Abstract
Androgen signalling is a critical driver of male development. Fetal steroid signalling can be dysregulated by a range of environmental insults and clinical conditions. We hypothesised that poor adult male health was partially attributable to aberrant androgen exposure during development. Testosterone was directly administered to developing male ovine fetuses to model excess prenatal androgenic overexposure associated with conditions such as polycystic ovary syndrome (PCOS). Such in utero androgen excess recreated the dyslipidaemia and hormonal profile observed in sons of PCOS patients. 1,084 of 15,134 and 408 of 2,766 quantifiable genes and proteins respectively, were altered in the liver during adolescence, attributable to fetal androgen excess. Furthermore, prenatal androgen excess predisposed to adolescent development of an intrahepatic cholestasis-like condition with attendant hypercholesterolaemia and an emergent pro-fibrotic, pro-oxidative stress gene and protein expression profile evident in both liver and circulation. We conclude that prenatal androgen excess is a previously unrecognised determinant of lifelong male metabolic health.
Similar content being viewed by others
Introduction
Prenatal programming of health and disease is driven by maternal/fetal endocrine alterations1,2, maternal nutritional status3 and maternal/fetal chemical insults4. During development, interactions between endogenous hormones, underlying endocrine disease and/or environmental exposure to toxicants, such as endocrine disrupting compounds (EDC)5 impact upon postnatal health, and thus the incidence of chronic disease6, with attendant economic burdens7,8.
The linkage between adult health and altered prenatal androgen exposure is robust. Androgens, secreted by developing testes9 and other organs10 promote masculinisation in target tissues expressing androgen receptors. In addition to pathologies of insufficient androgen exposure, for example hypospadias11, excess exposure may also occur, for example, in offspring of hyperandrogenaemic women with polycystic ovary syndrome (PCOS) or congenital adrenal hyperplasia12. Human data and animal experimentation has demonstrated aetiological contributions of prenatal androgen overexposure to the development of PCOS in female offspring13, linking the reproductive phenotype with metabolic phenotypes, including insulin resistance (IR), obesity and dyslipidaemia. This is underscored by morphometric markers of increased prenatal androgens in women with PCOS and their daughters14,15, and male range umbilical cord blood androgen concentrations in daughters of women with PCOS16. Indeed animal models designed to recreate PCOS-associated pathologies utilise over-exposure of female fetuses/pregnant dams to androgenic steroids1,17,18,19,20,21,22,23,24,25.
Male offspring from human PCOS pregnancies also develop hyperinsulinaemia and dyslipidaemia, predicting potentially increased risk of cardiovascular disease26,27,28. Elevated cholesterol is an early marker of metabolic dysfunction in pubertal sons of PCOS patients29. Early metabolic changes in PCOS sons are likely to be independent of gonadotropin and sex steroid levels although they have elevated anti-Müllerian hormone (AMH) concentrations during childhood30.
We hypothesised that androgen excess during male fetal development increases the risk of developing liver dysfunction and dyslipidaemia. The aims of the study were, in an animal model of prenatal androgen excess: 1. Determine whether dyslipidaemia observed in PCOS-patient male relatives may have early-life-environment contributions. 2. Comprehend hepatic mechanisms behind prenatally programmed dyslipidaemia. 3. Determine circulating proteins reflective of prenatally programmed liver dysfunction prior to overt disease symptom development.
We discovered that fetal androgen excess was linked to adolescent male hepatic dysfunction/dyslipidaemia, elucidated mechanisms underpinning fetal androgen-excess programming of postnatal hepatic dysfunction, and identified circulating markers reflective of such programming in adolescence.
Results
Prenatal androgen treatment replicates human male PCOS-offspring characteristics
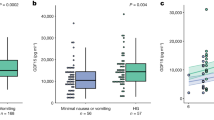
Circulating biomarkers characteristic of the sons of women with PCOS were examined in male sheep from control and prenatally androgen treated pregnancies. AMH and insulin were significantly elevated in adolescent prenatally androgen treated males (P < 0.05), but testosterone concentrations were comparable with control offspring (Fig. 1a). Plasma from prenatally androgen treated sheep therefore replicated circulating biomarkers of PCOS patients first degree male relatives26,27,28,29,30.
Prenatally androgen exposed males exhibit characteristics typical of pre- and post-pubertal sons of PCOS women. (a) Plasma characterisation: Samples from control (C; n = 14) and prenatal androgen excess (PA; n = 14) were collected at 6 months postnatal age from male offspring and assessed for common characteristics of pre- and pubertal sons of PCOS women. Prenatally androgen treated male offspring had significantly elevated circulating AMH, insulin and triglycerides, with no alteration in terms of testosterone concentrations. (b) Plasma cholesterol profiles: Prenatally androgen treated male offspring had significantly elevated total, LDL and HDL cholesterol concentrations as compared to vehicle control animals. Differences were analysed by unpaired, two-tailed t-test. (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001). Box plot whiskers are lowest and highest observed values, box is the upper and lower quartile, with median represented by line in box.
Evidence of dyslipidaemia
Adolescent prenatally androgen treated males had increased plasma triglycerides (0.90 ± 0.08 mmol/l vs 0.58 ± 0.07 mmol/l; P < 0.05) (Fig. 1a) and elevated total cholesterol concentrations (3.27 ± 0.25 nmol/l vs 1.7 ± 0.16 mmol/l; P < 0.0001) (ovine reference range 1.3–1.96 mmol/l31) compared with control offspring indicative of hypercholesterolaemia (Fig. 1b). Both LDL cholesterol (P < 0.001) and HDL cholesterol (P < 0.0001) concentrations were increased in adolescent prenatally androgen treated males when compared to controls (Fig. 1b).
Prenatal programming of hepatic lipid metabolism
The hepatic transcriptome analysis revealed 15,134 expressed genes for analysis after low count transcript removal, with 1,084 differentially expressed genes in adolescent life between control and prenatally androgen treated livers (FDR adjusted P < 0.05). Hepatic proteome analysis quantified and identified 2766 proteins, with 408 differentially expressed in adolescent life between control and prenatally androgen treated livers (P < 0.05). Of the 449 plasma proteins quantified and identified, 96 were differentially expressed in adolescent prenatally androgen treated males compared to control animals (P < 0.05). Collectively, bioinformatics analysis indicated the prenatally-programmed dysregulation of pathways involved in lipid metabolism, metabolism of cholesterol, sterol synthesis, cholesterol synthesis and lipid homeostasis, together with prediction of altered cholesterol concentrations (Table 1), manifesting in plasma as hypercholesterolaemia (Fig. 1b).
Prenatally programmed dysregulation of hepatic cholesterol synthesis, uptake and secretion pathways
Analysis of differentially expressed hepatic genes and proteins between control and prenatally androgen treated males revealed a consistent reduction in the cholesterol synthesis pathway (Fig. 2). When genes involved in the hepatic import pathways were analysed the significant differences pointed to an increased potential for cholesterol uptake (Fig. 2). Within the liver there was also increased mRNA encoding genes critical for esterification and assembly of lipoproteins with increased apolipoprotein expression. In addition, gene expression of cholesterol and phospholipid efflux potential into the bile was increased (Table 2). There is thus transcriptional evidence for augmented hepatic pathways involved in cholesterol efflux. However, the hypercholesterolemia in prenatally androgen treated males cannot be explained by increased hepatic cholesterol synthesis or reduced uptake into the liver.
Prenatally androgen exposed male offspring develop hepatic dysregulation of cholesterol and bile acid synthesis. Differentially expressed hepatic genes and proteins between control (C; n = 14) and prenatal androgen excess (PA; n = 14) adolescent offspring relevant to hepatic cholesterol homeostasis. To increase confidence in findings from RNAseq analysis a limited subset of genes was examined by qRT-PCR. In all cases differential expression was similar between RNAseq and qRT-PCR results. Fold change is PA compared to control animals. P values cited are nominal values, false discovery rate adjusted P values (FDR) were calculated as described by Benjamini and Hochberg (1995). Red denotes decreased, green denotes increased in response to prenatal androgen excess.
Dysregulation of the enterohepatic circulation
While there was evidence of increased capacity for cholesterol and phospholipid secretion into the bile, the same was not true for the secretion of bilirubin, bile acids and organic substrates (Fig. 3), predicting increased risk of bilirubin and bile acids accumulation within the liver. Adolescent prenatally androgen treated males had significantly increased hepatic bilirubin content as compared with adolescent controls (P < 0.001) (Fig. 4), with foci of accumulation reminiscent of human cholestasis histopathology. Prenatally androgen treated males also exhibited markedly higher circulating concentrations of bilirubin as compared to control pregnancy male offspring during adolescence (3.8 ± 0.44 µmol/l vs 2.27 ± 0.22 µmol/l; P < 0.01) (Fig. 4).
