Abstract
Asymptomatic oropharyngeal carriage of Neisseria meningitidis peaks in adolescence and young adulthood. Following a meningococcal disease outbreak at a U.S. college, we profiled the oropharyngeal microbiomes of 158 students to identify associations between bacterial community composition and meningococcal carriage or risk factors for carriage, including male gender, smoking, and frequent social mixing. Metagenomic shotgun sequencing identified 268 bacterial taxa at the genus or species level, with Streptococcus, Veillonella, and Rothia species being most abundant. Microbiome composition showed weak associations with meningococcal carriage and risk factors for carriage. N. meningitidis abundance was positively correlated with that of Fusobacterium nucleatum, consistent with hypothesized propionic acid cross-feeding. Additional species had positive abundance correlations with N. meningitidis, including Aggregatibacter aphrophilus, Campylobacter rectus, Catonella morbi, Haemophilus haemolyticus, and Parvimonas micra. N. meningitidis abundance was negatively correlated with unidentified Veillonella species. Several of these species are commonly found in dental plaque, while N. meningitidis is primarily found in the pharynx, suggesting that ecological interactions extend throughout the oral cavity. Although risk factors for meningococcal carriage do not strongly impact most bacterial species in the oropharynx, variation in the upper respiratory tract microbiome may create conditions that are more or less favorable for N. meningitidis carriage.
Similar content being viewed by others

Introduction
Neisseria meningitidis is a transient commensal of the human pharynx that also causes life-threatening diseases including meningitis and bacteremia1. Meningococcal carriage prevalence peaks in late adolescence and early adulthood2, corresponding to an increased risk of disease in this age group1. Risk factors for meningococcal carriage among adolescents and young adults in the United States include male gender, smoking, and frequent social mixing3. However, the influence of the pharyngeal microbiota on meningococcal carriage has been largely unexplored. A variety of bacterial species colonize the pharynx and other upper respiratory tract (URT) sites4,5, but there is currently little information about how they interact with N. meningitidis or how their populations are influenced by traditional risk factors for meningococcal carriage.
Understanding N. meningitidis ecology requires identification of microbial species that co-colonize the pharynx. Culture-based studies target particular species, often from the oropharynx, where N. meningitidis is most readily cultured6. These studies have provided evidence of competitive exclusion between N. lactamica and N. meningitidis in the oropharynx. This evidence includes the low prevalence of N. lactamica carriage among age groups with highest prevalence of N. meningitidis carriage7,8, and the reduced probability of isolating N. meningitidis from oropharyngeal swabs following inoculation with N. lactamica9.
Sequence-based, culture-independent approaches provide a more comprehensive assessment of the oropharyngeal bacterial community10, enabling both the discovery of unanticipated relationships and the evaluation of new hypotheses using previously collected data. For instance, N. meningitidis can utilize propionic acid as a carbon source, prompting the hypothesis that N. meningitidis growth is promoted by the presence of anaerobic bacteria that produce this short fatty acid by fermentation11. This cross-feeding hypothesis was initially evaluated with 16S rRNA gene data from bacteria in salivary specimens, identifying correlations between the proportional abundances of Neisseria spp. and two genera of anaerobic propionic acid producers, Fusobacterium and Porphyromonas. Further analysis of 16S rRNA genes showed that the average proportional abundance of both Fusobacterium spp. and Porphyromonas spp. increases through adolescence, in parallel to meningococcal carriage in the nasopharynx11.
Interpretation of 16S rRNA gene sequences can be limited by difficulty discriminating between N. meningitidis and other Neisseria species12,13. However, shotgun metagenomic sequencing can readily discriminate among the Neisseria species that are present in different parts of the oral cavity14. Furthermore, shotgun metagenomic sequencing enables identification of genes encoding metabolic enzymes in a bacterial community15.
To investigate how carriage of N. meningitidis among young adults may be influenced by variation in the bacterial community, we performed metagenomic shotgun sequencing on DNA obtained from oropharyngeal swabs collected from college students following a serogroup B meningococcal (MenB) disease outbreak in Providence, Rhode Island3. We evaluated the proportional abundances of each bacterial species in the oropharynx and identified species that were associated with meningococcal carriage or risk factors for meningococcal carriage.
