Abstract
Chronic obstructive pulmonary disease (COPD) is induced by cigarette smoking and characterized by inflammation of airway tissue. Since smokers with COPD have a higher risk of developing lung cancer than those without, we hypothesized that they carry more mutations in affected tissue. We called somatic mutations in airway brush samples from medium-coverage whole genome sequencing data from healthy never and ex-smokers (n = 8), as well as from ex-smokers with variable degrees of COPD (n = 4). Owing to the limited concordance of resulting calls between the applied tools we built a consensus, a strategy that was validated with high accuracy for cancer data. However, consensus calls showed little promise of representing true positives due to low mappability of corresponding sequence reads and high overlap with positions harbouring known genetic polymorphisms. A targeted re-sequencing approach suggested that only few mutations would survive stringent verification testing and that our data did not allow the inference of any difference in the mutational load of bronchial brush samples between former smoking COPD cases and controls. High polyclonality in airway brush samples renders medium-depth sequencing insufficient to provide the resolution to detect somatic mutations. Deep sequencing data of airway biopsies are needed to tackle the question.
Similar content being viewed by others
Introduction
Exposure to cigarette smoke is the major risk factor for both lung cancer and chronic obstructive pulmonary disease (COPD)1. COPD diagnosis predates as many as 70% of lung cancer cases2, but the frequent co-existence is not only due to common causative agents. In fact, COPD also seems an independent risk factor for lung cancer as positive associations remain after accounting for smoking and co-occurring respiratory diseases3,4. While chronic inflammation in the lung could explain the direct link between COPD and lung cancer5, mitochondrial dysfunction has also been proposed to be one of the driving mechanisms6. The suggestive higher burden of mitochondrial reactive oxygen species in cells of the respiratory system in COPD patients, shown e.g. for airway smooth muscle cells7, could result in increased mutagenesis8. The same consequence may also be attributed to the downregulation of DNA repair genes in COPD cases9. An elevated mutational load in cells of the respiratory tract may then increase the risk of impairing cell cycle control genes and eventually transform cells to the point of forming malignant clones with the propensity to develop bronchial carcinomas.
The approach for detecting the somatic mutational burden in a particular tissue is well established for cancer. The standard way is to carry out whole genome sequencing (WGS) of a tumour sample as well as of a sample from a reference tissue (usually blood) to at least intermediate read depth (≈10–40x, used synonymously with medium-coverage WGS) and then simultaneously search for differences between the data of the two tissues. Examples of published genome-wide mutational loads in non-cancerous tissue are however scarce10,11,12. This is mainly because the described approach has appeared less promising beyond cancer, since the presumed lack of cell populations of recent clonal origin and the usual presence of a variety of cell types in available bulk tissue samples may limit a particular mutation to very few cells and to stay hence below the detection threshold. Nevertheless, WGS of intermediate read depth proved suitable for somatic mutation calling (SMC) in prostate tissue distant to clinically overt tumours, whereby many mutations were found that were neither present in the tumour nor in the blood representing the germline genome13. This indicates clonal expansion adjacent to a tumour in morphological normal prostate tissue and has also been observed in several other tissues including lung14. Furthermore, the number of different clonal lineages in a tissue may also depend on age as was shown for blood in which an expansion of pre-cancerous clonal cell populations was detected in an essential part of the elderly population15.
Little is known about the distribution and division rate of progenitor cells in the lung epithelium of healthy individuals and COPD patients, and consequently about the existence of cell populations of recent clonal origin in this tissue. Data from smokers, however, have suggested intensified consolidation of clones in the airways of smokers16, potentially carrying a high burden of persistent mutagenic alterations. Similarly, hyperplasia of airway epithelial cells has been reported to be a typical feature in the early COPD pathogenesis17. Therefore, we leveraged medium-coverage WGS data of lower lobe epithelial lung tissue, isolated from airway brushings, and blood as reference tissue from 12 elderly individuals to identify somatic single-nucleotide mutations (SSMs) and compared the total mutational load with respect to smoking status (ex-smokers vs. never smokers) and presence of COPD (within ex-smokers). For a subset of ex-smokers, upper lobe brushings were also analysed in order to reveal temporal information of mutational development. Verification analysis of the mutation calls resulted in a surprisingly small set of confirmed SSMs arguing for a highly polyclonal cell mixture in brush samples.
Methods
Study sample and bronchoscopy
Subjects 1–9, including three COPD cases as well as three ex-smoking and three never smoking controls (Table 1), were recruited from the EvA (Emphysema vs. Airway disease) study18, all providing blood and bronchial brushings from a lower lobe of the cancer-free lung by bronchoscopy. COPD was diagnosed by post-bronchodilation spirometry with a ratio between forced expiratory volume in one second and forced vital capacity (FEV1/FVC) <0.7. The cellular material of bronchial brushings represented an area of ≈9 cm2 of the airways and contained over 90% airway epithelial cells. Subjects 10–12, one COPD case and two healthy ex-smokers, were later selected from the same cohort, providing each two lung brush samples, one from a lower and one from an upper lobe of the lung, in addition to blood (Table 1). A more detailed description of the selection of the individuals and the collection of lung brush samples is given in Supplementary Methods. All individuals provided informed consent, and ethical approval was obtained from the respective local Ethics Committees (Ethics Committee of Philipps-University of Marburg, Albert-Ludwigs-University of Freiburg, Medizinische Hochschule Hannover, all Germany; Medical Research Council Budapest, Hungary; Ethics Committee of the Province of Ferrara, Italy; Ethics Committee of the National Tuberculosis and Lung Diseases Research Institute Warszawa, Poland; and NHS Research Ethics Committee of Leicestershire, Northamptonshire and Rutland Research Ethics Committee 2, UK). The central approval was obtained from the Ethic Committee of the University Hospital Munich, document # 400-07. All methods were performed in accordance with the principles of the Declaration of Helsinki.