Prenatally androgen exposed develop hepatic dysregulation of cholesterol and bile trafficking.: Enterohepatic circulation. Differentially expressed hepatic genes and proteins between control (C; n = 14) and prenatal androgen excess (PA; n = 14) adolescent offspring relevant to hepatic regulation of cholesterol, bilirubin, bile acids and organic substrates trafficking. False discovery rate (RNAseq and proteomics; FDR) was determined by Benjamini and Hochberg (1995) method. Data are fold change in PA animals relative to vehicle treated animals. To increase confidence in findings from RNAseq analysis a limited subset of genes was examined by qRT-PCR. In all cases differential expression was similar between RNAseq and qRT-PCR results. In a small number of selected cases, where a protein has either particular relevance as part of a subset of proteins with related function, or where its gene was found to be differentially regulated, we have reported fold changes of those which were not statistically significant after FDR control (nominal and adjusted P values cited in all such cases).
Prenatally androgen exposed male offspring develop increased hepatic bilirubin accumulation, increased circulatory bilirubin concentrations, and increased circulating ALP, without any significant alterations circulating bile acid concentration. Plasma samples collected at 6 months postnatal age from male offspring were assayed for bilirubin, bile acids and ALP; incidence of bilirubin accumulation in liver was determined histologically after oxidisation to biliverdin via Fouchet’s reagent (indicated by black/green punctate staining, upper panels). Increased circulating total bilirubin, hepatic bilirubin accumulation incidence and plasma ALP were noted in PA males. Representative images of sections from each prenatal treatment group are shown (x20 magnification). Differences were analysed by unpaired, two-tailed t-test in plasma, and Chi-square in the case of incidence of hepatic bilirubin accumulation (*P < 0.05; **P < 0.01, ***P < 0.001). Box plot whiskers are lowest and highest observed values, box is the upper and lower quartile, with median represented by line in box.
Bile acids directly interact with several nuclear receptors within the liver. There were clear effects on genes that mediate bile acid action and homeostasis in prenatally androgen treated males (Fig. 3). While it is difficult to predict the net outcome of these changes it is clear that hepatic bile acid synthesis (via. CYP7A1) is likely to be reduced (Fig. 2). Bioinformatic analysis of the hepatic proteome predicted altered bile acid synthesis (Table 1). There was no difference in total circulating bile acids in prenatally androgen treated males (12.79 ± 2.36 µmol/l) as compared to control pregnancy offspring (9.39 ± 1.67µmol/l) (P = 0.16) (Fig. 4). Plasma ALP, known to increase during cholestasis, was increased in PA males (892.9 ± 83.8 U/L) as compared to control males (535.5 ± 67.8 U/L) during adolescence (P < 0.01) (Fig. 4). Bioinformatic analysis of the prenatally androgen treated male hepatic transcriptome predicted intrahepatic cholestasis and analysis of the hepatic proteome also predicted cholestasis (Table 1).
Reduced hepatic detoxification potential
Hepatic transcriptome analysis identified the potential for liver damage in prenatally androgen treated males. Liver damage, drug-induced liver disease and hepatocyte proliferation pathways were identified by bioinformatics analysis (Table 1). Hepatic proteome bioinformatics also predicted an increase of oxidative stress in livers of adolescent prenatally androgen treated males, accompanied by decreased hydrogen peroxide metabolism and increased synthesis of reactive oxygen species (Table 1). Moreover, there was a prediction of altered concentration of glutathione and increased cell death (Table 1). Bioinformatic analysis of the plasma proteome indicated increased biosynthesis of hydrogen peroxide, metabolism of reactive oxygen species and synthesis of reactive oxygen species (Table 1). There was a general reduction in the expression of hepatic genes and proteins relevant to both liver detoxification potential and liver reactive oxygen species (ROS) detoxification as a consequence of prenatal androgen excess in males (Table 2). Overall the emergent picture is one of reduced hepatic detoxification and potential liver damage.
Early liver damage
There was no difference between prenatally androgen treated males (14.86 ± 1.67 U/L) and control males (19.46 ± 2.52 U/L) (P = 0.13) in concentrations of plasma alanine aminotransferase (ALT), a marker of hepatocellular damage. This is despite the observation, mentioned above, that ALP was significantly increased in prenatally androgen treated animals. However, the plasma transcriptome predicted fibrosis, liver lesion, hepatic injury and liver cirrhosis (Table 1). We interrogated expression of differentially expressed genes and proteins relevant to liver fibrosis as a potential consequence of liver damage, cell death and proliferation (Table 3). The resulting data suggest that these males have an increased potential for hepatic fibrosis.
Systemic effects
Bioinformatic analysis of the profile of altered plasma proteins in postnatal, adolescent, prenatally androgen treated males was consistent with an increased risk of overall increased morbidity and mortality. Altered pathways included diabetes mellitus, diabetic complications, occlusion of artery and atherosclerosis (Table 1). Indeed, postnatal, adolescent, prenatally androgen treated males had increased levels of fasting plasma insulin (0.55 ± 0.27 ng/ml vs 0.37 ± 0.10 ng/ml; P < 0.05) (Fig. 1A), consistent with a prenatally programmed reduction in health-span in adult males.
Hepatic expression of selected genes in fetal life
Hepatic expression of selected genes (altered in postnatal life by prenatal androgen excess) was investigated in fetal males, on day 90 of pregnancy, exposed to increased levels of androgens and compared to fetal controls. There was no difference in the expression of HMGCS1, HMGCR, NR1H4, RXRG, GSS, GSTM1, GSTO1, GPX4, GPX7, XDH however expression of CYP7A1 and ABCC2 was significantly decreased in prenatally androgenised males (P < 0.05; Fig. 5).
Examination of differentially expressed genes identified in adult life as consequential of prenatal androgen excess by qPCR in fetal livers (day 90 of gestation) during androgen excess treatment. Control (C; n = 10) and prenatal androgen excess (PA; n = 6) male fetal livers were examined by qPCR in terms of genes identified as differentially expressed during adolescence attributable to prenatal androgen status. Statistical testing by unpaired, two tailed Student t test, (*P < 0.05). Of all genes examined, only CYP7A1 and ABCC2 mRNA were depressed in fetal life (in addition to postnatal life) by fetal androgen excess. Box plot whiskers are lowest and highest observed values, box is the upper and lower quartile, with median represented by line in box.
Plasma markers of altered liver function during adolescence
We examined the plasma proteome to determine potential readouts of this pattern of dyslipidaemia associated with reduced hepatic cholesterol synthesis and cholestasis. The plasma proteome highlighted intra-hepatic changes associated with cholesterol homeostasis, detoxification potential, hepatic damage and fibrosis (Table 4). Based upon the observation of reduced expression of CYP7A1 and ABCC2 in both fetal and postnatal livers from prenatal androgen excess pregnancies (Figs. 5,2 and 3 respectively), we performed correlation analyses between these two permanently differentially expressed genes, circulating cholesterol, and plasma biomarkers of cholesterol homeostasis, ROS detoxification and liver damage/fibrosis (Table 4). Based upon strength of correlation, whilst all plasma protein markers showed strong correlation with circulating cholesterol, the panel of fibrotic proteins, COL1A1, COL1A2, COL2A1 and COL5A1 were the most consistent in terms of robust fold changes in the circulation and association with reduced expression of CYP7A1 and ABCC2 genes in the liver (Table 4). This indicated potential utility of circulating monomeric collagen peptides as markers of developing hepatic dysfunction.
Discussion
Suboptimal intrauterine environments can ‘programme’ adverse postnatal health outcomes. First degree male relatives of hyperandrogenaemic PCOS patients exhibit dyslipidaemia and hyperinsulinaemia26,27,28,32. We investigated whether excess developmental androgen exposure could explain postnatal male dyslipidaemia using an outbred clinically-realistic large-animal model, previously shown to parallel female offspring (fetal androgen excess models and human PCOS daughters) pancreatic function outcomes23,24,25,33. Adolescent prenatally androgenised ovine males had increased circulating concentrations of total cholesterol, HDL-C, LDL-C, TG, AMH and insulin, independent of body weight, adiposity and contemporary testosterone concentrations, thus faithfully reproducing the dyslipidaemic / endocrine profile of first degree male relatives of PCOS patients26,27,28,29,30. Neither these observations, nor those discussed below, were recreated by in utero estrogenic excess, underscoring the androgen-specific nature of the alterations reported (see Supplementary data set, Fig. 3), and decreasing the likelihood of metabolism of androgen to estrogen as a component mechanism driving observed effects.
