between different admixture categories and their individual width, expecting wider CI in the assignment of admixed individuals due to difficulties in estimating parental allele frequencies57,66,94.
In order to test the robustness of the assignment values under varying levels of genotyping errors and missing data, we simulated increasing levels of allelic dropout (ADO) and missing data (number of missing loci) for the 1200 simulated parental and admixed genotypes (both at 39 and 12 STRs) in Gimlet 1.3.396, assuming 10%, 20% and 30% for both parameters, then re-ran the assignment tests in Parallel Structure55 with the same settings.
The software NewHybrids58 was used to compute the posterior probabilities that each genotype belongs to each of the following five classes: wild and domestic parentals (PW and PD), first (F1) and second (F2) generation hybrids, and first backcrosses of F1 with wolves (BC1W). Posterior distributions were evaluated running four independent replicates of NewHybrids58 with 105 iterations of the Monte Carlo Markov chains, following a burn-in period of 104 iterations, without any individual or allele frequency prior information, and using “Jeffreys-like” or “Uniform” priors for both mixing proportions and allele frequencies58.
Criteria for the definition of admixture thresholds and assignment error rates
We tried to identify the most appropriate q-thresholds that were able to distinguish between pure, older admixed and recent admixed individuals (Table 1), while minimizing the risk of both type I (actually pure individuals erroneously identified as admixed animals) and type II (admixed individuals falsely identified as pure animals) errors12,16,26,33.
Therefore, we estimated the “performance” of different q-thresholds with intervals of 0.005, spanning from 0.500 to 0.999. In particular, each performance was computed as the product between the “mean efficiency”, which is the ratio of the number of admixed individuals correctly identified on the total number of admixed individuals actually included in the sample, and the “accuracy”, defined as the number of admixed individuals correctly assigned to a certain simulated admixture class on the total number of individuals actually belonging to that class38.
Each q-threshold was tested between groups of simulated individuals at increasing levels of admixture (e.g. PW vs. BC8W to BCW1, F2 & F1; PW & BC8W vs. BC7W to BCW1, F2 & F1; PW, BC8W & BC7W vs. BC6W to BCW1, F2 & F1; PW, BC8W, BC7W & BC6W vs. BC5W to BCW1, F2 & F1; PW, BC8W, BC7W, BC6W & BC5W vs. BC4W to BCW1, F2 & F1, etc.), considering a minimum performance of 0.90 for any combination to be retained38.
Application of the identified admixture thresholds to the management classification of tested samples
The selected q-thresholds were finally applied to classify the 39-STR canid genotypes obtained from the carcasses of 569 putative wolves (236 females and 333 males) collected from 1987 to 2019 throughout the whole wolf distribution range in Italy26,33,47,52. Extraction, amplification and post-amplification procedures were carried out in separate rooms reserved to low-template DNA samples following protocols described in Randi et al.33, Fabbri et al.52 and Caniglia et al.56. To check for the occurrence of allelic dropout and false alleles, samples were independently analysed twice for each locus. Negative (no DNA) and positive (samples with known genotypes) PCR controls were used to check for laboratory contaminations. Genotypes were accepted as reliable only when ADO and missing data were less than 10%33,52,56.
Assignments of the 39-STR canid genotypes were further integrated with the information derived from uniparental markers (mtDNA control region and four Y-linked STRs) and from the functional melanistic deletion at the β-defensin CBD103 gene (corresponding to the K-locus), which were used to provide the directionality of the hybridization and determine the presence of the atypical dog-derived black coat coloration18,31,33,52.
Based on the assignment results, both simulated (pure and admixed) and real 39-STR canid genotypes were classified into three appropriate management classes (Table 1): “operational pure individuals” (including pure wolves and admixed individuals with a negligible dog ancestry, that do not require management actions), “introgressed individuals” (likely old admixed individuals with a marginal domestic ancestry that only require low priority management actions, such as further investigations) and “operational hybrids” (recent admixed individuals with a clearly detectable dog ancestry, that should be targeted by high priority management operations such as sterilization or captivation).
Data availability
The majority of the data generated and analysed during the current study are presented within the published article or in Supplementary information files. The raw data are available from the corresponding author on reasonable request.
References
Abbott, R. J., Barton, N. H. & Good, J. M. Genomics of hybridization and its evolutionary consequences. Mol. Ecol. 25, 2325–2332 (2016).