Results
The oropharynx harbors a diverse bacterial community
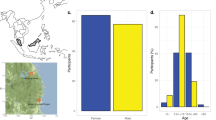
Following four cross-sectional oropharyngeal carriage surveys of undergraduate students, 158 oropharyngeal swabs were selected for sequencing from a pool of 705 swabs collected during the third and fourth rounds from students who were between the age of 18 and 23 years, had not used antibiotics in the past 30 days, and had received both a quadrivalent meningococcal conjugate vaccine (MenACWY) and at least two doses of MenB-FHbp vaccine at least two weeks prior to swab collection. The 158 oropharyngeal swabs included 75 N. meningitidis culture-positive swabs (64 nongroupable, 6 serogroup E, and 5 serogroup B). The culture-positive and culture-negative swabs had similar composition with respect to the students’ carriage risk factors: gender, smoking, and frequent social mixing (Table 1; see methods for details).
A total of 268 bacterial taxa were identified among the metagenomic sequences from the 158 oropharyngeal swabs; 239 of these taxa are species that are represented in the reference genome database, while 29 of the taxa are “unclassified” species belonging to different genera. The total proportional abundance of unclassified species ranges from 1.4% to 46.2% of the bacterial community (mean of 19.5%).
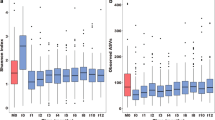
The proportional abundance of taxa varied substantially across specimens (Fig. 1). While five genera (Streptococcus, Veillonella, Rothia¸ Prevotella, and Actinomyces) were found in all 158 specimens, their proportional abundance varied by orders of magnitude between specimens (Fig. 1A). For instance, the most abundant genus, Streptococcus, ranged from 2.4% to 66%. Three species from two genera were detected in all 158 specimens: Rothia mucilaginosa, Streptococcus salivarius, and Streptococcus parasanguinis (Fig. 2). Clustering of swabs according to bacterial community composition placed 135 (85%) into a single group (Fig. 1, Supplemental Fig. 1). The effective number of taxa (exponential Shannon entropy) within the oropharyngeal specimens (13.4) is within the range of values calculated from taxonomic profiles of oral body sites generated by the Human Microbiome Project (HMP) using comparable methods (Supplemental Table 1). The taxonomic composition is most similar to that of the tongue dorsum (Supplemental Figs. 2 and 3; Mann-Whitney U, p < 0.001).
Bacterial species composition of 158 oropharyngeal swabs, clustered by species composition. (A) Abundance of each genus that accounts for over 10% of any swab. Average abundance is in parentheses in legend. (B) Heatmap showing the 100 most common bacterial taxa. Cells are color coded on a logarithmic scale to show the proportional abundance of each species (row) in each swab (column). In both panels, swabs are clustered according to average Bray-Curtis distances of their taxonomic profiles, represented by the dendrogram on the top. Taxa in panel B are clustered according to the Bray-Curtis distance of their abundance in each swab, shown by the dendrogram on the left. Colored portions of the dendrogram indicate groups for which distances are <70% of the maximum distance. Neisseria meningitidis is emphasized with bold font.
Abundance of dominant species among 158 oropharyngeal swabs. Curves show number of swabs in which each species was found at the given abundance; the y-intercept is equal to the number of swabs in which the species was identified. The twenty species shown have an abundance greater than 10% in one or more specimens, and a mean abundance greater than 1% overall (shown in parentheses in legend).
Bacterial community composition is not associated with student traits
We tested whether the bacterial community differed between swabs based on N. meningitidis culture results, three carriage risk factors (the student’s gender, smoking behavior, and social mixing behavior), and two other potentially influential factors (the student having symptoms of URT infection in the prior two weeks, and the study round in which the swab was collected). The effective number of communities (exponential Shannon beta diversity) was no lower for pairs of swabs with the same values for any of these variables than for pairs of swabs with different values (Table 2, Supplemental Figs. 1 and 4). None of the six tests passed the 5% Bonferroni family-wise error rate, which required uncorrected p-values to be less than 0.83%.
Likewise, only two significant correlations were identified between the proportional abundance of each taxon and variables describing the oropharyngeal specimen, using a multivariate linear model with a false discovery limit of 25%. The proportional abundance of N. meningitidis in the metagenomic data was significantly higher among swabs that were positive for N. meningitidis by culture (p = 3.4 × 10−4); the median proportional abundance was 3.57 × 10−4 in the Nm culture-negative swabs (99% CI, 1.72 × 10−4–9.65 × 10−4) and 1.48 × 10−3 in the Nm culture-positive swabs (99% CI, 4.92 × 10−4–3.98 × 10−3). S. salivarius was associated with females (p = 3.8 × 10−4); the median abundance was 6.0 × 10−2 among males (99% CI, 4.2 × 10−2–8.7 × 10−2) and 9.8 × 10−2 among females (99% CI, 7.4 × 10−2–1.7 × 10−1). No associations were identified with the remaining variables (study round, URT infections, smoking, or social mixing).
