Abstract
The spread of multidrug resistance via mobile genetic elements is a major clinical and veterinary concern. Pathogenic Escherichia coli harbour antibiotic resistance and virulence genes mainly on plasmids, but also bacteriophages and hybrid phage-like plasmids. In this study, the genomes of three E. coli phage-like plasmids, pJIE250-3 from a human E. coli clinical isolate, pSvP1 from a porcine ETEC O157 isolate, and pTZ20_1P from a porcine commensal E. coli, were sequenced (PacBio RSII), annotated and compared. All three elements are coliphage P1 variants, each with unique adaptations. pJIE250-3 is a P1-derivative that has lost lytic functions and contains no accessory genes. In pTZ20_1P and pSvP1, a core P1-like genome is associated with insertion sequence-mediated acquisition of plasmid modules encoding multidrug resistance and virulence, respectively. The transfer ability of pTZ20_1P, carrying antibiotic resistance markers, was also tested and, although this element was not able to transfer by conjugation, it was able to lysogenize a commensal E. coli strain with consequent transfer of resistance. The incidence of P1-like plasmids (~7%) in our E. coli collections correlated well with that in public databases. This study highlights the need to investigate the contribution of phage-like plasmids to the successful spread of antibiotic resistant pathotypes.
Similar content being viewed by others
Introduction
Escherichia coli is a ubiquitous Gram-negative rod commonly found as a commensal inhabitant of the mammalian gut, but also capable of causing a wide range of diseases (from diarrhea to urinary-tract infections, meningitis and sepsis) in both humans and animals1. The global spread of multidrug resistance (MDR) in pathogenic E. coli limits treatment options, imposing a significant burden on health and veterinary agencies worldwide2. Successful dissemination of E. coli pathotypes depends on the acquisition and carriage of niche-specific traits that allow for stable colonization and persistence1,3. The genes encoding such traits are often associated with mobile genetic elements (MGEs), such as plasmids and bacteriophages4,5,6.
In E. coli, MDR spreads predominantly through horizontal gene transfer of antimicrobial resistance genes (ARGs), residing on large self-mobilizable plasmids and often co-localized in large complex resistance loci with transposons and insertion sequences (IS)6,7. Virulence genes can also be found on the same plasmids as ARGs8,9. A crucial evolutionary step in the emergence of highly virulent MDR E. coli clones, a cause of major disease outbreaks worldwide, was the acquisition of MDR plasmids10,11,12,13. However, other elements of the mobile genome, such as bacteriophages, also play a role in the dissemination of these adaptive traits5,14,15, and a better understanding of their diversity and of the extent of their contribution to pathotype adaptation is critical16,17.
Improvements in sequencing technologies, particularly availability of long-read sequencing platforms, have allowed the identification of numerous elements combining bacteriophage and plasmid modules in enterobacteria15,18. Many of these phage-like plasmids are derivatives of phages P1 or P7, closely-related temperate coliphages of the family Myoviridae19. These prophages infect a wide range of enteric Gram-negative bacteria, including E. coli, and are unusual in being commonly found as free circular plasmids stably maintained in bacterial cells without integration in the chromosome. Based on the presence of a plasmid replicon and plasmid maintenance genes, these elements were classified as belonging to plasmid incompatibility group (Inc) Y19,20,21 (Fig. 1a). Their stability and versatility is enhanced by the presence of a complex immunity circuitry encoded by three loci (ImmI, ImmC and ImmT22) and of a variable ‘C-segment’ in the tail fiber-encoding operon19. The immunity functions provide protection from infection by foreign DNA and competing phages; the C-segment works like a plasmid shufflon, inverted by a site-specific recombinase (Cin in P1, located 5′ to the C-segment), which confers tail fiber variability and, consequently, flexible phage host range19,23 (Fig. 1a).
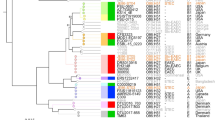
Comparison of P1-like genomes with coliphage P1 (NC_005856). (a) P1 genome mod749::IS5 c1.100 thermosensitive mutant19. RD, regions of difference from P1 genome in P1variants. (b) Low G + C content regions in the P1 backbone associated with recombination hot-spots (RD1-5). (c) pJIE250-3, from clinical human E. coli ST405. The region between the P1-like operons for ‘head processing’ (prt, pro) and ‘lysis’ (lydE, lydD, lyz) includes insertion of IS609 (positions 18,574–19,913; cleavage sites: left, TTAT; right, TCAA). (d) pSvP1, from porcine ETEC, with IS-mediated rearrangements leading to acquisition of F-type ETEC plasmid sequence including enterotoxin genes. From the end of ssb to the start of the ‘C segment’ (18,833–30,870), pSvP1 is closely related to P7 (96% overall nucleotide identity) with the addition of an IS10-like element (20,058–21,386) and IS609-like sequence (25,544–26,881). (e) pTZ20_1P, from commensal porcine E. coli, with plasmid fragments carrying antibiotic resistance genes (ARG). IS-mediated rearrangements led to the inversion of the P1 segment between tub and lpa, and deletion of P1-associated orfs (~11 kb encompassing the C-segment, tail fibers, ‘base plate and tail tube’ modules). The intergenic region between position 48,153 and 49,085 has 99% nucleotide identity to that of recently sequenced P1variants, but not P1 itself, and closer identity to P7 than P1 prevails in the ‘plasmid replication’, ‘lytic replication’ and ‘antirepressor’ operons. Between positions 89,842–89,967, corresponding to the P1 ant1/2 overlapping orfs (immunity determinants), there is a small 126-bp gap. Schematics of genome sequences generated using SnapGene Viewer 4.1.4 (GSL Biotech; available at snapgene.com), where colors of coding sequences indicate different functional modules. Genome comparisons generated using Easyfig visualization tool53. Blue/grey blocks between P1 and each of the three P1-like plasmids schematics represent regions of conserved synteny with varying pairwise nucleotide identity according to BLASTn [scale bars for direct matches from 100% (dark blue) to 65% (light blue); grey indicates matches in reverse orientation].
