Abstract
The influence that Burkholderia cenocepacia adaptive evolution during long-term infection in cystic fibrosis (CF) patients has on cell wall morphology and mechanical properties is poorly understood despite their crucial role in cell physiology, persistent infection and pathogenesis. Cell wall morphology and physical properties of three B. cenocepacia isolates collected from a CF patient over a period of 3.5 years were compared using atomic force microscopy (AFM). These serial clonal variants include the first isolate retrieved from the patient and two late isolates obtained after three years of infection and before the patient’s death with cepacia syndrome. A consistent and progressive decrease of cell height and a cell shape evolution during infection, from the typical rods to morphology closer to cocci, were observed. The images of cells grown in biofilms showed an identical cell size reduction pattern. Additionally, the apparent elasticity modulus significantly decreases from the early isolate to the last clonal variant retrieved from the patient but the intermediary highly antibiotic resistant clonal isolate showed the highest elasticity values. Concerning the adhesion of bacteria surface to the AFM tip, the first isolate was found to adhere better than the late isolates whose lipopolysaccharide (LPS) structure loss the O-antigen (OAg) during CF infection. The OAg is known to influence Gram-negative bacteria adhesion and be an important factor in B. cenocepacia adaptation to chronic infection. Results reinforce the concept of the occurrence of phenotypic heterogeneity and adaptive evolution, also at the level of cell size, form, envelope topography and physical properties during long-term infection.
Similar content being viewed by others
Introduction
The Gram-negative opportunistic bacterial pathogens Pseudomonas aeruginosa and Burkholderia cepacia complex (Bcc) exhibit extensive genetic and phenotypic heterogeneity during persistent infection and evolution in the lungs of cystic fibrosis (CF) patients over the years1,2,3,4. The molecular mechanisms underlying adaptation to the lung and genotypic and phenotypic diversification have been intensively studied in the more prevalent CF pathogen P. aeruginosa3,4. However, Bcc lung infections in CF are highly feared because they are associated with poor prognosis and increased risk of death due to rapid lung function deterioration and, in certain cases to a necrotizing pneumonia, bacteraemia, and sepsis, (the cepacia syndrome)5,6,7,8.
During long-term lung infection in CF patients, P. aeruginosa and Bcc bacteria face multiple selective pressures in the highly challenging, fluctuating, and stressful environment of the patients´ airways, in particular due to antimicrobial therapy, the action of the host immune system and of other members of the microbiome and the decrease of oxygen availability as the result of lung function deterioration9,10. Under those stresses, several genetic changes accumulate in the initial infecting bacterial strain leading to phenotype and genotype heterogeneity. CF bacterial pathogens phenotypic diversification can be recognized in terms of colony morphology diversity11,12,13,14,15,16,17 and variation of clinically relevant phenotypes such as antibiotic resistance11,17,18,19,20, ability to form biofilms16,21,22,23,24, virulence potential14,25,26,27, among many others12,17,28,29,30,31,32. Remarkably, such phenotypic heterogeneity within human hosts has important clinical implications. For example, antimicrobial susceptibility diversity within the bacterial population isolated from an individual sputum sample may affect the treatment of life-threatening infections given that the results from antimicrobial testing carried out on single isolates randomly collected can be a poor predictor of the clinical outcome of antibiotic therapy7,18,19.
Bacterial cell envelope plays a central role in cell physiology and the alteration of surface properties can implicate the variation of phenotypes that play a crucial role in the pathogenesis of infectious diseases, such as antibiotic resistance and biofilm formation28,32,33. However, very few bacterial species have been on the focus of studies related to cell surface physical properties33,34,35 and information on the diversification and adaptive evolution at the level of Bcc bacteria cell wall mechanical properties during CF chronic lung infections is missing. In this context, over the last years atomic force microscopy (AFM) emerged as an essential tool for understanding the nanomechanics of live systems36,37,38. Hence, the objective of the present study was to obtain this knowledge by studying cell surface morphology and mapping the mechanical properties of Burkholderia cenocepacia clonal variants isolated from the lungs of a CF patient during long term infection using AFM. The B. cenocepacia isolates examined are from a collection of 11 serial clonal variants obtained from the same CF patient over a period of 3.5 years, from the onset of infection until the patient’s death11,39. The clonal variants tested were: IST439, the first isolate retrieved; IST4113, obtained three years later after an exacerbation with the patient hospitalization and treatment with intravenous therapy with gentamicin and ceftazidime and found to be highly resistant to different classes of antimicrobials; and IST4134, obtained 3 months later, just before the patient’s death with cepacia syndrome11,39,40. These isolates were picked at random from selective agar plates obtained in the major Portuguese CF Center at Hospital de Santa Maria during consultation routines. The clinical isolates examined are of high interest in the context of this study because they were previously characterized by phenotypic11, transcriptomic40, proteomic27,41 and metabolic profiling42. Results on the comparison of the virulence potential of these isolates using non-mammalian infection models and of their ability to modulate dendritic cell function are also available25,43. The two late variants were found to have lost the ability to produce the OAg molecule of the lipopolysaccharide44 present in the early isolates and to be more internalized by dendritic cells and show improved survival within dendritic cells when compared to the initial isolate43. Inflammatory cytokines were highly expressed in all the sequential clonal isolates but this pro-inflammatory trait was more pronounced in dendritic cells infected with the late variants compared with the isolate retrieved at the first stages of infection43.
Results of the present study, in which AFM cell wall morphology and mechanical properties of these three sequential B. cenocepacia clonal variants were studied, reinforce the concept of the occurrence of phenotypic variation and adaptive evolution also at the level of cell size, form, envelope topography and physical properties during long-term infection.
Results
B. cenocepacia morphology and surface roughness evolution during long-term CF lung infection
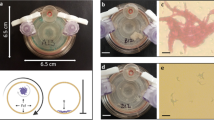
The cell morphology and topography of the three B. cenocepacia clonal variants were examined using AFM in both planktonic and biofilm forms. The images for individual cells were obtained in air and in liquid environment. Concerning cell topography in air (Fig. 1), the easiest way to clearly visualize individual cells, the three clonal variants examined in their planktonic form exhibit the porous network architecture of the cell wall previously reported by others45,46. However, late variants IST4113 and IST4134 show features, characterized by string-like formations on the surface of the bacteria, not found in the early isolate IST439. Specifically, the last isolate retrieved from the patient, IST4134, displays longer and well-organized string-like structures that span the entire length of the cell while IST4113 exhibits shorter and less organized structures (Fig. 1(b,c)). However, the same structures could not be observed in the biofilm images Fig. 1(a). The surface roughness of the cells examined in air in their planktonic form was assessed by defining longitudinal cross sections along the cell surface, as shown in Fig. 1(b,f). The first isolate, IST439, was found to be smoother, with average roughness (root mean square of the cross sections) of 0.9 ± 0.1 nm. The late variants showed an average roughness of 1.0 ± 0.1 nm and 2.0 ± 0.1 nm respectively. Alternatively, we have also measured the roughness by taking the root mean square over an area defined on the top of the cell after subtracting the cell envelope. The resulting values for the first, second and third isolate were respectively 1.7 nm, 2.0 nm and 2.5 nm with 0.5 nm standard deviation. We find the first method less prone to errors because it is easier to separate the cell roughness from the cell contour/envelope - nonetheless, both methodologies indicate the same trend. The surface roughness from the images obtained in liquid was impossible to visualize due to the poor resolution of the images seemingly caused by some mobility of either the cell or the cell surface. We have used both contact and tapping mode but the results were similar.