‘N. meningitidis’ abundance is correlated with the propionic acid producer, Fusobacterium nucleatum
Catanazzi et al. proposed that N. meningitidis growth in the oral cavity is promoted by the presence Fusobacterium spp. or Porphyromonas spp., which produce propionic acid by anaerobic fermentation11. Of the Fusobacterium or Porphyromonas species detected by metagenomic sequencing, only F. nucleatum had a significant compositional correlation with N. meningitidis (r = 0.22, p < 0.001). None of the remaining three Fusobacterium species (F. gonidiaformans, F. necrophorum, F. periodonticum) nor the four Porphyromonas taxa (P. catoniae, P. endodontalis, P. gingivalis, Porphyromonas oral taxon 279) had statistically significant compositional correlations with N. meningitidis (Supplemental Table 2).
Functional analysis of gene sequences in the metagenomic data indicated the potential for propionic acid production through the L-1,2-propanediol degradation pathway; no other pathway for fermentation to propionic acid was confidently identified. The L-1,2-propanediol degradation pathway was identified in 57 specimens; it was attributed to F. nucleatum in 18 specimens, to F. periodonticum in 27 specimens, and to both F. nucleatum and F. periodonticum in 12 specimens. The normalized abundance of genes for the L-1,2-propanediol degradation pathway was not correlated with the proportional abundance of N. meningitidis (Spearman r = −0.11, p = 0.16).
‘N. meningitidis’ abundance is correlated with other species
To identify potential interactions between N. meningitidis and any of the 267 taxa, we used three different correlation coefficients (Pearson’s, Spearman’s, and compositional) to test for correlations in proportional abundance between taxa. Six different taxa had significant positive correlations with N. meningitidis according to all three tests: Aggregatibacter aphrophilus, Campylobacter rectus, Catonella morbi, Haemophilus aegyptius, Haemophilus haemolyticus, and Parvimonas micra (Table 3). Haemophilus aegyptius is commonly described as a biogroup of H. influenzae rather than a separate species16; the Pearson correlation of H. influenzae and N. meningitidis abundances was not significant (r = 0.06; p = 0.46), but the Spearman (r = 0.34, p = 1.0 × 10−5) and compositional correlations (r = 0.17, p = 0.004) were significant. The only taxon for which all three tests identified a significant negative correlation with N. meningitidis was unclassified Veillonella, representing one or more Veillonella spp. that could not be identified at the species level. Notably, N. lactamica did not have a significant correlation with N. meningitidis (Supplemental Table 2).
Discussion
To better understand the ecology of N. meningitidis, we examined the oropharyngeal microbiome among young adults at a U.S. college where 20%–24% of surveyed students carried meningococcal bacteria, as detected via culture3. Previous DNA-based profiling of the oropharyngeal microbiome has used amplicon sequencing of 16S rRNA genes for genus-level analysis of bacterial communities4,17,18. The major oropharyngeal bacterial genera identified here among college students were similar to those described previously among adult populations such as United States active duty military personnel17 and a portion of the German National Cohort study18. The most prominent differences were a high abundance of Rothia spp. and a low abundance of Leptotrichia spp. among this college population. Shotgun metagenomic sequencing obtains species-level resolution for most of the bacterial community, but published shotgun metagenomic sequence data for the oropharyngeal microbiome is limited; comparison of these oropharyngeal communities’ profiles to the profiles published by the Human Microbiome Project showed that they were most similar to the communities of the tongue dorsum.
The overall bacterial community profile was not associated with any of the meningococcal carriage risk factors that we tested. Only one bacterial species was associated with any student trait tested here; S. salivarius was detected in all swabs but was more abundant among females. While we did not identify any association between smoking and bacterial taxon abundance, previous research has found that tobacco smoking is associated with differences in bacterial species abundance in oral rinse19 and nasopharyngeal20 specimens. These studies differed from our analysis of oropharyngeal specimens in several respects, including the examination of different body sites, older populations, and 16S rRNA gene sequencing rather than shotgun metagenomic sequencing. Furthermore, the oral rinse microbiome study of Bornigen et al.19 specified the intensity of cigarette use, finding that most of the differences in taxon abundance were primarily due to the comparison between regular cigarette smokers and cigarette non-smokers. In contrast, our smoking variable included regular cigarette smokers, occasional cigarette smokers (in the past 30 days), and marijuana smokers. The inclusion of a broader set of behaviors as “smoking” may have diminished the apparent association between smoking and bacterial taxon abundance.