The unique lifestyle of IncY phage-like plasmids facilitates transduction events24, and, in the era of multidrug resistant virulent clones, these elements may play an as-yet-underestimated role in the transfer of resistance and virulence genes. The genomes of parents P1 and P7 contain signature mobile elements, e.g. IS1 in P1; IS903 and Tn2 with blaTEM-1b (ampicillin resistance) in P7, and phage-like plasmids carrying ARGs linked to transposable elements have been sequenced from enteric bacteria isolated worldwide. P1/P7-like elements carrying extended spectrum β-lactamase genes have been identified in human clinical E. coli (e.g. blaSHV-225) and Klebsiella pneumoniae strains (e.g. blaCTX-M-1526), and in Salmonella from pigs (blaCTX-M-2727), and carrying colistin resistance (mcr-1) in E. coli from humans18 and animals28,29, pointing at clinical and livestock reservoirs.
Here, we have carried out detailed comparisons of the genomes of three P1-like elements from Australian clinical and veterinary E. coli collections to identify genomic traits that may indicate the adaptive capacities and ease of spread/persistence of these elements: pJIE250-3, with no accessory genes, from an MDR E. coli isolated from a hospitalized human, and pTZ20_1P carrying ARGs, and pSvP1 carrying ETEC virulence genes, both from porcine E. coli.
Results
Genomes of P1-like plasmids
For all three phage-like elements PacBio sequencing and assembly generated one contig. E. coli TZ20_1P does not carry any plasmids other than pTZ20_1P, while JIE250, host of pJIE250-3, carries an MDR F-type plasmid30 and SvETEC, host of pSvP1, carries MDR IncHI2 and IncI1 plasmids31,32 and an FII: (F10:A-:B)-type plasmid encoding additional virulence factors and tetracycline resistance32. pJIE250-3, pSvP1 and pTZ20_1P are all variants of coliphage P1 (Fig. 1; Table 1), although some operons in each genome have closer identity to P7 (GenBank AF503408; Fig. 1c–e). pJIE250-3 appears to be a P1 derivative that has lost lytic replication functions and does not contain any cargo genes. In pSvP1 and pTZ20_1P, the P1-like backbone is interrupted by different IS-mediated insertions of plasmid fragments: a large section from an ETEC F-type plasmid carrying virulence determinants in the former, and plasmid backbone plus modules carrying ARGs in the latter.
Common regions of difference (RD)
Comparative analysis with the chosen P1 reference genome19 (P1 mod749::IS5 c1.100 mutant, GenBank NC_005856) revealed five hot-spots of sequence variation common to all three elements (RD-1 to RD-5; Fig. 1a), mainly associated with boundaries between regions of average G+C content and AT-rich loci (Fig. 1b). However, in each RD, changes from the P1 backbone were unique for each element (Fig. 1c–e; Table 1). The res and mod genes (restriction modification function; RD-1) and IS1 of P1 (RD-2) are missing from all three elements, replaced by orfs unique to each, but generally highly similar to sequences identified in other phage-like plasmids from E. coli (Table 1). Between RD-1 and RD-2, the ‘C1 modulation’ (implicated in control of lysogeny), ‘antirestriction’, ‘head processing’, ‘viral architecture’, and ‘lysis’ operons largely match those of P1 or P719 (Fig. 1c–e), except for IS609 inserted in pJIE250-3 and pSvP1 (Fig. 1c,d; Table 1). RD-3 corresponds to the P1 C-segment and adjacent genes, encoding tail structures (fibers, baseplate, tail tube), the ImmI-associated region (immunity function: sim operon, icd, ant1/2;), kilA and the ‘lytic replication’ module (repL, oriL, rlfA and B)19,22,33 (Fig. 1a). pJIE250-3 has only one tail fiber gene and all other P1 modules are missing (Fig. 1c). In pSvP1 and pTZ20_1P, RD-3 includes IS-mediated insertions of plasmid backbone fragments and associated resistance or virulence genes, with consequent deletion of various segments of the P1 backbone (Fig. 1d,e; Table 1). A gene related, but not identical, to cin of P1, encoding the invertase responsible for tail switching, was identified in each element (Table 1). However, this gene was truncated (pSvP1; pTZ20_1P) or in reverse orientation (pJIE250-3), when compared to P1-cin, and/or its target (C-segment) was greatly modified (pJIE250-3 and pTZ20_1P), likely impairing functionality of the switch (Table 1).
RD-4 corresponds to the P1 plasmid replicon (including parAB and repA) and flanking regions (Fig. 1). pJIE250-3 is almost identical to P1 in this region except for a short insertion 5′ of oriR, containing two orfs. In both pSvP1 and pTZ20_1P, the main changes here involve sequences encoding incompatibility determinants, with potential modification of replicon function (Table 1). RD-5 is located between pap and lxr, where in all three elements some of the genes encoding head morphogenesis in P1 are modified (closer identity to P7) or deleted (Fig. 1; Table 1). Additionally, in pTZ20_1P two short hypothetical orfs and a gene encoding an antirepressor (ant) are found inserted in the P1 backbone between pacB and c1, as in other phage-like plasmids (e.g. RCS47, NC_042128; Fig. 1e).
Unique features of P1-like plasmids
pJIE250-3
In pJIE250-3, five additional orfs are inserted between the c8 and mat genes at an AT-rich junction, near a cluster of 4× 4-bp direct repeats (DR) (ATTG; 1,957–1,982), close to the insertion site of Tn2 in P7. A similar segment was found in nine other E. coli plasmids (e.g. CP032890.1, CP019262.1 and MG825383.1, best matches with >99.5% nt identity) and in the chromosome of E. coli ST274734 (CP007393.1, >99.4% nt identity), and includes genes encoding a putative transcriptional regulator of metabolic functions as well as a putative resolvase.
