Abstract
While the link between diet-induced changes in gut microbiota and lipid metabolism in metabolic syndrome (MetS) has been established, the contribution of host genetics is rather unexplored. As several findings suggested a role for the lysosomal lipid transporter Niemann-Pick type C1 (NPC1) in macrophages during MetS, we here explored whether a hematopoietic Npc1 mutation, induced via bone marrow transplantation, influences gut microbiota composition in low-density lipoprotein receptor knockout (Ldlr−/−) mice fed a high-fat, high-cholesterol (HFC) diet for 12 weeks. Ldlr−/− mice fed a HFC diet mimic a human plasma lipoprotein profile and show features of MetS, providing a model to explore the role of host genetics on gut microbiota under MetS conditions. Fecal samples were used to profile the microbial composition by 16 s ribosomal RNA gene sequencing. The hematopoietic Npc1 mutation shifted the gut microbiota composition and increased microbial richness and diversity. Variations in plasma lipid levels correlated with microbial diversity and richness as well as with several bacterial genera. This study suggests that host genetic influences on lipid metabolism affect the gut microbiome under MetS conditions. Future research investigating the role of host genetics on gut microbiota might therefore lead to identification of diagnostic and therapeutic targets for MetS.
Similar content being viewed by others
Introduction
Metabolic syndrome (MetS) is a complex disorder that identifies centrally obese patients at increased risk for developing cardiovascular disease (CVD) and diabetes mellitus type 21,2. Due to the continuous increased prevalence of obesity, MetS is considered a global epidemic, putting an enormous pressure on health care services3. Next to insulin resistance and hypertension, another central abnormality observed within MetS are disturbances in lipid metabolism, resulting in dyslipidemia1,4. Dyslipidemia is characterized by increased plasma triglyceride-rich lipoproteins, decreased plasma high-density lipoprotein (HDL) and increased plasma low-density lipoprotein (LDL), leading to MetS-associated pathologies4.
Next to disturbances in plasma lipid levels, alterations in intracellular lipid metabolism have also been observed in MetS. While mutations in the gene encoding for the lysosomal membrane protein Niemann-Pick type C1 (NPC1) are well-known to induce a rare lysosomal storage disease, recent findings have also linked a dysfunctional NPC1 protein to the development of obesity and MetS5,6. Specifically, heterozygous Npc1 mutation carriers showed a 4.8 higher risk for developing morbid adult obesity7. Moreover, identification of several Npc1 small nucleotide polymorphism (SNPs) via genome-wide association5 and confirmation studies8,9,10 have further highlighted the link between Npc1 variants (allele frequency ~38%) and obesity in the European population. Mechanistically, a dysfunctional NPC1 protein leads to lysosomal lipid accumulation, thereby disturbing lipid metabolism11. Likewise, the phenomenon of lipid accumulation within the endo-lysosomal compartment of cells has also been observed in pathologies associated with MetS12. While lysosomal dysfunction in proximal tubular cells contributed to obesity-related kidney diseases13, lysosomal lipid storage within macrophages is a well-described mechanism disturbing general lipid metabolism and results in the induction of a chronic low-grade inflammatory response in non-alcoholic steatohepatitis14,15 and atherosclerosis12,16,17,18. These findings imply a role for macrophage NPC1 in disturbing lipid homeostasis in MetS.
Relevantly, several studies have shown a link between the composition of the gut microbiota and lipid parameters such as LDL, triglyceride and HDL levels19,20,21,22,23. Moreover, modulating the gut microbiota by transferring the microbiota of lean donors has been shown to improve MetS in patients24, indicating the gut microbiota as a promising target to tackle metabolic disorders. However, the underlying mechanisms through which the gut microbiota modulates host’s metabolism remain largely elusive25. Gaining knowledge into the exact interactions between environment and host genetics on gut microbiota and associated pathophysiological processes is therefore essential to improve our diagnostic and therapeutic approaches to tackle metabolic pathologies.
The current study investigated whether a hematopoietic Npc1 mutation influences the gut microbiota in low-density lipoprotein receptor knockout (Ldlr−/−) mice fed a high-fat, high-cholesterol (HFC) diet for 12 weeks. Ldlr−/− mice fed a HFC diet mimic a human plasma lipoprotein profile and show features of MetS26, providing a model to explore the role of host genetics on gut microbiota under MetS conditions. For this purpose, Ldlr−/− mice received bone marrow from Niemann-Pick type C1 mutant (Npc1mut) mice. This study suggests that host genetic influences related to lipid metabolism shift the gut microbiota composition and increases microbial richness and diversity under MetS conditions. Variations in plasma lipid levels correlated with microbial diversity and richness as well as with several bacterial genera. Future research investigating the role of host genetics on gut microbiota can therefore lead to identification of novel diagnostic and therapeutic targets for MetS.
Results
Hematopoietic Npc1 mutation disturbs intracellular and circulating lipid levels
To confirm successfulness of the bone marrow transplantation, intracellular and circulating lipid levels were measured in Npc1mut-tp Ldlr−/− mice. Subjecting Npc1mut-tp Ldlr−/− mice to a HFC diet for 12 weeks resulted in a strong increase in hepatic cholesterol levels, while a reduction was observed for hepatic triglyceride levels (Table 1). Moreover, due to the hematopoietic Npc1 mutation, hepatic lipids accumulated in the lysosomal fraction of hepatic macrophages (Fig. 1A,B and Supplementary Figs S1 and S2). Additionally, plasma cholesterol, triglyceride and free fatty acid levels decreased significantly, indicating a decrease of circulating lipids in Npc1mut-tp Ldlr−/− mice (Table 1). In line with these data, hepatic gene expression levels of ATP-binding cassette sub-family G member 1(Abcg1), Niemann-Pick type C2 (Npc2) and ATP-binding cassette transporter A1 (Abca1) increased in Npc1mut-tp Ldlr−/− mice. In spite of increased hepatic cholesterol levels (Table 1), these observations indicate increased hepatic cholesterol efflux signaling, which is likely a compensation mechanism to overcome the accumulation of cholesterol in lysosomes. Overall, these data show that a hematopoietic Npc1 mutation induces lysosomal accumulation of lipids inside liver macrophages and disturbs general lipid metabolism in Ldlr−/− mice on a HFC diet.