Patterns of co-occurrence among microbial species can provide insights into ecological relationships, such as competition and mutualism21. Among the proposed ecological relationships of N. meningitidis is competition with N. lactamica in the pharynx, resulting in lower meningococcal carriage among individuals colonized with N. lactamica9. In this study, no significant association between N. lactamica and N. meningitidis abundances was identified. This is perhaps due to the low prevalence of N. lactamica carriage among older adolescents7,8, which was reflected in the detection of N. lactamica in only 18 of the 158 swabs. Ultimately, the metagenomic data indicate that competition with N. lactamica was not a substantial contributor to the variation in N. meningitidis carriage within this population.
N. meningitidis is capable of using propionic acid as a carbon source, and therefore may benefit from cross-feeding with propionic acid producers11. Studies based on 16S rRNA gene sequencing have shown that in saliva, the average proportional abundance of the anaerobic propionic acid producing genera Fusobacterium and Porphyromonas increases through adolescence, in parallel to meningococcal carriage in the nasopharynx11. The metagenomic data of the current study improved on prior 16S rRNA gene studies by estimating species-level abundances for all three genera, thereby identifying a significant positive correlation between N. meningitidis and F. nucleatum. The metagenomic data also confirmed the presence of F. nucleatum genes encoding a fermentation pathway that produces propionic acid; however, the abundance of genes for this pathway within the full microbial community was not correlated with N. meningitidis abundance.
Interactions between F. nucleatum and N. meningitidis may be limited by their tendency to grow in different locations within the URT – the gingival crevice for F. nucleatum22 and the pharynx for N. meningitidis. However, interactions may be mediated by saliva23, in which propionic acid levels can reach 3.3 mM24. The correlations in our metagenomic data suggest that N. meningitidis may interact with other taxa that are thought to grow in dental plaque and the gingival crevice, particularly A. aphrophilus, C. rectus, and C. morbi25,26,27,28; H. haemolyticus is also commonly found in dental plaque, as well as the pharynx29.
Veillonella species (for which a negative correlation with N. meningitidis was found) are commonly found in dental plaque, buccal mucosa, and the tongue26. Of note, like members of the genera Fusobacterium and Porphyromonas, members of Veillonella are anaerobic producers of propionic acid. While many Veillonella genomes encode genes for fermentation to propionic acid (e.g. V. atypica)30, these genes were not confidently identified in our metagenomic data. In contrast to members of Fusobacterium and Porphyromonas, members of Veillonella are abundant in the oral cavity throughout life26 and have been found to be more abundant among younger adolescents than older adolescents31.
The significant correlations identified between the proportional abundances of N. meningitidis and other bacterial species suggests possible beneficial or antagonistic interactions between these species and N. meningitidis. However, the bacterial species abundance is also affected by factors that were not examined in this study, such as oral and gastrointestinal health19,32. Such unmeasured and untested factors could influence the correlations between the abundances of N. meningitidis and other species. For instance, the negative correlation between N. meningitidis and Veillonella abundances could reflect the tendency for their abundances to change with age.
As sequencing technology and associated analysis tools advance, expanded investigation of URT ecology will be possible, providing context for studies of N. meningitidis transmission and host interactions. To understand meningococcal carriage and disease, further investigations will need to incorporate longitudinal observations of the microbiome as N. meningitidis carriage is gained or lost, particularly alongside medical interventions such as vaccination and antimicrobial treatments. Sampling from additional locations of the URT5 may help explain the correlations that we detected among the abundances of certain bacterial species that typically grow in dental plaque and the abundance of N. meningitidis. Examination of the nasopharynx is also important, since it is known to harbor N. meningitidis6, yet its bacterial community is dominated by different genera than the oropharynx (e.g. Staphylococcus, Corynebacterium, and Moraxella17). Further microbiome profiling can identify the large-scale structure of microbial communities, and guide more detailed studies to evaluate bacterial physiology and micron-scale interactions in situ. Ultimately, understanding the URT microbiome will improve knowledge for a variety of diseases, ranging from oral cancer19 to bacterial infections4.