Plasmid sequence in pSvP1
Five copies of ISEc84 (ISEc84.1- ISEc84.5), a member of the IS91-family (rolling circle transposition)6,35, are present in RD-3 of pSvP1 and rearrangements mediated by ISEc84 seem to have been largely responsible for modification of the P1 backbone here (Fig. 1d; Fig. 2). The P1-like backbone (pmgB to upfA) between oppositely-oriented ISEc84.1 and ISEc84.2 is inverted, presumably as a result of IS-mediated recombination (Fig. 2). ISEc84.2 is followed by a fragment of the left end (inverted repeat, IRL) of IS3 interrupted by IS100, then a region that includes several IS (complete and partial) within a large segment (~41.5 kb; at position 67,105) matching F-type plasmids of porcine ETEC [e.g. p14ODTX (MG904993.1; 94,167 bp); pUMNK88_Ent (CP002732.1; 81,475 bp)36] (Fig. 1d). Addition of this plasmid backbone segment has led to acquisition of a new plasmid replicon (repFIB), endotoxin genes (stb, encoding the heat stable enterotoxin II, and eltAB, encoding the heat labile enterotoxin), and a toxin-antitoxin stability system. Several other IS, including ISEc84.3, are found within this insertion exactly as in the ETEC plasmids (Fig. 2). The acquired plasmid sequence ends at ISEc84.4, flanking another partial IS3. This arrangement suggests incorporation of a plasmid-derived circular molecule containing ISEc84 within IS3 into a P1-like backbone (Fig. 2) and concomitant deletion of the P1 DNA methylation operon, phage tRNAs, DNA helicase and part of the ‘baseplate/tail tube’ module (between upfA and 24; Fig. 1a,d).
Insertion sequences and IS-mediated rearrangements in the pSvP1 genome. IS10 (IRR, 21,385–21,336, and IRL, 20,057–20,106; 9-bp DR, TGCTCTGCA) interrupts the P7 porin precursor gene nmpC, and at 40,334 IS2 interrupts the orf related to P1 sit (5-bp direct repeats, ACCAA). IS186B (IS4 family; 10-bp direct repeats, GGATCTCTCC) is inserted between bplB and the ‘lytic replication’ module, causing deletion of P1-like sequence associated with lytic replication (rlfA, rlfB) and with putative morphogenetic function (pmgF). ISEc84.1 interrupts a P1 pmgB homolog, but the remaining part of pmgB lies adjacent to ISEc84.2 in the opposite orientation. As shown, reversing the segment between ISEc84.1 and ISEc84.2 and removing an ISEc84 would regenerate a complete pmgB gene. ISEc84 insertion mediated acquisition of a large fragment of ETEC plasmid containing multiple IS elements. The region between ISEc84.2 and ISEc84.4 corresponds to ETEC plasmid sequence containing multiple IS elements. IS3 fragments (green) flanking these ISEc84 suggest insertion of a circular molecule carrying ISEc84 inserted in IS3 (as shown) by recombination in a copy of ISEc84 in the P1-like backbone. ISEc84 and IS91 may also have been responsible for the acquisition of a region related to E. coli IncY MDR plasmid pR15_MCR-1 (95% nucleotide identity; GenBank MK256965.1), containing CDSs involved in restriction modification (type I) and DNA methylation with deletion of several P1 pmg genes (putative morphogenetic function). Schematic not to scale.
Plasmid sequence in pTZ20_1P
pTZ20_1P contains a sul3 type class 1 integron in which intI1 is truncated by IS26 (ΔintI1509) and the cassette array is dfrA12-gcuF-aadA2-cmlA-aadA1-qacI, conferring resistance to trimethoprim, streptomycin and spectinomycin, chloramphenicol and quaternary ammonium compounds. Beyond sul3 (sulphonamide resistance), the mefB gene is truncated by IS26 (ΔmefB260). Another resistance region includes blaTEM-1b (ampicillin resistance), aac(3)-IV (gentamicin and tobramycin resistance) and aph(4)-Ia (hygromycin resistance) genes associated with complete and truncated IS and fragments of various Tn3-family transposons (Fig. 3). This set of resistance genes fully accounts for the antibiotic resistance phenotype of TZ20_1P.
Multiple antibiotic resistance regions and rearrangements in pTZ20_1P. (a) Schematic of the complete pTZ20_1P genome showing insertion points of plasmid regions containing resistance genes, transposons and IS. Two oppositely-oriented copies of part of Tn1722 (marked by grey chevrons) are found 79,484 bp apart and the 5-bp immediately adjacent to the IRL and IRR of Tn1722 are reverse complements of one another (AACTA; TGATT). (b) Reversing the region between the Tn1722 fragments (to mimic homologous recombination) regenerates a complete Tn1722, with matching 5-bp direct repeats (TGATT) marking the insertion. Similarly, reversing a 2,182 bp segment between two inversely-oriented copies of IS26 (to mimic intramolecular transposition by IS2660) results in an IS26 flanked by matching 8-bp direct repeats. This, presumably ancestral, version of pTZ20_1P corresponds to a P1-like backbone with three separate insertions (1) an 39,980 bp region bounded by directly oriented copies of IS26, (2) IS26, and (3) Tn1722. Almost identical sequence after the leftmost IS26 through the sul3-type integron to IRL of Tn21 (1) is found in a few Salmonella ssp. and E. coli plasmids in INSDC databases. Tn21 truncates Tn1722, which is also truncated by the IRR end of Tn5393, with novel boundaries in both cases. Tn5393 is truncated by ISEc59, an IS6 family element, with another copy of IS26 flanking a 2,447 bp region containing the aph(4)-Ia and aac(3)-IV genes. The IRR end of IS26 truncates the IRL end of Tn2, leaving an intact blaTEM-1b gene. IRR of Tn2 is immediately followed by 49 bp of Tn5393 IRL, 106 bp of the IRL end of Tn1722 and 177 bp of the IRL end of IS1. The 14,126 bp region between IS1Δ and the next IS26 corresponds to finO-traX-traI-traD-traT-traG-traHΔ genes matching (~98% identity) several F-type plasmids in INSDC, some of which have the same boundary with IS1. Diagrams not to scale.