Hepatic phenotype of HFC-fed Ldlr−/− mice carrying a hematopoietic Npc1 mutation. (A) Representative images of hepatic staining for hematoxylin and eosin (H&E), showing lysosomal lipid accumulation and granuloma formation in Npc1mut-transplanted Ldlr−/− mice. The left panel shows a normal view of the liver under high cholesterol conditions, with the green dots indicating lipid droplets. The right panel show clustering of foamy macrophages, referred to as granulomas (indicated by the asterisks). (B) Representative electron microscopy image of hepatic macrophages carrying a hematopoietic Npc1 mutation in HFC-fed Ldlr−/− mice, confirming the massive accumulation of lipids. The left panel shows a single Kupffer cell (K), surrounded by 2 hepatocytes (Hep.). The right panel shows the granuloma (surrounded by the dashed line), which is surrounded by Kupffer cells (K).
Microbial richness and diversity are increased and linked to lipid metabolism
To investigate the influence on microbial richness and diversity, α-diversity metrics were calculated. Both the observed microbial richness (observed number of OTUs) as well as the estimated richness as indicated by the Chao 1 index, was significantly higher in Ldlr−/− mice carrying the hematopoietic Npc1 mutation (Fig. 2A,B). Likewise, microbial biodiversity, measured via the Shannon index (Fig. 2C) was significantly increased in Npc1mut-tp mice. To further define which variables are linked to microbial richness and diversity in this mouse model, Spearman correlation analysis was performed on all variables and the Chao 1 (richness) or Shannon (biodiversity) index. Plasma free fatty acids levels were negatively correlated with microbial richness (r = −0.47; p = 0.03) (Fig. 2D). Concomitantly, plasma total cholesterol levels showed the same inverse correlation with microbial richness (r = −0.41; p = 0.07) (Fig. 2E), though this correlation did not reach statistical significance. Furthermore, biodiversity correlated positively with hepatic gene expression levels of the intracellular cholesterol transporter Niemann-Pick type C2 (Npc2) (r = 0.47; p = 0.04) (Fig. 2F) and negatively with plasma total cholesterol levels (r = −0.56; p = 0.007) (Fig. 2G). Together, these data demonstrate that a hematopoietic Npc1 mutation in Ldlr−/− mice increases microbial richness and biodiversity, which is correlated to changes in lipid metabolism.
Alpha diversity metrics in Ldlr−/− mice on HFC diet carrying a hematopoietic Npc1 mutation compared to wildtype. (A–C) The number of OTUs (A), Chao 1 index (B) and Shannon index (C) indicate increased diversity in Npc1mut-transplanted Ldlr−/− mice. (D–G) Spearman correlation between alpha diversity metrics and lipid parameters indicate a link between lipid metabolism and gut microbiota diversity. *Indicates p < 0.05. All errors are SEM.
The microbial community structure is related to the hematopoietic Npc1 mutation but the latter is not linked to enterotypes
The dissimilarity in the microbial community composition (beta-diversity) of stool samples was assessed using the Bray-Curtis (BC) dissimilarities. Principal coordinate analysis (PCoA) based on Bray-Curtis dissimilarities showed that samples could be partly separated based upon the hematopoietic Npc1 mutation (Fig. 3A). Moreover, the within-group distance in microbial community structure (i.e. average Bray-Curtis dissimilarity between samples from the same treatment group) was significantly smaller than the between-group distance, indicating that the microbiota composition of animals within the same treatment group is more similar compared to animals from different treatment groups (Fig. 3B). Enterotype analyses revealed two enterotypes driven by Allobaculum, Lactobacillus and S24-7 (Fig. 3C). However, the clustering was not associated with the bone marrow-specific Npc1 mutation.
Principal Coordinate Analyses (PCoAs) based on Bray-Curtis distance and assortment of gut microbial communities into enterotypes for HFC-fed Ldlr−/− mice carrying a hematopoietic Npc1wt or Npc1mut mutation. (A) PCoA-plot based on unweighted UniFrac metrics for all samples showed that samples could be partly separated based upon the hematopoietic Npc1 mutation. (B) Bray-Curtis distances were calculated within Npc1wt-transplanted and Npc1mut-transplanted HFC-fed Ldlr−/− mice (average pairwise distance in microbiota composition between samples of the same experimental group) and between Npc1wt -transplanted and Npc1mut -transplanted HFC-fed Ldlr−/− mice (average pairwise distance between samples of different experimental groups), confirming the separation based on the hematopoietic Npc1 mutation. (C) Between-class analysis visualizing the two distinct enterotypes that could be identified based upon PAM clustering of Bray-Curtis distances, with closed dots (Npc1mut) and squares (Npc1wt) representing individual mice and numbered white rectangles marking the center of each enterotype. Enterotypes were mainly driven by differential abundances in Allobaculum, Lactobacillus and S24-7, however the clustering was not associated with the bone marrow-specific Npc1 mutation. *Indicates p < 0.05.