Methods
Specimen collection
Oropharyngeal swabs were collected during four cross-sectional carriage surveys performed alongside a MenB-FHbp (factor H binding protein) vaccination campaign in response to the outbreak, as described by Soeters, et al.3. A total of 1,121 undergraduate students participated in the third and fourth study rounds (September 2015 and March 2016), during which one prong of a bifurcated swab was preserved for sequencing in 1 ml skim milk/tryptone/glucose/glycerin medium (STGG) on ice33, then frozen at −80 °C. The other prong was used for culture-based detection of N. meningitidis by plating on a modified Thayer-Martin (MTM) agar plate (BD BBL, Franklin Lakes, New Jersey) that was incubated at 37 °C with 5% CO2 and checked for growth at 24, 48, and 72 hours3. Colonies with typical Neisseria morphology were subcultured onto a blood agar plate (BD BBL), and N. meningitidis cultures were identified using Gram staining (BD BBL), oxidase testing (Hardy Diagnostics, Santa Maria, California), sodC rt-PCR, and API Neisseria-Haemophilus strip testing (bioMerieux, Durham, North Carolina)34. N. meningitidis isolates were serogrouped by slide agglutination34. Participants completed a short, self-administered questionnaire including potential meningococcal carriage risk factors, including gender, smoking (tobacco or marijuana in the past 30 days), frequent social mixing (visits to bars, clubs, or parties at least once per week), and recent URT infections (symptoms in the previous two weeks).
The carriage evaluation3 was approved by the Rhode Island Department of Health Institutional Review Board (IRB) (#IRB 2015 – 02) and the Providence College IRB (#15-062), and was determined to be public health non-research by the CDC National Center for Immunization and Respiratory Diseases (#2015 6436) and did not require CDC IRB approval. All participants provided written informed consent to participate in the carriage evaluation3 and for their oropharyngeal specimens to be stored for future research. This microbiome study used de-identified specimens and questionnaire data and was determined to not involve human subjects (#2016 6651) and did not require CDC IRB approval.
DNA extraction and sequencing
DNA was extracted from 200 μL of STGG medium using QIAamp DNA Microbiome Kit (Qiagen) as per manufacturer protocol. DNA concentrations were measured using a Qubit 2.0 fluorometer with dsDNA High Sensitivity reagents (Invitrogen, Grand Island, NY, USA). The eluted DNA concentration ranged from 0.2 to 5 ng/μL.
Sequencing libraries were prepared from 2 ng of DNA using Nextera XT DNA library preparation kit following the manufacturer’s protocol (Illumina, San Diego, CA, USA). The resulting libraries were sequenced on three Rapid-Runs of an Illumina HiSeq 2500 (CDC Genomic Core facility) to obtain 250 bp paired-ends reads. Sixteen specimens with exceptionally low numbers of reads (<100,000 reads) were sequenced again on an Illumina MiSeq using a fresh library preparation.
Metagenomic sequence analysis
Human DNA was removed by mapping reads to the hg38 reference genome using Bowtie2 (v2.2.9; local alignment mode)35, retaining a median of 94.8% (interquartile range: 85.7–98.4%) of reads with a minimum of 336,841 read pairs per specimen. Unmapped reads were submitted to SRA BioProject PRJNA428766. PCR duplicates were removed with bbtools-clumpify (v37.41)36, then Nextera adapter sequences and low-quality base calls were trimmed from reads with cutadapt (v1.8.3; options n = 5, trim-n = True, m = 50)37. The proportional abundance of bacterial species was estimated using MetaPhlAn2 (v2.2.0)38, which is a precise method for species detection (i.e. it rarely reports the presence of species that are not in a specimen)39. This precision is achieved by identifying marker genes that clearly distinguish different species, and if a sufficient number of these marker genes cannot be detected to confidently identify a species, MetaPhlAn2 conservatively limits its reporting to the genus or a group of species (e.g., Streptococcus mitis/oralis/pneumoniae). Data from the Human Microbiome Project (HMP) was retrieved as MetaPhlAn2 abundance files from the HMP website40.
Community taxonomic diversity of both the HMP specimens and the specimens from this study was calculated from MetaPhlAn2 taxonomic profiles using an exponential transformation of Shannon entropy to describe the effective number of taxa within a specimen (alpha) and the effective number of distinct communities in a collection of specimens (beta)41. Alpha diversity approaches a lower limit of 1 when a single species dominates the community; alpha diversity is equal to the number of observed taxa in a specimen when taxa are equally abundant in a specimen. Beta diversity approaches a lower limit of 1 when specimens have identical taxonomic profiles; beta diversity equals the number of specimens in the collection when there are no shared taxa between the specimens.