Sample preparation and whole genome sequencing
Details about the DNA extraction, library preparation, WGS and post-processing of sequencing data from blood and brush samples are given in Supplementary Methods. Sequence reads were mapped to Human Reference Genome (hg19/GRCh37 decoy) using the Genome Multitool Mapper 2 (GEM2)19. The average mean coverage across samples was 23.7 (Supplementary Table S1), showing better results for samples of blood compared to airway tissue of the lower lobe (26.7 vs. 19.9, P = 0.02) and for samples acquired at the second stage of the project as opposed to the first stage (28.8 vs. 21.2, P = 0.005, Wilcoxon Rank-Sum tests). Since most SMC tools require a minimal positional read depth of ≈10 in both the affected and the reference tissue in order to evaluate the presence of a SSM, we defined the SMC power as the number of bp with a read depth of ≥10 in both tissues. Except for subject 2, whose brush sample was of substandard quality, the SMC power exceeded 2 Gb for all participants (Supplementary Table S1). We checked pair-wise relatedness (VCFtools v.0.1.12) between blood and lower lobe airway tissue based on germline variants called with SAMtools v.0.1.1920. All values were between 0.344 (subject 2) and 0.450 (subjects 11 and 12), leaving only subject 2 marginally below the recommended threshold for identity (0.354). The lower lobe brush sample of subject 10 showed unusual coverage peaks in IGHM and the presence of reads from Epstein-Barr virus (EBV) as is further described in Supplementary Methods and Supplementary Fig. S1.
Strategy of somatic mutation calling
SMC was done on processed files of aligned sequence data (BAM format) after first ruling out the presence of large copy number alterations (ploidy deviations from 2) with Control-FREEC21. We applied three of the most popular SMC tools for cancer, all in standard mode: MuTect, v.1.1.422, Strelka, v.1.0.1423 and VarScan 2, v.2.3.224 and only concentrated on SSM calls. Rather than independently calling variants in affected and reference tissue against a reference genome, all three tools process the sequencing data of the two given tissues simultaneously and search for positions where the reference tissue matches the reference genome (variant allele frequency, VAF, ≈0) and the affected tissue shows an elevated VAF. Unlike tools jointly modelling genotypes and therefore requiring VAFs close to 0.5 or 1.0 to detect SSMs, MuTect and Strelka jointly model allele frequencies and are therefore considered particularly appropriate to call low-frequency SSMs25. In MuTect’s core algorithm a somatic variant model must fit better than models assuming artefacts or heterozygous SNPs as the reason for the presence of variant alleles. Strelka calls SSMs based on probabilities that VAFs differ in the two tissues and that the reference tissue has a homozygous genotype as in the reference genome. VarScan 2 uses a heuristic approach applying thresholds for the number and frequency of the variant allele in order to be called a germline variant and separates SSMs with the help of statistical tests comparing the affected with the reference tissue. In this case, the accuracy of calling low-frequency SSMs depends on the careful selection of a minimal VAF threshold in the affected tissue, which in turn is influenced by the available sequencing depth. Importantly, the availability of merely medium-coverage sequence data implied that mutations needed to be present in a substantial fraction of the cells in the sample of interest in order to be detectable (e.g. VarScan 2 required a VAF ≥ 0.15 in the applied setting). Further details about the performance of the SMC tools in benchmarking studies and the applied settings are given in Supplementary Methods. Finally, the tools also differ in the application and stringency of built-in filters that evaluate scores of base and read quality, of mapping quality of reads as well as for strand bias of alleles, which may all be related to sequencing or mapping errors. Post-calling filters can also be used to reduce the influence of such errors, but in order to avoid defining thresholds for post-calling filters, we chose to intersect the individual SSM results provided by each caller. Rather than unifying the results or considering calls detected by at least two of the three callers, we chose this conservative strategy in order to end up with a small, but presumably precise set of true positives. Intersections are represented with eulerAPE26 in area-proportional Venn-diagrams using ellipses. In individuals recruited at the second stage, similarity between the calls from upper and lower lobe brushings was assessed with the Dice similarity coefficient, SDice = 2*C/(N1 + N2), where C is the intersection between N1 and N2.
Call annotation
Resulting SSM calls were annotated with dbSNP v.13727 and snpEff v3.628. The UCSC Table Browser29 was used to determine overlap with DNase I hypersensitive sites (DHS, track: DNase Clusters) and repetitive regions (track: RepeatMasker). Structural annotation was based on ENSEMBL considering one category per call. Intermutation distances (IMDs) were used to determine clustering of SSM calls and were represented in Rainfall-plots with the help of in-house R scripts.
Validation of the calling strategy
We validated our calling strategy by applying the same mapping and SMC tools and versions to a medulloblastoma (MB) and chronic lymphocytic leukemia (CLL) case, comparing for both a cancer sample with ≈40x WGS to a blood sample with ≈30x WGS data. These two cases had already been extensively investigated in a benchmarking study of different SMC pipelines30 and curated sets of verified mutations in the malignant samples were available. Calculation of sensitivity (recall) and precision (positive predictive value) of different SMC procedures was solely based on the predicted position (without considering mutation type). We further assessed the impact of a filtering script to the calls of VarScan 2 as recommended by the authors31. As the combination of the mapping software and the SMC tool had sometimes proven crucial for the overall accuracy of a pipeline30, we also carried out sensitivity analyses with GEM3, a meanwhile updated version of GEM2, as well as with the popular aligner BWA-mem32. We finally included MuTect 2, a completely revised version, which had in the meantime been integrated in GATK v.4.x and had performed well in a benchmark study33. Recall-Precision scatter plots were built with the function “scatterplot” in the package “car” in R 3.3.1.
Confidence of mutation calls
Estimation of the likelihood of a SSM call to represent a true mutation was based on the overlap with known common or low-frequent (minor allele frequency, MAF, ≥1%) single nucleotide polymorphisms (SNPs) and the localisation in a genomic region difficult to map. For the former criterion, we annotated against an updated version of dbSNP (v.147) with the help of UCSC Table Browser (track “Common SNPs(147)”, representing about 0.5% of the genome), while for the latter, tracks “RepeatMasker”, a filter that leaves only ≈45% of the genome marked as non-repetitive, and “Mappability” (table “CRG Align 50”, a filter for uniqueness in a 50-bp window that assigns 77% of the genome to unique) were used. A call was set to high confidence if it did not coincide with a SNP, did not fall into a repetitive region and was unique based on the 50mer mappability filter. Medium-confidence assignment required the same SNP criterion, but respective positions could either be in a non-unique or repetitive region. The coincidence of a SSM call with a SNP or with a position in a non-unique as well as repetitive region built the low-confidence segment. We did not include in those confidence definitions positions coinciding with rare SNPs or SNPs with unknown frequencies in population-based studies due to potential unverified entries in recent versions of dbSNP (the track “All SNPs(147)” assigns over 5% of the genome to a polymorphism). Neither did we include read depth nor VAF filters for either of the tissues, as all three mutation callers already consider these parameters.