To determine whether the hematopoietic Npc1 mutation or other variables had an additional effect on the gut microbiota community structure, we subsequently performed distance-based redundancy analysis (dbRDA); a constrained ordination technique. Constrained ordination techniques attempt to explain differences in microbial composition between samples by differences in explanatory variables (e.g. hematopoietic Npc1 mutation). In db-RDA, the information from explanatory variables is combined with the eigenvalues obtained from the PCoA. Permutational multivariate analysis of variance (PERMANOVA) was used to determine significant dissimilarity within bacterial communities and multiple explanatory variables. The hematopoietic Npc1 mutation (p = 0.034, explained variance = 18.9%) as well as hepatic gene expression levels of the cholesterol transporter Abcg1 (p = 0.041, explained variance = 20.8%) showed a relationship with the microbial community structure (Fig. 4A,B), whereas Abca1, Npc2, plasma total cholesterol, plasma total triglycerides, plasma free fatty acids, plasma cupper oxLDL, plasma EO6, cage number, liver total cholesterol and liver total triglycerides did not.
Distance-based Redundancy Analysis (db-RDA) showing the relationship of the hematopoietic deficiency for Npc1 and Abcg1 to the microbial community structure. The plots represent a dbRDA ordination based upon the Bray-Curtis distance including (A) Npc1wt- and Npc1mut -transplanted Ldlr−/− mice with the hematopoietic Npc1 mutation as explanatory variable. This panel confirms the separation of the microbiota based on the hematopoietic Npc1 mutation. (B) Plasma cholesterol, plasma triglycerides, liver triglycerides, plasma free fatty acids, cage number, liver cholesterol, plasma EO6, plasma CuOxLDL and hepatic gene expression of Abcg1, Npc2, Npc1 and Abca1 as explanatory variables and samples colored to hepatic expression of Abcg1. Multivariate PERMANOVA analysis including all explanatory variables demonstrated that both Npc1 and Abcg1 were significantly associated with the microbial community structure, while all other explanatory variables were not when mutually adjusting for each other. This plot further confirms that increased Npc1 and Abcg1 expression levels are associated with similar changes in the microbial community structure. MDS1 represents the unconstrained axis. CAP1 and CAP2 represent, respectively, the first and second constrained axes used in the CAP (canonical analysis of principal coordinates).
Hematopoietic Npc1 mutation shifts gut microbiota composition in HFC-fed Ldlr −/− mice
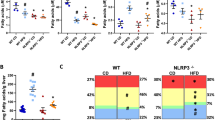
Based on 16S rRNA gene sequencing, the dominant phyla/genera were selected for further analysis, leading to in depth analysis of 4 phyla and 44 genera (Fig. 5A,B). Firmicutes, Bacteroidetes and, to a lesser extent, Actinobacteria and Proteobacteria were the dominant observed phyla (Fig. 5A). The relative abundance of the bacterial phyla was not significantly different between Npc1wt- and Npc1mut-tp mice (Fig. 5A), however, within the Firmicutes phylum significant differences were observed in the relative abundance of several bacterial genera. Specifically, Staphylococcus spp. (p = 0.008; q = 0.34) and unclassified Mogibacteriaceae spp. (p = 0.008; q = 0.34) showed significantly higher levels in relative abundance in the Npc1mut-tp mice compared to Npc1wt-tp mice, whereas the abundance of Allobaculum spp. (p = 0.01, q = 0.34) was significantly lower (Fig. 5B–E). Relevantly, the unclassified Mogibacteriaceae spp. correlated with hepatic gene expression levels of Abcg1 (r = 0.57; p = 0.009) (Fig. 5F) and plasma total triglyceride levels (r = −0.41; p = 0.07), though the latter association was not statistically significant (Fig. 5G). Furthermore, the genus Allobaculum spp. showed a positive correlation with plasma free fatty acid levels (r = 0.52; p = 0.02) (Fig. 5H). Thus, a hematopoietic Npc1 mutation alters the composition of gut microbiota in HFC-fed Ldlr−/− mice, which are correlated with changes in lipid metabolism.
Taxonomy at phylum and genus level for Npc1wt- and Npc1mut -transplanted HFC-fed Ldlr−/− mice. (A,B) Overview of the mean relative abundance at phylum level. (C) An overview of the mean relative abundance at genus level is given. Significant differences between the groups at genus level are presented in separate box-whiskers for Staphylococcus spp. (D), unclassified Mogibacteriaceae spp. (E) and Allobaculum spp. (F). (G–I) Spearman correlation between lipid parameters and relative abundances of unclassified Mogibacteriaceae spp. and Allobaculum spp. *Indicates p < 0.05 and ** p < 0.01.
Discussion
Whereas increasing evidence has linked diet-induced changes in gut microbiota to perturbations in lipid metabolism in MetS, the contribution of host genetics to this relation is rather unexplored. This study suggests that host genetic influences on lipid metabolism such as a hematopoietic Npc1 mutation affect the gut microbiome under MetS conditions.