Cell topography and surface roughness. AFM images of the studied clonal variants (i - IST439, ii - IST4113 and iii - IST4134) show (a) topography of biofilm; (b) topography of planktonic and (c) corresponding deflection images in air; (d) topography of planktonic and (e) corresponding deflection in liquid samples; (f) representative roughness profiles (marked by the black lines) for the same variants. The late variants exhibit distinct string-like surface features, whereas both the images and roughness profile of the IST439 variant show a smoother surface.
Concerning cell dimensions measurements, the late variants, IST4113 and IST4134, were found to be significantly smaller in length than the early isolate IST439, but no significant differences were found between the lengths of late variants, as shown on Fig. 2(i). Similar significant differences of the cell length were observed both in air and liquid, however the absolute values registered in liquid environment were smaller than those in air (Fig. 2(i)). Additionally, either in air or in liquid environment, the height of the cells also decreases from the first to the last clonal isolate by about 30% (Fig. 2(iv)). Overall, the observed trend points towards an evolution of the cell shape from rod to a more rounded shape, with the ratio width/length increasing from around 0.52/0.62 (air/liquid) for the early isolate to about 0.68/0.75 (air/liquid) and 0.65/0.86 (air/liquid) for the second and third isolates, respectively (Fig. 2(iii)). The AFM images of the biofilm (Fig. 1(a)) show cell dimensions consistent with the reduction of the cell size and increase of the ratio width/length (Fig. 2(iii, panel b)) observed for the individual cells. However, due to the fact that cells are tightly packed it is more difficult to accurately determine cell dimensions in biofilm.
Cell morphology. Distribution of the cell dimensions (i - length, ii - width, iii - width/length ratio and iv - height) measured with AFM for N individual cells of the studied clonal variants (blue circle - IST439, red square - IST4113 and green triangle - IST4134), for (a) planktonic and (b) biofilm growth conditions both measured in air and for the (c) planktonic form measured in liquid environment. An increase of the width/length ratio and a decrease of cell height, as well as the evolution of the cell shape during long-term infection from a rod-like to a more cocci-like morphology can be observed. The results of the Mann-Whitney u-test (*P < 0.05, **P ≤ 0.005, ****P ≤ 0.0005, ns not significant) are indicated.
Surface and mechanical properties evolution during long-term CF lung infection
Cell elasticity was examined at selected points (inset of Fig. 3(a–c)) along the cell surface and the resulting force-distance curves were compared with the Sneddon contact mechanics model (Fig. 3(a–c)), as described in the Methods section. All these measurements were performed in liquid environment because the measurement of the mechanical properties of dried cells is questionable47,48,49,50,51. In fact, when measurements were made in air, the apparent Young’s modulus of the cell surface was about 40 times larger.
Elasticity and adhesion studied by AFM in liquid samples. (a–c) Example of indentation curves and respective fits using Sneddon model for the first, second and third clonal variants with insets showing the respective 3D maps and the selection of points at the top of the cell; (d,e) distribution of young modulus and adhesion forces respectively, for each isolate; (f) a retract curve obtained for the first isolate illustrating the adhesion measurement. The results of the Mann-Whitney u-test (*P < 0.05, ***P ≤ 0.001, ****P ≤ 0.0005, ns not significant) are indicated.
Concerning the elasticity, and in particular the apparent Young’s modulus of the cell surface measured in liquid environment, the values show a significant decrease from the early isolate to the late variant (Fig. 3(d)), but the highly antibiotic resistant clonal variant IST411311 exhibited the highest values. Concerning the adhesion of the bacteria surface to the Si3N4 AFM tip (Fig. 3(e)), the first isolate was found to adhere better than the late isolates whose lipopolysaccharide (LPS) structure loss, during CF infection, the O-antigen (OAg) present in the early isolate44. No significant differences were found between the adhesion of the two late isolates missing the OAg, Fig. 3(e).
Growth curves of the B. cenocepacia clonal variants under aerobic or microaerophilic conditions
The growth curves of the three clonal variants examined were compared in the same Lysogeny broth (LB) medium under aerobic and microaerophilic conditions, at 37 °C (Fig. 4). The general conclusion is that under aerobic conditions, the growth performance of the first isolate is slightly better (higher specific growth rate and higher final biomass concentration attained) than the late isolates, with the highly antibiotic resistant intermediary isolate (IST4113) exhibiting the slowest and less efficient growth. This behaviour contrasts with the growth performance observed under microaerophilic conditions which are conditions closer to those expected to occur in the CF patient lung, especially at late stages of disease progression and very low values of Forced expiratory volume in one second (FEV1)52. Although the reported differences are small, the early isolate consistently exhibited, under oxygen limitation, the lowest specific growth rate while the last isolate showed the more rapid growth and efficient biomass production suggesting that the late isolates are better adapted to the CF lung. Moreover, under microaerophilic conditions, the growth curves of the 3 isolates exhibit a pattern consistent with diauxic growth in the complex LB medium, a behaviour observed before (unpublished data).
Growth under oxygen limitation. Evolution of the optical density values during the incubation time in the same medium (LB) for the three clonal variants examined under (a) aerobic condition and (b), microaerophilic conditions. Experimental values and error bars represent the mean and the estimated standard deviation, respectively, for three independent growth experiments. These results indicate that the late clonal variant grows better under microaerophilic conditions whereas the early IST439 grows better under aerobic conditions.
Biofilm growth of B. cenocepacia clonal variants
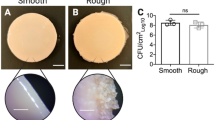
The quantification of biofilms in terms of biomass using crystal violet staining after 4 and 6 hours of incubation is consistent with the specific growth rates and final biomass attained by the three clonal variants, grown under microaerophilic conditions in the planktonic lifestyle (Figs. 4 and 5). In fact, the late isolates produce more rapidly immature biofilms of higher biomass, formed after 4–6 hours. However, the relative biomass of the mature biofilms formed after 24–48 h of incubation is consistent with the level of exopolysaccharide produced by the three variants, as reported before for the same isolates/growth medium11, being maximal for the intermediary isolate.
Biofilm formation. Quantification of the biofilm formed at different incubation times, based on the OD600nm values of crystal violet-stained biomass. Bar height and error bars represent the mean and standard deviation obtained from three independent growth experiments with 7 measurements each. The Mann-Whitney u-test (*P < 0.05, ***P ≤ 0.001, ****P ≤ 0.0005, ns not significant) values are indicated.