Gompert, Z. & Buerkle, C. A. What, if anything, are hybrids: enduring truths and challenges associated with population structure and gene flow. Evol. Appl. 9, 909–923 (2016).
Wayne, R. K. & Shaffer, H. B. Hybridization and endangered species protection in the molecular era. Mol. Ecol. 25, 2680–2689 (2016).
Gómez-Sánchez, D. et al. On the path to extinction: inbreeding and admixture in a declining grey wolf population. Mol. Ecol. 27, 3599–3612 (2018).
McFarlane, S. E. & Pemberton, J. M. Detecting the true extent of introgression during anthropogenic hybridization. Trends Ecol. Evol. xx, 1–12, https://doi.org/10.1016/j.tree.2018.12.013 (2019).
Lavrenchenko, L. A. & Bulatova, N. S. The role of hybrid zones in speciation: a case study on chromosome races of the house mouse Mus domesticus and common shrew Sorex araneus. Biol. Bull. Rev. 6, 232–244 (2016).
Irisarri, I. et al. Phylogenomics uncovers early hybridization and adaptive loci shaping the radiation of Lake Tanganyika cichlid fishes. Nat. Commun. 9, 3159 (2018).
Allendorf, F. W. & Luikart, G. Conservation and the genetics of populations. Blackwell Publishing, Malden, Massachusetts, USA (2007).
Brennan, A., Cross, P. C. & Creel, S. Managing more than the mean: using quantile regression to identify factors related to large elk groups. J. Appl. Ecol. 52, 1656–1664 (2015).
Grabenstein, K. C. & Taylor, S. A. Breaking barriers: causes, consequences, and experimental utility of human-mediated hybridization. Trends Ecol. Evol. 33, 198–212 (2018).
Rhymer, J. M. & Simberloff, D. Extinction by hybridization and introgression. Annu. Rev. Ecol. Syst. 27, 83–109 (1996).
Allendorf, F. W., Leary, R. F., Spruell, P. & Wenburg, J. K. The problems with hybrids: setting conservation guidelines. Trends Ecol. Evol. 16, 613–622 (2001).
Crispo, E., Moore, J. S., Lee-Yaw, J. A., Gray, S. M. & Haller, B. C. Broken barriers: human-induced changes to gene flow and introgression in animals. BioEssays 33, 508–518 (2011).
Bohling, J. H. Strategies to address the conservation threats posed by hybridization and genetic introgression. Biol. Conserv. 203, 321–327 (2016).
Todesco, M. et al. Hybridization and extinction. Evol. Appl. 9(7), 892–908 (2016).
Donfrancesco, V. et al. Unravelling the Scientific Debate on How to Address Wolf-Dog Hybridization in Europe. Front. Ecol. Evol. 7, 175, https://doi.org/10.3389/fevo.2019.00175 (2019).
Schumer, M. et al. Natural selection interacts with recombination to shape the evolution of hybrid genomes. Science 360, 656–660, https://doi.org/10.1126/science.aar3684 (2018).
Anderson, T. M. et al. Molecular and evolutionary history of melanism in North American gray wolves. Science 323, 1339–1343 (2009).
Witzenberger, K. A. & Hochkirch, A. Ex situ conservation genetics: a review of molecular studies on the genetic consequences of captive breeding programmes for endangered animal species. Biodivers. Conserv. 20, 1843–1861 (2011).
Boscari, E. et al. Species and hybrid identification of sturgeon caviar: a new molecular approach to detect illegal trade. Mol. Ecol. Resour. 14, 489–498 (2014).
Trouwborst, A. The EU Habitats Directive and wolf conservation and management on the Iberian Peninsula: a legal perspective. Galemys 26, 15–30 (2014).
Peltola, T. & Heikkilä, J. Outlaws or protected? DNA, hybrids, and biopolitics in a Finnish wolf-poaching case. Soc. Anim. 26(2), https://doi.org/10.1163/15685306-12341509 (2018).
Godinho, R. et al. Real-time assessment of hybridization between wolves and dogs: combining noninvasive samples with ancestry informative markers. Mol. Ecol. Resour. 15, 317–328 (2015).
Pilot, M. et al. Widespread, long-term admixture between grey wolves and domestic dogs across Eurasia and its implications for the conservation status of hybrids. Evol. Appl. 11, 662–680, https://doi.org/10.1111/eva.12595 (2018).