Experimental verification of mutation calls
From each confidence segment, sets of 30 positions were selected randomly, but with the fix inclusion of all calls pointing either to non-synonymous SSMs or effecting a splice site (based on ENSEMBL, N = 4) and the requirement of at least one call from each individual being present. Calls lying on the Y-chromosome (N = 9) were not considered. Details about the primer design, the polymerase chain reactions (PCR) for amplifying the 90 selected positions, the concatenation with subsequent sequencing and data processing are given in Supplementary Methods. Four PCRs failed and successful concatenation of products of the remaining 86 reactions was confirmed by gel electrophoresis for each sample (length above 2 kb). After sequencing the concatenated products, SAMtools v.1.2 (mpileup) was used to determine coverage (upper limit: 150,000) as well as allele frequencies at positions of the targeted SSM calls (orphan reads were allowed and base quality scores of ≥30 were required). For each sample, VAF values at each call position were converted to a presence-absence matrix according to the following pattern. VAFs below 1% supported the reference allele. VAFs between 20 and 80% suggested heterozygous positions and those over 80% homozygous positions for the variant allele. VAFs between 5 and 20% at positions with good coverage (>1000) suggested the subclonal presence of the variant allele (and therefore possibly a SSM). All other VAFs (i.e. those between 1 and 5%, as well as those between 5% and 20% at positions with suboptimal coverage) were manually inspected, and decision whether the value supported the reference or the variant allele was taken according to results for the same position in other samples. The number of confirmed mutations was extrapolated to account for the total number of SSM calls per confidence category. Confidence of the non-verified SSM calls in the highest category was then re-assessed by utilizing three additional dichotomized features, which were read depth in reference and affected tissue (≥20 in both tissues vs. rest), VAF in reference tissue (>0 vs. 0), as well as presence of a SNP, either rare or of unknown frequency, at the respective position (UCSC Table Browser track “All SNPs(147)”, but not “Common SNPs(147)”).
Statistical analysis
Statistical calculations were carried out with STATA 11.
Results
Somatic mutation calling
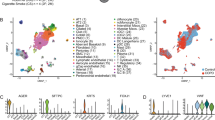
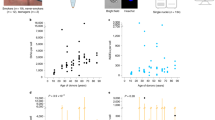
We observed large differences between the individual results of the three SMC tools. While VarScan 2 and MuTect called more than 2800 SSMs in the lower lobe lung brush sample of each individual, the unfiltered outcome of Strelka was one order of magnitude smaller. Although similar in size, the overlaps between the two permissive calling tools did not coincide with the results of the restrictive caller, resulting in very small intersections between the three callers (Supplementary Fig. S2). Overlapping SSM calls ranged from 43 (subject 1) to 493 (subject 10) and amounted to a total of 1651 (1645 unique ones) when only considering lower lobe samples (Fig. 1). Strikingly, neither brushings from COPD patients, nor those from ex-smokers with normal lung function did unambiguously show elevated numbers of SSM calls when compared to those of never smokers. Subject 10, already mentioned for some unusual sequencing coverage at immunoglobulin loci (Supplementary Methods and Supplementary Fig. S1A), was the only individual with a clearly increased number of SSMs in the intersected results. Moreover, IMDs were particularly low in this sample, an issue far less notable in the brush sample of the upper lobe of the same individual (Supplementary Fig. S3) or in any other brush sample. Although clustering of mutations (kataegis) is a well-described phenomenon in certain cancers34, the observed pattern favoured a technical problem since the clustering was distributed over the whole genome. Accordingly, similarities between intra-individual calls in upper and lower lobe airway brushings were smaller in this subject than in the other two with this type of data (Supplementary Table S2). Nevertheless, we decided not to exclude the deviant sample at this stage owing to two reasons. First, the increased number and unusual distribution of calls could have been related to a local EBV infection (Supplementary Methods and Supplementary Fig. S1B), which can enhance mutational loads35. Second, comparing the uncorrected number of SSM calls would only be informative if the SMC power was equal between the subjects. When comparing the numbers relative to the positions covered by enough reads to determine the presence of SSMs, subject 10 was no longer an outlier. More importantly, this adjustment did not alter the main counter-intuitive finding, namely that there were no clear differences between the two groups of former smokers (with and without COPD) and never smokers in terms of the total somatic mutational burden suggested by state-of-the-art SMC tools (Table 2).
Validation of the applied strategy in cancer
In order to validate the applied strategy to call SSMs, we carried out the same procedure with two cancer cases (one CLL and one MB case) from which we had fully verified somatic mutation call sets30. The strategy of taking the intersection of the individual results of the three callers proved accurate (Supplementary Fig. S4A). Compared to intersections of only two tools, the loss in recall was marginal and clearly compensated by a material gain in precision. Replacing the aligner GEM2 with the newer version GEM3 improved results for Strelka and respective intersections in the MB data set, but no such effect was found in the CLL data set. Further sensitivity analyses with BWA-mem confirmed that the intersection strategy of three tools led to higher precision than alternative strategies, while the recall remained high (Supplementary Fig. S4B). Improvements from MuTect to MuTect 2 were clearly visible in the individual results, but were not essential in intersections. Therefore, we did not consider applying these updated mapping or calling tools to the sequencing data of the brushings (EvA data set).
