Abstract
Human activities have resulted in the loss of over 90% of sharks in most ocean basins and one in four species of elasmobranch are now listed at risk of extinction by the IUCN. How this collapse will affect the ability of populations to recover in the face of continued exploitation and global climate change remains unknown. Indeed, important ecological and biological information are lacking for most shark species, particularly estimates of genetic diversity and population structure over a range of spatial scales. Using 15 microsatellite markers, we investigated genetic diversity and population structure in gray reef sharks over their Indo-Pacific range (407 specimens from 9 localities). Clear genetic differentiation was observed between the Indian and the Pacific Ocean specimens (FST = 0.145***). Further differentiation within the Pacific included a West and East cleavage as well as North-Central and South-Central Pacific clusters. No genetic differentiation was detected within archipelagos. These results highlight the legacy of past climate changes and the effects of large ocean expanses and circulation patterns on contrasting levels of connectivity at global, regional and local scales. Our results indicate a need for regional conservation units for gray reef sharks and pinpoint the isolation and vulnerability of their French Polynesian population.
Similar content being viewed by others
Introduction
Anthropogenic pressures, including over-fishing and habitat destruction, have resulted in the loss of over 90% of sharks and large predatory fishes across all ocean basins1,2,3. Collapse of shark populations has had profound effects on ecosystem functioning and resilience4,5,6 and potentially threatens their capacity to maintain demographic connectivity and genetic diversity across their ranges7. Reef sharks are of particular concern as they are a primarily reef-associated group that plays an important structural role in coral reef ecosystems as apex or meso-predators8,9,10. Recent work has reported striking declines in reef shark densities associated with the size of adjacent human populations8. More information on basic evolutionary and demographic processes are urgently needed to ensure conservation of individual reef shark populations and the ecosystem services that they provide across the world’s tropical and sub-tropical oceans9,10.
The application of molecular DNA techniques in conservation biology is well established7. Molecular tools are, in many cases, the only means available to estimate essential parameters for populations, and this is certainly the case for rare and elusive species. Genetic methods have been used to infer dispersal patterns, demographic trends and important reproductive traits in many sharks. Indeed, mitochondrial sequences and microsatellite markers have revealed genetic structure and recent declines in whale sharks11, a strong trans-Pacific break and the need for regional conservation units in the Galapagos shark12, and panmixia throughout the Indo-Pacific range of the highly migratory tiger shark13. Furthermore, reef shark studies have demonstrated significant genetic structure and variable demographic histories for blacktip reef sharks over their Indo-Pacific range14, and female blacktip reef sharks were shown to be philopatric in French Polynesia15. Genetic differentiation is also apparent in whitetip reef sharks across the Indo-Pacific but populations are more homogeneous at the regional scale16. Taken together, these studies illustrate that spatial and temporal variability in genetic differentiation among populations often reflect important components of the movement ecology of elasmobranch species.
The gray reef shark (Carcharhinus amblyrhynchos) is a reef-associated species distributed throughout the Indian Ocean and West and Central Pacific17,18. Gray reef sharks are among the most abundant reef sharks in the Indo-Pacific and can comprise up to 50% of the upper trophic level biomass on coral reefs in some areas19. Yet possible collapses of gray reef shark populations have been documented, even on reefs with relatively low human impacts20. Without a pelagic larval phase in their life cycle, population connectivity in gray reef sharks occurs exclusively through movements of adults and juveniles21,22,23,24. These movements and residency patterns are complex and vary among reefs and life stages and between sexes22,23,24. Gray reef sharks have been shown to change territory often at some locations (Rangiroa, French Polynesia25) while at other places they have been documented to stay up to two years within the same territory (the Great Barrier Reef24,26, GBR). Low residency was also recorded on the semi-isolated reefs of the Great Barrier Reef23 whereas on isolated habitats, gray reef sharks seem to exhibit longer residency on a single reef27,28. The few genetic studies undertaken to date on gray reef sharks have shown high migration frequency along the Great Barrier Reef29, a barrier to gene flow across large expanses of ocean, and an isolation-by-distance pattern (when the genetic distance between populations is proportional to the geographic distance between these populations) along the Australian continental shelf30. So far, no study has investigated the genetic composition of gray reef shark populations across the species’ range. Therefore, while reef characteristics such as geography and oceanographic context likely play an important role, more specific factors regulating dispersal of gray reef sharks remain unknown.
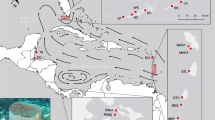
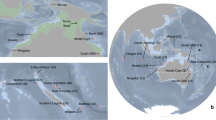
Here, we used 15 microsatellite markers to investigate the genetic structure and diversity of the gray reef shark populations from French Polynesia and the Line islands to the western Indian Ocean (Fig. 1; Table S1). We focused efforts on contrasting local and regional levels of gene flow in the Central and West Pacific. Population structure will likely be strongly influenced by behavioral trade-offs between an inherent ability to roam over large distances (high gene flow potential) while also displaying site-fidelity and residency behavior (low gene flow potential).