Though Npc1 was previously identified in a human GWAS for obesity5, our study is the first to imply its direct impact on the gut microbiome under MetS conditions. This observation suggests that host-derived genetic disturbances in lipid metabolism can impact the gut microbiome. Indeed, an increasing amount of reports have established the capacity of the host’s genetic profile to shape the gut microbiome, thereby supporting our findings27,28. Also, the observation that a hematopoietic Npc1 mutation impacts the gut microbiota under MetS conditions confirms the previously established link between lipid metabolism and gut microbiota composition. However, in contrast to genetic manipulation of lipid metabolism, the link between lipid metabolism and gut microbiota has mainly been established based on studies wherein dietary components (i.e. high fat diet, prebiotics) altered lipid metabolism via influences on gut microbiota29,30. Therefore, in addition to gut microbiota-mediated influences of dietary compounds on lipid metabolism, our study implies that also host genetic factors that influence lipid metabolism can impact the gut microbiome. However, as we employed a HFC diet to both experimental conditions, the observed impact of the gut microbiota of the Npc1 mutation is likely also influenced by this diet. Indeed, previous reports have indicated that a high fat diet influences the composition of the gut microbiota31,32. Therefore, in spite of the observed changes in gut microbiota composition, to incontrovertibly demonstrate that the hematopoietic Npc1 mutation influences gut microbiota composition, it is essential to investigate the gut microbiota of Npc1mut–tp Ldlr−/− mice receiving a normal chow diet.
Our results also shed new light on the association between overall health status and the gut bacterial richness and diversity. Here, under HFC conditions, induction of the pathological hematopoietic Npc1 mutation elevated bacterial richness and diversity. However, this is opposed to the concept that increased bacterial richness and diversity reflect ecosystem stability and resilience and is associated with overall health33,34,35,36,37. Yet, our apparent discordant finding on the association between disease and richness/diversity has also been observed by Kasai et al. who observed increased levels of bacterial diversity in obese compared to lean subjects38. Moreover, Vandeputte et al. associated stool consistency, potentially due to long colonic transit time39,40, with higher gut bacterial richness. Therefore, the increased bacterial richness and diversity observed in the pathological experimental group in the current study might have been influenced by stool consistency, hampering the view on bacterial richness and diversity as a marker for overall health status. Of note, while it is generally accepted that reduced plasma lipids are beneficial for health41, in the current model the reduced plasma lipids are a consequence of the accumulation of lipids inside macrophages (due to the Npc1 mutation). Therefore, the observed reduction in plasma lipids in our model is not considered beneficial for the health status, as these mice develop inflammation in the liver42 and arteries43.
Among the shifted gut bacteria after hematopoietic Npc1 mutation, we identified reduced relative abundances of the genus Allobaculum spp. Several studies have previously shown that Allobaculum spp. abundance is reduced in response to dietary fat intake44,45 and correlates with variations in plasma HDL concentrations46 and body weight47. In line, hepatic Abcg1 and Abca1 expression was increased in Npc1mut-tp mice, suggesting changes in lipid metabolism via lysosomal lipid-mediated activation of the transcription factor liver X receptor48,49. Therefore, our finding is consistent with the proposed view that Allobaculum spp. can be considered beneficial for the physiology of the host29. Next, in a prospective cross-sectional study, Staphylococcus spp. abundance was positively correlated with energy intake in obese children50. Here, we found an increased abundance of Staphylococcus spp. in HFC-fed Ldlr−/− mice carrying the hematopoietic Npc1 mutation. As such, Staphylococcus spp. might be involved in regulating the host’s lipid metabolism. Finally, increased relative abundances were observed of unclassified Mogibacteriaceae spp. in this study. Relevantly, a recent study by Harach et al. correlated levels of unclassified Mogibacteriaceae spp. with amyloid beta 42 levels in the brain of mice suffering from Alzheimer’s disease51. As multiple reports have previously linked a dysfunctional NPC1 protein to Alzheimer’s disease52,53,54, relative abundance changes in unclassified Mogibacteriaceae spp. might be directly related to mutations in Npc1 and deserve further investigation. Of note, to ensure the causality between mutations in Npc1 and changes in gut microbiota composition, host responses should also be investigated in the future by means of fecal transplantation experiments. As such, the results in the current study do not provide information on causality, but are rather of associative nature.
Overall, this study suggests the impact of host genetic influences on lipid metabolism such as a hematopoietic Npc1 mutation on the gut microbiome in the context of MetS, confirming the link between lipid metabolism and gut microbiota. In addition, in contrast to previous studies, our findings suggest that increased microbial richness and diversity do not necessarily associate with improved health. Despite the significant associations between microbial richness and diversity and several lipid parameters, these Spearman correlations were rather low, potentially due to low sample size. As such, further research is necessary to confirm these associations. Finally, future research investigating the role of host genetics on gut microbiota is therefore warranted as it will increase our understanding of the function of the gut microbiome in human physiology and pathology.
Methods
Experimental design
Niemann-Pick type C1nih mutant (Npc1mut) mice (a kind gift from Prof. Dr. Lieberman from University of Michigan Medical School) were backcrossed into a C57BL/6J background for more than 10 generations. Npc1mut and Ldlr−/− mice were housed under standard conditions and had access to food and water ad libitum. Experiments were performed according to Dutch regulations and approved by the Committee for Animal Welfare of Maastricht University (approval number DEC 2011-133)42,43. Specifically, mice were housed in individually ventilated cages (IVC) in a conventional specified pathogenic free (SPF) facility where a 12 hr day/night cycle was handled using a dusk transition phase. Room temperature was kept constant at 21 °C. Corncob bedding material was used in the cages and animals were fed 10 mm food pellets that were irradiated with 25 kGray.