No DR indicative of insertion were identified adjacent to any mobile elements. However, sequences of the expected DR length that are reverse complements of one another are found adjacent to the terminal IR of two Tn1722 fragments (5 bp) and two different copies of IS26 (8 bp). This suggests inversions, consistent with the comparison between pTZ20_1P and P1 (Fig. 3). Reversing these generates a (presumably ancestral) version of pTZ20_1P with three distinct insertions in the P1 backbone. One comprises a 39,980 bp region bookended by directly-oriented copies of IS26 and containing the integron and other resistance genes, plus 14,126 bp matching part of the tra region of several F-type plasmids in INSDC databases (Fig. 3). This suggests IS26-mediated transfer of part of a plasmid to a P1-like backbone. The other two are separate insertions of IS26 and of a complete Tn1722 directly into the P1-like backbone, each flanked by DR. The truncated cin gene and the packase-encoding gene pacA (88% nt identity to P1) flank the plasmid insertion (Figs. 1c and 3).
pTZ20_1P self-transfer ability
Induction, lysogeny and conjugation ability were tested using pTZ20_1P, as this element carries resistance markers and no additional plasmids were detected in its host, simplifying selection strategies. We evaluated whether pTZ20_1P is inducible, treating exponential growth phase cultures with mitomycin C or UV light. To confirm the presence of the induced phage-like element in the filtered suspension, we performed PCR for the replication genes repA (plasmid) and repL (phage), and linking Tn1722 (plasmid) and the pacA gene (phage) (Fig. S1). PCR amplification was successful in all induced cultures, except samples exposed to UV light for 20 s (shortest exposure time; Supplementary Fig. S1a). No amplicons were obtained from DNA extracted from an un-induced filtered suspension of E. coli TZ20_1P. We tested the ability of pTZ20_1P to lysogenize different commensal E. coli host strains. Lysogens were only detected for E. coli WH17, not for E. coli J53 or WGNB13. PCR amplification of marker genes and whole genome sequencing (WGS) of lysogens confirmed the acquisition of pTZ20_1P (Supplementary Fig. S1b). The capacity of pTZ20_1P to transfer by conjugation was also tested, but no transconjugants were obtained, as expected due the absence of a complete conjugative transfer operon in the inserted plasmid segment (Fig. 3).
Incidence of P1-like plasmids in Australian E. coli
In silico screening of available genome sequences for the P1-associated repL (phage) and repA (plasmid) replication genes, identified both in 8/117 (6.8%) isolates from a collection of porcine commensal E. coli, but only in 3/328 (1%) isolates from human clinical collections of MDR E. coli and Klebsiella pneumoniae. However, BLASTn searches of the INSDC database (accessed September 2018) using the same targets returned 54 entries containing both genes (repA >98% identity; repL >97% identity). Using the search term ‘plasmids Escherichia coli’ to query genome entries in the NCBI database, we found 1429 sequences annotated as ‘plasmids’. Of these, ~7% had characteristics indicative of phage sequences (e.g. in 97/1429 (6.7%) t-RNA presence), a frequency comparable to that in our local collections, and 22 of these were also identified by the BLASTn screening. Screening for pSvP1 in the genome of sister strain E. coli O157 734/3 showed the presence of this element in the Australian lineage since 199531.
Discussion
In the era of rising MDR, large conjugative plasmids have been recognized as the main vehicles of antibiotic resistance maintenance and transfer, particularly in Enterobacteria. Phages and phage-like plasmids, however, may also have a prominent role in the dissemination of accessory adaptive traits5,15,37. Enterobacteria phage P1 is known to infect and lysogenize E. coli, being maintained within the cell as an autonomous low-copy number plasmid19. It does not contain cargo genes of clinical interest and, being a prophage, has not been considered for therapeutic applications. Therefore, although P1 was discovered in E. coli over 50 years ago19,22, research efforts have mainly focussed on its properties as a molecular biology tool rather than as an active element of the accessory genome22,38,39. However, recent advances in sequencing technologies (e.g. PacBio) have facilitated the detection of numerous P1-like variants, though detailed characterization has been limited to a few carrying ARGs of clinical relevance18,25,27,29. In this study, we defined the fine diversity between three P1-like plasmids from E. coli, isolated from different reservoirs (animal and human), with the intention of identifying modifications that may associate with pathogen adaptation.
Analysis of pJIE250-3, pTZ20_1P and pSv1P revealed three different variants of P1 sharing common traits. pJIE250-3 is an example of a P1-derivative that has lost its lytic replicon, but retained P1 plasmid functions augmented by acquisition of additional plasmid-related orfs (e.g. transcriptional regulators). pTZ20_1P and pSv1P are genetic mosaics of P1-like elements and plasmid segments with ARGs and virulence determinants, respectively. The genomes of all three phage-like plasmids are defined by different features unique to each, and none has an exact match in INSDC databases. However, each genetic locus is shared with at least two other phage-like plasmids, highlighting the variability of phage-like MGEs, testament to their recognised transduction ability19,39. Whether this variability is a product of random reassortment due to generalized transduction or a product of the adaptive strategy of different pathotypes to their specialized environment is yet to be determined.
The variable regions (RD) distinguishing each element are associated with the same limited number of P1 genetic loci of low G+C content, where P1 genes previously described as recent acquisitions (e.g. res-mod, sim, rfl) are located19. These hot-spots tend to be related to host-range determinants (C-segment; tail fibers), replication functions (modification of lytic or plasmid replicons), and immunity encoding regions (ImmI specifically). Inspection of other P1/P7-like plasmids described in the literature confirms that these RD tend to be shared among elements isolated from different E. coli worldwide18,25,28,29. All three P1-like plasmids have a modified tail fiber operon when compared to P1, with an altered cin sequence, and lack a complete C-segment locus (tail tip switch40). Cin, a P1/P7 specific invertase, is responsible for the production of virions with different host specificity from the parent, a property presumably lost in these P1 variants, with possible consequent restriction of host range. In pJIE250-3, this feature is combined with loss of repL (essential for lytic replication), likely preventing self-mobilization of this element from its host, an E. coli clinical strain carrying multidrug resistance on a large F-type conjugative plasmid30. However, pJIE250-3 has retained P1 plasmid loci and acquired several additional genes with putative metabolic/DNA processing functions, which could allow for better stability in a fixed genomic background.
In P1, the integrity of the plasmid replicon (iterons), partitioning module and plasmid addiction systems, including res-mod (notably absent in the three variants), is important to ensure stable plasmid maintenance. The absence of res-mod may decrease protection from entry of foreign DNA, but there are indications that the additional genes acquired by the P1variants may provide functions facilitating adaptation to local environments (e.g. in pSvP1, acquisition of a type I restriction modification system, in pJIE250-3 transcriptional regulators etc.). In pSvP1, plasmid functions are also likely enhanced through acquisition of repFIB and additional stability and maintenance modules in the inserted plasmid segment.