Critical appraisal of the somatic mutation calls
Despite the successful validation of our SMC strategy in cancer cases, we further assessed some characteristics of the 1651 suggested SSMs in lower lobe brushings across all subjects and compared them with those of the (true and false positive) calls in the two cancer cases when applying the same tools. Calls did not show material differences in terms of chromosomal distribution, structural annotation, overlap with DHS or repetitive regions, as well as mutation type (Fig. 2). As 9 of the 12 individuals in the EvA data set were former smokers, a higher proportion of C > A substitutions in lung epithelial tissue (as compared to malignant brain or blood tissue) could have been anticipated34, but sample size was obviously too limited to draw conclusions from the absence of such an association. There were other features of the calls in the EvA data set that deviated considerably. Variant alleles peaked at markedly lower frequencies (median VAF ranged from 0.22 for subject 12 to 0.35 for subject 2) than in MB (0.45) or CLL (0.44). This is not unexpected since clonal heterogeneity is likely much higher in non-cancerous samples. However, results from the two cancer cases suggested that the reliability of calls with decreased VAF was reduced. The false positive proportion of the SSM calls shared between the three callers showed median VAFs of 0.23 (MB) and 0.29 (CLL), whereas the true positive calls barely deviated from 0.5 (median VAFs of 0.47 and 0.46 for MB and CLL, respectively). Furthermore, median coverage of the calls in the EvA data set was lower, and for only 53.1% of the called SSMs, coverage at the respective position was at least 20 in both, the reference and the affected tissue. Respective numbers in the cancer cases were 92.9% (MB) and 77.9% (CLL). The feature creating most doubts about the certainty of calls in the EvA data set was, however, the high overlap with polymorphisms in dbSNP v.137 (55.7%). This was markedly higher than for calls from the MB (13.5%) and CLL sample (11.7%), and the latter proportions even decreased further when including only the confirmed calls (to 5.3% and 4.6%, respectively). Especially at suboptimal sequencing depth of the respective position in blood, the chance for a heterozygous SNP to appear absent in blood, but not in lung (which would hence result in a wrong SSM call), may be substantial. We defined then three confidence levels for our calls based on location in genomic regions difficult to assess (i.e. high repetitivity or low mappability) or of polymorphisms in an updated version of dbSNP (Methods). 21.9% of the 1651 SSM calls overlapped with a common or low-frequent SNP. This is lower than stated in Fig. 2 as only SNPs with a MAF > 1% were considered here. Only 14.8% of calls were neither in a repetitive nor in a non-unique region, while 33.3% were in both. Taken together, less than 10% of the suggested SSMs (N = 156) were assessed as high confidence, whereas N = 711 and N = 784 were assigned to the middle- and low-confidence category. Finally, in the three individuals from which upper and lower lobe brushings were available, ≈15–20% of the SSM calls were shared between the two sampled locations (Supplementary Table S2). While such calls may appear more robust (mutational events that occurred early in lung development), they can also be explained by the aforementioned uncertainty of correctly assigning the reference allele to the blood sample, which could lead to faulty calls in both, upper and lower lobe brushings, at once. Indeed, only four of the 46 calls in this group were classified as highly confident (≈9%).
Characteristics of somatic single-nucleotide mutation calls in lung brushings of subjects from the EvA study (N = 12) as well as in tumour tissue of two cancer cases based on the same calling procedure. Coverages and VAFs were based on the Strelka results. MB = medulloblastoma, CLL = chronic lymphocytic leukemia, DHS = DNase I hypersensitive sites, IQR = interquartile range, VAF = variant allele frequency.
Verification of selected mutation calls
The 90 selected calls (30 for each confidence category) are shown in Supplementary Table S3. For four positions representing calls in the high- (N = 1), medium- (N = 1) and low-confidence (N = 2) category, no functioning PCR could be established. The sequencing depth of the remaining positions in the verification approach is shown in Supplementary Table S4 across all 27 tissue samples. One position did not show any coverage in any of the samples and was excluded as a faulty primer pair not designed to include the position of the respective SSM call was used. In general, we achieved very high coverage for the selected positions in the concatenated amplicons (overall median coverage was ≈27,000), but due to the design of the experiment, median coverage per position over the tissue samples varied strongly (minimum of 87, maximum of >150,000). Although positional coverages of up to 100 (red colour) or 1000 (orange) may be considered of limited value for a verification attempt, it is noteworthy that we detected reads for the remaining 85 candidate positions across all 27 samples, meaning that products of all PCRs had been present in the final ligations. This includes those from samples with low concentration of DNA stock solution (subject 4, bronchial tissue) or for which DNA from former library preparations had to be amplified due to missing stock solution (lower lobe bronchial tissue of subjects 1, 2 and 12, Supplementary Methods). There were only very few positions at which we could not assess the likelihood of the presence of a variant allele, and these did not concern the subjects in which the mutation was actually suspected.
Overall, VAF results for many positions did not support the presence of a variant allele in the brushings of the respective individuals, indicating considerable false positive calls in the WGS-based SMC results (Table 3 and more detailed in Supplementary Tables S5 to S7). Grouped by confidence category, this finding accounted for 21 of 28 (75.0%), 21 of 29 (72.4%) and 12 of 28 (42.9%) of SSM calls included in the verification analysis. There was only one case where verification of the SSM call seemed successful, showing a clearly higher VAF in the brush than in the blood sample at the respective position (Supplementary Table S5, position 5 in subject 6). For other positions, the detected VAF in the respective sample was slightly elevated, but either limited coverage or comparisons with the same position in samples not suspected to carry the mutation nevertheless favoured the exclusive presence of the reference allele (N = 1, 6 and 3 for high-, medium- and low-confidence calls, respectively; indicated by a red 0 in Supplementary Tables S5 to S7). Germline heterozygous variants are expected to exhibit VAFs very close to 0.5 in blood and airway brushings as, unlike for SSMs, neither polyclonality nor infiltration of other cell types reduces this frequency. Not surprisingly, we detected such cases (indicated by a black 1) particularly in the low-confidence category (N = 9, 32.1%) as calls overlapping with common and low-frequent SNPs had been classified there. Furthermore, we observed several cases where results for the respective position in the targeted sample supported the subclonal presence of a variant allele, but those findings were not tissue-specific (N = 5, 1 and 4 for high-, medium- and low-confidence calls, respectively; indicated as a red or green 1). They may either point to allele-specific sequencing or mapping bias, especially if deviant VAFs were present in samples of several individuals, or to postzygotic mosaicism if only detected in one individual. The 84 non-verified calls (out of 85 with valid verification results) also included three calls that were made in lower and upper lobe brushings of the same individual. We note here that even very high coverage observed in the verification analysis did sometimes not prevent us from results which were difficult to interpret. As even calls in the high-confidence category proved faulty, we retrospectively assessed more stringent criteria for defining a high-confidence call. Of the 27 high-confidence calls that could not be verified, 9 would have been down-graded if a WGS read depth filter of 20 in both, blood and brush sample, had been applied, 5 would not have met the requirement of 0 variant alleles in blood, and 10 would not have been considered if overlap with any entry in dbSNP v.147 (i.e. even if of unknown MAF in the general population) had been taken into account. Only 7 out of 28, including the one that was verified, would have ended up in the high confidence category if such still rather permissive criteria had been applied in a combined way.