Hypercholesterolemia in prenatally androgen treated males contrasted with downregulated cholesterol biosynthesis pathway genes (HMGCS1, HMGCR, MVD, FDPS, SQLE, CYP51A1, HSD17B7, NSDHL and DHCR7). Such endogenously suppressed de novo cholesterol synthesis, as a negative feedback response to increased circulating cholesterol34, provides biologically consistent insight into why statin treatment is less effective35,36 in some individuals regarding cholesterol normalisation.
Since we did not find evidence of increased cholesterol synthesis, we focused on cholesterol absorption and excretion. Experimental control and prenatally androgen treated animals were fed a standard diet, with no evidence of increased calorific intake in prenatally androgen treated males, either in terms of feeding behaviour or body weight/adiposity. Adolescent prenatally androgen-treated males had increased expression of hepatic chylomicron remnant receptors LRP1 and LRP5, suggestive of increased hepatic uptake of diet-derived cholesteryl esters37,38,39. However, perhaps as a response to increased cholesterol uptake, these prenatally androgen treated animals also had increased expression of ABCG5 and ABCG8 that positively correlated with cholesterol concentrations (r = 0.53; P = 0.002). ABCG5 and ABCG8 are the heterodimer transporter pair that determine clearance of free cholesterol into bile40,41. Furthermore, increased biliary secretion of cholesterol underpins predisposition to gallstone formation42,43,44, in turn increasing risk of extrahepatic cholestasis45.
Adolescent prenatally androgen treated males also displayed increased concentrations of plasma apolipoproteins, notably APOA4, APOC3, APOD and APOM, in agreement with their dyslipidaemic profile, and in turn strongly correlated with circulating cholesterol. APOA4, a component of HDL and chylomicrons, was also over-represented in liver. APOA4, primarily synthesised in small intestine with only minor amounts derived from liver, reflects dietary lipid absorption46,47, and is increased with progression of NAFLD severity48. Clearly, elevated plasma APOA4 likely indicates increased small intestinal lipid absorption in prenatally androgen treated animals, supported by a positive relationship between circulating cholesterol and plasma APOA4, increased ABCG5/8, and absence of relationship between plasma APOA4 protein and hepatic APOA4 mRNA (r 0.19, P > 0.05) or protein (r 0.27, P > 0.05). Increased intestinal cholesterol absorption, in combination with decreased cholesterol synthesis, is strongly associated with elevated cardiovascular risk49,50. This, in tandem with bioinformatic analysis of plasma protein profiles defining atherosclerosis as a disease risk, reinforces the premise of prenatal excess androgen exposure as a factor in male lifelong health.
Whilst we observed no alterations in circulating bile acid concentrations, our prenatally androgen treated males had decreased hepatic CYP7A1 expression. CYP7A1 encodes the rate-limiting enzyme in the classic pathway of cholesterol metabolism to bile acids51 (responsible for 90% of bile acid synthesis in humans52). Altered CYP7A1 expression negatively correlated with total cholesterol (r−0.9, P = 0.004), underscoring the likelihood of decreased metabolism of cholesterol into bile acids, thus offering opportunity for increased secretion / absorption, as witnessed by hepatic ABCG5/8 profiles and plasma APOA4 increase respectively. Pertinently, CYP7A1 gene knockout mice have increased total- and LDL-cholesterol53, and the human CYP7A1 homozygous deletion mutation is associated with hypercholesterolemia, HMG-CoA reductase inhibitor resistance and premature coronary disease54. Conversely, CYP7A1 overexpression is characterized by elevated bile acid pool, increased hepatic cholesterol synthesis and lowered serum cholesterol55,56. Resveratrol can induce CYP7A1 expression, increase bile acid pool, reduce hypercholesterolemia57 and attenuate liver injury / cholestasis58,59, suggesting a potential intervention for prenatally programmed hepatic dysfunction.
Efficient bile acid synthesis is critical for cholesterol homeostasis. Both dyslipidaemia and cholestasis, here predicted by IPA bioinformatics and evidenced by hepatic and circulatory endpoints, may have fetal origins60. CYP7A1 epigenetic modulation during development is an offspring outcome of maternal protein restriction61,62, while intra-uterine calorie restriction causes decreased CYP7A1 expression during fetal and postnatal life, resulting in male-specific hypercholesterolemia61,62. We noted that CYP7A1 was also significantly depressed in fetal livers from prenatally androgen treated males compared to gestational age matched controls. No changes in fetal expression of de novo cholesterol synthesis genes were observed, suggesting decreased cholesterol synthesis in adolescents may be consequential of the novel concept of decreased CYP7A1 expression ‘programmed’ by androgen excess in utero.
Adolescent prenatally androgen treated males had increased hepatic expression of NR1I2 (PXR) and VDR, but decreased NR1H4 (FXR). This in turn may be reflective of altered bile acid signaling63,64,65,66, due to differential receptor regulation by primary and secondary bile acids63,65,67 (partially dependent upon bile acid hydrophobicity68,69,70). Hepatocyte retention of hydrophobic bile acids is causative of liver damage in cholestasis68. Furthermore, the role of FXR in bile acid homeostasis, liver regeneration71, and cholestatic disease72,73 has received significant attention, not least in terms of being a therapeutic target74,75,76. Therefore, this finding of reduced FXR expression is particularly interesting.
ABCC2 mRNA, encoding MRP2, which mediates canalicular excretion and detoxification of a broad range of compounds, including bilirubin and xenobiotics77, was decreased in the livers of our adolescent prenatally androgen treated males. An absence of functional MRP2 from the canalicular membrane is associated with conjugated hyperbilirubinemia and ‘dark pigment’ deposition in hepatocytes in the hereditary ABCC2 deficiency Dubin-Johnson syndrome (DJS)78,79, Eisai hyperbilirubinemic rats80,81 and knockout mouse strains82. These are in close concordance with our observations of increased plasma bilirubin concentration and hepatic accumulation in our prenatally androgen treated males. Decreased ABCC2 may be functionally compensated for by upregulation of ABCC3 (MRP3) and ABCC4 (MRP4) in DJS patients and animal models of cholestasis82,83,84,85. However, prenatally androgen treated males showed no compensatory increase in ABCC3, while ABCC4 expression was downregulated. Our data indicated an extremely strong correlation between depressed ABCC2 and NR1H4 mRNA (encoding FXR) expression (r 0.91, P < 0.0001). This is in keeping with FXR’s regulatory role over ABCC286,87. Adolescent prenatally androgenised males also had decreased expression of RDX mRNA (encoding radixin), which is required for MRP2 anchoring in the canalicular membrane88,89. RDX-deficient mice show increasing serum bilirubin concentrations, culminating in mild liver injury by 8 weeks postnatal age90. Although attenuated MRP2 is not of itself a major cause of cholestasis, it is decreased in patients with cholestasis91,92,93, in turn leading to clinical presentations including fibrosis, cirrhosis, liver failure or hepatobiliary malignancy94. In addition, our prenatally androgen treated male offspring also displayed increased plasma ALP, in the absence of increased ALT, a common characteristic of cholestasis95,96. We conclude, therefore, that an outcome of fetal androgen excess is increased male cholestatic disease risk.
Adolescent prenatally androgen treated males had decreased hepatic expression of GSH synthetase (GSS), glutathione reductase (GSR), (implying altered oxidative stress response97); and glutathione S-transferase genes (GSTM1, GSTM4 and GSTO1) (involved in conjugation prior to transport via MRP efflux pumps98). Decreased GSH levels are associated with disease states99. Such reduced detoxification potential in prenatally androgen treated males was accompanied by increased PXR expression. PXR activates phase I drug metabolizing P450 enzymes, phase II drug conjugation enzymes, and phase III drug transporters, including MRP2100,101,102. However, as discussed above, in our study, ABCC2 and phase II enzyme expression were decreased. PXR (and FXR) form heterodimers with retinoid X receptors RXRA, RXRB and RXRG103. Our adolescent prenatally androgen treated males had markedly decreased hepatic RXRG expression, which in turn may have impacted detoxification enzyme transcript expression. It is noteworthy that several studies in humans and animal models report decreased glutathione in cholestasis104,105,106,107. In agreement with our results, expression of proteins UGT2B7 (protein), GSTM1 (mRNA and protein) and GSTM4 (protein) are reduced in obstructive cholestasis105, and we extend this panel to include GST01 (mRNA and protein). A similar expression profile of nuclear receptors to that which we observe, with increased PXR, VDR, PPARA and decreased FXR expression105, and downregulation of CYP7A1 expression, was also reported in human obstructive cholestasis108. Prenatally androgen treated males also had decreased hepatic glutathione peroxidase 4 (GPX4), peroxiredoxin 5 (PRDX5) and thioredoxin (TXN), antioxidant enzymes that protect cells against oxidative damage109,110,111. In terms of consequential changes linked to altered transcripts/proteins, increased expression of glutathione peroxidase 7 (GPX7), an oxidative stress sensor112, in our prenatally androgen treated males, in combination with decreased plasma catalase (CAT), glutathione peroxidase 3 (GPX3) and superoxide dismutase 3 (SOD3), yields an antioxidant enzyme profile strongly consistent with chronic liver diseases113.