Discussion
During long term infection, the genetic adaptation of bacteria of the Burkholderia cepacia complex (Bcc) to the challenges of multiple selective pressures occurring in the cystic fibrosis (CF) airways is known to take place1,2,53,54,55. The present study provides the first insights into the adaptive evolution of these bacteria at the level of cell size, form, envelope topography and physical properties during long-term infection. However, it cannot be guaranteed that the properties reported here are identical to those exhibited by bacteria grown in the lungs and measured in their native environment. Given the highly relevant information gathered over the years on the three sequential B. cenocepacia variants examined here, it is possible to speculate on how the variation in surface properties can impact phenotypes of clinical relevance in the pathogenesis of infectious diseases, such as adhesion, resistance to antibiotics, biofilm formation and growth efficiency in the CF environment.
Former studies have shown that in B. cenocepacia the O-antigen (OAg) of the lipopolysaccharide (LPS), occurring in the outermost layer of the cell, is lost or modified during persistent infection of the lungs32,44. The loss or modification of the OAg appears to play an important role during the infection process, in particular in the colonization step (adherence) and ability to overcome host defence mechanisms32,56,57. Recently, we have shown that the tendency of the most prevalent and feared species B. cenocepacia and B. multivorans to lose the OAg during chronic infection is higher than the one of the rarely found B. cepacia and B. contaminants that keep the OAg even during decades of infection58. Moreover, B. cenocepacia recA lineage IIIA strains, as it is the case of the isolates examined in the present study known to lead to particularly destructive infections, exhibit the most frequent OAg loss, compared with lineage IIIB58. Concerning the clonal isolates tested in the present study, the OAg is only present in the early isolate IST43944. Mutations within the OAg cluster of the serial isolates examined were identified but none of them could definitely be associated to OAg loss44. In the present study, we found that the ability of the earlier isolate to adhere to the AFM Si3N4 tip, when assessed in liquid environment, was significantly higher than the estimated ability of the late variants, lacking the OAg whose adhesion values were similar. These results are consistent with the concept that the variability in the LPS OAg affects bacterial adherence and colonisation and the ability to evade the host’s defence mechanisms being an important factor in B. cenocepacia adaptation to chronic infection32,44,58,59,60.
In agreement with the characterization of Gram-negative sacculus61,62, images in air have clearly revealed tube-like features not regularly spaced mostly in the plane of the sacculus and roughly perpendicular to the long axis of the cell, for the two late isolates, but not for the early isolate. These formations are short and organized randomly in IST4113 cell surface but well developed in IST4134. These features do not seem to be associated to the division septum and its equatorial rings, as previously reported in studies in which the sacculus of Escherichia coli and Bacillus subtilis63,64, of Streptococci and Enterococci65,66 and Staphylococcus aureus67 was analysed. With very few exceptions, the chemical and biological bacterial components studied that contribute to cell mechanics are related with the peptidoglycan layer of the cell envelope and changes in its structure33,34,68. Apparently, the peptidoglycan is made up of circumferential oriented bands of material interspersed with a more porous network45. Peptidoglycan is the largest component of the bacterial cell wall determining the shape and preserving its integrity62. Its elastic nature helps withstand stretching forces caused by bacterial turgor pressure. The reported bands were proposed to define regions with different availability for insertion of new peptidoglycan45,46. Interestingly, indentation tests performed in liquid show that the surface elasticity modulus, decreases significantly from the early isolate to the last clonal variant. However, the highly antibiotic resistant intermediary isolate IST411311 expressed maximal values. Higher rigidity and increased elasticity was recently reported to be associated with a lower outer membrane permeability which may lead to the reduction of antibiotic diffusion into the cells69.
The suggested remodelling of cell surface of the three clonal variants examined in this study during long term infection was already anticipated based on the results of the comparison of genomic expression of these same clonal variants using transcriptomic and quantitative proteomic analyses27,40,41. These studies have shown differences in the level of expression of genes/proteins involved in the biogenesis of cell envelope and outer membrane in the three variants, among the several hundred of genes found to be differentially transcribed in the late isolates compared to the early isolate. These genome-wide expression results reflect a marked reprogramming of genomic expression at different levels27,40,41, including the alteration of bacterial cell surface that contributes to the intrinsic and acquired resistance of Bcc bacteria to antibiotics. Remarkably, it was found that the late isolates are significantly more resistant to a wide range of antibiotics, with isolate IST4113 displaying the higher resistance levels7,11,27,40. Recent genomic studies strengthened the concept that cell wall remodelling relates with the alteration of bacterial mechanical properties33,35. For example, E. coli mutants deleted for genes encoding proteins associated with cell-wall synthesis exhibit different stiffness defects35 and the accumulation of the peptidoglycan D-Alanine residues is tightly regulated in P. aeruginosa since their accumulation reduces peptidoglycan cross-linking and cell stiffness33.
In this study, we clearly observed a consistent and progressive pattern of decrease of the height and the increase of the width/length ratio of B. cenocepacia cells during long term infection, both in air and in liquid environments. Although the B. cenocepacia clonal isolates examined may not be representative of the expected population heterogeneity present at each isolation time in the CF lung, the consistency of the pattern strongly suggests that B. cenocepacia underwent convergent evolution towards the minimization of bacterial size during infection. Moreover, besides the decrease of the size of B. cenocepacia cells during infection, the bacterium underwent a cell shape evolution from the typical rod form of the species to a cell morphology closer to the spherical form of cocci. The referred pattern was observed for both the planktonic and biofilm growth mode. This same pattern was described before for two nasopharyngeal bacterial pathogens during adaptation to human mucosa and the authors hypothesized that this transition was selected to reduce the cell surface sensitivity to immune attacks given that the ratio surface/volume is smaller than that of bacilli70. Consistent with this hypothesis, several studies have shown the relevance of cell surface size when bacterial cells are facing immune attacks, small microbial size allowing a more efficient evasion of host defences71,72,73. Microbial cell size appears to be an important pathogenesis factor and minimization of bacterial size was demonstrated to be a mechanism used for example by Streptococcus pneumoniae to circumvent complement-mediated killing by the host73. The cell shape modification from rods to cocci-like form has been very-recently reported to occur in response to antibiotic stress in multi-drug resistant E. coli74 and has been genetically and biochemically demonstrated to occur during the prolonged antibiotic selective pressure that is extensively and aggressively administered to CF patients chronically infected with P. aeruginosa bacteria71. Remarkably, in the particular case of the CF patient from whom the examined isolates were obtained, the clinical situation was significantly deteriorated being hospitalized and submitted to intensive intravenous antibiotic therapy immediately before IST4113 isolation11,39.