To generate myeloid Npc1mut deficient Ldlr−/− mice, a bone marrow transplantation was performed. Twenty-two week-old female Ldlr−/− mice received one week before and four weeks after irradiation antibiotic water containing 100 mg/l neomycin (Gibco, Breda, the Netherlands) and 6 * 104 U/l polymycin B Sulphate (Gibco, Breda, the Netherlands). One day before and on the day of the transplantation, Ldlr−/− mice were lethally irradiated with 6 Gray of γ-radiation in the morning, thus receiving 12 Gray in total. Lethally irradiated Ldlr−/− mice were then injected with 1 * 107 bone marrow cells donated from either Npc1mut mice or wildtype littermate controls (Npc1wt). In order to fully ensure bone marrow replacement, mice had a nine-week recovery period. After nine weeks of recovery, transplanted (-tp) mice received an HFC diet, containing 21% butter, 0.2% cholesterol, 46% carbohydrates and 17% casein (diet 1635; Scientific Animal Food and Engineering, Villemoissonsur-Orge, France) for 12 weeks. We further highlight that the same mouse model as well as the described lipid measurements were used in previously published manuscripts, where administration of dietary cholesterol was essential to induce lysosomal lipid-induced inflammation42,43,55. As we anticipated only subtle changes56 in the gut microbiome due to the Npc1 mutation being restricted to the hematopoietic line, we did not cohouse both genotypes. Though the influence of cage-effects cannot be completely excluded, the following measures were taken to minimize the impact of housing conditions: the litters were separated per genotype after weaning, mice of similar age were used, the entire experiment was performed at once (so not over different time points), food and autoclaving parameters were standardized, only 1 person handled the mice throughout the entire experiment, mice were kept in cages in the same room, in the same rack and finally, mice of each genotype were put in four different cages. Finally, the between-genotype Bray-Curtis dissimilarity was significantly increased to the between-cage Bray-Curtis dissimilarity ((p = 0.03 compared to WT cages and p = 0.04 compared to Npc1mut cages), providing further evidence that the observed effects in the current study are at least partly due to the Npc1 mutation. Liver tissue was isolated and snap-frozen in liquid nitrogen and stored at −80 °C or fixed in 4% formaldehyde/PBS. The biochemical determination of plasma cholesterol and liver triglyceride levels, electron microscopy, RNA isolation, complementary DNA synthesis, quantitative polymerase chain reaction are described extensively42,43,57. Liver cholesterol levels were quantified as described previously58.
Sample collection and processing
Mice were housed in groups of 4 in separate cages, which were kept in the same room throughout the study. Fecal samples were snap-frozen with liquid nitrogen and stored at −80 °C. DNA isolation was done using the FavorPrep Stool DNA Isolation Mini Kit (FASTI 001-1) according to manufacturer’s protocol.
Sequencing
Amplicon libraries and sequencing was performed according to previously published protocols19,59,60. Briefly, the hypervariable region V4 of the 16S rRNA gene was PCR amplified from each DNA sample using forward primer 515 F and reverse primer 806 R as described previously19. Illumina MiSeq paired end sequencing of equimorlarly pooled amplicons was subsequently used to determine the bacterial composition of fecal samples. Custom scripts were used to remove primer sequences, align paired end reads and quality-filter the sequencing reads. Details can be found in Fu et al.19 and Bonder et al.59. Sequencing data of all samples have been deposited in the European Bioinformatics Institute (EBI) database under primary accession code (ENA) PRJEB31646 and secondary accession code ERP114028.
Data analysis and statistics
Sequences were clustered into Operational Taxonomic Units (OTUs) using UCLUST (version 1.2.22q) at 97% similarity. For taxonomic assignments of OTUs, a TaxMan61 filtered and truncated version of the full Greengenes reference database version 13.5 was used. OTUs that did not cluster against any of the sequences in the reference database were suppressed.
A total of 1,503,804 paired-end sequences, with a sequencing depth ranging from 15,637–117,585 reads/sample, were clustered in a total of 13,312 OTUs. To further reduce spurious OTUs and normalize for sequencing depth, data were rarefied to 10,000 reads/sample and OTUs containing less than 5 reads or occurring in only a single sample were discarded, resulting in a final number of 2.695 OTUs. Downstream analysis were conducted in QIIME version 1.962 and R version 3.1.3.
Microbial data-analyses and statistics
Taxonomic composition
Differences in the relative abundance of bacterial phyla and genera between Npc1wt-tp and Npc1mut-tp Ldlr−/− mice were compared using the non-parametric t-test Metastats while controlling for multiple comparisons by means of the False Discovery Rate (q = 0.15; 750 permutations). Relations between bacterial phyla/genera and other continuous variables were analyzed via Spearman correlation using GraphPad Prism, version 6.0 for Windows. Though correlations between lipid parameters and gut microbiota abundances were calculated on pooled data to increase the power, it was verified whether both groups showed the same effect.
Microbial richness & diversity and microbial community structure
The following metrics of species richness and diversity within communities (alpha-diversity) were determined: observed OTUs (observed richness), Chao1 index (estimated richness), and Shannon diversity index. Alpha diversity metrics between the wild-type and mutant animals were compared by Mann Whitney tests using GraphPad Prism, version 6.0 for Windows. Beta-diversity, or diversity shared across samples was determined by the Bray-Curtis dissimilarity (BC) at a rarefaction depth of 10,000 seq/sample (Supplementary Fig. S3). Clustering of samples was visualized using Principal Coordinate analysis (PCoA) followed by distance-based redundancy analysis (db-RDA), a constrained extension of PCoA. Db-RDA was performed in R63 package vegan using the capscale function. To determine to what extend the hematopoietic Npc1 mutation contributed in explaining the microbial community structure, we used variance partitioning with distance-based redundancy analysis (db-RDA) of Bray-Curtis distances. In a subsequent db-RDA, the following data were additionally included as explanatory variables to examine whether they had a significant impact on the microbial community structure: liver cholesterol, liver triglycerides, plasma cholesterol, plasma triglycerides, plasma free fatty acids, plasma cupper oxLDL, plasma EO6, cage number and hepatic gene expression of Abcg1, Npc2 and Abca1.