In order to comprehend early life underpinnings of decreased hepatic detoxifying potential, we assessed expression of ABCC2 and selected detoxifying enzymes in livers from control and prenatally androgen treated fetuses. Interestingly, only ABCC2 was downregulated in fetal life by androgen excess, indicative that decreased detoxification potential may be secondary to reduced ABCC2 expression.
Male cholestasis due to anabolic androgenic steroid abuse114 is mechanistically associated with increased oxidative stress115 and deficiency in hepatic canalicular transporters116. Whether this phenomenon is male specific in humans is unknown, but in male rodents, raised testosterone can decrease MRP2 mRNA and protein, with no effect observed in females117. We suggest individualised approaches to treatment, accounting for the stage of cholestatic disease development118 may also benefit from understanding of early life contributions to disease development and/or susceptibility.
Early stage markers of different liver diseases, ideally with prognostic value, are acutely required118. The utility of new molecular techniques for biomarker discovery is demonstrated by shotgun plasma proteomic assessment of NAFLD48. Furthermore, the most robust patho-biochemical signatures identified in a murine model of obstructive cholestasis include increased bilirubin and COL1A1119,120, which we also note elevated in circulation here. We observed increased hepatic expression of collagen genes, and increased collagen proteins in plasma of prenatally androgen treated males. We also noted elevated expression of hepatic transforming growth factor beta induced protein (TGFBI) (also increased in NAFLD plasma)48, SMAD-3 and -7, FGF7 (which correlates with hepatic fibrosis severity121,122 and suppresses CYP7A1 during fibrosis development123), FGFR2 and FGFR3. Collectively, this indicated a pro-tissue remodelling/pro-fibrotic hepatic environment developing. In light of collagen-1 as a potential prognostic marker of fibrosis in biliary atresia124, we suggest that in combination with circulating cholesterol and bilirubin, COL1A1, COL1A2, COL2A1 and COL5A1 may constitute circulating markers of early stage, pre-symptomatic cholestatic disease development in at risk individuals.
We recognise three major limitations to this work. Only a single postnatal time point was studied; however, we see no suggestions here that the condition we describe was likely to spontaneously resolve in later life. We acknowledge species difference between humans and sheep, and that a single dose of testosterone was used, nonetheless, our data closely parallels human clinical outcomes (dyslipidaemia and hormonal alterations) of hyperandrogenaemic pregnancy conditions, and mechanistic investigations performed in laboratory rodent models of hepatic disease
In conclusion, the legacy of male fetal androgenic excess is the development of a cholestasis-like condition in adolescence, accompanied by dyslipidaemia and increased fibrotic potential. We report fetal androgen excess as a risk factor for poor male hepatic/metabolic health.
Materials and Methods
Statistical analysis
All experiments were carried out on n = 14 male offspring per group, except in the case of a limited estrogenic overexposure comparison (see Supplementary Materials), and in the case of fetal tissue analyses (actual numbers described in methods and results sections). No animals or data were excluded from any of the analyses described. RNAseq and proteomic data was examined by pairwise comparisons, using Benjamini and Hochberg (1995) false discovery rate control. In cases of single gene analyses, all data sets were normality tested prior to further analysis (Shapiro-Wilk test), and logarithmically transformed if necessary. For comparing means of two treatment groups with equal variances, unpaired, two-tailed Student’s t test was used accepting P < 0.05 as significant. Bonferroni multiple test correction was utilised in the case of data subjected to t-test. A Chi-square test was used for testing categorical variables. Correlation was assessed by calculation of Pearson product-moment co-efficient. Statistical analysis software package R (version 3.4.0) was utilised for all RNAseq and proteomic analyses, and the remainder of statistical analysis was performed using GraphPad Prism 8.0 software (GraphPad Prism Software, San Diego, CA, USA). All P values are cited in full.
Study approval
All studies were approved by the UK Home Office and conducted under approved Project Licence PPL 60/4401, reviewed by The University of Edinburgh Animal Research Ethics Committee.
Animals and tissue collection
All animal based components of the study were conducted under natural lighting conditions with no environmental manipulations. Animal husbandry, experimental protocols and tissue collection were performed as are exactly as previously described23,24. Briefly, mature Scottish Greyface ewes were fed in order to achieve a comparable body condition score (2.75–3) prior to estrous cycle synchronisation. After a synchronised mating (Texel ram), animals were allocated at random to one of two experimental groups – vehicle control or testosterone propionate (TP) exposed. Treatment occurred on day 62 and day 82 of gestation (post-sexual differentiation, designed to maintain elevated testosterone concentrations over experimental period23,24). Anaesthesia was induced on day 62 of gestation by initial sedation via administration of 10 mg Xylazine i.m (‘Rompun’, Baylor plc Animal Health Division, UK), followed by 2 mg/kg ketamine (i.v, Keteset, Fort Dodge Animal Health, UK). All downstream procedures were conducted under surgical aseptic conditions. TP was dissolved in vegetable oil (100 mg/ml) and a 200 μl volume injected (20 G Quinke spinal needle, BD Biosciences) via ultrasound guidance into the fetal flank (intraperitoneal injection). Control fetuses received 200 μl vegetable oil vehicle alone. In order to account for potential aromatisation of testosterone to estrogen, and thus incorrect interpretation of effects being attributable to androgenic excess, an additional treatment group of estrogenic overexposure (n = 8) was created by exposing fetuses, in an identical manner to androgen excess exposure, by injection of 200 µl of 20 mg/ml DES (diethylstilbesterol), this lower dose as compared to testosterone being selected on the basis of binding affinity of DES as compared to the naturally occurring estradiol receptor ligand estradiol-17β125. These procedures were repeated on d82 of gestation. Immediately after surgical procedure completion all pregnant ewes were given prophylactic antibiotics (Streptacare, Animalcare Ltd, UK, 1 ml/25 kg) and were then monitored during recovery; no adverse effects of these procedures were observed.
Fetal tissue collection was as previously described, performed on day 90 of gestation (day 147 is term)23,24.
Offspring were lambed and reared conventionally (weaned at 3 months postnatal age). At birth, and during postnatal life, we saw no phenotypic evidence of any deleterious effects of the treatment regimens used. In female children of PCOS sufferers, early puberty (Tanner 1 developmental stage) is when measurable metabolic perturbations (hyperinsulinaemia) begin to manifest, hence the 6 months postnatal age studied here, as an equivalent of the human observations126. Prior to sacrifice at 6 months of age, all animals received a bolus injection of glucose (10 g glucose in 20 ml saline), and 15 minutes post injection sacrifice was achieved via barbiturate overdose. Animals were sacrificed in a random order with respect to treatments. Tissues were recovered immediately, and snap frozen prior to storage at −80 °C until downstream analysis.
In both fetal and postnatal tissue collection, liver sampling occurred from the same lobe (right posterior), in approximately the same area, and was immediately snap frozen, then stored at −80 °C until further processing in the case of nucleic acid and protein extraction, or fixed and processed as previously described23,24. These procedures were repeated over two breeding seasons, maintaining all conditions identically, to ensure sufficient numbers of male offspring for analysis.
RNA and protein extraction
RNA and protein were extracted and purified using Qiagen AllPrep kits, following manufacturer’s instructions. All samples were randomised within a larger study set to control for potential batch effects. RNA concentrations were determined using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific, UK), with Agilent Bioanalyser analysis utilised for RNA quality control – all samples processed for downstream analysis recorded a RIN value of >7.5. RNA and protein extracts were stored at −80 °C until further analysis.
RNA sequencing transcriptomic analyses
1 μl of ERCC (External RNA Controls Consortium) spike in controls were added to 500 ng of RNA sample to permit library quality assessment and estimation of lowest limit of detection127. One of the two ERCC mixes were selected at random for each sample. Libraries were prepared with the Illumina TruSeq Stranded mRNA kit, using fetal-vehicle treated (control) liver samples (n = 14) and fetal-testosterone propionate treated liver samples (n = 14), sampled at 6 months postnatal age. Sequencing was performed on the NextSeq. 500 High Output v2 kit (75 cycles) on the Illumina NextSeq. 500 platform producing 75 bp single end reads. Raw sequencing data is available from the ArrayExpress database (http://www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-8032. To assess quality of sequencing data, reads were analysed with FastQC (version 0.11.3)128. To remove any lower quality and adapter sequences, TrimGalore! (version 0.4.0)129 was used to filter the reads (phred quality score threshold of 30). To remove the ERCC reads, all reads were aligned to the ERCC reference genome using HISAT2 (version 2.1.0)130. These alignments were processed using SAMtools (version 1.2)131, reads were counted using featureCounts (part of the sub read version 5.0.1 package)132 and analysed using the R package erccdashboard (version 1.6.0)133.