Results
Genetic diversity
Mean allele numbers ranged from 4.7 to 15.5 (Table 1). Private allele numbers per sampling site ranged from 0 to 18 while there was a total of 143 alleles found exclusively in the Pacific Ocean and 10 alleles recovered only from the western Indian Ocean. Observed and expected heterozygosity values ranged from 0.7120 to 0.8500 (Table 1). Inbreeding coefficients were significant for the Australian Great Barrier Reef, French Polynesia and the Phoenix archipelago samples (Table 1).
Genetic differentiation between ocean basins
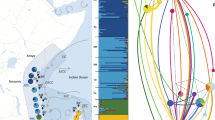
The Fst pairwise comparison between western Indian Ocean and Pacific Ocean populations was high and significant (Fst = 0.145***). Therefore, all individual Fst values including the Indian Ocean were significant and high (from 0.145 to 0.161, Table 2). Furthermore, the first axis of the Principal Coordinates Analysis (PCoA) clearly differentiated the western Indian Ocean samples from the samples collected in the Pacific Ocean (Fig. 2). This differentiation was also clear in the Bayesian clustering analysis which suggested two major clusters between the western Indian and Pacific basins (K = 2; Fig. 3a). Finally, the test of isolation-by-distance, when considering the entire dataset, was significant (p < 0.001).
Bayesian Structure plots showing the most likely number of clusters (K) partitioning the dataset when including (a) all 407 specimens from the Indian and the Pacific Ocean, K = 2; and (b) the Pacific specimens only, K = 3. The most likely number of clusters were determined using the Evanno’s method (Evanno et al. 2005).
Genetic differentiation within the Pacific Ocean
Of the total 45 pairwise Fst comparisons only 5 were not significant, and those were always between close localities such as New Caledonia vs. Chesterfield, Society vs. Tuamotu, or for localities with few specimens (Tuvalu N = 4, Table 2). Regarding the PCoA, the second axis clearly separated West Pacific samples (Australia, Chesterfield and New Caledonia) and Central Pacific samples (Tuvalu, Phoenix, Palmyra and French Polynesia, Fig. 2). The Bayesian clustering, focusing on Pacific specimens only, also revealed Central and West Pacific groups but showed further segregation within the Central Pacific samples: Phoenix, Tuvalu and Palmyra grouped together while French Polynesia was separated in another group with a distinct genetic composition (Fig. 3b). At the scale of an archipelago (Phoenix) or two close archipelagos (Society and Tuamotu), few pairwise Fst comparisons were significant (Supp Data Tables S2, S3). Finally, the test of isolation-by-distance was also significant when considering the Pacific Ocean samples only (i.e. removing the western Indian Ocean samples), with or without Tuvalu (N = 4; p < 0.01 and p < 0.001, respectively).
Discussion
Population genetic structure of gray reef sharks throughout their Indo-Pacific range revealed contrasting levels of connectivity across global, regional and local scales. The effects of the Indo-Pacific barrier are clear between the Indian and the Pacific Ocean samples. Gray reef sharks also showed regional genetic differentiation between the West and Central Pacific, likely generated by the large expanses of open water between the regions. We also found some level of genetic differentiation between the North and South-Central Pacific, suggesting potential effects of oceanic current circulation on the population structure of gray reef sharks. We found no significant genetic differentiation on small spatial scales within the Phoenix archipelago or within French Polynesia. However, we detected a pattern of isolation-by-distance among populations of both oceans and within the Pacific. Overall, the genetic structure found in gray reef sharks appears higher than that of reef fishes, suggesting that adult movements of this shark limit dispersal capabilities compared to the pelagic larval stage of teleost coral reef fishes31.