Associations of taxon abundance with specimen characteristics were calculated with MaAsLin (v0.0.4, default parameters)42, which applies a multivariate linear model to associate the abundance of each taxon with the available metadata. MaAsLin adjusts for multiple hypothesis testing by first applying a Bonferroni correction for each taxon’s association with each value of each metadata variable, then calculating a False Discovery Rate (FDR) over all taxa and metadata variables, reporting associations with FDR ≤ 25%. Metadata variables included in the multivariate linear model were: study round; N. meningitidis culture results; gender; smoking; social mixing; URT infection; and sequencing run.
Compositional correlations calculated with SparCC (v0.1)43 were used to assess correlated taxon abundances. Calculations were performed using pseudo-counts that were generated by multiplying proportional abundances by 1,000; this is a conservative scaling factor, as the proportional abundance table for each specimen was generated from at least 336,841 non-human read pairs and contained at least one taxon with proportional abundance under 0.001. Tests for previously predicted correlations between N. meningitidis and Fusobacterium spp. or Porphyromonas spp. were considered significant if the compositional correlation was more extreme than 99.375% of the correlation coefficients from permutated matrices (two-tailed, 1,000 permutations; Bonferroni adjustment for 8 tests with α ≤ 5%). When testing for associations that had not been previously hypothesized, a Benjamini-Hochberg FDR44 was applied to account for the large number of comparisons (267), and specificity was increased by using both Pearson and Spearman correlations to confirm associations (two-tailed t-test)21, controlling for FDR ≤ 25% for each test.
A functional interpretation of metagenome data was performed on each specimen using HUMAnN2 v2.915 (default parameters, UniRef90 database). The potential for propionic acid production was assessed based on the presence of genes from each of the five pathways within the “Fermentation to Propanoate” class on the MetaCyc website30: “L-1,2-propanediol degradation pathway” (PWY-7013), “L-alanine fermentation to propanoate and acetate” (PROPFERM-PWY), “L-glutamate degradation VIII” (PWY-5088), and “pyruvate fermentation to propanoate” (P108-PWY and PWY-5494). The abundance of each pathway was measured at the community level and normalized to “copies per million” reads using the human2_renorm_table script. Confident detection of pathways required the coverage value to exceed 0.5.
Data analysis was performed with SciPy45 with scikit-bio version 0.5.2.
Data availability
The dataset supporting the conclusions of this article is available in the Short Read Archive repository, BioProject PRJNA428766.
References
MacNeil, J. R., Blain, A. E., Wang, X. & Cohn, A. C. Current epidemiology and trends in meningococcal disease-United States, 1996–2015. Clin Infect Dis 66, 1276–1281, https://doi.org/10.1093/cid/cix993 (2018).
Christensen, H., May, M., Bowen, L., Hickman, M. & Trotter, C. L. Meningococcal carriage by age: a systematic review and meta-analysis. Lancet Infect Dis 10, 853–861, https://doi.org/10.1016/s1473-3099(10)70251-6 (2010).
Soeters, H. M. et al. Meningococcal carriage evaluation in response to a serogroup B meningococcal disease outbreak and mass vaccination campaign at a college-Rhode Island, 2015–2016. Clin Infect Dis 64, 1115–1122, https://doi.org/10.1093/cid/cix091 (2017).
de Steenhuijsen Piters, W. A., Sanders, E. A. & Bogaert, D. The role of the local microbial ecosystem in respiratory health and disease. Philos. Trans. Royal Soc. B 370, https://doi.org/10.1098/rstb.2014.0294 (2015).
Proctor, D. M. & Relman, D. A. The Landscape Ecology and Microbiota of the Human Nose, Mouth, and Throat. Cell Host Microbe 21, 421–432, https://doi.org/10.1016/j.chom.2017.03.011 (2017).
Roberts, J., Greenwood, B. & Stuart, J. Sampling methods to detect carriage of Neisseria meningitidis; literature review. J Infect 58, 103–107, https://doi.org/10.1016/j.jinf.2008.12.005 (2009).
Diallo, K. et al. Pharyngeal carriage of Neisseria species in the African meningitis belt. J. Infect. 72, 667–677, https://doi.org/10.1016/j.jinf.2016.03.010 (2016).