Bassi, E. et al. Trophic overlap between wolves and free-ranging wolf × dog hybrids in the Apennine Mountains, Italy. Glob. Ecol. Conserv. 9, 39–49 (2017).
Galaverni, M. et al. Disentangling timing of admixture, patterns of introgression and phenotypic indicators in a hybridizing wolf population. Mol. Biol. Evol. 34, 2324–2339 (2017).
Coulson, T. et al. Modeling effects of environmental change on wolf population dynamics, trait evolution, and life history. Science 334, 1275–1278 (2011).
Grossen, C. et al. Introgression from domestic goat generated variation at the major histocompatibility complex of alpine ibex. PLoS Genetics 10(6), e10044385 (2014).
Adams, J. R., Lucash, C., Schutte, L. & Waits, L. P. Locating hybrid individuals in the red wolf (Canis rufus) experimental population area using a spatially targeted sampling strategy and faecal DNA genotyping. Mol. Ecol. 16(9), 1823–1834 (2007).
Bohling, J. H. & Waits, L. P. Assessing the prevalence of hybridization between sympatric Canis species surrounding the red wolf (Canis rufus) recovery area in North Carolina. Mol. Ecol. 20, 2142–2156 (2011).
Caniglia, R. et al. Black coats in an admixed wolf × dog pack is melanism an indicator of hybridization in wolves? Eur. J. Wildl. Res. 59, 543–555 (2013).
Nussberger, B., Wandeler, P. & Camenisch, G. A SNP chip to detect introgression in wildcats allows accurate genotyping of single hairs. Eur. J. Wildl. Res. 60, 405–410 (2014).
Randi, E. et al. Multilocus detection of wolf × dog hybridization in Italy, and guidelines for marker selection. PLoS One 9, e86409 (2014).
Afonso, E., Goydadin, A. C., Giraudoux, P. & Farny, G. Investigating hybridization between the two sibling bat species Myotis myotis and M. blythii from guano in a natural mixed maternity colony. PLoS One 12, e0170534–e0170534 (2017).
Mekonnen, A. et al. Population genetic structure and evolutionary history of Bale monkeys (Chlorocebus djamdjamensis) in the southern Ethiopian Highlands. BMC Evol. Biol. 18(1), 1–15, https://doi.org/10.1186/s12862-018-1217-y (2018).
Steyer, K., Tiesmeyer, A., Muñoz-Fuentes, V. & Nowak, C. Low rates of hybridization between European wildcats and domestic cats in a human-dominated landscape. Ecol. Evol. 8, 2290–2304, https://doi.org/10.1002/ece3.3650 (2018).
Dufresnes, C. et al. Two decades of non-invasive genetic monitoring of the grey wolves recolonizing the Alps support very limited dog introgression. Sci. Rep. 9, 148, https://doi.org/10.1038/s41598-018-37331-x (2019).
Vähä, J. P. & Primmer, C. R. Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol. Ecol. 15, 63–72 (2006).
Randi, E. Detecting hybridization between wild species and their domesticated relatives. Mol. Ecol. 17, 285–293 (2008).
Karlsson, S., Diserud, O. H., Moen, T. & Hindar, K. A standardized method for quantifying unidirectional genetic introgression. Ecol. Evol. 4(16), 3256–3263 (2014).
Malde, K. et al. Whole genome resequencing reveals diagnostic markers for investigating global migration and hybridization between minke whale species. BMC Genomics 18, 1–11 (2017).
van Wyk, A. M. et al. Quantitative evaluation of hybridization and the impact on biodiversity conservation. Ecol. Evol. 7, 320–330 (2017).
Wringe, B. F., Stanley, R. R. E., Jeffery, N. W., Anderson, E. C. & Bradbury, I. R. Hybrid detective: a workflow and package to facilitate the detection of hybridization using genomic data in R. Mol. Ecol. Resour. 17, e275–e284 (2017).
vonHoldt, B. M. et al. A genome-wide perspective on the evolutionary history of enigmatic wolf-like canids. Genome Res. 21, 1294–1305 (2011).
Stronen, A. V. & Paquet, P. C. Perspectives on the conservation of wild hybrids. Biol. Conserv. 167, 390–395 (2013).