The western Indian Ocean population of the gray reef shark is clearly separated from the rest of the samples collected in the Pacific Ocean (Table 2, Figs 2 and 3A). Momigliano et al.30 also demonstrated genetic differentiation between the Pacific Ocean samples of gray reef sharks and samples from the Chagos Archipelago in the North Indian Ocean. However, more samples in the northern and the eastern Indian Ocean are needed to pinpoint the precise area of the genetic break between Indian and Pacific populations. Significant genetic differentiation between Indian and Pacific oceans populations has also been identified in many fish species32,33, including three other coastal shark species. The scalloped hammerhead shark Sphyrna lewini34, the blacktip shark Carcharhinus limbatus35 and the whitetip shark Triaenodon obesus36 all demonstrated a significant genetic break across the Indo-Pacific barrier; however, neither tiger sharks13 or whale sharks11 show a similar discontinuity across this barrier. The most likely explanation for the genetic break in reef fish, including sharks, is the effect of the low sea-level stands during glacial periods (low sea-level, about 100 m from current level, during glacial periods)33,36. Moreover, the large oceanic distances are now likely hindering widespread genetic exchange between the gray reef shark populations of each ocean basin as large stretches of deep open water have been suggested to create a barrier to gene flow for this species30. The strong genetic differentiation that we found (Table 2, Figs 2 and 3A) is even more noteworthy as sharks have slow rates of evolution compared to mammals and bony fish due to their slow metabolic rates37. From the current sampling, it is not clear if these populations are still capable of interbreeding or if they should be considered as distinct sister species given the numerous private alleles (Table 1). For the Indo-West Pacific marine biogeographic area, a contact zone is known to be located at Christmas and Cocos (Keeling) islands for many reef fishes38. The Cocos population from Momigliano et al.30 seems to show a mixed genetic make-up but future gray reef shark sampling should focus on this area and on northern areas in Indonesia to definitively assess the status of the Indian and the Pacific populations (i.e. sister species or intraspecific genetic differentiation).
At a regional scale, gray reef shark samples from the Pacific Ocean were clustered in distinct groups, with the Coral Sea specimens forming one cluster (Eastern Australia, Chesterfield and New Caledonia), the North-Central specimens forming another cluster (Tuvalu, Phoenix and Palmyra) and the South-Central specimens (French Polynesia) comprising a third cluster (Fig. 3b). Additionally, Tuvalu, Phoenix and Palmyra populations were more closely related to each other than Palmyra and French Polynesia (Fig. 3b), whereas the Line islands (Palmyra) and French Polynesia are geologically and geographically linked with the intersection of the Line islands seamount chain with the Tuamotu plateau. Genetic differentiation has previously been reported between Central and West Pacific teleost (bony) reef fish populations39,40 as well as in the sedentary whitetip sharks16. Large ocean expanses between these two regions likely reduce the probability of long-distance dispersal. Schultz et al.41 revealed moderate genetic differentiation (FST = 0.07) between Australian and French Polynesian populations of the Sicklefin lemon shark Negaprion acutidens and proposed a stepping stone mechanism for rare long-distance dispersal events between West and Central Pacific islands. Additionally, genetic differentiation between the Line Islands and French Polynesia has been demonstrated in the three-spot damselfish, Dascyllus trimaculatus42 and the brown surgeonfish Acanthurus nigrofuscus43. This genetic differentiation on either side of the equator in the Central Pacific likely arose due to isolation maintained by the major currents in the zone (Equatorial Counter Current and North Equatorial Current)44,45. A similar pattern of genetic differentiation in gray reef sharks suggests that dispersal seems to be dependent, to some extent, on the ocean circulation and the major currents in the area likely limit migration of sharks across this boundary. Gray reef sharks are viviparous and therefore, contrary to most reef fishes, population connectivity occurs through movements of adults and juveniles rather than dispersal of eggs and larvae. It is therefore likely that the same barriers to dispersal are acting for species both with and without larval dispersal. However, gray reef sharks have strong and complex reef residency behaviors23,24 that, when combined with the large expanse of deep open ocean between the regions, apparently reduces the frequency of long-distance movements. Thus, the genetic structure of gray reef sharks appears somewhat higher than that of reef fishes, that are more homogenous across their range39,42,43. We did identify a pattern of isolation-by-distance in our dataset when including all samples, samples from the Pacific only, and when removing the small size sample (Tuvalu), indicating that dispersal probability in gray reef sharks is proportional to geographic distance among populations. Similar patterns of isolation-by-distance were described by Momigliano et al.30 and highlight the low probability of adult gray reef sharks migrating long distances across expanses of open ocean.
At a local scale, we collected sufficient numbers of samples to investigate genetic differentiation at the scale of a single archipelago (Phoenix archipelago, about 500 km in diameter) or several close archipelagos (French Polynesia, 500 km to 1000 km). No significant differences among reefs were found in either comparison. Momigliano et al.29 similarly detected no genetic differentiation in gray reef sharks over the 1200 km of the Great Barrier Reef system. In contrast, blacktip reef sharks show differentiation both among and within the archipelagos of French Polynesia14,46. Blacktip reef sharks must, therefore, either home to natal reefs at much higher frequencies, or move among reefs at much lower frequencies, than gray reef sharks. Interestingly, inbreeding coefficients from Eastern Australia, Phoenix, Society and Tuamotu samples were significant, suggesting somewhat limited exchange among reefs that went undetected by more traditional statistical analyses based on Fst. Overall, our results suggest that a small number of gray reef sharks make infrequent long-distance movements between reefs across open water. White et al.28 came to a similar conclusion based on a small sample of satellite-tagged gray reef sharks at Palmyra Atoll in the North tropical Pacific. Two from a total of six sharks were detected outside of the 200 NM limit MPA (Marine Protected Area) boundary surrounding Palmyra Atoll for 3% and 46% of their daily location estimates. The shark that traveled the furthest reached a maximum linear distance of 908 km from the 12NM MPA boundary surrounding Palmyra Atoll28.