Gold, R., Goldschneider, I., Lepow, M. L., Draper, T. F. & Randolph, M. Carriage of Neisseria meningitidis and Neisseria lactamica in infants and children. J. Infect. Dis. 137, 112–121 (1978).
Deasy, A. M. et al. Nasal Inoculation of the Commensal Neisseria lactamica Inhibits Carriage of Neisseria meningitidis by Young Adults: A Controlled Human Infection Study. Clin Infect Dis 60, 1512–1520, https://doi.org/10.1093/cid/civ098 (2015).
Li, K., Bihan, M. & Methe, B. A. Analyses of the stability and core taxonomic memberships of the human microbiome. PLoS One 8, e63139, https://doi.org/10.1371/journal.pone.0063139 (2013).
Catenazzi, M. C. et al. A large genomic island allows Neisseria meningitidis to utilize propionic acid, with implications for colonization of the human nasopharynx. Mol Microbiol 93, 346–355, https://doi.org/10.1111/mmi.12664 (2014).
Bennett, J. S., Watkins, E. R., Jolley, K. A., Harrison, O. B. & Maiden, M. C. Identifying Neisseria species by use of the 50S ribosomal protein L6 (rplF) gene. J Clin Microbiol 52, 1375–1381, https://doi.org/10.1128/jcm.03529-13 (2014).
Walcher, M., Skvoretz, R., Montgomery-Fullerton, M., Jonas, V. & Brentano, S. Description of an unusual Neisseria meningitidis isolate containing and expressing Neisseria gonorrhoeae-Specific 16S rRNA gene sequences. J Clin Microbiol 51, 3199–3206, https://doi.org/10.1128/jcm.00309-13 (2013).
Albanese, D. & Donati, C. Strain profiling and epidemiology of bacterial species from metagenomic sequencing. Nat. Commun. 8, 2260, https://doi.org/10.1038/s41467-017-02209-5 (2017).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968, https://doi.org/10.1038/s41592-018-0176-y (2018).
Norskov-Lauritsen, N. Classification, identification, and clinical significance of Haemophilus and Aggregatibacter species with host specificity for humans. Clin. Micro. Rev. 27, 214–240, https://doi.org/10.1128/cmr.00103-13 (2014).
Hang, J. et al. Composition and variation of respiratory microbiota in healthy military personnel. PLoS One 12, e0188461, https://doi.org/10.1371/journal.pone.0188461 (2017).
Akmatov, M. K. et al. Determination of nasal and oropharyngeal microbiomes in a multicenter population-based study - findings from Pretest 1 of the German National Cohort. Sci. Rep. 7, 1855, https://doi.org/10.1038/s41598-017-01212-6 (2017).
Bornigen, D. et al. Alterations in oral bacterial communities are associated with risk factors for oral and oropharyngeal cancer. Sci. Rep. 7, 17686, https://doi.org/10.1038/s41598-017-17795-z (2017).
De Boeck, I. et al. Comparing the Healthy Nose and Nasopharynx Microbiota Reveals Continuity As Well As Niche-Specificity. Front. Microbiol. 8, 2372, https://doi.org/10.3389/fmicb.2017.02372 (2017).
Weiss, S. et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J 10, 1669–1681, https://doi.org/10.1038/ismej.2015.235 (2016).
Krieg, N. R. et al. Bergey’s manual of systematic bacteriology. Vol. 4 The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes (Springer, New York, 2010).
Proctor, D. M. et al. A spatial gradient of bacterial diversity in the human oral cavity shaped by salivary flow. Nat. Commun. 9, 681, https://doi.org/10.1038/s41467-018-02900-1 (2018).
Park, Y. D., Jang, J. H., Oh, Y. J. & Kwon, H. J. Analyses of organic acids and inorganic anions and their relationship in human saliva before and after glucose intake. Arch. Oral. Biol. 59, 1–11, https://doi.org/10.1016/j.archoralbio.2013.10.006 (2014).
Brenner, D. J., Krieg, N. R., Staley, J. T. & Garrity, G. M. Bergey’s manual of systematic bacteriology. Vol. 2. The proteobacteria. Part B, the gammaproteobacteria (ed. James T. Staley) (Springer, New York, 2005).
De Vos, P. et al. Bergey’s manual of systematic bacteriology. Vol. 3, The firmicutes (Springer, New York, 2009).