Negative impact on broad host-range could be associated with modification of the complex P1 immunity circuitry protecting the prophage by exclusion of both foreign phage and transducing DNA19. P1 has three immunity encoding regions, Imm I, T and C22,33. ImmC, with C1 repressor of lytic function, and ImmT, implicated in C1 modulation, promote plasmid (prophage) maintenance versus entry into the lytic cycle. ImmI (sim, c4, icd, ant1/2) has been ascribed a specific role in permitting P1 and P7 coexistence (with minor nucleotide differences between P1 and P7 responsible for their reported heteroimmunity) and adding flexibility to ImmC functions33,41. P1 and P7 are heteroimmune relatives known to readily recombine42, as evidenced in the genomes of our elements, but in pJIE250-3 and pSvP1, the ImmI region is modified, possibly compromising superinfection immunity functions. However, interestingly in pJIE250-3 the absence of ImmI is associated with the concomitant loss of repL, and in pSvP1 with the acquisition of homologs of P1 genes with putative immunity functions (icd and ant1/2 in RD-1).
Here, we showed that P1-like elements in E. coli can co-exist with large plasmids carrying complex resistance regions, can pick up different cargo genes (MDR and virulence), and move into a different host. In the lysogenized commensal E. coli WH17, F-type replicons were detected and the MDR transferred as part of pTZ20_1P could potentially transfer to these. There have also been reports of transfer of non-conjugative plasmids between E. coli cells by transformation mediated by P1 phage lysis43,44,45, indicating that multiple modes of interaction and spread of these elements may shape the adaptive potential of a bacterial population44. Both pJIE250-3 and pSvP1 were found in the same cell as large plasmids and, provided that they are capable of productive lysis45 (less likely for pJIE250-3 lacking the repL gene), could promote or enhance plasmid movement by these mechanisms. As the mammalian gut is an excellent niche for ARG exchange via horizontal transfer, tracking these elements may become crucial in responding to the threat of rising MDR in enteric pathogens.
In pTZ20_1P, IS26-mediated transposition events likely led to the acquisition of the multidrug resistance region by the P1-like backbone, with IS26 truncating both intI1 (IS26-ΔintI1509) and mefB (IS26-ΔmefB260). The presence of an intact PC promoter allows expression of the dfrA12 gene in the cassette array, as confirmed by the strain’s antibiotic resistance phenotype (trimethoprim resistance), even though IS26-ΔintI1509 may prevent the integration of new gene cassettes into the array. The IS26-ΔmefB260 signature is frequently found on enterobacterial plasmids in combination with sul3, including in porcine and human E. coli4,31,46, while IS26-ΔintI1509 is much less common. IS26-ΔmefB260 and IS26-ΔintI1509 together represent a valid signature for tracking this specific class 1 integron in different hosts and environments.
The elements described here are examples of how P1 can evolve in different E. coli genomic backgrounds by modification of replication and host range properties in association with acquisition of cargo genes, showing that P1/P7-like plasmids may not only be capable of generalized transduction, but also be specific vehicles for spread and maintenance of virulence and antimicrobial resistance in both clinical and livestock production settings. MGEs coexisting within the same host interact with each other, affecting reciprocal stability, and their cooperative interactions should be considered when defining bacterial adaptive strategies.
Methods
Bacterial strains
Three E. coli isolates belonging to different sequence types (ST) were host to the three P1-like plasmids described here. E. coli TZ20_1P, ST372, phylogroup B2, serotype O6:H31, was isolated in January 2017 at the Elizabeth MacArthur Agricultural Institute (EMAI), Menangle, NSW, Australia, from faecal material from a healthy four-week-old piglet not previously treated with antimicrobials. E. coli SvETEC ST4245, phylogroup C, serotype O157, was also isolated from faecal material from a diarrhoeic piglet, in 200832. E. coli JIE250, ST405, phylogroup B2, is a human isolate, part of a large clinical collection of Enterobacteriaceae30. This isolate was shown to carry a large conjugative F-type plasmid with a complex MDR region including multiple antibiotic resistance genes (e.g. blaCTX-M-15), transposons and IS47.
PacBio sequencing, annotation and bioinformatic analysis of MGEs genomes
Genomic DNA was isolated and purified from bacterial cultures grown overnight using the Mo Bio Powersoil® DNA Isolation kit (Mo Bio, Carlsbad, CA, USA) or the DNAesy Blood and Tissue kit (Qiagen, Hilden, Germany), according to manufacturer’s instructions. Long-read sequencing was performed on the three E. coli isolates on a PacBio RSII Instrument at the Ramaciotti Centre for Genomics (UNSW, Sydney, Australia). Polishing and assembly of sequenced reads was performed at the Ramaciotti Centre using HGAP and CANU, and plasmids were closed using Circlator48. Errors in PacBio assemblies were checked and curated by alignment with Illumina short reads from whole genome sequencing of the E. coli hosts. P1-like genomes were first annotated using RASTtk49, then manually curated using BLAST functions50, SnapGene (GSL Biotech; available at snapgene.com) and Geneious v.9.1 (https://www.geneious.com). Plasmid virulence and resistance regions were also annotated using web-based software [Center for Genomic Epidemiology, www.genomicepidemiology.org; GalileoTM AMR (formerly MARA), galileoamr.arcbio.com/mara/51]. All allelic variants of IS2652 detected in pTZ20_1P are referred to as IS26 throughout. Comparisons with the reference P1 genome (P1 mod749::IS5 c1.100 mutant; GenBank NC_005486)19 were visualized using EasyFig.53.