The need to consider genetic diversity in conservation of sharks was recently re-emphasized7,47. In our study, observed heterozygosity values were comparable for all the samples analyzed and also to those from the gray reef sharks of the Great Barrier Reef (0.79 to 0.829). These values were higher than in blacktip reef sharks28, where observed heterozygosity ranged from 0.47 to 0.67 but comparable to that of other shark species also analyzed with microsatellite markers7. Noticeably, heterozygosity levels did not correlate with the level of isolation of a reef, as all reefs show similar values. Thus, we found no evidence for recent genetic bottlenecks, as suggested for other reef shark species28. In our study, we further revealed clear genetic groupings across the Pacific region, with genetic differentiation among broader geographic clusters (West vs Central Pacific and North vs South Central Pacific; Fig. 3b). Gray reef sharks should thus be divided into regional conservation units, as are many marine mammals and sea turtles48,49. Similarly, 3 to 4 conservation units were recently suggested for the Galapagos sharks across their Pacific range12. For reef sharks, a strong tendency for site fidelity and residency behavior should also be taken into consideration when delineating conservation units. However, at the scale of an archipelago (Phoenix) or several closely related archipelagos (French Polynesia), gray reef sharks appear to represent quasi-panmictic units with free movements of individuals across reefs. Juhel et al.50 also showed that human-linked behavioral alterations should be considered in management strategies to ensure the persistence of gray reef shark populations. Furthermore, rare long-distance dispersal across open ocean barriers seem to be an important process driving genetic diversity and structure in gray reef sharks. Open ocean movements through “high-seas” corridors therefore represent an important component of the life history of at least some coastal sharks that functions to reduce local inbreeding. “High-seas” are currently the subject of much debate at the United Nations, and the first-ever “high-seas” conservation treaty is expected to be finalized in 2020. Momigliano et al.28 went so far as to suggest a move from discontinuous networks of MPAs on the GBR to MPAs connected via protected corridors to better protect these sharks with larger home ranges. A conservation plan for these gray reef shark populations should therefore include several conservation units with the potential for migration corridors in the high seas that would serve to connect regional reefs within each conservation unit. The Indian Ocean gray reef sharks clearly represent a conservation unit on their own while another conservation unit can be identified for the West Pacific and two others for the North and South-Central Pacific populations (Fig. 3a,b).
Finally, French Polynesia appeared isolated from the rest of the gray reef shark populations including archipelagos in the Central Pacific (Fig. 3b). The isolation of the French Polynesian atolls has been demonstrated in several species, with a complex legacy of past climate effects and rare contemporaneous migrations43,51. This genetic isolation has direct implications for conservation and resilience of sharks in the region and French Polynesia should be considered as a separate conservation unit. Blacktip reef sharks also showed signals of recent bottleneck in Moorea, likely due to the growing anthropogenic pressure in these populated islands14. Furthermore, gray reef sharks in the Tuamotu archipelago seem to show peculiarities, suggesting that this population may warrant a high conservation priority. Indeed, their extremely high number at Fakarava Atoll was recently demonstrated to produce an inverted trophic pyramid, maintained through subsidies linked to spawning aggregations of teleosts52. Finally, as French Polynesia was recently shown as a likely past glacial refuge for green sea turtles53,54 and as a projected refuge under future warm conditions55,56, the gray reef sharks from this region are particularly important in a conservation context as they represent a genetically-distinct group and are likely to be less impacted by warmer conditions in this refuge.
Material and Methods
Ethical statement
All samples were collected in agreement with local legislations. No permit was required for DNA collection in New Caledonia. Research on sharks in French Polynesia was approved under Arrêté N° 9524 issued by the Ministère de la Promotion des Langues, de la Culture, de la Communication et de l’Environnement of the French Polynesian government on 30 October 2015, and Arrêté N° 5129 issued by the Ministère de la Promotion des Langues, de la Culture, de la Communication et de l’Environnement of the French Polynesian government on 22 June 2016. Samples in the Phoenix Islands Protected Area were collected under relevant permits from the Government of Kiribati and sampling protocols approved by Woods Hole Oceanographic Institution Institutional Animal Care and Use Committee (IACUC) ID number 18417. Sampling protocols for Palmyra Atoll collections were certified by the IACUC, University of California, Santa Barbara, Protocol no. 856 (date of IACUC approval: 5/31/2012) under U.S. Fish and Wildlife Service special use permits (Permit Numbers #12533–14011, #12533–13011, #12533–12011, #12533–11007, #12533–10011, #12533–09010, #12533–08011, and #12533–07006). Samples from the Great Barrier Reef were provided by collaborators from Australia in accordance with local regulations.