The most recent release of the sheep Ovis aries reference genome was downloaded from NCBI (https://www.ncbi.nlm.nih.gov/genome?term = ovis%20aries)134 along with the equivalent annotation file, and reads were aligned to this reference using HISAT2 (version 2.1.0)130 with the parameter for stranded library preparation used. SAMtools (version 1.2)131 was used to process the alignments and reads were counted at gene locations using featureCounts (part of the sub read version 5.0.1 package)132 utilising the parameter to split multi-mapped reads as a fraction across all genes that they align to. Pairwise gene comparisons were carried out with edgeR (version 3.16.5)135 with all genes with CPM (count per million) value of more than one in six kept for analysis, and all other genes removed as low count genes, leaving 15134 genes for analysis. P values were adjusted using the Benjamini-Hochberg procedure, with a false discovery rate (FDR) set at q < 0.05136.
Liver and plasma protein quantification
Hepatic and plasma proteins were identified and quantified using a Q Exactive Plus hybrid quadrupole Orbitrap mass spectrometer fitted with an EASY-Spray nano-ESI source (Thermo Scientific) as previously described137. Briefly, 10 micrograms of proteins were diluted in 100 μl of 50 mM NH4HCO3 (BioUltra grade, Sigma Aldrich), and were reduced, alkylated and digested with trypsin overnight according to the PRIME-XS protocol (http://www.primexs.eu/protocols/Public-Documents/04%2D%2D-Protocols/PRIME-XS-Protocol-NPC-In-Solution Digestion.pdf/). The equivalent of 2 μg of peptides (assuming no losses) were analyzed by liquid chromatography tandem mass spectrometry (LC-MS/MS). All peptide matching searches were performed against FASTA file of the ovine, bovine, swine, equine, and caprine proteomes (canonical and isoform sequences retrieved from Uniprot on the 22nd of February 2018). Match between runs was used to identify peptide signals lacking MS/MS information. Protein intensities across samples were normalized using the maxLFQ algorithm138. Normalised protein intensities for all proteins were then extracted from the results file for the downstream statistical analyses. Pairwise protein comparisons were carried out using limma139 for those proteins that yielded normalised intensities in at least 75% of the compared samples. P values were adjusted using the Benjamini-Hochberg procedure136. Mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE140 partner repository with the dataset identifier PXD014050.
Quantitative PCR
Quantitative PCR was performed exactly as previously described23, utilising Genorm analysis (PrimerDesign Ltd, UK) in order to identify a panel of two stable housekeeping genes – thus the geometric mean of ATPsynth and RPL19 was used as the normalisation reference. Negative controls consisted of an RT-ve and a template negative reaction. Primers were designed in house using the Primer3Plus bioinformatics software tool, were synthesised by Eurofins MWG Operon, Germany, and were validated prior to use (Supplementary Table 1), or were outsourced and designed by PrimerDesign Ltd, (UK).
Bioinformatic analysis
IPA (Ingenuity Pathway Analysis, Qiagen) was used to perform initial screens of affected pathways, and to identify potential disease states associated with the differentially expressed genes (DEG’s) and differentially expressed proteins (DEP’s). Pre-filtering of data uploaded to IPA was based upon FDR adjusted significance, using q < 0.05 as cut-off.
IPA P values of <0.05 (Right-tailed Fishers exact test) delivered information regarding association of significant DEG with biological processes, and directional change with regards to such biological pathways in health and disease generated where possible. Given smaller numbers of proteins identified, in order to generate meaningful bioinformatics analyses, we relaxed the stringency of input by using nominal P values of <0.05.
Plasma analyte determination
Concentrations of fasting plasma total cholesterol, high density lipoprotein (HDL), Triglyceride (TG) total bilirubin, alanine aminotransferase (ALT) and alkaline phosphatase (ALP) were obtained using commercial assay kits (Alpha Laboratories Ltd., Eastleigh, UK) as per manufacturer’s instruction, using a Cobas Mira automated analyser (Roche Diagnostics Ltd, UK). Low density lipoprotein (LDL) was calculated by the Friedewald equation141. For all assays within-run and intra-batch precisions were <4% CV and <5% CV, respectively. Fasting plasma bile acid concentration was measured using Total Bile Acid Assay Kit (STA-631; Cell Biolabs, INC., San Diego, US), as per the manufacturer’s instructions with all samples assayed in duplicate. The assay sensitivity was 0.39 µM and intra-assay CV was <4%. Plasma insulin was measured using the ALPCO Ovine Insulin Elisa kit (80-INSOV-E01; American Laboratory Products Company, Salem, US) as per the manufacturer’s instructions, with all samples measured in duplicate. The assay sensitivity was 0.15 ng/ml and intra-assay CV was <4.5%. Testosterone was measured by in-house ELISA. Briefly, 96-well plates (Greiner Bio-One GmbH, Germany) were coated with 100 µl of donkey anti rabbit IgG (Jackson ImmunoResearch Inc, USA) (1:500 in ELISA coating buffer, 100 mM Sodium Bicarbonate, pH 9.6) and incubated (4 °C, 12 hours). After washing twice (0.05 M Tris/HCl + 0.05% Tween 20, pH 7.4), 250 μl of Blocking buffer was added and incubated for 1 h (room temperature with shaking), then washed twice. Standards (0–24.3 ng/ml), samples and controls (20 µl per well) were added to each well, followed by 80 µl of Testosterone-HRP conjugate (Astra Biotech GmbH, Germany) at 1:20,000 in assay buffer (PBS pH 7.4, 0.1%BSA with 250 ng/ml cortisol (to displace any testosterone bound to binding globulins)), followed by 50 μl of Rabbit anti Testosterone −19 antibody (AMS Biotechnology, USA) (1:200,000). Plates were incubated (room temperature, 2 hours) then washed 5 times. 120 µl of substrate solution (3,3,5,5-Tetramethylbenzidine, Millipore Corporation, USA) was added and incubated (room temperature, in darkness, 20 min). Reaction was stopped by adding 80 µl of 2 N H2SO4 solution, and. plates read at 450 nm. The inter-assay CV for low and high testosterone QC pools were 11.4 and 9.1% respectively; intra-assay CV’s were 8.9 and 5.6%, with detection limit of 0.1 ng/ml. Dihydrotestosterone cross-reacted 20.4%, all other tested steroids showed <0.2% cross-reactivity.
Hepatic bilirubin content
Wax embedded tissue sections (5 μm; 2 per sample), were mounted on to charged slides (Superfrost Plus, Thermoscientific, Epsom, UK), then rehydrated through a graded alcohol series. Halls Bilirubin stain was then performed, using Fouchets and Van Giesons reagents (Sigma Aldrich, Poole, Dorset) following an online protocol (https://webpath.med.utah.edu/HISTHTML/MANUALS/BILE.PDF). Assessment by light microscopy was performed by two independent operators, both blinded to treatment. Scoring was semiquantitive, categorical based, where 0 = no bilirubin deposits, 1 = mild and 2 = intense bilirubin deposits).
References
Dumesic, D. A., Abbott, D. H. & Padmanabhan, V. Polycystic ovary syndrome and its developmental origins. Rev. Endocr. Metab. Disord. 8, 127–141 (2007).
Franks, S., McCarthy, M. I. & Hardy, K. Development of polycystic ovary syndrome: involvement of genetic and environmental factors. Int. J. Androl. 29, 278–290 (2006).
Warner, M. J. & Ozanne, S. E. Mechanisms involved in the developmental programming of adulthood disease. Biochem. J. 427, 333–347 (2010).
Treviño, L. S. & Katz, T. A. Endocrine disruptors and developmental origins of nonalcoholic fatty liver disease. Endocrinology 159, 20–31 (2018).
Fowler, P. A. et al. Impact of endocrine-disrupting compounds (EDCs) on female reproductive health. Moll. Cell. Endocrinol. 355, 231–239 (2012).
Bergman, Å. et al. Science and policy on endocrine disrupters must not be mixed: a reply to a ‘common sense’ intervention by toxicology journal editors. Environ. Health 12, 69–72 (2013).
Health and Environment Alliance (HEAL), https://www.env-health.org (2019).