Transfer of pTZ20_1P
Phage induction and lysogenization of commensal E. coli
E. coli TZ20_1P was grown in lysogeny broth (LB; Becton Dickinson, Franklin Lakes, NJ, US), with vigorous shaking at 37 °C, to OD600 0.3, and treated with mitomycin C at 0.05, 0.1, 0.15, or 0.2 μg/mL, or exposed to UV light for 20 s, 45 s or 1 min (UV Stratalinker 1800 (230 Vac, 2 A, 50 Hz), Stratagene, CA, US). Induced cultures were incubated at 37 °C for 3 h with gentle shaking and centrifuged at 4,000 x g for 20 min to remove bacterial cells debris. Supernatants were filtered and concentrated using 0.22 μm Amicon Ultra-15 filters (Sigma-Aldrich, St. Louis, MO, USA). Suspensions were stored overnight at 4 °C prior to lysogenization assays. Three previously characterised commensal E. coli strains (WGNB13 and WH1754 and J5355), which are streptomycin susceptible but carry other resistance genes suitable for counter-selection, were used as recipients to test the lysogenic ability of pTZ20_1P. Recipient strains grown to OD600 0.6 in LB were pelleted by centrifugation and resuspended in LB supplemented with 1 mM CaCl2. Bacteria were mixed (1:1) with phage lysates, and the mix was incubated in LB broth (static conditions, 40 min) or on LB agar plates (overnight) at 37 °C. E. coli recipients were also mixed with un-induced E. coli TZ20_1P, as negative controls. The mixtures were plated on LB agar supplemented with Str 25 μg/mL (selection for phage-like plasmid) and incubated at 37 °C for 24 h and 48 h. All transfer experiments were performed in triplicate.
Characterization of lysogenic E. coli
To confirm that the suspension obtained after induction contained pTZ20_1P and its consequent stable acquisition by recipient strains, we performed PCR amplification of two regions: (1) the plasmid replication gene repA (repA-fw, AAAGCCGAGGGTTACGATGA, and repA-rev, ATGATACGGTTTTGCTCGCC; amplicon size 542 bp; this study), and (2) the phage lytic replication gene, repL, unique to the P1 component (RepL-fw and RepL-rev25; amplicon size 489 bp). To unequivocally confirm the identity of the transferred element, we also amplified the region linking plasmid (Tn1722) and phage (pacA) modules of this element (Tn1722, CAGACTGGAAGACGGGAAGT; pacA TCAGCCATTTCAGCCACAAC; amplicon size 428 bp; this study). Phage DNA was isolated from filtered induced suspensions by treatment with DNAse (10 mg/mL; 30 mins) and RNAse (100 mg/mL; 30 mins), followed by extraction and purification using the Wizard DNA Clean-up System/kit (Promega, Madison, WI, USA) following manufacturer’s instructions.
The genome of one representative E. coli lysogen (E. coli WH17 mitomycin C_0.2 μg/mL) was sequenced to confirm pTZ20_1P acquisition. Bacterial DNA was extracted and purified using the ISOLATE II genomic DNA kit (Bioline, London, UK). WGS was performed on the Illumina NextSeq (paired-end 150 bp × 2) platform at the Pathogen Genomics Unit, Centre for Infectious Disease and Microbiology – Public Health, Westmead Hospital (NSW, Australia). Briefly, total DNA concentration was quantified using Quant-it PicoGreen dsDNA Assay Kit (Invitrogen, Carlsbad, CA, USA) and 1 ng/µl of DNA was used to prepare DNA libraries using the Nextera XT Library Preparation Kit and Nextera XT v2 Indexes (Illumina, San Diego, CA, USA). Multiplexed libraries were sequenced using paired end 150 bp chemistry on the NextSeq. 500 NCS v2.0 (Illumina). Error rates were calculated using PhiX Sequencing Control v3 for each run. Demultiplexing and FastQC generation was performed automatically by BaseSpace (Illumina). Alignment between the lysogenic genome (raw reads) and the original pTZ20_1P PacBio sequence was generated and visualised using Bowtie256 v2.3.0, SAMtools57 v1.4.1, and Tablet58 v1.19.05.28.
Conjugation assay
To assess the ability of pTZ20_1P to transfer via a plasmid-related mobilization mechanism, we performed conjugation assays using E. coli EC JM109 RifR/NalR, resistant to rifampicin (Rif) and nalidixic acid (Nal) as recipient8. A loopful each of donor (E. coli TZ20_1P) and recipient cultures were mixed thoroughly in saline (500 μL), and the mating mix was plated onto LB agar and incubated overnight at 37 °C. The mating lawn was resuspended in 1 mL saline and serial dilutions spotted in triplicate onto LB agar plates supplemented with Nal (30 μg/mL, Nal30 - recipient selection) and Nal30 plus ampicillin (100 μg/mL, Amp100) or streptomycin (25 μg/mL, Str25) to detect transconjugants (Nal30Amp100 or Nal30Str25, selection for pTZ20_1P), and LB only agar (total bacterial count). Donor and recipient were also separately plated on LB Nal30Amp100, LB Nal30Str25 and LB agar only as controls. Conjugation assays were also performed in the presence of E. coli HB101 containing the helper plasmid pRK600 with chloramphenicol resistance8,59. Donor and recipient, grown independently under the same conditions and plated onto LB supplemented with Str25 and Nal30 respectively, were used as controls.
Occurrence of phage-like plasmids in Australian E. coli collections and NCBI databases
The occurrence of P1-like plasmids in available sequences was determined by querying the NCBI bacterial genomes database and by in silico BLAST-type searches with repA and repL sequences (minimum percentage identity 95%) of representative collections of local E. coli genomes.
Data availability
The complete sequences of pTZ20_1P, pJIE250-3, and pSvP1 have been deposited in GenBank (NCBI) under accession numbers MN510447, MN510445, and MN510446 respectively. The complete raw read dataset for lysogenic E. coli WH17 mitomycin C_0.2 μg/mL containing pTZ20_1P has been deposited in the SRA (NCBI) database under Bioproject PRJNA565856 (SRR10127725). All other data generated or analysed during this study are included in this published article (and its Supplementary Information Files).
References
Kaper, J. B., Nataro, J. P. & Mobley, H. L. T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2, 123–140 (2004).
Roca, I. et al. The global threat of antimicrobial resistance: science for intervention. New Microbes New Infect. 6, 22–29 (2015).
Johnson, J. R. Virulence factors in Escherichia coli urinary tract infection. Clin. Microbiol. Rev. 4, 80–128 (1991).
Brilhante, M., Perreten, V. & Donà, V. Multidrug resistance and multivirulence plasmids in enterotoxigenic and hybrid Shiga toxin-producing/enterotoxigenic Escherichia coli isolated from diarrheic pigs in Switzerland. Vet J 244, 60–68 (2019).
Brown-Jaque, M., Calero-Cáceres, W. & Muniesa, M. Transfer of antibiotic-resistance genes via phage-related mobile elements. Plasmid 79, 1–7 (2015).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 31, e00088–17 (2018).
Carattoli, A. Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 2227–2238 (2009).