In summary, the verification analysis could only confirm one predicted SSM. Extrapolating this result would leave us with only a handful true SSMs of the 1651 calls, leaving over 99% as presumably false positives. High confidence in the overlapping output of common SMC tools is only warranted if stringent requirements in terms of sequencing coverage, complexity of the region and overlap with data bases of polymorphisms are met.
Discussion
Motivated by the fact that several studies called numerous somatic mutations in healthy tissue by sequencing affected and reference tissue to an intermediate read depth13,14, we applied here a similar approach for the genome-wide calling of SSMs in airway brushings from COPD cases and controls. The number of included individuals was obviously small and at best appropriate to formulate a trend respective to a possible difference in mutation numbers between cases and controls, but the strong assumption of detecting at least differing numbers between former and never smoking controls served as a proof-of-concept of the methodology. However, we were unable to detect a robust number of SSMs in this study, neither in lung brushings of former smokers in general, nor specifically in COPD patients. While we first seek evidence for a biological interpretation of this observation, we concentrate later on limitations of the applied methodology and discuss alternative ways to tackle this task.
As it has never been investigated in a genome-wide manner, it could be argued that, years after smoking cessation, negative selection of strongly mutated lung progenitor cells eventually leads to a harmonization of mutational levels between former and never smokers, which would explain our failure to detect such a difference in samples of airway epithelial cells. There is, however, compelling evidence for a higher number of accumulated mutations in the airways of ex-smokers. First, investigations of microsatellite instability in former smokers at selected chromosomal sites support such a notion36. Second, the mutational load in tumours of (current and former) smokers is clearly elevated compared to tumours of never smokers37,38. Third, the risk of developing lung cancer in former smokers never reverts to the level of never smokers in epidemiological studies39,40, which could be due to a greater number of persistent mutations and hence a higher likelihood to affect cancer driver genes. Moreover, the complete failure to detect any mutational mark in airway epithelial cells of brush samples does not match available results from other tissues. Estimates of 600 SSMs for blood10 and a few thousands for sun-exposed epidermis41 were reported. Numbers for internal organs often derive from tumour-adjacent tissue and also lie in the range of several hundred13,14. Data from lung tumours suggest around 10 per Mb in smokers32, a similar load in former smokers38 and one order of magnitude fewer in never smokers37. Since a large part of the mutational burden is assumed to be present before tumour initiation42, we may expect cumulative SSM numbers in the range of hundreds in samples of airway epithelial cells of elderly never smokers, similar to blood or internal organs. Such a number would also be obtained by assuming an intrinsic mutation rate of about three per cell division42 and a cell division rate of around once per several months for airway epithelial cells43. With SSM estimates of 150 per year due to smoking one cigarette pack per day37, the mutational burden in bronchial tissue of former smokers may hence be well in the thousands. Taken together, methodological reasons must explain the lack of verifiable SSMs in our study.
In the absence of a standard approach how to estimate the genome-wide mutational load in healthy tissues, we selected here a procedure that was strongly based on established methods for tumours (medium-coverage WGS of bulk samples from target and reference tissue of the same individual). Three different callers were chosen due to the fact that concordance between different tools is sometimes low25,30,33. Our conservative strategy of only considering the overlap was believed to lead to a set with high precision as was confirmed in a validation approach with two cancer cases. The resulting number of SSM calls of around 100 per brush sample was indeed smaller than anticipated, but could have represented lower bounds due to the conservative strategy and the limited SMC power in several individuals. However, an assessment of the confidence of calls and a subsequent verification approach corroborated the suspicion of having picked up mainly false positives. Hence, stringent application of criteria like minimal read depth, permitted number of variant alleles in reads of the reference tissue and low-complex or blacklisted regions remain essential and cannot entirely be replaced by intersecting results from different callers in order to reduce false positives to a negligible number.
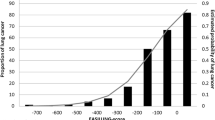
A far more crucial issue is the large number of false negatives in this study. Intersecting results derived from different calling tools is admittedly prone to result in many false negatives in the sense that results can only become as sensitive as the least sensitive caller performs33. However, all three callers have shown reasonable sensitivity in benchmarking studies44 as well as in our validation with cancer data. The ability to detect a variant allele at a specific position in bulk samples depends on the proportion of reads supporting the variant. This proportion is related to the number of cells carrying the variant allele with respect to the total number of cells. Its minimal required value to reliably call the presence of a variant allele also varies according to the available read depth at that position (Fig. 3). In tumour biopsies, the relative number of cells sharing a particular mutation is generally high as recent clonal expansion has taken place, even in the case of a substantial amount of non-malignant tissue or intratumoural heterogeneity45. This becomes even more obvious when a minority clone outgrows other divergent clones, e.g. by positive selection owing to treatment. Consequently, read depth requirements are modest, and common SMC tools are accurate when using the typical minimal VAF of about 0.15 in order to call a mutation (i.e. ≈30% of the sampled cells must carry the mutation). For non-pathological tissue, it is generally unknown how many cells originate from a recent specific progenitor cell and form an ancestral cell lineage. In blood, depletion of cell lineages is common in the elderly15, and a sample of a centenarian suggested the presence of only two stem cell populations, which explains the ability to call hundreds of SSMs, albeit with high sequence coverage (60×)10. On the other hand, studies of selected genes in biopsies of the epidermis found clonal unit sizes per mutation of ≈0.2 mm2 (around 2000 cells)41 or even less46. Proportions of affected cells were so small that ultra-deep sequencing (500×) was necessary, a procedure which is still prohibitively costly for whole genomes. Interestingly, skin fibroblast samples contained SSMs with >0.1 VAF, but those may represent events that arose early in skin development47. Large clonal cell populations of recent origin must also be widespread in tumour-adjacent tissue of several internal organs, in order to explain the detection of several hundred SSMs that were neither in blood nor in the tumour close-by using medium-coverage sequencing13,14. The fact that biopsies were gained in close proximity to tumour tissue may, however, point to field effects, i.e. the presence of molecular abnormalities in tissue that appears histologically normal. The airway epithelium contains facultative progenitor cells that mainly proliferate in response to injury48, which could be smoking- or COPD-related. Indeed, smokers have shown higher basal cell division rates leading to clonal consolidation16, and basal cell hyperplasia is one of the earliest lesions in the pathogenesis of COPD17. In addition, and unlike lung biopsies10,49, brush samples contain very small quantities of immune cells50 (confirmed in our cytospins), all enhancing the chance of finding mutations confined to reasonably large cell populations.