Pilot, M. et al. Genome-wide signatures of population bottlenecks and diversifying selection in European wolves. Heredity 112, 428–442 (2014).
Montana, L. et al. Combining phylogenetic and demographic inferences to assess the origin of the genetic diversity in an isolated wolf population. PLoS One 12, e0176560 (2017).
Galaverni, M., Caniglia, R., Fabbri, E., Milanesi, P. & Randi, E. One, no one, or one hundred thousand: how many wolves are there currently in Italy? Mammal Res. 61, 13–24 (2016).
Caniglia, R., Fabbri, E., Greco, C., Galaverni, M. & Randi, E. Forensic DNA against wildlife poaching: Identification of a serial wolf killing in Italy. Forensic Sci. Int. Genet. 4, 334–338 (2010).
Imbert, C. et al. Why do wolves eat livestock? Factors influencing wolf diet in northern Italy. Biol. Conserv. 195, 156–168 (2016).
Lorenzini, R., Fanelli, R., Grifoni, G., Scholl, F. & Fico, R. Wolf-dog crossbreeding: “Smelling” a hybrid may not be easy. Mamm. Biol. 79, 149–156 (2014).
Fabbri, E. et al. From predation to management: monitoring wolf distribution and understanding depredation patterns from attacks on livestock. Hystrix, Ital. J. Mammal. 29, 101–110, https://doi.org/10.4404/hystrix-00070-2018 (2018).
Godinho, R. et al. Genetic evidence for multiple events of hybridization between wolves and domestic dogs in the Iberian Peninsula. Mol. Ecol. 20, 5154–5166 (2011).
Nielsen, E. E. G., Bach, L. A. & Kotlicki, P. Hybridlab (version 1.0): a program for generating simulated hybrids from population samples. Mol. Ecol. Notes 6, 971–973 (2006).
Besnier, F. & Glover, K. A. Parallel Structure: a R package to distribute parallel runs of the population genetics program Structure on multi-core computers. Plos One 8, e70651 (2013).
Caniglia, R., Fabbri, E., Galaverni, M., Milanesi, P. & Randi, E. Noninvasive sampling and genetic variability, pack structure, and dynamics in an expanding wolf population. J. Mammal. 95, 41–59 (2014).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–59 (2000).
Anderson, E. C. & Thompson, E. A. A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160, 1217–1229 (2002).
Laikre, L., Schwartz, M. K., Waples, R. S. & Ryman, N. Compromising genetic diversity in the wild: unmonitored large-scale release of plants and animals. Trends Ecol. Evol. 25, 520–529 (2010).
Mattucci, F. et al. Genetic structure of wildcat (Felis silvestris) populations in Italy. Ecol. Evol. 3(8), 2443–245 (2013).
Iacolina, L. et al. Hotspots of recent hybridization between pigs and wild boars in Europe. Sci. Rep. 8, 17372, https://doi.org/10.1038/s41598-018-35865-8 (2018).
Nussberger, B., Currat, M., Quilodran, C. S., Pronta, N. & Keller, L. F. Range expansion as an explanation for introgression in European wildcats. Biol. Conserv. 218, 49–56, https://doi.org/10.1016/j.biocon.2017.12.009 (2018).
Alexander, D. H. & Lange, K. Enhancements to the Admixture algorithm for individual ancestry estimation. BMC Bioinformatics 12, 246 (2011).
Brisbin, A. et al. PCAdmix: principal components-based assignment of ancestry along each chromosome in individuals with admixed ancestry from two or more populations. Hum. Biol. 84(4), 343–364 (2012).
Caniglia, R. et al. Wolf outside, dog inside? The genomic make-up of the Czechoslovakian Wolfdog. BMC Genomics 19, 533, https://doi.org/10.1186/s12864-018-4916-2 (2018).
Verardi, A., Lucchini, V. & Randi, E. Detecting introgressive hybridization between free-ranging domestic dogs and wild wolves (Canis lupus) by admixture linkage disequilibrium analysis. Mol. Ecol. 15, 2845–2855 (2006).
Iacolina, L. et al. Y-chromosome microsatellite variation in Italian wolves: a contribution to the study of wolf-dog hybridisation patterns. Mamm. Biol. 75, 341–347 (2010).
Kusak, J. et al. Wolf-dog hybridization in Croatia. Vet. Arh. 88, 375–395, https://doi.org/10.24099/vet.arhiv.170314 (2018).