Many Gram-negative pathogens alter their characteristic rod-shaped forms to smaller coccoid-like forms after incubation for days to weeks in fresh or salt water and in nutrient poor environments bacteria tend to be much smaller in size than those grown in laboratory cultures71. Free-living cells tend to be smaller in nutrient poor environments because the acquisition of nutrients relies on diffusion and capture of molecules at the surface of the cell. In E. coli, cell size was found to be reduced by a factor of 3 in response to nutrient starvation75, E. coli adjusting size and growing larger and faster in nutrient-rich media compared with nutrient-poor media75,76,77,78. During the continuous and rapid deterioration of lung function, as the disease progresses, the oxygen concentration levels in the CF airways suffer a marked decrease3,26. Responses of P. aeruginosa to oxygen limitation indicate that this species growth in the CF lung is by aerobic respiration79 and the same metabolism was proposed for B. cenocepacia11,40,80. According to the hospital records, when the early isolate IST439 was obtained, the FEV1 value (the forced expiratory value in the first second) was 22% but no further values of FEV1 are available due to the subsequent severe deterioration of pulmonary function11,52. The fact that the late variants appear to grow more efficiently under microaerophilic conditions, while the early isolate exhibits the most efficient growth when in aerobiosis, supports the hypothesis of an adaptation of the late variants to severe oxygen depletion. The more adapted growth of the late isolates to oxygen-limitation is also consistent with the biomass increase of the biofilms resulting from growth during the first hours (4 and 6 hours) following initial bacteria adhesion. After 12–48 hours of growth, with the maturation of the biofilms formed, other mechanisms take over as it is the case of exopolysaccharide (EPS) production capacity11,23,24, the biomass concentration of the biofilms formed correlating well with the levels of EPS produced by each clonal variant11. Differences observed between IST4113 and IST4134 growth curves under both microaerophilic and aerobic conditions are likely the result of IST113 resistance to multiple antibiotics, resistance to which a fitness cost is associated81,82.
In summary, independently of the selective pressures that drive B. cenocepacia cell size and shape alterations during chronic infection of the lungs, it is likely that the adaptive evolution registered in this study may lead to a better performance under limiting oxygen concentration, to more efficient nutrient acquisition and to evasion of the host complement deposition, favouring persistent infection and pathogenesis. The positive correlation observed between cell shape change and elasticity modulus indicates that elasticity of the cell wall may play a key role in this adaptation process. Results from former genome wide expression analyses and extensive phenotyping of the isolates here examined have provided clues that strongly suggest a genetic adaptation to the challenges exerted by the immune system, antimicrobial therapy and nutrient and oxygen availability27,40,41. The shape and size evolution observed in this study is considered part of such metabolic reprogramming that leads to B. cenocepacia persistence in the CF lung27,40,41. Understanding the underlying adaptation mechanisms is essential also for an improved therapeutic outcome of long term infections in CF patients.
Methods
Bacterial strains and growth conditions
The three Burkholderia cenocepacia clonal variants examined in this study (IST439, IST4113 and IST4134) were recovered, as part of the hospital routine, from the sputum of a CF patient under surveillance at the major Portuguese CF Center in the Hospital de Santa Maria, Centro Hospitalar Lisboa Norte (CHLN) EPE, from 1999 to 200211,25,39. Studies involving these isolates were approved by CHLN ´ ethics committee and the anonymity of the patient was preserved. Informed consent was also obtained from all participants and/or their legal guardians. All the methods were performed in accordance with the relevant guidelines and regulations. Bacterial cultures are stored at −80 °C in 1:1 (v/v) glycerol. Bacterial growth was carried out in Lysogeny Broth, Lennox (LB; Conda, Pronadisa), at 37 °C and 250 rpm, or in LB agar plates obtained by supplementation of LB with 2% agar (Iberagar, Portugal). LB medium at 37 °C was also used in biofilm experiments.
Preparation of the AFM samples
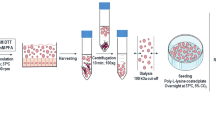
Bacterial cells used for AFM analysis were deposited onto the gelatin coated mica for the observations and measurements done in liquid environment and onto freshly cleaved mica surfaces for the observations done in air. B. cenocepacia isolates were cultured overnight in LB medium, at 37 °C with shaking at 250 rpm, and then sub-cultured until mid-exponential phase. Bacterial planktonic cells in suspension were collected by centrifugation and washed three times with phosphate buffer saline (PBS). For observations and measurements in liquid environment, the gelatin-coated mica was prepared and the bacterial immobilization was done as described before69,83,84. Two types of gelatin with different concentrations (0.25%, 0.5% and 1% (w/v)), porcine gelatin Sigma G-6144 and G-2625 and bovine gelatin Sigma G-9382, were tested83. Briefly, a gelatin solution was prepared by dissolving 0.25 g, 0.5 g or 1 g gelatin in 100 ml of deionized water at 90 °C and cooled to 60–70 °C prior to vertically dipping several discs of the freshly cleaved mica into the solution. Following optimization, gelatin G-6144 was found to allow the best immobilization effectiveness and used thereafter. The gelatin-G-6144-coated mica surfaces were supported on edge on a paper towel and then air dried overnight. 20–40 μl of the bacterial suspension in PBS (108 CFU/ml) was applied onto a gelatin-coated mica surface after being sonicated in ultrasonic bath (40 kHz, 19 W – Branson, Model 200, NL) for ~5 min84. The sample was allowed to rest for 10–20 min before it was rinsed in PBS and imaged in the liquid cell of the AFM.
For the images/observations taken in air, the deposition of bacteria on the mica surfaces was carried out as described before85,86 and used for other Gram-negative bacteria45,48,87, with few modifications. In brief and during the optimization of the protocol, PBS and deionized water were tested to prepare the bacterial suspension aliquots prior to deposition on mica. Given that the samples prepared with PBS formed aggregates at the freshly cleaved-mica surface, whereas those prepared with deionized water did not, 10 μl of those aliquots in ddH2O, for observation done in air, with a final bacterial concentration of 108 CFU/ml, were immobilized onto freshly cleaved mica surfaces and left to rest for 15–20 min. The mica surfaces were rinsed twice with deionized water to detach the weakly adherent and the non-adherent cells and allowed to dry before AFM analysis for another 15–20 min. The biofilm samples were prepared for AFM observation as described before88 with few modifications. The microtiter plates were incubated without shaking at 37 °C for 12 hours. The unattached planktonic bacteria were washed twice with sterile saline solution by pipetting in and out. The remaining biofilms were fixed by 10% formalin in PBS for 10 min. Next, the plates were inverted to remove all the fixation solution and the fixed biofilms were washed twice with sterile saline solution by pipetting in and out. Finally, the biofilm was resuspended in ddH2O, and 10 µl of the suspended biofilm was deposited into the freshly cleaved micas and air-dried for AFM imaging.