Representation of mutant clones in the bronchial epithelium. Coloured cylinders represent different cellular clones of recent origin in the airway epithelium, each containing cells that share the same somatic mutations. Such a clone (shown for the one in salmon at the bottom) may contain a mixture of cell types, such as differentiated secretory cells (shown with granules in the cytoplasm), ciliated cells (depicted with cilia at the outer surface) and unstratified basal cells (square shaped), all sitting on the basement membrane (line at the very bottom). Such a cellular arrangement is named pseudostratified columnar epithelium and is typical for the airways. Clone size may vary, but presumably lies in the sub-millimeter range, which is much smaller than the diameter and the length of a brush used in bronchoscopy (shown on the right). Consequently, a brush sample contains many different clones and the proportion of cells carrying the same somatic mutations is very small and falls below the detection limit. Successful somatic mutation calling requires sampling a smaller area (i.e. coming closer to the detection limit by harvesting a higher proportion of cells carrying the same somatic mutations) and/or sequencing to a higher read depth (i.e. lowering the detection limit).
The observed absence of basically any SSMs in our brush samples point however to major limitations of our approach. While biopsies typically originate from areas (sections) of 1–5 mm2 13,41,47, brushings have a >100x higher and less cohesive area sampled (Fig. 3 and Supplementary Methods). Assuming similar sizes of clonal units, even ultra-deep sequencing approaches41 may struggle to compensate for this loss of resolution. In terms of COPD, we must further acknowledge that by random sampling of one location in the airways, we may not necessarily have hit a spot severely affected by the disorder. Furthermore, it is conceivable that clonal sizes due to pre-cancerous metaplastic processes increase into a more detectable range in later stages of the disease or with very old age. If recently expanded clones are absent and mutations thus only confined to few cells, even small biopsies could be insufficient to detect a robust number of them. Such an interpretation underlies most likely the finding of no10 or very few SSMs51 in post-mortem brain samples with sequencing depth up to 100×. The very few detected mutations could represent early events similar to our study, in which their occurrence was most likely limited to the small time window in embryonic lung development after the separation of mesodermal (forming components of the blood) and endodermal tissue (forming epithelia, e.g. that of the respiratory tract). The few overlapping mutations between brushings from upper and lower lobe lung within the same individual would also belong to this type (assuming successful verification). In contrast, for SSMs appearing postnatally, including all smoking-related ones, brush sampling, in spite of its easy execution, is an inadequate detection strategy, at least in combination with medium-coverage sequencing where about 30% of sampled cells are required to be affected.
Recently, more sophisticated methods to find rare SSMs in healthy tissues emerged52,53. Single adult stem cells, or single differentiated cells via the detour of reprogramming them into induced pluripotent stem cells (iPSC)47, can be amplified in culture, and consequently mutations become clonally expanded as well. WGS at intermediate read depth is then sufficient to detect them because their VAFs lie very close to 0.5 unless representing events during cell proliferation in culture. Even post-mitotic neurons could be assessed by amplification of single-cell DNA before application of WGS12. Several studies using such novel methods have reported unexpected findings. Small differences in mutation rates between tissues of quiescent organs and high-turnover tissues were for instance reported by independent methods11,52. This challenges the concept backed by tumour data that cell division rate is a better indicator of the mutational load in a tissue than age34,54,55. Mutational loads seems also higher than suggested from sequencing bulk tissue samples although stochastic effect can play a role if single cells are analysed individually. For example, genetically reprogrammed skin fibroblasts from children contained on average over 1000 SSMs in spite of sampling locations not exposed to the sun, and only few could be attributed to the reprogramming process47. Postnatally non-dividing neurons also showed several hundred mutations already one year after birth and further SSM accumulation correlated with age during adulthood12. Amplification artifacts and the high error rate have so far been considered major limitations of single-cell WGS, but as results were in good agreement with those using other novel methodology52,56, the accumulation of SSMs must also occur spontaneously or transcription-associated. Taking these observations together, it is fair to assume that the mutational load in airway epithelial single cells of elderly people lies well above 1000, even in the absence of former or current smoking exposure, and the need to standardize procedures is obvious.
In summary, it remains elusive why cigarette smokers that retain normal lung function are at lower risk of developing lung cancer than those who develop airflow obstruction. In this work, serving as a pilot study by tackling this question from a mutational point of view, the mutational load in samples of bronchial epithelial cells could not be accurately estimated due to sizes of clonal cell populations below the detection limit of the applied methodology. The synthesis with results from similar work further implies that i) sequencing to an intermediate read depth is rarely sufficient to illuminate the mutational load in samples of healthy tissue; ii) airway brushing by bronchoscopy is an inappropriate sampling strategy given the relatively large area from which cells are gained, possibly not even surmountable by deep-sequencing data; iii) tissue directly adjacent to cancer does not serve as a valuable proxy for healthy tissue as the cell lineage composition might resemble more closely that of a tumour. Consequently, the question whether COPD influences lung cancer risk via the accumulation and persistence of somatic mutations8 remains up for debate. Although the typically non-reversible processes in the lung of COPD patients like hypoxia, inflammation, tissue damage and remodeling indeed represent conditions favorable to mutagenesis, they may also act on the better survival of emerging tumour cells. For instance, COPD-related symptoms seem to leave methylation signatures on immune genes57 and lung cancer associated genes58, which may in turn facilitate tumour growth in a manner independent of mutagenesis. The mutational burden in bronchial epithelial tissue is maybe currently best estimated via amplifying single stem cells in organoid cultures. Such approaches may help to decipher whether differences of the somatic mutational burden in airway epithelial tissue between ex-smokers and never smokers as well as between COPD patients and non-obstructive individuals with similar smoking history explain disparity in lung cancer risk, a question that remains intriguing.
Data availability
Processed sequencing data from this study have been deposited in BAM-format at the European Genome-phenome archive (EGA) under accession number EGAS00001003406. Additional information for reproducing the results is available upon reasonable request.
References
Celli, B. R. Chronic obstructive pulmonary disease and lung cancer: common pathogenesis, shared clinical challenges. Proceedings of the American Thoracic Society 9, 74–79, https://doi.org/10.1513/pats.201107-039MS (2012).
Young, R. P. et al. COPD prevalence is increased in lung cancer, independent of age, sex and smoking history. The European respiratory journal 34, 380–386, https://doi.org/10.1183/09031936.00144208 (2009).