Kalinowski, S. T. The computer program Structure does not reliably identify the main genetic clusters within species: simulations and implications for human population structure. Heredity 106(4), 625–632 (2011).
Wang, J. The computer program structure for assigning individuals to populations: easy to use but easier to misuse. Mol. Ecol. Resour. 17, 981–990 (2017).
Oliveira, R., Godinho, R., Randi, E., Ferrand, N. & Alves, P. C. Molecular analysis of hybridization between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv. Genet. 9, 1–11 (2008).
Sanz, N., Araguas, R. M., Fernández, R., Vera, M. & Garcìa-Marìn, J. L. Efficiency of markers and methods for detecting hybrids and introgression in stocked populations. Conserv. Genet. 10, 225–236 (2009).
Grobler, J. P. et al. Management of hybridization in an endemic species: decision making in the face of imperfect information in the case of the black wildebeest - Connochaetes gnou. Eur. J. Wildl. Res. 57, 997–1006 (2011).
Pacheco, C. et al. Spatial assessment of wolf-dog hybridization in a single breeding period. Sci. Rep. 7, 42475 (2017).
Vähä, J. P., Erkinaro, J., Falkegård, M., Orell, P. & Niemelä, E. Genetic stock identification of Atlantic salmon and its evaluation in a large population complex. Can. J. Fish. Aquat. Sci. 74, 327–338 (2016).
Vaz Pinto, P., Beja, P., Ferrand, N. & Godinho, R. Hybridization following population collapse in a critically endangered antelope. Sci. Rep. 6, 18788 (2016).
vonHoldt, B. M. et al. Identification of recent hybridization between gray wolves and domesticated dogs by SNP genotyping. Mamm. Genome 24, 80–88 (2013).
Oliveira, R., Godinho, R., Randi, E. & Alves, P. C. Hybridization versus conservation: are domestic cats threatening the genetic integrity of wildcats (Felis silvestris silvestris) in Iberian Peninsula? Philos. Trans. R. Soc. Lond. B. Biol. Sci. 363, 2953–2961 (2008).
Canu, A., Mattioli, L., Santini, A., Apollonio, M. & Scandura, M. “Video-scats”: combining camera trapping and non-invasive genotyping to assess individual identity and hybrid status in gray wolf. Wildllife Biol. 4(1), 1–10 (2017).
Mattioli, L., Canu, A., Passilongo, D., Scandura, M. & Apollonio, M. Estimation of pack density in grey wolf (Canis lupus) by applying spatially explicit capture-recapture models to camera trap data supported by genetic monitoring. Front. Zool. 15, 38, https://doi.org/10.1186/s12983-018-0281-x (2018).
Santostasi, N. L. et al. Use of hidden Markov capture–recapture models to estimate abundance in the presence of uncertainty: application to the estimation of prevalence of hybrids in animal populations. Ecol. Evol. 9, 744–755, https://doi.org/10.1002/ece3.4819 (2019).
Salvatori, V., Godinho, R., Braschi, C., Boitani, L. & Ciucci, P. High levels of recent wolf × dog introgressive hybridization in agricultural landscapes of central Italy. Eur. J. Wildl. Res. 65, 73, https://doi.org/10.1007/s10344-019-1313-3 (2019).
Galaverni, M. et al. Monitoring wolves (Canis lupus) by non-invasive genetics and camera trapping: a small-scale pilot study. Eur. J. Wildl. Res. 58, 47–58 (2012).
Loh, P., Lipson, M., Patterson, N., Moorjani, P. & Pickrell, J. K. Inferring admixture histories of human populations. Genetics 193, 1233–1254 (2013).
Kraus, R. H. S. et al. A single-nucleotide polymorphism-based approach for rapid and cost-effective genetic wolf monitoring in Europe based on noninvasively collected samples. Mol. Ecol. Resour. 15, 295–305 (2015).
Norman, A. J. & Spong, G. Single nucleotide polymorphism-based dispersal estimates using noninvasive sampling. Ecol. Evol. 5, 3056–3065 (2015).
von Thaden, A. et al. Assessing SNP genotyping of noninvasively collected wildlife samples using microfluidic arrays. Sci. Rep. 7, 10768 (2017).