AFM observations and measurements
Cell samples were analysed using a PicoSPM LE system of Molecular Imaging in a liquid cell containing PBS and in air at room temperature. Bruker MLCT-F microlevers with nominal cantilever stiffness of 0.6 N/m and nominal tip radius of 20 nm were used for all experiments. Images obtained in air were taken in contact mode whereas in liquid environment both contact and tapping modes were used depending on which proved best. To measure cell surface roughness 2.5 × 2.5 μm2 (approximately) images with 512 × 512 pixels were obtained. Twelve representative bacteria of each isolate were selected for roughness measurements.
For nanomechanics and adhesion measurements in liquid environment force spectroscopy maps, consisting of 32 × 32 approach/retract force-distance (FD) curves, were obtained over an area slightly larger than the cell footprint. The maximum cantilever deflection was set constant in all experiments, yielding a maximum applied force of roughly 15 nN. The tip-sample approach speed was also set constant, to 0.4 μm/s. To reduce bias due to different cantilevers being used on different populations, each cantilever was used to measure 2–3 bacteria of each isolate population and the order in which the different isolates was measured was randomized89,90. In total, about 40 bacteria of each population were measured in liquid environment and 14 cantilevers were used. Nanomechanical analysis was performed using a custom-made software. For each grid, we selected only curves obtained at the top of the bacteria (above 85% of the total height of the bacteria), as shown in the inset of Fig. 3(a–c). The contact part of the approach curves was analysed according to Sneddon contact model36 that establishes a relationship between load and indentation and from which it is possible to extract the reduced Young’s modulus Fig. 3(d). We have estimated the adhesion force Fig. 3(e) from the minimum of the retract part of the curves as illustrated in Fig. 3(f). For each cell, all curves obtained at the top of the cell were analyzed and the median was kept, then for each isolate sample we rejected values further away from 3 standard deviations. To determine the apparent Young’s modulus of the cell we assumed a non-deformable tip and a Poisson’s ratio of 0.536,91. We assumed nominal values for the tip radius and used Sader method to calibrate the cantilever spring constant92.
Growth curves
The growth curves of the three clonal variants examined under aerobic and microaerophilic conditions were monitored by measuring culture optical density at 640 nm (OD640). Cells were grown in LB medium at 37 °C in shaking flasks (100 ml with 30 ml of liquid medium) in an orbital shaker at 250 rpm (for aerobic growth) or standing in a microaerophilic atmosphere, containing 5–8% oxygen and 12–15% carbon dioxide, generated in sealed jars using the GENbox microaerator (bioMérieux, Marcy L’Etoile, France). Results are from three independent growth experiments.
Biofilm formation assays
Biofilm formation assays were based on a described methodology11,93. Overnight liquid cultures of each CF isolate were transferred to LB medium and grown at 30 °C with orbital agitation until the mid-exponential phase was reached. The cultures were subsequently diluted to a standardized culture OD640 of 0.5, and 20 μl of this cell suspension was used to inoculate the wells of a 96-well polystyrene microtiter plate (Greiner Bio-One) containing 180 μl of LB medium. Wells containing sterile growth medium were used as negative controls. Plates were incubated at 37 °C from 4 to 48 h without agitation. For biofilm quantification, the culture media and unattached bacterial cells were removed from the wells by careful rinsing with water (three times, 200 μl for each rinse). Adherent bacteria were stained with 200 μl of a 1% crystal violet solution for 15 min at room temperature (50 ml of the solution was prepared by adding 1% [wt/vol] crystal violet in 10 ml of 95% ethanol to 40 ml of water containing 0.4 g of ammonium oxalate). After three gentle rinses with 200 μl of water each time, the dye associated with the attached cells was solubilized in 200 μl of 95% ethanol and the biofilm was quantified by measuring the absorbance of the solution at 600 nm (A600nm) in a microplate reader.
Statistics
A non-parametric Mann-Whitney u-test was used to determine statistical significance of the observed variations (GraphPad Prism 7; GraphPad Software, CA). P ≤ 0.05 was considered statistically significant.
Data availability
All datasets generated for this study are included in the manuscript.
References
Lee, A. H. et al. Phenotypic diversity and genotypic flexibility of Burkholderia cenocepacia during long-term chronic infection of cystic fibrosis lungs. Genome Research 27, 650–662, https://doi.org/10.1101/gr.213363.116 (2017).
Lieberman, T. D. et al. Genetic variation of a bacterial pathogen within individuals with cystic fibrosis provides a record of selective pressures. Nature Genetics 46, 82–87, https://doi.org/10.1038/ng.2848 (2014).
Winstanley, C. et al. Pseudomonas aeruginosa Evolutionary Adaptation and Diversification in Cystic Fibrosis Chronic Lung Infections. Trends in Microbiology 24, 327–337, https://doi.org/10.1016/j.tim.2016.01.008 (2016).
Govan, J. R. et al. Evolving epidemiology of Pseudomonas aeruginosa and the Burkholderia cepacia complex in cystic fibrosis lung infection. Future Microbiology 2, 153–164, https://doi.org/10.2217/17460913.2.2.153 (2007).
Mahenthiralingam, E. et al. The multifarious, multireplicon Burkholderia cepacia complex. Nature Reviews Microbiology 3, 144–156, https://doi.org/10.1038/nrmicro1085 (2005).
Drevinek, P. & Mahenthiralingam, E. Burkholderia cenocepacia in cystic fibrosis: epidemiology and molecular mechanisms of virulence. Clinical Microbiology and Infection 16, 821–830, https://doi.org/10.1111/j.1469-0691.2010.03237.x (2010).
Coutinho, C. P. et al. Long-term colonization of the cystic fibrosis lung by Burkholderia cepacia complex bacteria: epidemiology, clonal variation, and genome-wide expression alterations. Frontiers in Cellular and Infection. Microbiology 1, 12, https://doi.org/10.3389/fcimb.2011.00012 (2011).
Jones, A. M. et al. Burkholderia cenocepacia and Burkholderia multivorans: influence on survival in cystic fibrosis. Thorax 59, 948–951, https://doi.org/10.1136/thx.2003.017210 (2004).
Cullen, L. & McClean, S. Bacterial Adaptation during Chronic Respiratory Infections. Pathogens 4, 66–89, https://doi.org/10.3390/pathogens4010066 (2015).
Döring, G. et al. Differential adaptation of microbial pathogens to airways of patients with cystic fibrosis and chronic obstructive pulmonary disease. FEMS Microbiology Reviews 35, 124–146, https://doi.org/10.1111/j.1574-6976.2010.00237.x (2011).
Coutinho, C. P. et al. Burkholderia cenocepacia phenotypic clonal variation during a 3.5-year colonization in the lungs of a cystic fibrosis patient. Infection and Immunity 79, 2950–2960, https://doi.org/10.1128/IAI.01366-10 (2011).
Silva, I. N. et al. Mucoid morphotype variation of Burkholderia multivorans during chronic cystic fibrosis lung infection is correlated with changes in metabolism, motility, biofilm formation and virulence. Microbiology 157, 3124–3137, https://doi.org/10.1099/mic.0.050989-0 (2011).