Venturini, C., Beatson, S. A., Djordjevic, S. P. & Walker, M. J. Multiple antibiotic resistance gene recruitment onto the enterohemorrhagic Escherichia coli virulence plasmid. FASEB J. 24, 1160–1166 (2010).
McKinnon, J., Roy Chowdhury, P. & Djordjevic, S. P. Genomic analysis of multidrug-resistant Escherichia coli ST58 causing urosepsis. Int. J. Antimicrob. Agents 52, 430–435 (2018).
Johnson, J. R., Johnston, B., Clabots, C., Kuskowski, M. A. & Castanheira, M. Escherichia coli sequence type ST131 as the major cause of serious multidrug-resistant E. coli infections in the United States. Clin. Infect. Dis. 51, 286–294 (2010).
Rasko, D. A. et al. Origins of the E. coli strain causing an outbreak of hemolytic-uremic syndrome in Germany. N. Engl. J. Med. 365, 709–717 (2011).
Petty, N. K. et al. Global dissemination of a multidrug resistant Escherichia coli clone. Proc. Natl. Acad. Sci. 111, 5694–5699 (2014).
Navarro-Garcia, F. Escherichia coli O104:H4 Pathogenesis: an enteroaggregative E. coli/Shiga toxin-producing E. coli explosive cocktail of high virulence. Microbiol. Spectr. 2(6), EHEC–008–2013 (2014).
Canchaya, C., Fournous, G. & Brüssow, H. The impact of prophages on bacterial chromosomes. Mol. Microbiol. 53, 9–18 (2004).
Balcazar, J. L. Bacteriophages as vehicles for antibiotic resistance genes in the environment. PLoS Pathog. 10(7), e1004219 (2014).
Dionisio, F., Zilhãob, R. & Gama, J. A. Interactions between plasmids and other mobile genetic elements affect their transmission and persistence. Plasmid 102, 29–36 (2019).
Smalla, K., Cook, K., Djordjevic, S. P., Klümper, U. & Gillings, M. Environmental dimensions of antibiotic resistance: assessment of basic science gaps. FEMS Microbiol. Ecol. 94, 12 (2018).
Zhou, W., Liu, L., Feng, Y. & Zong, Z. A P7 phage-like plasmid carrying mcr-1 in an ST15 Klebsiella pneumoniae clinical isolate. Front. Microbiol. 9, 11 (2018).
Łobocka, M. B. et al. Genome of bacteriophage P1. J. Bacteriol. 186, 7032–7068 (2004).
Meyer, J., Stålhammar-Carlemalm, M., Streiff, M., Iida, S. & Arber, W. Sequence relations among the IncY plasmid p15B, P1, and P7 prophages. Plasmid 16, 81–89 (1986).
Hedges, R. W., Jacob, A. E., Barth, P. T. & Grinter, N. J. Compatibility properties of P1 and φAMP prophages. Molec. Gen. Genet. 141, 263–267 (1975).
Yarmolinsky, M. B. Bacteriophage P1 in retrospect and in prospect. J. Bacteriol. 186, 7025–7028 (2004).
Iida, S., Meyer, J., Kennedy, K. E. & Arber, W. A site-specific, conservative recombination system carried by bacteriophage P1. Mapping the recombinase gene cin and the cross-over sites cix for the inversion of the C segment. EMBO J. 1, 1445–1453 (1982).
Schneider, C.L. Bacteriophage-mediated horizontal gene transfer: Transduction. In Bacteriophages; Harper, D., Abedon, S., Burrowes, B., McConville, M., Eds; Springer: Cham, Switzerland (2017).
Billard-Pomares, T. et al. Characterization of a P1-like bacteriophage carrying an SHV-2 extended-spectrum β-lactamase from an Escherichia coli strain. Antimicrob. Agents Chemother. 58, 6550–6557 (2014).
Shin, J. & Ko, K. S. A Plasmid bearing the bla CTX-M-15 gene and phage P1-Like sequences from a sequence type 11 Klebsiella pneumoniae isolate. Antimicrob. Agents Chemother. 59, 6608–6610 (2015).
Yang, L. et al. Characterization of a P1-like bacteriophage carrying CTX-M-27 in Salmonella spp. resistant to third generation cephalosporins isolated from pork in China. Sci. Rep. 7, 40710 (2017).
Zhang, C. et al. A phage-like IncY plasmid carrying the mcr-1 gene in Escherichia coli from a pig farm in China. Antimicrob. Agents Chemother. 61, e02035–16 (2017).
Li, R., Xie, M., Lv, J., Wai-Chi Chan, E. & Chen, S. Complete genetic analysis of plasmids carrying mcr-1 and other resistance genes in an Escherichia coli isolate of animal origin. J. Antimicrob. Chemother. 72, 696–699 (2017).
Zong, Z., Partridge, S. R., Thomas, L. & Iredell, J. R. Dominance of bla CTX-M within an Australian extended-spectrum beta-lactamase gene pool. Antimicrob. Agents Chemother. 52, 4198–4202 (2008).
Wyrsch, E. R. et al. Complete sequences of multiple-drug resistant IncHI2 ST3 plasmids in Escherichia coli of porcine origin in Australia. Front. Sustain. Food Syst. 3, 18 (2019).
Wyrsch, E. et al. Comparative genomic analysis of a multiple antimicrobial resistant enterotoxigenic E. coli O157 lineage from Australian pigs. BMC Genomics 16, 1382 (2015).
Heinrich, J., Velleman, M. & Schuster, H. The tripartite immunity system of phages P1 and P7. FEMS Microbiol. Rev. 17, 121–126 (1995).
Xavier, B. B. et al. Complete genome sequences of nitrofurantoin-sensitive and -resistant Escherichia coli ST540 and ST2747 strains. Genome Announc. 2, e00239–14 (2014).
Del Pilar Garcillán-Barcia, M., Bernales, I., Mendiola, M. V. & de la Cruz, F. Single-stranded DNA intermediates in IS91 rolling-circle transposition. Mol. Microbiol. 39, 494–501 (2001).
Shepard, S. M. et al. Genome sequences and phylogenetic analysis of K88- and F18-positive porcine enterotoxigenic Escherichia coli. J. Bacteriol. 194, 395–405 (2012).