Denholm, R. et al. Is previous respiratory disease a risk factor for lung cancer? American journal of respiratory and critical care medicine 190, 549–559, https://doi.org/10.1164/rccm.201402-0338OC (2014).
Papi, A. et al. COPD increases the risk of squamous histological subtype in smokers who develop non-small cell lung carcinoma. Thorax 59, 679–681, https://doi.org/10.1136/thx.2003.018291 (2004).
Mantovani, A., Allavena, P., Sica, A. & Balkwill, F. Cancer-related inflammation. Nature 454, 436–444, https://doi.org/10.1038/nature07205 (2008).
Ng Kee Kwong, F. et al. Is mitochondrial dysfunction a driving mechanism linking COPD to nonsmall cell lung carcinoma? European respiratory review: an official journal of the European Respiratory Society 26, https://doi.org/10.1183/16000617.0040-2017 (2017).
Wiegman, C. H. et al. Oxidative stress-induced mitochondrial dysfunction drives inflammation and airway smooth muscle remodeling in patients with chronic obstructive pulmonary disease. The Journal of allergy and clinical immunology 136, 769–780, https://doi.org/10.1016/j.jaci.2015.01.046 (2015).
Anderson, G. P. & Bozinovski, S. Acquired somatic mutations in the molecular pathogenesis of COPD. Trends in pharmacological sciences 24, 71–76, https://doi.org/10.1016/S0165-6147(02)00052-4 (2003).
Sauler, M. et al. The DNA Repair Transcriptome in Severe COPD. The European respiratory journal, https://doi.org/10.1183/13993003.01994-2017 (2018).
Holstege, H. et al. Somatic mutations found in the healthy blood compartment of a 115-yr-old woman demonstrate oligoclonal hematopoiesis. Genome research 24, 733–742, https://doi.org/10.1101/gr.162131.113 (2014).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264, https://doi.org/10.1038/nature19768 (2016).
Lodato, M. A. et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science, https://doi.org/10.1126/science.aao4426 (2017).
Cooper, C. S. et al. Analysis of the genetic phylogeny of multifocal prostate cancer identifies multiple independent clonal expansions in neoplastic and morphologically normal prostate tissue. Nature genetics 47, 367–372, https://doi.org/10.1038/ng.3221 (2015).
Yadav, V. K., DeGregori, J. & De, S. The landscape of somatic mutations in protein coding genes in apparently benign human tissues carries signatures of relaxed purifying selection. Nucleic acids research 44, 2075–2084, https://doi.org/10.1093/nar/gkw086 (2016).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. The New England journal of medicine 371, 2477–2487, https://doi.org/10.1056/NEJMoa1409405 (2014).
Teixeira, V. H. et al. Stochastic homeostasis in human airway epithelium is achieved by neutral competition of basal cell progenitors. eLife 2, e00966, https://doi.org/10.7554/eLife.00966 (2013).
Shaykhiev, R. & Crystal, R. G. Early events in the pathogenesis of chronic obstructive pulmonary disease. Smoking-induced reprogramming of airway epithelial basal progenitor cells. Annals of the American Thoracic Society 11(Suppl 5), S252–258, https://doi.org/10.1513/AnnalsATS.201402-049AW (2014).
Ziegler-Heitbrock, L. et al. The EvA study: aims and strategy. The European respiratory journal 40, 823–829, https://doi.org/10.1183/09031936.00142811 (2012).
Marco-Sola, S., Sammeth, M., Guigo, R. & Ribeca, P. The GEM mapper: fast, accurate and versatile alignment by filtration. Nature methods 9, 1185–1188, https://doi.org/10.1038/nmeth.2221 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425, https://doi.org/10.1093/bioinformatics/btr670 (2012).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature biotechnology 31, 213–219, https://doi.org/10.1038/nbt.2514 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817, https://doi.org/10.1093/bioinformatics/bts271 (2012).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome research 22, 568–576, https://doi.org/10.1101/gr.129684.111 (2012).
Xu, H., DiCarlo, J., Satya, R. V., Peng, Q. & Wang, Y. Comparison of somatic mutation calling methods in amplicon and whole exome sequence data. BMC genomics 15, 244, https://doi.org/10.1186/1471-2164-15-244 (2014).
Micallef, L. & Rodgers, P. eulerAPE: drawing area-proportional 3-Venn diagrams using ellipses. PloS one 9, e101717, https://doi.org/10.1371/journal.pone.0101717 (2014).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic acids research 29, 308–311 (2001).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92, https://doi.org/10.4161/fly.19695 (2012).
Karolchik, D. et al. The UCSC Table Browser data retrieval tool. Nucleic acids research 32, D493–496, https://doi.org/10.1093/nar/gkh103 (2004).
Alioto, T. S. et al. A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing. Nature communications 6, 10001, https://doi.org/10.1038/ncomms10001 (2015).
Koboldt, D. C., Larson, D. E. & Wilson, R. K. Using VarScan 2 for Germline Variant Calling and Somatic Mutation Detection. Current protocols in bioinformatics 44, 15.4.1–17, https://doi.org/10.1002/0471250953.bi1504s44 (2013).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. ArXiv e-prints 1303, http://adsabs.harvard.edu/abs/2013arXiv1303.3997L (2013).
Cai, L., Yuan, W., Zhang, Z., He, L. & Chou, K. C. In-depth comparison of somatic point mutation callers based on different tumor next-generation sequencing depth data. Scientific reports 6, 36540, https://doi.org/10.1038/srep36540 (2016).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421, https://doi.org/10.1038/nature12477 (2013).
Liang, Q. et al. Integrative identification of Epstein-Barr virus-associated mutations and epigenetic alterations in gastric cancer. Gastroenterology 147, 1350–1362 e1354, https://doi.org/10.1053/j.gastro.2014.08.036 (2014).
Wistuba, I. I. et al. Molecular damage in the bronchial epithelium of current and former smokers. Journal of the National Cancer Institute 89, 1366–1373 (1997).
Alexandrov, L. B. et al. Mutational signatures associated with tobacco smoking in human cancer. Science 354, 618–622, https://doi.org/10.1126/science.aag0299 (2016).
Shi, J. et al. Somatic Genomics and Clinical Features of Lung Adenocarcinoma: A Retrospective Study. PLoS medicine 13, e1002162, https://doi.org/10.1371/journal.pmed.1002162 (2016).