Enterotyping analyses
Enterotype analyses were performed as described previously by Armougham and colleagues64. Bray-Curtis (BC) distances were calculated for the genus-level relative abundance profiles. We used the R63 package “vegan: Community ecology”, version 2.2-1 by Oksanen et al. from 2011 for calculating the Bray-Curtis distances65.
To cluster the samples based on these distance metrics, we used the Partitioning Around Medoids (PAM) clustering algorithm in the R package “Cluster analysis basics and extensions”, version 2.0.1 ed. by Maechler et al. from 201266. The optimal number of clusters was chosen based on Calinski-Harabasz (CH) index and validated by the silhouette index67 using the R package ‘clusterSim’.
An optimal number of 2 clusters was identified based on the CH index, which was confirmed by the silhouette index although clustering was weak (SI 0.28).
Subsequent visualization and identification of relevant taxa was conducted for analyses based on BC distance. Between-class analysis (BCA) was performed to plot the samples using the R package “Analysis of Ecological Data: Exploratory and Euclidean Methods in Environmental Sciences” version 1.7.2. by Dray et al. from 2015. The similarity percentage analysis (SIMPER)68 was used to identify taxa contributing to similarity within- and dissimilarity between groups. Similar analyses have also been described previously60.
References
Kassi, E., Pervanidou, P., Kaltsas, G. & Chrousos, G. Metabolic syndrome: definitions and controversies. BMC Med 9, 48, https://doi.org/10.1186/1741-7015-9-48 (2011).
Hotamisligil, G. S. Inflammation and metabolic disorders. Nature 444, 860–867, https://doi.org/10.1038/nature05485 (2006).
Boudreau, D. M. et al. Health care utilization and costs by metabolic syndrome risk factors. Metab Syndr Relat Disord 7, 305–314, https://doi.org/10.1089/met.2008.0070 (2009).
Ruotolo, G. & Howard, B. V. Dyslipidemia of the metabolic syndrome. Curr Cardiol Rep 4, 494–500 (2002).
Meyre, D. et al. Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nat Genet 41, 157–159, https://doi.org/10.1038/ng.301 (2009).
Jelinek, D., Castillo, J. J., Heidenreich, R. A. & Garver, W. S. The C57BL/6J Niemann-Pick C1 mouse model with decreased gene dosage is susceptible to increased weight gain when fed a high-fat diet: Confirmation of a gene-diet interaction. Gene 568, 112–113, https://doi.org/10.1016/j.gene.2015.05.025 (2015).
Liu, R. et al. Rare Loss-of-Function Variants in NPC1 Predispose to Human Obesity. Diabetes 66, 935–947, https://doi.org/10.2337/db16-0877 (2017).
Sandholt, C. H. et al. Studies of metabolic phenotypic correlates of 15 obesity associated gene variants. PLoS One 6, e23531, https://doi.org/10.1371/journal.pone.0023531 (2011).
Cotsapas, C. et al. Common body mass index-associated variants confer risk of extreme obesity. Hum Mol Genet 18, 3502–3507, https://doi.org/10.1093/hmg/ddp292 (2009).
Berndt, S. I. et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat Genet 45, 501–512, https://doi.org/10.1038/ng.2606 (2013).
Miller, W. L. & Bose, H. S. Early steps in steroidogenesis: intracellular cholesterol trafficking. J Lipid Res 52, 2111–2135, https://doi.org/10.1194/jlr.R016675 (2011).
Hendrikx, T., Walenbergh, S. M., Hofker, M. H. & Shiri-Sverdlov, R. Lysosomal cholesterol accumulation: driver on the road to inflammation during atherosclerosis and non-alcoholic steatohepatitis. Obes Rev 15, 424–433, https://doi.org/10.1111/obr.12159 (2014).
Yamamoto, T. et al. High-Fat Diet-Induced Lysosomal Dysfunction and Impaired Autophagic Flux Contribute to Lipotoxicity in the Kidney. J Am Soc Nephrol, https://doi.org/10.1681/ASN.2016070731 (2016).
Bieghs, V. et al. Specific immunization strategies against oxidized low-density lipoprotein: a novel way to reduce nonalcoholic steatohepatitis in mice. Hepatology 56, 894–903, https://doi.org/10.1002/hep.25660 (2012).
Ioannou, G. N. The Role of Cholesterol in the Pathogenesis of NASH. Trends Endocrinol Metab 27, 84–95, https://doi.org/10.1016/j.tem.2015.11.008 (2016).
Settembre, C. & Ballabio, A. Lysosome: regulator of lipid degradation pathways. Trends Cell Biol 24, 743–750, https://doi.org/10.1016/j.tcb.2014.06.006 (2014).
Weber, K. & Schilling, J. D. Lysosomes integrate metabolic-inflammatory cross-talk in primary macrophage inflammasome activation. J Biol Chem 289, 9158–9171, https://doi.org/10.1074/jbc.M113.531202 (2014).
Xu, X. et al. Lysosomal cholesterol accumulation in macrophages leading to coronary atherosclerosis in CD38(-/-) mice. J Cell Mol Med 20, 1001–1013, https://doi.org/10.1111/jcmm.12788 (2016).