Sample collection
A total of 407 sharks were sampled at 9 locations in the South West Indian Ocean, and the West and Central Pacific (Fig. 1, see Table S1 for details). Fin clips were collected from sharks sampled by hooks and lines. Sharks were brought to the surface and put in tonic immobility along the side of small boats and restrained with a tail line while cutting a small tissue sample from one of the fins and then immediately released.
Molecular data
Total DNA was extracted from fin clips using the PureGene protocol (Qiagen, Hilden, Germany). A total of 15 microsatellite markers developed for gray reef sharks were amplified using the primers and cycling parameters from Momigliano et al.57. PCR products were sent for sequencing to an external private company (GenoScreen, Lille, France) to be run on an Applied Biosystems 3730 sequencer and were genotyped employing GeneMapper software version 4.0 (Applied Biosystems, Foster City, CA, USA).
Data analyses
The presence of null alleles, scoring errors and large allele drop-out was verified using Micro-checker v2.2.358. The mean number of alleles (A), observed (Ho) and expected (He) heterozygosity, the inbreeding coefficient (Fis) and the fixation index (Fst) were computed in Genetix v4.05.0259. The number of private alleles (Ap) was computed in GenAlEx v6.560. The correction for multiple tests of Benjamini & Hochberg61 was applied.
To investigate population structure, a Principal Coordinates Analysis (PCoA) on populations was first computed in GenAlEx. Additionally, Structure v2.362 was used to search for the most likely number of clusters. This analysis was run with no priors by location. After initial runs, the parameters were set as follows: a burn-in period of 100 000 iterations followed by 500 000 recorded iterations for K = 1 to K = 9 clusters and 15 iterations per K values. We ran a first analysis with all specimens and a second analysis with the specimens from the Pacific Ocean only. In this last analysis, we used sampling locations as prior (model LOCPRIOR63), known to be able to reveal subtle differentiations. The most probable number of clusters present in this dataset was determined using the Evanno’s ΔK approach64 and computed with Structure Harvester online65.
We also tested for a pattern of isolation-by-distance using a Mantel test in Genetix between the matrix of genetic distances [Fst ⁄ (1 – Fst)]66 and the matrix of log coastal geographical distances (in km) of each pair of localities. Significance was tested using a random permutation procedure implemented in Genetix (5 000 permutations). This analysis was performed on the whole dataset and on the Pacific samples only with or without the site with small sample size (Tuvalu, N = 4).
Data availability
The dataset consisting of 407 multilocus genotypes is provided in Table S4.
References
Nadon, M. O. et al. Re-creating missing population baselines for Pacific reef sharks. Conserv. Biol. 26, 493–503, https://doi.org/10.1111/j.1523-1739.2012.01835.x (2012).
Worm, B. et al. Global catches, exploitation rates, and rebuilding options for sharks. Mar. Pol. 40, 194–204, https://doi.org/10.1016/j.marpol.2012.12.034 (2013).
Dulvy, N. K. et al. Extinction risk and conservation of the world’s sharks and rays. eLife 3, 34, https://doi.org/10.7554/eLife.00590 (2014).
Myers, R. A., Baum, J. K., Shepherd, T. D., Powers, S. P. & Peterson, C. H. Cascading effects of the loss of apex predatory sharks from a coastal ocean. Science 315, 1846–1850, https://doi.org/10.1126/science.1138657 (2007).
Baum, J. K. & Worm, B. Cascading top-down effects of changing oceanic predator abundances. J. Anim. Ecol. 78, 699–714, https://doi.org/10.1111/j.1365-2656.2009.01531.x (2009).
Estes, J. A. et al. Trophic downgrading of planet earth. Science 333, 301–306, https://doi.org/10.1126/science.1205106 (2011).
Domingues, R. R., Hilsdorf, A. W. S. & Gadig, O. B. F. The importance of considering genetic diversity in shark and ray conservation policies. Conserv. Genet. 19, 501–525, https://doi.org/10.1007/s10592-017-1038-3 (2018).
Osgood, G. J. & Baum, J. K. Reef sharks: recent advances in ecological understanding to inform conservation. J. Fish Biol. 87, 1489–1523, https://doi.org/10.1111/jfb.12839 (2015).
Frisch, A. J. et al. Reassessing the trophic role of reef sharks as apex predators on coral reefs. Coral Reefs 35, 459–472, https://doi.org/10.1007/s00338-016-1415-2 (2016).
Roff, G. et al. The ecological role of sharks on coral reefs. Trends Ecol. Evol. 31, 395–407, https://doi.org/10.1016/j.tree.2016.02.014 (2016).