Hunt, P. A., Sathyanarayana, S., Fowler, P. A. & Trasande, L. Female reproductive disorders, diseases, and dosts of exposure to endocrine disrupting chemicals in the European Union. J. Clin. Endocrinol. Metab. 101, 1562–1570 (2016).
Wilson, J. D., George, F. W. & Griffin, J. E. The hormonal control of sexual development. Science 211, 1278–1284 (1981).
O’Shaughnessy, P. J. et al. Alternative (backdoor) androgen production and masculinization in the human fetus. PLoS Biol. 17, e3000002 (2019).
Thankamony, A., Pasterski, V., Ong, K. K., Acerini, C. L. & Hughes, I. A. Anogenital distance as a marker of androgen exposure in humans. Andrology 4, 616–625 (2016).
Hakim, C., Padmanabhan, V. & Vyas, A. K. Gestational hyperandrogenism in developmental programming. Endocrinology 158, 199–212 (2017).
Filippou, P. & Homburg, R. Is foetal hyperexposure to androgens a cause of PCOS? Hum. Reprod. Update 23, 421–432 (2017).
Sánchez-Ferrer, M. L. et al. Presence of polycystic ovary syndrome is associated with longer anogenital distance in adult Mediterranean women. Hum. Reprod. 32, 2315–2323 (2017).
Barrett, E. S. et al. Anogenital distance in newborn daughters of women with polycystic ovary syndrome indicates fetal testosterone exposure. J. Dev. Orig. Health Dis. 9, 307–314 (2018).
Barry, J. A. et al. Umbilical vein testosterone in female infants born to mothers with polycystic ovary syndrome is elevated to male levels. J. Obstet. Gynaecol. 30, 444–446 (2010).
Abbott, D. H., Barnett, D. K., Bruns, C. M. & Dumesic, D. A. Androgen excess fetal programming of female reproduction: a developmental aetiology for polycystic ovary syndrome? Hum. Reprod. Update 11, 357–374 (2005).
Roland, A. V., Nunemaker, C. S., Keller, S. R. & Moenter, S. M. Prenatal androgen exposure programs metabolic dysfunction in female mice. J. Endocrinol. 207, 213–223 (2010).
Abbott, D. H. et al. Experimentally induced gestational androgen excess disrupts glucoregulation in rhesus monkey dams and their female offspring. Am. J. Physiol. Endocrinol. Metab. 299, E741–E751 (2010).
Padmanabhan, V., Veiga-Lopez, A., Abbott, D. H., Recabarren, S. E. & Herkimer, C. Developmental programming: impact of prenatal testosterone excess and postnatal weight gain on insulin sensitivity index and transfer of traits to offspring of overweight females. Endocrinology 151, 595–605 (2010).
Hogg, K., Wood, C., McNeilly, A. S. & Duncan, W. C. The in utero programming effect of increased maternal androgens and a direct fetal intervention on liver and metabolic function in adult sheep. PLoS ONE 6, e24877 (2011).
Hogg, K. et al. Enhanced thecal androgen production is prenatally programmed in an ovine model of polycystic ovary syndrome. Endocrinology 153, 450–461 (2012).
Rae, M. et al. The pancreas is altered by in utero androgen exposure: implications for clinical conditions such as polycystic ovary syndrome (PCOS). PLoS ONE 8, e56263 (2013).
Ramaswamy, S. et al. Developmental programming of polycystic ovary syndrome (PCOS): prenatal androgens establish pancreatic islet α/β cell ratio and subsequent insulin secretion. Sci. Rep. 6, 27408 (2016).
Nicol, L. E. et al. Abnormal infant islet morphology precedes insulin resistance in PCOS-like monkeys. PLoS ONE 9, e106527 (2014).
Sam, S., Coviello, A. D., Sung, Y.-A., Legro, R. S. & Dunaif, A. Metabolic phenotype in the brothers of women with polycystic ovary syndrome. Diabetes Care 31, 1237–1241 (2008).
Baillargeon, J. P. & Carpentier, A. C. Brothers of women with polycystic ovary syndrome are characterised by impaired glucose tolerance, reduced insulin sensitivity and related metabolic defects. Diabetologia 50, 2424–2432 (2007).
Recabarren, S. E. et al. Metabolic profile in sons of women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 93, 1820–1826 (2008).
Crisosto, N. et al. Reproductive and metabolic features during puberty in sons of women with polycystic ovary syndrome. Endocr. Connect. 6, 607–613 (2017).
Recabarren, S. E. et al. Pituitary and testicular function in sons of women with polycystic ovary syndrome from infancy to adulthood. J. Clin. Endocrinol. Metab. 93, 3318–3324 (2008).
Jenkins, S. J., Green, S. A. & Clark, P. A. Clinical chemistry reference values of normal domestic animals in various age groups–as determined on the ABA-100. Cornell Vet. 72, 403–415 (1982).
Kaushal, R., Parchure, N., Bano, G., Kaski, J.-C. & Nussey, S. S. Insulin resistance and endothelial dysfunction in the brothers of Indian subcontinent Asian women with polycystic ovaries. Clin. Endocrinol. 60, 322–328 (2004).
Sir-Petermann, T. et al. Early metabolic derangements in daughters of women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 92, 4637–4642 (2007).
Jones, P. J. et al. Dietary cholesterol feeding suppresses human cholesterol synthesis measured by deuterium incorporation and urinary mevalonic acid levels. Arterioscler. Thromb. Vasc. Biol. 16, 1222–1228 (1996).
Goettsch, W. G. et al. Statins are less effective in common daily practice among patients with hypercholesterolemia: the REALITY-PHARMO study. Curr. Med. Res. Opin. 20, 1025–1033 (2004).
Kuver, R. The effects of statins on cholestasis: good, bad or indifferent? J Gastroenterol. Hepatol. 26, 1467–1469 (2011).
Cooper, A. D. Hepatic uptake of chylomicron remnants. J. Lipid Res. 38, 2173–2192 (1997).
Frazier-Wood, A. C. et al. The association between LRP-1 variants and chylomicron uptake after a high fat meal. Nutr. Metab. Cardiovasc. Dis. 23, 1154–1158 (2013).
Fujino, T. et al. Low-density lipoprotein receptor-related protein 5 (LRP5) is essential for normal cholesterol metabolism and glucose-induced insulin secretion. Proc. Natl. Acad. Sci. USA 100, 229–234 (2003).
Yu, L. et al. Overexpression of ABCG5 and ABCG8 promotes biliary cholesterol secretion and reduces fractional absorption of dietary cholesterol. J. Clin. Invest. 110, 671–680 (2002).
Graf, G. A. et al. ABCG5 and ABCG8 are obligate heterodimers for protein trafficking and biliary cholesterol excretion. J. Biol. Chem. 278, 48275–48282 (2003).
Wittenburg, H. et al. FXR and ABCG5/ABCG8 as determinants of cholesterol gallstone formation from quantitative trait locus mapping in mice. Gastroenterology 125, 868–881 (2003).
Buch, S. et al. A genome-wide association scan identifies the hepatic cholesterol transporter ABCG8 as a susceptibility factor for human gallstone disease. Nature 39, 995–999 (2007).
Rudkowska, I. & Jones, P. J. Polymorphisms in Abcg5/G8 transporters linked to hypercholesterolemia and gallstone disease. Nutr. Rev. 66, 343–348 (2008).
Mortelé, K. J., Wiesner, W., Cantisani, V., Silverman, S. G. & Ros, P. R. Usual and unusual causes of extrahepatic cholestasis: assessment with magnetic resonance cholangiography and fast MRI. Abdom. Imaging 29, 87–99 (2004).
Wang, F. et al. Apolipoprotein A-IV: a protein intimately involved in metabolism. J. Lipid Res. 56, 1403–1418 (2015).
Kalogeris, T. J., Rodriguez, M.-D. & Tso, P. Control of synthesis and secretion of intestinal apolipoprotein A-IV by lipid. J. Nutr. 127, 537S–543S (1997).
Niu, L. et al. Plasma proteome profiling discovers novel proteins associated with non-alcoholic fatty liver disease. Mol. Syst. Biol. 15, e8793 (2019).
Weingärtner, O., Lutjohann, D., Böhm, M. & Laufs, U. Relationship between cholesterol synthesis and intestinal absorption is associated with cardiovascular risk. Atherosclerosis 210, 362–365 (2010).
Matthan, N. R. et al. Alterations in cholesterol absorption/synthesis markers characterize Framingham Offspring Study participants with CHD. J. Lipid Res. 50, 1927–1935 (2009).