Zlosnik, J. E. & Speert, D. P. The role of mucoidy in virulence of bacteria from the Burkholderia cepacia complex: a systematic proteomic and transcriptomic analysis. The Journal of Infectious Diseases 202, 770–781, https://doi.org/10.1086/655663 (2010).
Zlosnik, J. E. et al. Mucoid and nonmucoid Burkholderia cepacia complex bacteria in cystic fibrosis infections. American Journal of Respiratory and Critical Care Medicine 183, 67–72, https://doi.org/10.1164/rccm.201002-0203OC (2011).
Deretic, V. et al. Conversion of Pseudomonas aeruginosa to mucoidy in cystic fibrosis: environmental stress and regulation of bacterial virulence by alternative sigma factors. Journal of Bacteriology 176, 2773–2780 (1994).
Drenkard, E. & Ausubel, F. M. Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature 416, 740–743, https://doi.org/10.1038/416740a (2002).
Moreira, A. S. et al. Burkholderia dolosa phenotypic variation during the decline in lung function of a cystic fibrosis patient during 5.5 years of chronic colonization. Journal of Medical Microbiology 63, 594–601, https://doi.org/10.1099/jmm.0.069849-0 (2014).
Rhodes, K. A. & Schweizer, H. P. Antibiotic resistance in Burkholderia species. Drug Resistance Updates 28, 82–90, https://doi.org/10.1016/j.drup.2016.07.003 (2016).
Lopez-Causape, C. et al. The problems of antibiotic resistance in cystic fibrosis and solutions. Expert Review of Respiratory Medicine 9, 73–88, https://doi.org/10.1586/17476348.2015.995640 (2015).
Leitao, J. H. et al. Variation of the antimicrobial susceptibility profiles of Burkholderia cepacia complex clonal isolates obtained from chronically infected cystic fibrosis patients: a five-year survey in the major Portuguese treatment center. European Journal of Clinical Microbiology & Infectious Diseases 27, 1101–1111, https://doi.org/10.1007/s10096-008-0552-0 (2008).
Sriramulu, D. D. et al. Microcolony formation: a novel biofilm model of Pseudomonas aeruginosa for the cystic fibrosis lung. Journal of Medical Microbiology 54, 667–676, https://doi.org/10.1099/jmm.0.45969-0 (2005).
Lee, B. et al. Heterogeneity of biofilms formed by nonmucoid Pseudomonas aeruginosa isolates from patients with cystic fibrosis. Journal of Clinical Microbiology 43, 5247–5255, https://doi.org/10.1128/JCM.43.10.5247-5255.2005 (2005).
Valentini, M. et al. Lifestyle transitions and adaptive pathogenesis of Pseudomonas aeruginosa. Current Opinion in Microbiology 41, 15–20, https://doi.org/10.1016/j.mib.2017.11.006 (2018).
Fazli, M. et al. Regulation of biofilm formation in Pseudomonas and Burkholderia species. Environmental Microbiology 16, 1961–1981, https://doi.org/10.1111/1462-2920.12448 (2014).
Moreira, A. S. et al. Variation of Burkholderia cenocepacia virulence potential during cystic fibrosis chronic lung infection. Virulence 8, 782–796, https://doi.org/10.1080/21505594.2016.1237334 (2017).
Lorè, N. I. et al. Cystic fibrosis-niche adaptation of Pseudomonas aeruginosa reduces virulence in multiple infection hosts. PLoS One 7, e35648, https://doi.org/10.1371/journal.pone.0035648 (2012).
Madeira, A. et al. Proteomic profiling of Burkholderia cenocepacia clonal isolates with different virulence potential retrieved from a cystic fibrosis patient during chronic lung infection. PLoS One 8, e83065, https://doi.org/10.1371/journal.pone.0083065 (2013).
Hill, P. J. et al. Modifications of Pseudomonas aeruginosa cell envelope in the cystic fibrosis airway alters interactions with immune cells. Scientific Reports 7, 4761, https://doi.org/10.1038/s41598-017-05253-9 (2017).
Sousa, S. A. et al. Burkholderia cepacia Complex: Emerging Multihost Pathogens Equipped with a Wide Range of Virulence Factors and Determinants. International Journal of Microbiology 2011, 1–9, https://doi.org/10.1155/2011/607575 (2011).
Warren, A. E. et al. Genotypic and phenotypic variation in Pseudomonas aeruginosa reveals signatures of secondary infection and mutator activity in certain cystic fibrosis patients with chronic lung infections. Infection and Immunity 79, 4802–4818, https://doi.org/10.1128/IAI.05282-11 (2011).
Mowat, E. et al. Pseudomonas aeruginosa population diversity and turnover in cystic fibrosis chronic infections. Ajrccm - American Journal of Respiratory and Critical Care Medicine 183, 1674–1679, https://doi.org/10.1164/rccm.201009-1430OC (2011).
Maldonado, R. F. et al. Lipopolysaccharide modification in Gram-negative bacteria during chronic infection. FEMS Microbiology Reviews 40, 480–493, https://doi.org/10.1093/femsre/fuw007 (2016).
Trivedi, R. R. et al. Mechanical Genomic Studies Reveal the Role of d-Alanine Metabolism in Pseudomonas aeruginosa Cell Stiffness. MBio 9, https://doi.org/10.1128/mBio.01340-18 (2018).
Auer, G. K. & Weibel, D. B. Bacterial Cell Mechanics. Biochemistry 56, 3710–3724, https://doi.org/10.1021/acs.biochem.7b00346 (2017).
Auer, G. K. et al. Mechanical Genomics Identifies Diverse Modulators of Bacterial Cell Stiffness. Cell Systems 2, 402–411, https://doi.org/10.1016/j.cels.2016.05.006 (2016).
Touhami, A. et al. Nanoscale mapping of the elasticity of microbial cells by atomic force microscopy. Langmuir 19, 4539–4543, https://doi.org/10.1021/la034136x (2003).
Muller, D. J. et al. Force probing surfaces of living cells to molecular resolution. Nature Chemical Biology 5, 383–390, https://doi.org/10.1038/nchembio.181 (2009).
Costa, L. et al. Spectroscopic investigation of local mechanical impedance of living cells. PLoS One 9, e101687, https://doi.org/10.1371/journal.pone.0101687 (2014).
Cunha, M. V. et al. Molecular analysis of Burkholderia cepacia complex isolates from a Portuguese cystic fibrosis center: a 7-year study. Journal of Clinical Microbiology 41, 4113–4120, https://doi.org/10.1128/JCM.41.9.4113-4120.2003 (2003).
Mira, N. P. et al. Genomic expression analysis reveals strategies of Burkholderia cenocepacia to adapt to cystic fibrosis patients’ airways and antimicrobial therapy. PLoS One 6, e28831, https://doi.org/10.1371/journal.pone.0028831 (2011).