Fu, J. et al. The Gut Microbiome Contributes to a Substantial Proportion of the Variation in Blood Lipids. Circ Res 117, 817–824, https://doi.org/10.1161/CIRCRESAHA.115.306807 (2015).
Caesar, R., Nygren, H., Oresic, M. & Backhed, F. Interaction between dietary lipids and gut microbiota regulates hepatic cholesterol metabolism. J Lipid Res 57, 474–481, https://doi.org/10.1194/jlr.M065847 (2016).
Caesar, R., Tremaroli, V., Kovatcheva-Datchary, P., Cani, P. D. & Backhed, F. Crosstalk between Gut Microbiota and Dietary Lipids Aggravates WAT Inflammation through TLR Signaling. Cell Metab 22, 658–668, https://doi.org/10.1016/j.cmet.2015.07.026 (2015).
Zhong, C. Y. et al. Microbiota prevents cholesterol loss from the body by regulating host gene expression in mice. Sci Rep 5, 10512, https://doi.org/10.1038/srep10512 (2015).
Zhao, L. The gut microbiota and obesity: from correlation to causality. Nat Rev Microbiol 11, 639–647, https://doi.org/10.1038/nrmicro3089 (2013).
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–916 e917, https://doi.org/10.1053/j.gastro.2012.06.031 (2012).
Sommer, F. & Backhed, F. The gut microbiota–masters of host development and physiology. Nat Rev Microbiol 11, 227–238, https://doi.org/10.1038/nrmicro2974 (2013).
Ishibashi, S. et al. Hypercholesterolemia in low density lipoprotein receptor knockout mice and its reversal by adenovirus-mediated gene delivery. J Clin Invest 92, 883–893, https://doi.org/10.1172/JCI116663 (1993).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799, https://doi.org/10.1016/j.cell.2014.09.053 (2014).
Bonder, M. J. et al. The effect of host genetics on the gut microbiome. Nat Genet 48, 1407–1412, https://doi.org/10.1038/ng.3663 (2016).
Everard, A. et al. Microbiome of prebiotic-treated mice reveals novel targets involved in host response during obesity. ISME J 8, 2116–2130, https://doi.org/10.1038/ismej.2014.45 (2014).
Turnbaugh, P. J., Backhed, F., Fulton, L. & Gordon, J. I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3, 213–223, https://doi.org/10.1016/j.chom.2008.02.015 (2008).
He, C. et al. High-Fat Diet Induces Dysbiosis of Gastric Microbiota Prior to Gut Microbiota in Association With Metabolic Disorders in Mice. Front Microbiol 9, 639, https://doi.org/10.3389/fmicb.2018.00639 (2018).
Brandsma, E. et al. A Proinflammatory Gut Microbiota Increases Systemic Inflammation and Accelerates Atherosclerosis. Circ Res 124, 94–100, https://doi.org/10.1161/CIRCRESAHA.118.313234 (2019).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638, https://doi.org/10.1126/science.1110591 (2005).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546, https://doi.org/10.1038/nature12506 (2013).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230, https://doi.org/10.1038/nature11550 (2012).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588, https://doi.org/10.1038/nature12480 (2013).
Mosca, A., Leclerc, M. & Hugot, J. P. Gut Microbiota Diversity and Human Diseases: Should We Reintroduce Key Predators in Our Ecosystem? Front Microbiol 7, 455, https://doi.org/10.3389/fmicb.2016.00455 (2016).
Kasai, C. et al. Comparison of the gut microbiota composition between obese and non-obese individuals in a Japanese population, as analyzed by terminal restriction fragment length polymorphism and next-generation sequencing. BMC Gastroenterol 15, 100, https://doi.org/10.1186/s12876-015-0330-2 (2015).
Vandeputte, D. et al. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 65, 57–62, https://doi.org/10.1136/gutjnl-2015-309618 (2016).
Roager, H. M. et al. Colonic transit time is related to bacterial metabolism and mucosal turnover in the gut. Nat Microbiol 1, 16093, https://doi.org/10.1038/nmicrobiol.2016.93 (2016).
Hardman, A. E. Physical activity, obesity and blood lipids. Int J Obes Relat Metab Disord 23(Suppl 3), S64–71 (1999).
Houben, T. et al. Blood-derived macrophages prone to accumulate lysosomal lipids trigger oxLDL-dependent murine hepatic inflammation. Sci Rep 7, 12550, https://doi.org/10.1038/s41598-017-13058-z (2017).
Jeurissen, M. L. J. et al. Prevention of oxLDL uptake leads to decreased atherosclerosis in hematopoietic NPC1-deficient Ldlr(-/-) mice. Atherosclerosis 255, 59–65, https://doi.org/10.1016/j.atherosclerosis.2016.10.038 (2016).
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57, 1470–1481, https://doi.org/10.2337/db07-1403 (2008).
Fleissner, C. K. et al. Absence of intestinal microbiota does not protect mice from diet-induced obesity. Br J Nutr 104, 919–929, https://doi.org/10.1017/S0007114510001303 (2010).
Martinez, I. et al. Diet-induced metabolic improvements in a hamster model of hypercholesterolemia are strongly linked to alterations of the gut microbiota. Appl Environ Microbiol 75, 4175–4184, https://doi.org/10.1128/AEM.00380-09 (2009).
Ravussin, Y. et al. Responses of gut microbiota to diet composition and weight loss in lean and obese mice. Obesity (Silver Spring) 20, 738–747, https://doi.org/10.1038/oby.2011.111 (2012).