Halpern, M. T., Gillespie, B. W. & Warner, K. E. Patterns of absolute risk of lung cancer mortality in former smokers. Journal of the National Cancer Institute 85, 457–464 (1993).
Ebbert, J. O. et al. Lung cancer risk reduction after smoking cessation: observations from a prospective cohort of women. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 21, 921–926, https://doi.org/10.1200/JCO.2003.05.085 (2003).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886, https://doi.org/10.1126/science.aaa6806 (2015).
Tomasetti, C., Vogelstein, B. & Parmigiani, G. Half or more of the somatic mutations in cancers of self-renewing tissues originate prior to tumor initiation. Proceedings of the National Academy of Sciences of the United States of America 110, 1999–2004, https://doi.org/10.1073/pnas.1221068110 (2013).
Snyder, J. C., Teisanu, R. M. & Stripp, B. R. Endogenous lung stem cells and contribution to disease. The Journal of pathology 217, 254–264, https://doi.org/10.1002/path.2473 (2009).
Wang, Q. et al. Detecting somatic point mutations in cancer genome sequencing data: a comparison of mutation callers. Genome medicine 5, 91, https://doi.org/10.1186/gm495 (2013).
Swanton, C. Intratumor heterogeneity: evolution through space and time. Cancer research 72, 4875–4882, https://doi.org/10.1158/0008-5472.CAN-12-2217 (2012).
Stahl, P. L. et al. Sun-induced nonsynonymous p53 mutations are extensively accumulated and tolerated in normal appearing human skin. The Journal of investigative dermatology 131, 504–508, https://doi.org/10.1038/jid.2010.302 (2011).
Abyzov, A. et al. One thousand somatic SNVs per skin fibroblast cell set baseline of mosaic mutational load with patterns that suggest proliferative origin. Genome research 27, 512–523, https://doi.org/10.1101/gr.215517.116 (2017).
Kotton, D. N. & Morrisey, E. E. Lung regeneration: mechanisms, applications and emerging stem cell populations. Nature medicine 20, 822–832, https://doi.org/10.1038/nm.3642 (2014).
Hogg, J. C. et al. The nature of small-airway obstruction in chronic obstructive pulmonary disease. The New England journal of medicine 350, 2645–2653, https://doi.org/10.1056/NEJMoa032158 (2004).
Hodge, S. J., Hodge, G. L., Holmes, M. & Reynolds, P. N. Flow cytometric characterization of cell populations in bronchoalveolar lavage and bronchial brushings from patients with chronic obstructive pulmonary disease. Cytometry. Part B, Clinical cytometry 61, 27–34, https://doi.org/10.1002/cyto.b.20020 (2004).
Nishioka, M. et al. Identification of somatic mutations in postmortem human brains by whole genome sequencing and their implications for psychiatric disorders. Psychiatry and clinical neurosciences 72(4), 280–294, https://doi.org/10.1111/pcn.12632 (2017).
Hoang, M. L. et al. Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing. Proceedings of the National Academy of Sciences of the United States of America 113, 9846–9851, https://doi.org/10.1073/pnas.1607794113 (2016).
Jager, M. et al. Measuring mutation accumulation in single human adult stem cells by whole-genome sequencing of organoid cultures. Nature protocols 13, 59–78, https://doi.org/10.1038/nprot.2017.111 (2018).
Tomasetti, C. & Vogelstein, B. Cancer etiology. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81, https://doi.org/10.1126/science.1260825 (2015).
Tomasetti, C., Li, L. & Vogelstein, B. Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention. Science 355, 1330–1334, https://doi.org/10.1126/science.aaf9011 (2017).
Bae, T. et al. Different mutational rates and mechanisms in human cells at pregastrulation and neurogenesis. Science 359(6375), 550–555, https://doi.org/10.1126/science.aan8690 (2017).
Wauters, E. et al. DNA methylation profiling of non-small cell lung cancer reveals a COPD-driven immune-related signature. Thorax 70, 1113–1122, https://doi.org/10.1136/thoraxjnl-2015-207288 (2015).
Bruse, S. et al. Increased methylation of lung cancer-associated genes in sputum DNA of former smokers with chronic mucous hypersecretion. Respiratory research 15, 2, https://doi.org/10.1186/1465-9921-15-2 (2014).
Acknowledgements
We thank Yohannes Tesfaigzi, Lovelace Respiratory Research Institute, Albuquerque, NM, USA, for helpful discussions. We are grateful to Raul Tonda for providing the script for the Rainfall-plots. The EvA study is an EU-funded project (#200506) under the Seventh Framework Programme (FP7). We acknowledge the support of the Spanish Ministry of Economy, Industry and Competitiveness (MEIC) through the Instituto de Salud Carlos III and the 2014–2020 Smart Growth Operating Program to the EMBL partnership. We also acknowledge the support of the Centro de Excelencia Severo Ochoa, and the Generalitat de Catalunya through the Departament de Salut, Departament d’Empresa i Coneixement and the CERCA Programme.
Author information
Authors and Affiliations
Contributions
I.G.G. initiated and directed the study. M.W., U.K., M.H.N., A.N., D.G., J.M.H., T.W., C.E.B., D.G.P., A.P., J.M.Q., T.G., M.S., P.B., I.B., B.D., D.S. and L.Z.H. contributed to the recruitment and examination of subjects. K.A.R, L.A., A.B., J.F.D. and M.G. participated in generating the sequencing data. G.A.T., S.D., F.C.G. performed the analyses. L.A., M.G., L.Z.H. and I.G.G. participated in supervising the data generation and analyses. G.A.T. and I.G.G. interpreted the results. G.A.T. drafted the manuscript. S.D., F.C.G, D.S., L.Z.H. and I.G.G. critically reviewed the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
T.G. reports personal fees from Astra Zeneca, Berlin-Chemie, Boehringer-Ingelheim, Chiesi, CSL-Behring, GSK, Novartis, and grants and personal fees from Grifols, outside the submitted work; and reports grants from the European Union during the conduct of the study. The other authors declare no competing financial and/or non-financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thun, GA., Derdak, S., Castro-Giner, F. et al. High degree of polyclonality hinders somatic mutation calling in lung brush samples of COPD cases and controls. Sci Rep 9, 20158 (2019). https://doi.org/10.1038/s41598-019-56618-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56618-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.