Madeira, A. et al. Quantitative proteomics (2-D DIGE) reveals molecular strategies employed by Burkholderia cenocepacia to adapt to the airways of cystic fibrosis patients under antimicrobial therapy. Proteomics 11, 1313–1328, https://doi.org/10.1002/pmic.201000457 (2011).
Moreira, A. S. et al. (1)H-NMR-Based Endometabolome Profiles of Burkholderia cenocepacia Clonal Variants Retrieved from a Cystic Fibrosis Patient during Chronic. Infection. Frontiers in Microbiology 7, 2024, https://doi.org/10.3389/fmicb.2016.02024 (2016).
Cabral, M. et al. Using dendritic cells to evaluate how Burkholderia cenocepacia clonal isolates from a chronically infected cystic fibrosis patient subvert immune functions. Medical Microbiology and Immunology 206, 111–123, https://doi.org/10.1007/s00430-016-0488-4 (2017).
Hassan, A. A. et al. Structure of O-Antigen and Hybrid Biosynthetic Locus in Burkholderia cenocepacia Clonal Variants Recovered from a Cystic Fibrosis Patient. Frontiers in Microbiology 8, 1027, https://doi.org/10.3389/fmicb.2017.01027 (2017).
Turner, R. D. et al. Cell wall elongation mode in Gram-negative bacteria is determined by peptidoglycan architecture. Nature Communication 4, 1496, https://doi.org/10.1038/ncomms2503 (2013).
Turner, R. D. et al. Atomic Force Microscopy Analysis of Bacterial Cell Wall Peptidoglycan Architecture. Methods in Molecular Biology 1440, 3–9, https://doi.org/10.1007/978-1-4939-3676-2_1 (2016).
Cerf, A. et al. Nanomechanical properties of dead or alive single-patterned bacteria. Langmuir 25, 5731–5736, https://doi.org/10.1021/la9004642 (2009).
Eaton, P. et al. Atomic force microscopy study of the antibacterial effects of chitosans on Escherichia coli and Staphylococcus aureus. Ultramicroscopy 108, 1128–1134, https://doi.org/10.1016/j.ultramic.2008.04.015 (2008).
Deng, Y. et al. Direct measurement of cell wall stress stiffening and turgor pressure in live bacterial cells. Physical Review Letters 107, 158101, https://doi.org/10.1103/PhysRevLett.107.158101 (2011).
Elbourne, A. et al. Bacterial-nanostructure interactions: The role of cell elasticity and adhesion forces. Journal of Colloid and Interface Science 546, 192–210, https://doi.org/10.1016/j.jcis.2019.03.050 (2019).
Francius, G. et al. Bacterial surface appendages strongly impact nanomechanical and electrokinetic properties of Escherichia coli cells subjected to osmotic stress. PLoS One 6, e20066, https://doi.org/10.1371/journal.pone.0020066 (2011).
Correia, S. et al. Infecção respiratória por bactérias do complexo cepacia: Evolução clínica em doentes com fibrose quística. Revista Portuguesa de Pneumologia 14, 5–26 (2008).
Lieberman, T. D. et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nature Genetics 43, 1275–1280, https://doi.org/10.1038/ng.997 (2011).
Silva, I. N. et al. Long-Term Evolution of Burkholderia multivorans during a Chronic Cystic Fibrosis Infection Reveals Shifting Forces of Selection. mSystems 1, e00029–00016, https://doi.org/10.1128/mSystems.00029-16 (2016).
Nunvar, J. et al. What matters in chronic Burkholderia cenocepacia infection in cystic fibrosis: Insights from comparative genomics. PLOS Pathogens 13, e1006762, https://doi.org/10.1371/journal.ppat.1006762 (2017).
King, J. D. et al. Review: Lipopolysaccharide biosynthesis in Pseudomonas aeruginosa. Innate Immunity 15, 261–312, https://doi.org/10.1177/1753425909106436 (2009).
Cunneen, M. M. et al. (eds Yuriy A. Knirel & Miguel A. Valvano) 339–370 (Springer Vienna, Vienna, 2011).
Hassan, A. A. et al. Burkholderia cepacia complex species differ in the frequency of variation of the lipopolysaccharide O-antigen expression during cystic fibrosis chronic respiratory infection. Frontiers in Cellular and Infection Microbiolog y 9, https://doi.org/10.3389/fcimb.2019.00273 (2019).
Saldias, M. S. et al. Burkholderia cenocepacia O antigen lipopolysaccharide prevents phagocytosis by macrophages and adhesion to epithelial cells. Journal of Medical Microbiology 58, 1542–1548, https://doi.org/10.1099/jmm.0.013235-0 (2009).
Ranf, S. Immune Sensing of Lipopolysaccharide in Plants and Animals: Same but Different. PLoS Pathogens 12, e1005596, https://doi.org/10.1371/journal.ppat.1005596 (2016).
Gan, L. et al. Molecular organization of Gram-negative peptidoglycan. PNAS, Proceedings of the National Academy of Sciences 105, 18953–18957, https://doi.org/10.1073/pnas.0808035105 (2008).
Turner, R. D. et al. Different walls for rods and balls: the diversity of peptidoglycan. Molecular Microbiology 91, 862–874, https://doi.org/10.1111/mmi.12513 (2014).
Yao, X. et al. Thickness and elasticity of gram-negative murein sacculi measured by atomic force microscopy. Journal of Bacteriology 181, 6865–6875 (1999).
Bramkamp, M. & van Baarle, S. Division site selection in rod-shaped bacteria. Current Opinion in Microbiology 12, 683–688, https://doi.org/10.1016/j.mib.2009.10.002 (2009).
Wheeler, R. et al. Super-resolution microscopy reveals cell wall dynamics and peptidoglycan architecture in ovococcal bacteria. Molecular Microbiology 82, 1096–1109, https://doi.org/10.1111/j.1365-2958.2011.07871.x (2011).
Dover, R. S. et al. Multiparametric AFM reveals turgor-responsive net-like peptidoglycan architecture in live streptococci. Nature. Communications 6, 7193, https://doi.org/10.1038/ncomms8193 (2015).
Turner, R. D. et al. Peptidoglycan architecture can specify division planes in Staphylococcus aureus. Nature Communication 1, 26, https://doi.org/10.1038/ncomms1025 (2010).
Persat, A. et al. The mechanical world of bacteria. Cell 161, 988–997, https://doi.org/10.1016/j.cell.2015.05.005 (2015).
Uzoechi, S. C. & Abu-Lail, N. I. Changes in cellular elasticities and conformational properties of bacterial surface biopolymers of multidrug-resistant Escherichia coli (MDR-E. coli) strains in response to ampicillin. The Cell Surface 5, 100019, https://doi.org/10.1016/j.tcsw.2019.100019 (2019).
Veyrier, F. J. et al. Common Cell Shape Evolution of Two Nasopharyngeal Pathogens. PLoS Genetics 11, e1005338, https://doi.org/10.1371/journal.pgen.1005338 (2015).