Vignaud, T. M. et al. Genetic structure of populations of whale sharks among ocean basins and evidence for their historic rise and recent decline. Mol. Ecol. 23, 2590–2601, https://doi.org/10.1111/mec.12754 (2014).
Pazmino, D. A. et al. Strong trans-Pacific break and local conservation units in the Galapagos shark (Carcharhinus galapagensis) revealed by genome-wide cytonuclear markers. Heredity 120, 407–421, https://doi.org/10.1038/s41437-017-0025-2 (2018).
Holmes, B. J. et al. Population structure and connectivity of tiger sharks (Galeocerdo cuvier) across the Indo-Pacific Ocean basin. R. Soc. Open Sci. 4, 9, https://doi.org/10.1098/rsos.170309 (2017).
Vignaud, T. M. et al. Blacktip reef sharks, Carcharhinus melanopterus, have high genetic structure and varying demographic histories in their Indo-Pacific range. Mol. Ecol. 23, 5193–5207, https://doi.org/10.1111/mec.12936 (2014).
Mourier, J. & Planes, S. Direct genetic evidence for reproductive philopatry and associated fine-scale migrations in female blacktip reef sharks (Carcharhinus melanopterus) in French Polynesia. Mol. Ecol. 22, 201–214, https://doi.org/10.1111/mec.12103 (2013).
Whitney, N. M., Robbins, W. D., Schultz, J. K., Bowen, B. W. & Holland, K. N. Oceanic dispersal in a sedentary reef shark (Triaenodon obesus): genetic evidence for extensive connectivity without a pelagic larval stage. J. Biogeogr. 39, 1144–1156, https://doi.org/10.1111/j.1365-2699.2011.02660.x (2012).
Compagno, L. J. V. FAO Species catalogue. Vol. 4. Sharks of the world. An annotated and illustrated catalogue of shark species known to date. Part 2. Carcharhiniformes. FAO Fish Synopsis 4, 251–655 (1984).
Wetherbee, B. M., Crow, G. L. & Lowe, C. G. Distribution, reproduction and diet of the gray reef shark Carcharhinus amblyrhynchos in Hawaii. Mar. Ecol.-Prog. Ser. 151, 181–189, https://doi.org/10.3354/meps151181 (1997).
Friedlander, A. M. et al. The real bounty: marine biodiversity in the Pitcairn Islands. PLoS One 9, 11, https://doi.org/10.1371/journal.pone.0100142 (2014).
Clua, E. & Vignaud, T. Possible collapse of reef shark populations in remote coral reef ecosystems in the Coral Sea (Western Pacific). Cybium 40, 51–59 (2016).
Speed, C. W., Field, I. C., Meekan, M. G. & Bradshaw, C. J. A. Complexities of coastal shark movements and their implications for management Mar Ecol-Prog Ser 408:275–U305, https://doi.org/10.3354/meps08581 (2010).
McKibben, J. N. & Nelson, D. R. Patterns of movement and grouping of gray reef sharks, Carcharhinus amblyrhynchos, at Enewetak, Marshall Islands. Bull. Mar. Sci. 38, 89–110 (1986).
Heupel, M. R., Simpfendorfer, C. A. & Fitzpatrick, R. Large-scale movement and reef fidelity of gray reef sharks. PLoS One 5, 5, https://doi.org/10.1371/journal.pone.0009650 (2010).
Espinoza, M., Heupel, M. R., Tobin, A. J. & Simpfendorfer, C. A. Residency patterns and movements of gray reef sharks (Carcharhinus amblyrhynchos) in semi-isolated coral reef habitats. Mar. Biol. 162, 343–358, https://doi.org/10.1007/s00227-014-2572-x (2015).
Nelson, D. R. & Johnson, R. H. Behavior of the reef sharks of Rangiroa, French Polynesia. National Geographic Society Research Report 12, 479–499 (1980).
Heupel, M. R. & Simpfendorfer, C. A. Importance of environmental and biological drivers in the presence and space use of a reef-associated shark. Mar. Ecol.-Prog. Ser. 496, 47–57, https://doi.org/10.3354/meps10529 (2014).
Barnett, A., Abrantes, K. G., Seymour, J. & Fitzpatrick, R. Residency and spatial use by reef sharks of an isolated seamount and its implications for conservation. PLoS One 7, 12, https://doi.org/10.1371/journal.pone.0036574 (2012).
White, T. D. et al. Assessing the effectiveness of a large marine protected area for reef shark conservation. Biol. Conserv. 207, 64–71, https://doi.org/10.1016/j.biocon.2017.01.009 (2017).
Momigliano, P., Harcourt, R., Robbins, W. D. & Stow, A. Connectivity in grey reef sharks (Carcharhinus amblyrhynchos) determined using empirical and simulated genetic data. Sci Rep 5, 9, https://doi.org/10.1038/srep13229 (2015).