Björkhem, I. et al. Differences in the regulation of the classical and the alternative pathway for bile acid synthesis in human liver. No coordinate regulation of CYP7A1 and CYP27A1. J. Biol. Chem. 277, 26804–26807 (2002).
Duane, W. C. & Javitt, N. B. 27-hydroxycholesterol: production rates in normal human subjects. J. Lipid Res. 40, 1194–1199 (1999).
Erickson, S. K. et al. Hypercholesterolemia and changes in lipid and bile acid metabolism in male and female cyp7A1-deficient mice. J. Lipid Res. 44, 1001–1009 (2003).
Pullinger, C. R. et al. Human cholesterol 7alpha-hydroxylase (CYP7A1) deficiency has a hypercholesterolemic phenotype. J. Clin. Invest. 110, 109–117 (2002).
Li, T., Francl, J. M., Boehme, S. & Chiang, J. Y. L. Regulation of cholesterol and bile acid homeostasis by the cholesterol 7α‐hydroxylase/steroid response element‐binding protein 2/microRNA‐33a axis in mice. Hepatology 58, 1111–1121 (2013).
Li, T. et al. Overexpression of cholesterol 7α-hydroxylase promotes hepatic bile acid synthesis and secretion and maintains cholesterol homeostasis. Hepatology 53, 996–1006 (2011).
Chen, Q., Wang, E., Ma, L. & Zhai, P. Dietary resveratrol increases the expression of hepatic 7α-hydroxylase and ameliorates hypercholesterolemia in high-fat fed C57BL/6J mice. Lipids Health Dis. 11, 56 (2012).
Wang, T. et al. Resveratrol effectively attenuates α-naphthyl-isothiocyanate-induced acute cholestasis and liver injury through choleretic and anti-inflammatory mechanisms. Acta Pharmacol. Sin. 35, 1527–1536 (2014).
Dolezelova, E. et al. Resveratrol modifies biliary secretion of cholephilic compounds in sham-operated and cholestatic rats. World J. Gastroenterol. 23, 7678–7692 (2017).
Calkins, K. & Devaskar, S. U. Fetal origins of adult disease. Curr. Probl. Pediatr. Adolesc. Health Care 41, 158–176 (2011).
Liu, X. et al. Maternal protein restriction induces alterations in hepatic tumor necrosis factor-α/CYP7A1 signaling and disorders regulation of cholesterol metabolism in the adult rat offspring. J. Clin. Biochem. Nutr. 55, 40–47 (2014).
Sohi, G., Marchand, K., Revesz, A., Arany, E. & Hardy, D. B. Maternal protein restriction elevates cholesterol in adult rat offspring due to repressive changes in histone modifications at the cholesterol 7alpha-hydroxylase promoter. Mol. Endocrinol. 25, 785–798 (2011).
Xie, W. et al. An essential role for nuclear receptors SXR/PXR in detoxification of cholestatic bile acids. Proc. Natl. Acad. Sci. USA 98, 3375–3380 (2001).
Staudinger, J. L. et al. The nuclear receptor PXR is a lithocholic acid sensor that protects against liver toxicity. Proc. Natl. Acad. Sci. USA 98, 3369–3374 (2001).
Makishima, M. et al. Vitamin D receptor as an intestinal bile acid sensor. Science 296, 1313–1316 (2002).
Hofmann, A. F. Detoxification of lithocholic acid, a toxic bile acid: relevance to drug hepatotoxicity. Drug Metab. Rev. 36, 703–722 (2004).
Makishima, M. et al. Identification of a nuclear receptor for bile acids. Science 284, 1362–1365 (1999).
Perez, M.-J. & Briz, O. Bile-acid-induced cell injury and protection. World J. Gastroenterol. 15, 1677–1689 (2009).
Attili, A. F., Angelico, M., Cantafora, A., Alvaro, D. & Capocaccia, L. Bile acid-induced liver toxicity: relation to the hydrophobic-hydrophilic balance of bile acids. Med. Hypotheses 19, 57–69 (1986).
Sârbu, C., Kuhajda, K. & Kevresan, S. Evaluation of the lipophilicity of bile acids and their derivatives by thin-layer chromatography and principal component analysis. J. Chromatogr. A 917, 361–366 (2001).
Li, G. & L Guo, G. Farnesoid X receptor, the bile acid sensing nuclear receptor, in liver regeneration. Acta Pharm. Sin. B 5, 93–98 (2015).
Cariello, M., Piccinin, E., Garcia-Irigoyen, O., Sabbà, C. & Moschetta, A. Nuclear receptor FXR, bile acids and liver damage: Introducing the progressive familial intrahepatic cholestasis with FXR mutations. BBA – Mol. Basis. Dis. 1864, 1308–1318 (2018).
Gomez-Ospina, N. et al. Mutations in the nuclear bile acid receptor FXR cause progressive familial intrahepatic cholestasis. Nat. Commun. 7, 10713 (2016).
Jonker, J. W., Liddle, C. & Downes, M. FXR and PXR: Potential therapeutic targets in cholestasis. J. Steroid Biochem. Mol. Biol. 130, 147–158 (2012).
Fiorucci, S. et al. Targeting FXR in cholestasis: hype or hope. Expert Opin. Ther. Targets 18, 1449–1459 (2014).
van de Wiel, S. M. W., Bijsmans, I. T. G. W., van Mil, S. W. C. & van de Graaf, S. F. J. Identification of FDA-approved drugs targeting the Farnesoid X Receptor. Sci. Rep. 9, 2193 (2019).
Jemnitz, K. et al. ABCC2/Abcc2: a multispecific transporter with dominant excretory functions. Drug Metab. Rev. 42, 402–436 (2010).
Kartenbeck, J., Leuschner, U., Mayer, R. & Keppler, D. Absence of the canalicular isoform of the MRP gene-encoded conjugate export pump from the hepatocytes in Dubin-Johnson syndrome. Hepatology 23, 1061–1066 (1996).
Dubin, I. N. & Johnson, F. B. Chronic idiopathic jaundice with unidentified pigment in liver cells; a new clinicopathologic entity with a report of 12 cases. Medicine (Baltimore) 33, 155–197 (1954).
Chen, C. et al. Up-regulation of Mrp4 expression in kidney of Mrp2-deficient TR- rats. Biochem. Pharmacol. 70, 1088–1095 (2005).
Johnson, B. M., Zhang, P., Schuetz, J. D. & Brouwer, K. L. R. Characterization of transport protein expression in multidrug resistance-associated protein (Mrp) 2-deficient rats. Drug Metab. Dispos. 34, 556–562 (2006).
Chu, X.-Y. et al. Characterization of mice lacking the multidrug resistance protein MRP2 (ABCC2). J. Pharmacol. Exp. Ther. 317, 579–589 (2006).
Donner, M. G. & Keppler, D. Up-regulation of basolateral multidrug resistance protein 3 (Mrp3) in cholestatic rat liver. Hepatology 34, 351–359 (2001).
Kuroda, M. et al. Increased hepatic and renal expressions of multidrug resistance-associated protein 3 in Eisai hyperbilirubinuria rats. J. Gastroenterol. Hepatol. 19, 146–153 (2004).
König, J., Rost, D., Cui, Y. & Keppler, D. Characterization of the human multidrug resistance protein isoform MRP3 localized to the basolateral hepatocyte membrane. Hepatology 29, 1156–1163 (1999).
Kast, H. R. et al. Regulation of multidrug resistance-associated protein 2 (ABCC2) by the nuclear receptors pregnane X receptor, farnesoid X-activated receptor, and constitutive androstane receptor. J. Biol. Chem. 277, 2908–2915 (2002).
Urquhart, B. L., Tirona, R. G. & Kim, R. B. Nuclear receptors and the regulation of drug-metabolizing enzymes and drug transporters: implications for interindividual variability in response to drugs. J. Clin. Pharmacol. 47, 566–578 (2007).
Kojima, H. et al. The role of radixin in altered localization of canalicular conjugate export pump Mrp2 in cholestatic rat liver. Hepatol. Res. 38, 202–210 (2008).
Rost, D., Kloeters-Plachky, P. & Stiehl, A. Retrieval of the rat canalicular conjugate export pump Mrp2 is associated with a rearrangement of actin filaments and radixin in bile salt-induced cholestasis. Eur. J. Med. Res. 13, 314–318 (2008).
Kikuchi, S. et al. Radixin deficiency causes conjugated hyperbilirubinemia with loss of Mrp2 from bile canalicular membranes. Nature 31, 320–325 (2002).
Shoda, J. et al. The expression levels of plasma membrane transporters in the cholestatic liver of patients undergoing biliary drainage and their association with the impairment of biliary secretory function. Am. J. Gastroenterol. 96, 3368–3378 (2001).