Mony, V. K., Benjamin, S. & O’Rourke, E. J. A lysosome-centered view of nutrient homeostasis. Autophagy 12, 619–631, https://doi.org/10.1080/15548627.2016.1147671 (2016).
Venkateswaran, A. et al. Control of cellular cholesterol efflux by the nuclear oxysterol receptor LXR alpha. Proc Natl Acad Sci USA 97, 12097–12102, https://doi.org/10.1073/pnas.200367697 (2000).
Bervoets, L. et al. Differences in gut microbiota composition between obese and lean children: a cross-sectional study. Gut Pathog 5, 10, https://doi.org/10.1186/1757-4749-5-10 (2013).
Harach, T. et al. Reduction of Abeta amyloid pathology in APPPS1 transgenic mice in the absence of gut microbiota. Sci Rep 7, 41802, https://doi.org/10.1038/srep41802 (2017).
Cupidi, C. et al. Role of Niemann-Pick Type C Disease Mutations in Dementia. J Alzheimers Dis 55, 1249–1259, https://doi.org/10.3233/JAD-160214 (2017).
Malnar, M., Hecimovic, S., Mattsson, N. & Zetterberg, H. Bidirectional links between Alzheimer’s disease and Niemann-Pick type C disease. Neurobiol Dis 72 Pt A, 37–47, https://doi.org/10.1016/j.nbd.2014.05.033 (2014).
Kresojevic, N., Dobricic, V., Svetel, M. & Kostic, V. Mutations in Niemann Pick type C gene are risk factor for Alzheimer’s disease. Med Hypotheses 83, 559–562, https://doi.org/10.1016/j.mehy.2014.08.025 (2014).
Jeurissen, M. L. et al. Myeloid DLL4 Does Not Contribute to the Pathogenesis of Non-Alcoholic Steatohepatitis in Ldlr−/− Mice. PLoS One 11, e0167199, https://doi.org/10.1371/journal.pone.0167199 (2016).
Laukens, D., Brinkman, B. M., Raes, J., De Vos, M. & Vandenabeele, P. Heterogeneity of the gut microbiome in mice: guidelines for optimizing experimental design. FEMS Microbiol Rev 40, 117–132, https://doi.org/10.1093/femsre/fuv036 (2016).
Bieghs, V. et al. Role of scavenger receptor A and CD36 in diet-induced nonalcoholic steatohepatitis in hyperlipidemic mice. Gastroenterology 138, 2477–2486, 2486 e2471-2473, https://doi.org/10.1053/j.gastro.2010.02.051 (2010).
Lutjohann, D. et al. High doses of simvastatin, pravastatin, and cholesterol reduce brain cholesterol synthesis in guinea pigs. Steroids 69, 431–438, https://doi.org/10.1016/j.steroids.2004.03.012 (2004).
Bonder, M. J. et al. The influence of a short-term gluten-free diet on the human gut microbiome. Genome Med 8, 45, https://doi.org/10.1186/s13073-016-0295-y (2016).
Mack, I. et al. Weight gain in anorexia nervosa does not ameliorate the faecal microbiota, branched chain fatty acid profiles, and gastrointestinal complaints. Sci Rep 6, 26752, https://doi.org/10.1038/srep26752 (2016).
Brandt, B. W., Bonder, M. J., Huse, S. M. & Zaura, E. TaxMan: a server to trim rRNA reference databases and inspect taxonomic coverage. Nucleic Acids Res 40, W82–87, https://doi.org/10.1093/nar/gks418 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, V., Austria. URL, http://www.R-project.org/ (2013).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180, https://doi.org/10.1038/nature09944 (2011).
Jari Oksanen, F. G. B., Michael Friendly, Roeland Kindt, Pierre Legendre, Dan McGlinn, Peter R. Minchin, R. B. O’Hara, Gavin L. Simpson, Peter Solymos, M. Henry H. Stevens, Eduard Szoecs, Helene Wagner, https://cran.r-project.org/web/packages/vegan/ (2011).
Maechler, M., Rousseeuw, P., Struyf, A., Hubert, M., Hornik, K. Cluster analysis basics and extensions. R package version 2.0.1. (2012).
PJ, R. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. Journal of Computational and Applied Mathematics, 53–65 (1987).
Clarke, K. R. Nonparametric Multivariate Analyses of Changes in Community Structure. Australian Journal of Ecology 18, 117–143, https://doi.org/10.1111/j.1442-9993.1993.tb00438.x (1993).
Acknowledgements
This research was supported by the CVON IN-CONTROL grant (CVON 2012-03) and the Netherlands Organisation for Scientific Research (NWO) (Vidi 016.126.327, ASPASIA 015.008.043 and TKI 40-41200-98-9306 to R.S.S. and Vidi 864.13.013 to J.F.).
Author information
Authors and Affiliations
Contributions
T.Ho., J.Pe., D.P.K. and R.S.S. the conception and design of the study; T.Ho, Y.Ol, IAMdR acquisition of data; T.Ho., J.Pe., M.JB. and J.Fu. (statistical) analysis and interpretation of data; T.Ho., J.Pe., M.H.H. and R.S.S. drafting the manuscript. All authors were involved in revising the paper critically and gave final approval of the version to be submitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Houben, T., Penders, J., Oligschlaeger, Y. et al. Hematopoietic Npc1 mutation shifts gut microbiota composition in Ldlr−/− mice on a high-fat, high-cholesterol diet. Sci Rep 9, 14956 (2019). https://doi.org/10.1038/s41598-019-51525-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-51525-x
This article is cited by
-
Host genetic control of gut microbiome composition
Mammalian Genome (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.