Brenner, D. J., Krieg, N. R., Staley, J. T. & Garrity, G. M. Bergey’s manual of systematic bacteriology. Vol. 2. The proteobacteria. Part C. The alpha-, beta-, delta-, and epsilonproteobacteria (ed. James T. Staley) (Springer, New York, 2005).
Mark Welch, J. L., Rossetti, B. J., Rieken, C. W., Dewhirst, F. E. & Borisy, G. G. Biogeography of a human oral microbiome at the micron scale. Proc. Nat. Acad. Sci. USA 113, E791–800, https://doi.org/10.1073/pnas.1522149113 (2016).
Kilian, M. Genus Haemophilus in Bergey’s manual of systematic bacteriology. Vol. 2. The proteobacteria. Part B, the gammaproteobacteria (eds.Don J. Brenner, Noel R. Krieg, James T. Staley, & George M. Garrity) (Springer, 2005).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res. 42, D459–471, https://doi.org/10.1093/nar/gkt1103 (2014).
Stahringer, S. S. et al. Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Res 22, 2146–2152, https://doi.org/10.1101/gr.140608.112 (2012).
Iaffaldano, L. et al. Oropharyngeal microbiome evaluation highlights Neisseria abundance in active celiac patients. Sci. Rep. 8, 11047, https://doi.org/10.1038/s41598-018-29443-1 (2018).
O’Brien, K. L. et al. Evaluation of a medium (STGG) for transport and optimal recovery of Streptococcus pneumoniae from nasopharyngeal secretions collected during field studies. J Clin Microbiol 39, 1021–1024, https://doi.org/10.1128/jcm.39.3.1021-1024.2001 (2001).
World Health Organization. Laboratory methods for the diagnosis of meningitis caused by Neisseria meningitidis, Streptococcus pneumoniae, and Haemophilus influenzae (World Health Organization, 2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Bushnell, B. BBTools software package. Available at: https://jgi.doe.gov/data-and-tools/bbtools/.
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12, https://doi.org/10.14806/ej.17.1.200.
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat Methods 12, 902–903, https://doi.org/10.1038/nmeth.3589 (2015).
McIntyre, A. B. R. et al. Comprehensive benchmarking and ensemble approaches for metagenomic classifiers. Genome Biol. 18, 182, https://doi.org/10.1186/s13059-017-1299-7 (2017).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature 550, 61–66, https://doi.org/10.1038/nature23889 (2017).
Jost, L. Partitioning diversity into independent alpha and beta components. Ecology 88, 2427–2439 (2007).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79, https://doi.org/10.1186/gb-2012-13-9-r79 (2012).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687, https://doi.org/10.1371/journal.pcbi.1002687 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. Royal Stat. Soc. (Methodological) 57, 289–300 (1995).
Jones, E., Oliphant, T., Peterson, P. & Others. SciPy: Open Source Scientific Tools for Python. Available at http://scipy.org (2001).
Acknowledgements
We would like to acknowledge assistance and advice provided by Sandeep Joseph, the CDC Biotechnology Core Facility, and the Rhode Island Meningococcal Carriage Evaluation Team. This work was supported by the Advanced Molecular Detection initiative at the Centers for Disease Control and Prevention. The funders of this research had no role in the design of the study, the collection, analysis, and interpretation of data, or in writing the manuscript. The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Centers for Disease Control and Prevention.
Author information
Authors and Affiliations
Contributions
A.C.R., C.B.K., L.D.R.R., A.C., H.M.S., M.J.W. and X.W. participated in the conception and design of this manuscript; C.B.K., L.D.R.R., H.M.S. and M.J.W. participated in the acquisition of the data; A.C.R., C.B.K., A.C. and X.W. participated in the analysis and interpretation of the data; A.C.R. wrote the manuscript; all authors provided critical review of the manuscript and approved its publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Retchless, A.C., Kretz, C.B., Rodriguez-Rivera, L.D. et al. Oropharyngeal microbiome of a college population following a meningococcal disease outbreak. Sci Rep 10, 632 (2020). https://doi.org/10.1038/s41598-020-57450-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-57450-8
This article is cited by
-
Distinctive features of the oropharyngeal microbiome in Inuit of Nunavik and correlations of mild to moderate bronchial obstruction with dysbiosis
Scientific Reports (2023)
-
Salivary proteome of aphthous stomatitis reveals the participation of vitamin metabolism, nutrients, and bacteria
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.