Chai, J. et al. Canalicular membrane MRP2/ABCC2 internalization is determined by Ezrin Thr567 phosphorylation in human obstructive cholestasis. J. Hepatol. 63, 1440–1448 (2015).
Kojima, H. et al. Disturbed colocalization of multidrug resistance protein 2 and radixin in human cholestatic liver diseases. J. Gastroenterol. Hepatol. 23, e120–8 (2008).
Hirschfield, G. M., Heathcote, E. J. & Gershwin, M. E. Pathogenesis of cholestatic liver disease and therapeutic approaches. Gastroenterology 139, 1481–1496 (2010).
Giannini, E. G., Testa, R. & Savarino, V. Liver enzyme alteration: a guide for clinicians. CMAJ 172, 367–379 (2005).
Siddique, A. & Kowdley, K. V. Approach to a patient with elevated serum alkaline phosphatase. Clin. Liver Dis. 16, 199–229 (2012).
Zitka, O. et al. Redox status expressed as GSH:GSSG ratio as a marker for oxidative stress in paediatric tumour patients. Oncol. Lett. 4, 1247–1253 (2012).
Board, P. G. & Menon, D. Glutathione transferases, regulators of cellular metabolism and physiology. Biochim. Biophys. Acta Gen. Subj. 1830, 3267–3288 (2013).
Ballatori, N. et al. Glutathione dysregulation and the etiology and progression of human diseases. Biol. Chem. 390, 191–214 (2009).
Ihunnah, C. A., Jiang, M. & Xie, W. Nuclear receptor PXR, transcriptional circuits and metabolic relevance. BBA – Mol. Basis. Dis. 1812, 956–963 (2011).
Sonoda, J. et al. Regulation of a xenobiotic sulfonation cascade by nuclear pregnane X receptor (PXR). Proc. Natl. Acad. Sci. USA 99, 13801–13806 (2002).
Gong, H. et al. Orphan nuclear receptor pregnane X receptor sensitizes oxidative stress responses in transgenic mice and cancerous cells. Mol. Endocrinol. 20, 279–290 (2006).
Evans, R. M. & Mangelsdorf, D. J. Nuclear receptors, RXR, and the big bang. Cell 157, 255–266 (2014).
Yang, H. et al. Dysregulation of glutathione synthesis during cholestasis in mice: Molecular mechanisms and therapeutic implications. Hepatology 49, 1982–1991 (2009).
Chai, J. et al. Hepatic expression of detoxification enzymes is decreased in human obstructive cholestasis due to gallstone biliary obstruction. PLoS ONE 10, e0120055 (2015).
Neuschwander-Tetri, B. A., Nicholson, C., Wells, L. D. & Tracy, T. F. Cholestatic liver injury down-regulates hepatic glutathione synthesis. J. Surg. Res. 63, 447–451 (1996).
Vendemiale, G., Grattagliano, I., Lupo, L., Memeo, V. & Altomare, E. Hepatic oxidative alterations in patients with extra-hepatic cholestasis. Effect of surgical drainage. J. Hepatol. 37, 601–605 (2002).
Chai, J. et al. Elevated hepatic multidrug resistance-associated protein 3/ATP-binding cassette subfamily C3 expression in human obstructive cholestasis is mediated through tumor necrosis factor alpha and c-Jun NH2-terminal kinase/stress-activated protein kinase-signaling pathway. Hepatology 55, 1485–1494 (2012).
Brigelius-Flohé, R. & Maiorino, M. Glutathione peroxidases. Biochim. Biophys. Acta Gen. Subj. 1830, 3289–3303 (2013).
Perkins, A., Nelson, K. J., Parsonage, D., Poole, L. B. & Karplus, P. A. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem. Sci. 40, 435–445 (2015).
Lu, J. & Holmgren, A. The thioredoxin antioxidant system. Free Radic. Biol. Med. 66, 75–87 (2014).
Chen, Y.-I., Wei, P.-C., Hsu, J.-L., Su, F.-Y. & Lee, W.-H. NPGPx (GPx7): a novel oxidative stress sensor/transmitter with multiple roles in redox homeostasis. Am. J. Transl. Res. 8, 1626–1640 (2016).
Czuczejko, J., Zachara, B. A., Staubach-Topczewska, E., Halota, W. & Kedziora, J. Selenium, glutathione and glutathione peroxidases in blood of patients with chronic liver diseases. Acta Biochim. Pol. 50, 1147–1154 (2003).
Elsharkawy, A. M. et al. Cholestasis secondary to anabolic steroid use in young men. BMJ 344, e468–e468 (2012).
Bond, P., Llewellyn, W. & Van Mol, P. Anabolic androgenic steroid-induced hepatotoxicity. Med. Hypotheses 93, 150–153 (2016).
Sherrif, E. Y. et al. Hepatotoxicity from anabolic androgenic steroids marketed as dietary supplements: contribution from ATP8B1/ABCB11 mutations? Liver Int. 33, 1266–1270 (2013).
Simon, F. R. et al. Hormonal regulation of hepatic multidrug resistance-associated protein 2 (Abcc2) primarily involves the pattern of growth hormone secretion. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G595–G608 (2006).
Jansen, P. L. M. et al. The ascending pathophysiology of cholestatic liver disease. Hepatology 65, 722–738 (2017).
Abshagen, K. et al. Pathobiochemical signatures of cholestatic liver disease in bile duct ligated mice. BMC Syst. Biol. 9, 83 (2015).
Lee, K. S., Buck, M., Houglum, K. & Chojkier, M. Activation of hepatic stellate cells by TGF alpha and collagen type I is mediated by oxidative stress through c-myb expression. J. Clin. Invest. 96, 2461–2468 (1995).
Steiling, H. et al. Activated hepatic stellate cells express keratinocyte growth factor in chronic liver disease. Am. J. Pathol. 165, 1233–1241 (2004).
Otte, J.-M. et al. Differential regulated expression of keratinocyte growth factor and its receptor in experimental and human liver fibrosis. Regul. Pept. 144, 82–90 (2007).
Sun, Z. et al. Fibroblast growth factor 7 inhibits cholesterol 7α-hydroxylase gene expression in hepatocytes. Biochem. Biophys. Res. Commun. 423, 775–780 (2012).
Nielsen, J. et al. Prognostic molecular markers in pediatric liver disease - are there any? Biochim. Biophys. Acta 1865, 577–586 (2019).
Kuiper, G. G. et al. Comparison of the ligand binding specificity and transcript tissue distribution of estrogen receptors alpha and beta. Endocrinology 138, 863–870 (1997).
Sir-Petermann, T. et al. Metabolic and reproductive features before and during puberty in daughters of women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 94, 1923–1930 (2009).
Baker, S. C. et al. The External RNA Controls Consortium: a progress report. Nat. Methods 2, 731–734 (2005).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics, http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2015).
Krueger, F. Trim Galore! Babraham Bioinformatics, http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ (2015).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Munro, S. A. et al. Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures. Nat. Commun. 5, 5125 (2014).
International Sheep Genomics Consortium. et al. The sheep genome reference sequence: a work in progress. Anim. Genet. 41, 449–453 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Filis, P. et al. Maternal smoking and high BMI disrupt thyroid gland development. BMC Med. 16, 194 (2018).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell Proteomics 13, 2513–2526 (2014).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 1–25 (2004).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Friedewald, W. T., Levy, R. I. & Fredrickson, D. S. Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin. Chem. 18, 499–502 (1972).
Acknowledgements
The authors wish to acknowledge Joan Docherty and the staff at the Marshall Building, University of Edinburgh for their excellent animal husbandry, and Dr Zeynab Heidari and Dr David Stead (University of Aberdeen) for the expertise in RNA sequencing and proteomic analysis respectively. This work was funded by Medical Research Council (MRC) project grant (G0901807) to WCD and MTR, and MRC project grant MR/P011535/1 (To MTR, PAF and WCD).
Author information
Authors and Affiliations
Contributions
Animal work was performed by M.T.R. and W.C.D. M.T.R., P.F., P.A.F., W.C.D., S.S., A.D. and K.S. designed experiments. K.S., P.F., S.S., J.T., S.M. and F.H. performed laboratory analyses and developed techniques. K.S., P.F., A.D., S.S., W.C.D., P.A.F. and M.T.R. analysed data sets and prepared manuscript.
Corresponding author
Ethics declarations
Competing interests
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript, and none of the authors have any competing financial interests to declare. The authors have declared that no conflict of interest exists.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Siemienowicz, K.J., Filis, P., Shaw, S. et al. Fetal androgen exposure is a determinant of adult male metabolic health. Sci Rep 9, 20195 (2019). https://doi.org/10.1038/s41598-019-56790-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56790-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.