Momigliano, P. et al. Genetic structure and signatures of selection in grey reef sharks (Carcharhinus amblyrhynchos). Heredity 119, 142–153, https://doi.org/10.1038/hdy.2017.21 (2017).
Mora, C. & Sale, P. F. Are populations of coral reef fish open or closed? Trends Ecol Evol 17, 422–428, https://doi.org/10.1016/s0169-5347(02)02584-3 (2002).
McMillan, W. O. & Palumbi, S. R. Concordant evolutionary patterns among Indo-West Pacific butterflyfishes. Proc. R. Soc. B-Biol. Sci. 260, 229–236, https://doi.org/10.1098/rspb.1995.0085 (1995).
Gaither, M. R. & Rocha, L. A. Origins of species richness in the Indo-Malay-Philippine biodiversity hotspot: evidence for the centre of overlap hypothesis. J. Biogeogr. 40, 1638–1648, https://doi.org/10.1111/jbi.12126 (2013).
Duncan, K. M., Martin, A. P., Bowen, B. W. & De Couet, H. G. Global phylogeography of the scalloped hammerhead shark (Sphyrna lewini). Mol. Ecol. 15, 2239–2251, https://doi.org/10.1111/j.1365-294X.2006.02933.x (2006).
Keeney, D. B. & Heist, E. J. Worldwide phylogeography of the blacktip shark (Carcharhinus limbatus) inferred from mitochondrial DNA reveals isolation of western Atlantic populations coupled with recent Pacific dispersal. Mol. Ecol. 15, 3669–3679, https://doi.org/10.1111/j.1365-294X.2006.03036.x (2006).
Briggs, J. C. Marine zoogeography. McGraw-Hill, New York (1974).
Martin, A. P. Substitution rates of organelle and nuclear genes in sharks: Implicating metabolic rate (again). Mol. Biol. Evol. 16, 996–1002, https://doi.org/10.1093/oxfordjournals.molbev.a026189 (1999).
Hobbs, J. P. A., Frisch, A. J., Allen, G. R. & Van Herwerden, L. Marine hybrid hotspot at Indo-Pacific biogeographic border. Biol. Lett. 5, 258–261, https://doi.org/10.1098/rsbl.2008.0561 (2009).
Planes, S. & Fauvelot, C. Isolation by distance and vicariance drive genetic structure of a coral reef fish in the Pacific Ocean. Evolution 56, 378–399 (2002).
Hoareau, T. B., Boissin, E. & Berrebi, P. Evolutionary history of a widespread Indo-Pacific goby: The role of Pleistocene sea-level changes on demographic contraction/expansion dynamics. Mol. Phylogenet. Evol. 62, 566–572, https://doi.org/10.1016/j.ympev.2011.10.004 (2012).
Schultz, J. K. et al. Global phylogeography and seascape genetics of the lemon sharks (genus Negaprion). Mol. Ecol. 17, 5336–5348, https://doi.org/10.1111/j.1365-294X.2008.04000.x (2008).
Bernardi, G., Holbrook, S. J. & Schmitt, R. J. Gene flow at three spatial scales in a coral reef fish, the three-spot dascyllus, Dascyllus trimaculatus. Mar. Biol. 138, 457–465, https://doi.org/10.1007/s002270000484 (2001).
Eble, J. A., Rocha, L. A., Craig, M. T. & Bowen, B. W. Not all larvae stay close to home: insights into marine population connectivity with a focus on the brown surgeonfish (Acanthurus nigrofuscus). Journal of marine biology 2011, article ID 518516, 12p., https://doi.org/10.1155/2011/518516 (2011).
Tabata, S. The general circulation of the Pacific Ocean and a brief account of the oceanographic structure of the North Pacific Ocean. Part I - circulation and volume transports. Atmosphere 13, 133–168 (1975).
Martinez, E., Ganachaud, A., Lefevre, J. & Maamaatuaiahutapu, K., Central South Pacific thermocline water circulation from a high‐resolution ocean model validated against satellite data: Seasonal variability and El Niño 1997–1998 influence. Journal of Geophysical Research, https://doi.org/10.1029/2008JC004824 (2009).
Vignaud, T., Clua, E., Mourier, J., Maynard, J. & Planes, S. Microsatellite analyses of blacktip reef sharks (Carcharhinus melanopterus) in a fragmented environment show structured clusters. PLoS One 8, 8, https://doi.org/10.1371/journal.pone.0061067 (2013).
Johri, S., Doane, M. P., Allen, L. & Dinsdale, E. A. Taking Advantage of the Genomics Revolution for Monitoring and Conservation of Chondrichthyan Populations. Diversity 11, 49, https://doi.org/10.3390/d11040049 (2019).