Ražen, N. et al. Long-distance dispersal connects Dinaric-Balkan and Alpine grey wolf (Canis lupus) populations. Eur. J. Wildl. Res. 62, 137, https://doi.org/10.1007/s10344-019-1313-3 (2016).
Nowak, S. et al. Sedentary but not dispersing wolves Canis lupus recolonizing western Poland (2001–2016) conform to the predictions of a habitat suitability model. Divers. Distrib. 23, 1353–1364, https://doi.org/10.1111/ddi.12621 (2017).
Hulva, P. et al. Wolves at the crossroad: Fission–fusion range biogeography in the Western Carpathians and Central Europe. Divers. Distrib. 24, 179–192, https://doi.org/10.1111/ddi.12676 (2018).
Ciucci, P., Lucchini, V., Boitani, L. & Randi, E. Dewclaws in wolves as evidence of admixed ancestry with dogs. Can. J. Zool. 81(12), 2077–2081 (2003).
Vaysse, A. et al. Identification of genomic regions associated with phenotypic variation between dog breeds using selection mapping. PLoS Genet. 7, e1002316 (2011).
Caniglia, R. et al. Big bad wolf or man’s best friend? Unmasking a false wolf aggression on humans. Forensic Sci. Int. Genet. 24, e4–e6 (2016).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664, https://doi.org/10.1101/gr.094052.109 (2009).
Valière, N. Gimlet: a computer program for analysing genetic individual identification data. Mol. Ecol. Notes 2, 377–379, https://doi.org/10.1046/j.1471-8286.2002.00228.x-i2 (2002).
Acknowledgements
We warmly thank E. Randi (University of Bologna), P. Ciucci (Univeristy of Rome La Sapienza), P. Genovesi (ISPRA) and W. Reggioni (Wolf Apennine Center, Apennino Tosco-Emiliano National Park), for their ideas and useful comments on a preliminary version of the manuscript. We are particularly grateful to all the colleagues who contributed to collect the dog and wolf samples used in this study, in particular to D. Bigi, C. Musto and M. Delogu (University of Bologna), L. Molinari, F. Moretti, M. Andreani and M. Canestrini (Wolf Apennine Center), F. Striglioni and N. Riganelli (Gran Sasso National Park), L. Mattioli (Tuscany Region), S. Luccarini (University of Sassari), F. Morimando (Univeristy of Siena), D. Berzi (Ischetus), C. Pedrazzoli, M. Mencucci and N. Cappai (Foreste Casentinesi National Park), E. Berti and R. Berti (CRAS Monte Adone). We are also indebted to M.A. Selvatici, A. Trentini, M. Cazzato and E. Morreale (ISPRA) for their support with the administrative practices needed for the publication of the manuscript.
Author information
Authors and Affiliations
Contributions
R.C., M.G., E.F. conceived, designed and planned the study. R.C. supervised the study. R.C., M.A., N.M., M.S., contributed reagents/materials/analysis tools. R.C., M.G., E.V., F.M., E.F. performed laboratory experiments. R.C., E.V., F.M., E.F. analysed the data. R.C., E.V., E.F. developed the analysis pipeline and provided the experimental supplies. R.C., M.G., M.S., E.F. wrote the manuscript. R.C., E.V., F.M., E.F. prepared figures and tables, and performed the elaborations. R.C., M.G., E.V., F.M., A.C., M.A., N.M., M.S., E.F. shared ideas to realize the manuscript, read, reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Caniglia, R., Galaverni, M., Velli, E. et al. A standardized approach to empirically define reliable assignment thresholds and appropriate management categories in deeply introgressed populations. Sci Rep 10, 2862 (2020). https://doi.org/10.1038/s41598-020-59521-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59521-2
This article is cited by
-
Possible origins and implications of atypical morphologies and domestication-like traits in wild golden jackals (Canis aureus)
Scientific Reports (2023)
-
Ex situ versus in situ Eurasian lynx populations: implications for successful breeding and genetic rescue
Conservation Genetics (2023)
-
A reduced SNP panel to trace gene flow across southern European wolf populations and detect hybridization with other Canis taxa
Scientific Reports (2022)
-
Genetic diversity and population structure of the grey wolf (Canis lupus Linnaeus, 1758) and evidence of wolf × dog hybridisation in the centre of European Russia
Mammalian Biology (2021)
-
Reliable wolf-dog hybrid detection in Europe using a reduced SNP panel developed for non-invasively collected samples
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.