Yang, D. C. et al. Staying in Shape: the Impact of Cell Shape on Bacterial Survival in Diverse Environments. Microbiology and Molecular Biology Reviews 80, 187–203, https://doi.org/10.1128/MMBR.00031-15 (2016).
Weiser, J. N. The battle with the host over microbial size. Current Opinion in Microbiology 16, 59–62, https://doi.org/10.1016/j.mib.2013.01.001 (2013).
Dalia, A. B. & Weiser, J. N. Minimization of bacterial size allows for complement evasion and is overcome by the agglutinating effect of antibody. Cell Host Microbe 10, 486–496, https://doi.org/10.1016/j.chom.2011.09.009 (2011).
Uzoechi, S. C. & Abu-Lail, N. I. The Effects of beta-Lactam Antibiotics on Surface Modifications of Multidrug-Resistant Escherichia coli: A Multiscale Approach. Microscopy and Microanalysis 25, 135–150, https://doi.org/10.1017/S1431927618015696 (2019).
Vadia, S. et al. Fatty Acid Availability Sets Cell Envelope Capacity and Dictates Microbial Cell Size. Current Biology 27, 1757–1767 e1755, https://doi.org/10.1016/j.cub.2017.05.076 (2017).
Yao, Z. et al. Regulation of cell size in response to nutrient availability by fatty acid biosynthesis in Escherichia coli. PNAS, Proceedings of the National Academy of Sciences 109, E2561–2568, https://doi.org/10.1073/pnas.1209742109 (2012).
Du Toit, A. Growth capacity and cell size. Nature Reviews Microbiology 17, 2, https://doi.org/10.1038/s41579-018-0124-y (2019).
Dai, X. et al. Sinorhizobium meliloti, a Slow-Growing Bacterium, Exhibits Growth Rate Dependence of Cell Size under Nutrient Limitation. mSphere 3, https://doi.org/10.1128/mSphere.00567-18 (2018).
Alvarez-Ortega, C. & Harwood, C. S. Responses of Pseudomonas aeruginosa to low oxygen indicate that growth in the cystic fibrosis lung is by aerobic respiration. Molecular Microbiology 65, 153–165, https://doi.org/10.1111/j.1365-2958.2007.05772.x (2007).
Pessi, G. et al. Response of Burkholderia cenocepacia H111 to Micro-Oxia. PLOS ONE 8, e72939, https://doi.org/10.1371/journal.pone.0072939 (2013).
Olivares Pacheco, J. et al. Metabolic Compensation of Fitness Costs Is a General Outcome for Antibiotic-Resistant Pseudomonas aeruginosa Mutants Overexpressing Efflux Pumps. MBio 8, https://doi.org/10.1128/mBio.00500-17 (2017).
Olivares, J. et al. Metabolic compensation of fitness costs associated with overexpression of the multidrug efflux pump MexEF-OprN in Pseudomonas aeruginosa. Antimicrob Agents Chemother 58, 3904–3913, https://doi.org/10.1128/AAC.00121-14 (2014).
Allison, D. P. et al. Bacterial immobilization for imaging by atomic force microscopy. Journal of Visualized Experiments. https://doi.org/10.3791/2880 (2011).
Liu, Y. & Camesano, T. A. In Microbial Surfaces Vol. 984 ACS Symposium Series Ch. 10, 163–188 (American Chemical Society, 2008).
Osiro, D. et al. Measuring bacterial cells size with AFM. Brazillian Journal of Microbiology 43, 341–347, https://doi.org/10.1590/S1517-838220120001000040 (2012).
Saar-Dover, R. et al. D-alanylation of lipoteichoic acids confers resistance to cationic peptides in group B. streptococcus by increasing the cell wall density. PLoS Pathogens 8, e1002891, https://doi.org/10.1371/journal.ppat.1002891 (2012).
Yang, C. H. et al. An engineered arginine-rich alpha-helical antimicrobial peptide exhibits broad-spectrum bactericidal activity against pathogenic bacteria and reduces bacterial infections in mice. Scientific Reports 8, 14602, https://doi.org/10.1038/s41598-018-32981-3 (2018).
Qin, Z. et al. Organic compounds inhibiting S. epidermidis adhesion and biofilm formation. Ultramicroscopy 109, 881–888, https://doi.org/10.1016/j.ultramic.2009.03.040 (2009).
Schillers, H. et al. Standardized Nanomechanical Atomic Force Microscopy Procedure (SNAP) for Measuring Soft and Biological Samples. Scientific Reports 7, 5117, https://doi.org/10.1038/s41598-017-05383-0 (2017).
Wagner, R. et al. Uncertainty quantification in nanomechanical measurements using the atomic force microscope. Nanotechnology 22, 455703, https://doi.org/10.1088/0957-4484/22/45/455703 (2011).
Butt, H. J. et al. Force measurements with the atomic force microscope: Technique, interpretation and applications. Surface Science Reports 59, 1–152, https://doi.org/10.1016/j.surfrep.2005.08.003 (2005).
Sader, J. E. et al. A virtual instrument to standardise the calibration of atomic force microscope cantilevers. Review of Scientific Instruments 87, 093711, https://doi.org/10.1063/1.4962866 (2016).
Cunha, M. V. et al. Studies on the involvement of the exopolysaccharide produced by cystic fibrosis-associated isolates of the Burkholderia cepacia complex in biofilm formation and in persistence of respiratory infections. Journal of Clinical Microbiology 42, 3052–3058, https://doi.org/10.1128/JCM.42.7.3052-3058.2004 (2004).
Acknowledgements
This work was supported by FCT – Portuguese Foundation for Science and Technology – project contract: PTDC/FIS-NAN/6101/2014. Funding received by iBB – Institute for Bioengineering and Biosciences and by BioISI – Biosystems and Integrative Sciences Institute – from FCT (UID/BIO/04565/2019 and UID/MULTI/04046/2019, respectively) is acknowledged, as well as FCT fellowships to AAH (BIOTECnico – PhD programme – PD/BD/113629/2015), MVV (PD/BD/105975/2014) and, MSR (SFRH/BPD/69201/2010).
Author information
Authors and Affiliations
Contributions
I.S.C. designed the experiment with contribution from M.S.R. A.A.H. prepared the AFM samples, examined the growth behaviour of the cells, and performed the biofilm formation assays, under the supervision of I.S.C. M.V.V., T.R. and A.A.H. performed the AFM measurements under the supervision of M.S.R. M.S.R. analysed the AFM indentation curves and A.A.H. analysed the roughness measurements. I.S.C. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hassan, A.A., Vitorino, M.V., Robalo, T. et al. Variation of Burkholderia cenocepacia cell wall morphology and mechanical properties during cystic fibrosis lung infection, assessed by atomic force microscopy. Sci Rep 9, 16118 (2019). https://doi.org/10.1038/s41598-019-52604-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52604-9
This article is cited by
-
Identification of Burkholderia cenocepacia non-coding RNAs expressed during Caenorhabditis elegans infection
Applied Microbiology and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.