Brown-Jaque, M. et al. Antibiotic resistance genes in phage particles isolated from human faeces and induced from clinical bacterial isolates. Int. J. Antimicrob. Agents 51, 434–442 (2018).
Westwater, C., Schofield, D. A., Schmidt, M. G., Norris, J. S. & Dolan, J. W. Development of a P1 phagemid system for the delivery of DNA into Gram-negative bacteria. Microbiology (Reading, Engl.) 148, 943–950 (2002).
Moore, S. D. Assembling new Escherichia coli strains by transduction using phage P1. Methods Mol. Biol. 765, 155–169 (2011).
Iida, S. et al. The bacteriophage P1 site-specific recombinase cin: recombination events and DNA recognition sequences. Cold Spring Harb. Symp. Quant. Biol. 49, 769–777 (1984).
Yun, T. & Vapnek, D. Electron microscopic analysis of bacteriophages P1, P1Cm, and P7. Determination of genome sizes, sequence homology, and location of antibiotic-resistance determinants. Virology 77, 376–385 (1977).
Iida, S. & Arber, W. Multiple physical differences in the genome structure of functionally related bacteriophages P1 and P7. Molec. Gen. Genet. 173, 249–261 (1979).
Keen, E. C. et al. Novel “superspreader” bacteriophages promote horizontal gene transfer by transformation. mBio 8, e02115–16 (2017).
Sugiura, C., Miyaue, S., Shibata, Y., Matsumoto, A. & Maeda, S. Bacteriophage P1vir-induced cell-to-cell plasmid transformation in Escherichia coli. AIMS Microbiol. 3, 784–797 (2017).
Kittleson, J. T., DeLoache, W., Cheng, H.-Y. & Anderson, J. C. Scalable plasmid transfer using engineered P1-based phagemids. ACS Synth. Biol. 1, 583–589 (2012).
Reid, C. J. et al. Porcine commensal Escherichia coli: a reservoir for class 1 integrons associated with IS26. Microb. Genom. 3, 1–13 (2017).
Partridge, S. R., Zong, Z. & Iredell, J. R. Recombination in IS26 and Tn2 in the evolution of multiresistance regions carrying bla CTX-M-15 on conjugative IncF plasmids from Escherichia coli. Antimicrob. Agents Chemother. 55, 4971–4978 (2011).
Hunt, M. et al. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 294 (2015).
Brettin, T. et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep. 5, 8365 (2015).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Partridge, S. R. & Tsafnat, G. Automated annotation of mobile antibiotic resistance in Gram-negative bacteria: the Multiple Antibiotic Resistance Annotator (MARA) and database. J. Antimicrob. Chemother. 73, 883–890 (2018).
Partridge, S. R. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol. Rev. 35, 820–855 (2011).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010 (2011).
Agyekum, A. et al. Predictability of phenotype in relation to common β-Lactam resistance mechanisms in Escherichia coli and Klebsiella pneumoniae. J. Clin. Microbiol. 54, 1243–1250 (2016).
Jacoby, G. A. & Han, P. Detection of extended-spectrum beta-lactamases in clinical isolates of Klebsiella pneumoniae and Escherichia coli. J. Clin. Microbiol. 34, 908–911 (1996).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Milne, I. et al. Using Tablet for visual exploration of second-generation sequencing data. Brief. Bioinformatics 14, 193–202 (2013).
Keen, N. T., Tamaki, S., Kobayashi, D. & Trollinger, D. Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene 70, 191–197 (1988).
He, S. et al. Insertion sequence IS26 reorganizes plasmids in clinically isolated multidrug-resistant bacteria by replicative transposition. mBio 6, e00762 (2015).
Iida, S. et al. Min DNA inversion enzyme of plasmid p15B of Escherichia coli 15T−: a new member of the Din family of site-specific recombinases. Mol. Microbiol. 4, 991–997 (1990).
Acknowledgements
The authors would like to acknowledge Alicia Arnott, Nathan Bachmann, Chayanika Biswas, Rajat Dhakal, Elena Martinez, Ranjeeta Menon, Rebecca Rockett, Rosemarie Sadsad, Verlaine Timms, Qinning Wang and Vitali Sintchenko at the Pathogen Genomics Unit, Centre for Infectious Disease and Microbiology – Public Health, Westmead Hospital, for their assistance with genome sequencing and bioinformatic analysis. PacBio sequencing was funded by a Research Grant from the Australian Society for Antimicrobials to SRP and JRI (pJIE250-3) and the Australian Research Council (Linkage grant LP150100912) to S.P.D (pSvP1 and pTZ20_1P). Dr. Toni Chapman, Dr. Graeme Eamens, Daniela Gaio, Linda Falconer and Shayne Fell are acknowledged for their management of and participation in the animal trial at the EMAI pig facility from which TZ20_1P originated. This project was partly funded by the Australian Centre for Genomic Epidemiological Microbiology (AusGEM), a collaborative partnership between the NSW Department of Primary Industries and the University of Technology Sydney. T.Z. is recipient of UTS International Research and UTS President’s Scholarships. C.V. is supported by a National Health and Medical Research Council grant (NHMRC GNT1107322).
Author information
Authors and Affiliations
Contributions
C.V. designed the study and planned all experiments, prepared sequencing samples (PacBio), performed bioinformatic comparisons and analysed all data, wrote the manuscript and prepared figures; T.Z. performed experiments, analyzed multidrug resistance regions, and participated in manuscript preparation; B.B. participated in performing transfer experiments; E.R.W. prepared sequencing samples (PacBio), and participated in bioinformatic analysis; J.I. contributed to experimental plan and edited manuscript; S.R.P. analysed plasmid insertions and multidrug resistance regions, wrote the manuscript and prepared figures; S.P.D. contributed to experimental plan design and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
S.R.P. co-developed the MARA annotation system, for which a patent application has been lodged with Dr. Guy Tsafnat (US patent application no. 16/1272,710), and acts as a consultant to Arc Bio, which has licensed the software as Galileo AMRTM. All other authors declare no potential conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Venturini, C., Zingali, T., Wyrsch, E.R. et al. Diversity of P1 phage-like elements in multidrug resistant Escherichia coli. Sci Rep 9, 18861 (2019). https://doi.org/10.1038/s41598-019-54895-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54895-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.