Baker, C. S. et al. Population structure of nuclear and mitochondrial DNA variation among humpback whales in the North Pacific. Mol. Ecol. 7, 695–707, https://doi.org/10.1046/j.1365-294x.1998.00384.x (1998).
Wallace, B. P. et al. Global conservation priorities for marine turtles. PLoS One 6, 14, https://doi.org/10.1371/journal.pone.0024510 (2011).
Juhel, J. B. et al. Isolation and no-entry marine reserves mitigate anthropogenic impacts on grey reef shark behavior. Sci Rep 9, 11, https://doi.org/10.1038/s41598-018-37145-x (2019).
Gaither, M. R., Toonen, R. J., Robertson, D. R., Planes, S. & Bowen, B. W. Genetic evaluation of marine biogeographical barriers: perspectives from two widespread Indo-Pacific snappers (Lutjanus kasmira and Lutjanus fulvus). J. Biogeogr. 37, 133–147, https://doi.org/10.1111/j.1365-2699.2009.02188.x (2010).
Mourier, J. et al. Extreme inverted trophic pyramid of reef sharks supported by spawning groupers. Curr. Biol. 26, 2011–2016, https://doi.org/10.1016/j.cub.2016.05.058 (2016).
Boissin, E., Neglia, V., Boulet Colomb D’hauterserre, F., Tatarata, M. & Planes, S. Evolutionary history of green turtle populations, Chelonia mydas, from French Polynesia highlights the putative existence of a glacial refugium. Mar. Biodivers. https://doi.org/10.1007/s12526-019-01001-6 (in press).
Jensen, M. P. et al. The evolutionary history and global phylogeography of the green turtle (Chelonia mydas). J. Biogeogr. 46, 860–870, https://doi.org/10.1111/jbi.13483 (2019).
van Hooidonk, R., Maynard, J. A. & Planes, S. Temporary refugia for coral reefs in a warming world. Nat. Clim. Chang. 3, 508–511, https://doi.org/10.1038/nclimate1829 (2013).
Freeman, L. A. Robust performance of marginal Pacific coral reef habitats in future climate scenarios. PLoS One 10, 16, https://doi.org/10.1371/journal.pone.0128875 (2015).
Momigliano, P., Robbins, W. D., Gardner, M. & Stow, A. Characterisation of 15 novel microsatellite loci for the grey reef shark (Carcharhinus amblyrhynchos). Conserv. Genet. Resour. 6, 661–663, https://doi.org/10.1007/s12686-014-0174-z (2014).
Van Oosterhout, C., Hutchinson, W. F., Wills, D. P. M. & Shipley, P. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538, https://doi.org/10.1111/j.1471-8286.2004.00684.x (2004).
Belkhir, K., Borsa, P., Chikhi, L., Raufaste, N. & Bonhomme, F. GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations, Laboratoire Génome, Populations, Interactions, CNRS UMR 5000, Université de Montpellier II, Montpellier (France). (1996–2004).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28, 2537–2539, https://doi.org/10.1093/bioinformatics/bts460 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate – a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B-Methodol. 57, 289–300 (1995).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332, https://doi.org/10.1111/j.1755-0998.2009.02591.x (2009).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620, https://doi.org/10.1111/j.1365-294X.2005.02553.x (2005).
Earl, D. A. & Vonholdt, B. M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361, https://doi.org/10.1007/s12686-011-9548-7 (2012).
Rousset, F. Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145, 1219–1228 (1997).
Acknowledgements
All of the following provided funding for the research presented here (no particular order after the first organization): Laboratoire d’Excellence CORAIL, Ministère de l’Ecologie du Développement Durable et de l’Energie, Ministère de l’Outre-Mer, Fonds Pacifique, IFRECOR, Délégation à la recherche de Polynésie, Agence Nationale de la Recherche and the Robertson Foundation. We also thank Andrew Chin, Jennifer Ovenden, Mark Meekan and Conrad Speed, Mael Imirizaldu, David Lecchini, Patrick Plantard, Jonathan Werry, Johann Mourier, Thomas Vignaud, Matis Jorge, Noémie Jublier and several other students for providing samples or for assistance with sampling. We are grateful to three anonymous reviewers who provided helpful comments.
Author information
Authors and Affiliations
Contributions
S.P. and E.B. designed the study. E.C., S.P., S.R.T., C.D.B. performed the sampling. Y.Z. performed the laboratory work. E.B. supervised the laboratory work, analyzed the data and drafted the manuscript. All co-authors read and improved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boissin, E., Thorrold, S.R., Braun, C.D. et al. Contrasting global, regional and local patterns of genetic structure in gray reef shark populations from the Indo-Pacific region. Sci Rep 9, 15816 (2019). https://doi.org/10.1038/s41598-019-52221-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52221-6
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.