Abstract
Stable isotope-resolved metabolomics (SIRM) provides information regarding the relative activity of numerous metabolic pathways and the contribution of nutrients to specific metabolite pools; however, SIRM experiments can be difficult to execute, and data interpretation is challenging. Furthermore, standardization of analytical procedures and workflows remain significant obstacles for widespread reproducibility. Here, we demonstrate the workflow of a typical SIRM experiment and suggest experimental controls and measures of cross-validation that improve data interpretation. Inhibitors of glycolysis and oxidative phosphorylation as well as mitochondrial uncouplers serve as pharmacological controls, which help define metabolic flux configurations that occur under well-controlled metabolic states. We demonstrate how such controls and time course labeling experiments improve confidence in metabolite assignments as well as delineate metabolic pathway relationships. Moreover, we demonstrate how radiolabeled tracers and extracellular flux analyses integrate with SIRM to improve data interpretation. Collectively, these results show how integration of flux methodologies and use of pharmacological controls increase confidence in SIRM data and provide new biological insights.
Similar content being viewed by others
Introduction
Metabolism underpins life. Cells take energy from their surroundings in the form of carbon- and nitrogen-containing compounds and return to their surroundings equal amounts of energy as heat and waste products. Although metabolism increases the level of exogenic disorder, it reduces local disorganization, thereby upholding cellular structure or enabling growth or adaptation to changing environmental conditions1. This requires joint regulation of catabolic and anabolic pathways, which must be fine-tuned to not only produce ATP but also to synthesize appropriate quantities of metabolites for synthesizing membranes, nucleic acids, and proteins. Although the interrelationships of metabolic pathways are clearly important to understand, it is difficult to assess how numerous pathways change in a biological sample.
Metabolomics evolved to address this issue. Although changes in metabolite levels can provide evidence of altered metabolic pathway activity, snapshot concentrations do not clearly resolve pathway flux. The incorporation of heavy isotope-labeled nutrient tracers, however, empowers metabolomics with the capacity to resolve differences in relative metabolic pathway flux and to assess the contribution of different nutrient sources to catabolic and anabolic pathways. For example, provision of 13C-labeled glucose (or other stably labeled nutrients) to cells, tissues, or organisms leads to time-dependent incorporation of 13C into metabolic intermediates or biosynthetic endproducts. The metabolite labeling patterns reflect shifts in the mass of metabolites and can be assigned as isotopologues or isotopomers. Assessed from such data are temporal differences in isotope enrichment, changes in the labeling pattern, or differences in the contribution of nutrients to a metabolite pool, which collectively provide intricate knowledge of the state of metabolism in cells or tissues2,3.
While useful, such stable isotope-resolved metabolomics (SIRM) experiments are not simple to perform. These experiments require knowledge and expertise in not only biochemistry and metabolism but in technical niches such as nuclear magnetic resonance (NMR), chromatography, and mass spectrometry (MS). In addition, the large amount of acquired data requires specialized software for spectral deconvolution, metabolite assignment, natural abundance removal, isotopologue assignment, normalization, and quantification. This means that expertise ranging from wet lab biology to in silico software development may be involved in the acquisition, assembly, and interpretation of stable isotope metabolomics data. In addition, the instrumentation and reagents are expensive, and the experiments in toto are time consuming. Therefore, optimization of experimental design and of communication between involved personnel is essential for obtaining valid data and interpreting results.
In this study, we demonstrate the importance of pharmacological controls for increasing confidence in metabolite assignments and data interpretation. The addition of specific inhibitors or activators of metabolic pathways permits cross-validation with other types of flux measurements and helps identify problems concerning metabolite detection or assignment. In particular, use of such controls facilitates constructive communication between technical personnel and biologists as well as assists in the identification of metabolic loci that could contribute to changes in cell phenotype. Metabolic flux configurations achieved using inhibitors or activators of key nodes of metabolism should allow for the development of an atlas of cell-specific metabolite enrichment patterns corresponding to a range of metabolic states, thereby improving data interpretation and increasing reproducibility within and between laboratories.
Results and Discussion
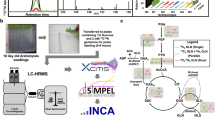
Stable isotope metabolomics experiments require knowledge of biology and metabolism, wet lab expertise, and expertise in mass spectrometry and bioinformatics (Fig. 1A). A list of terminology germane to stable isotope metabolomics is found in Table 1. Below, we discuss the major stages of a typical stable isotope metabolomics experiment.
General workflow and considerations for SIRM analyses. (A) Schematic illustrating the stages involved and expertise required in a typical SIRM experiment, including considerations at each stage; (B) Schematic illustrating assumptions regarding catabolic pathway and anabolic pathway flux; (C) Illustration of the dynamic and steady state isotopic labeling phases which must be considered in a typical SIRM experiment; and (D) Example of a 13C fractional enrichment plot of glutamate. Note that isotopologue analyses do not provide positional information of the isotopic atoms.
Stage 1 – Experimental design and execution
Biological expertise and knowledge of biochemistry are required for designing and executing SIRM experiments. This involves identifying tracer(s) relevant to the pathway(s) of interest and planning experiments to derive clean results, the latter of which should strive to minimize factors that obfuscate interpretation. Considerations for metabolic steady state and the time required for isotope enrichment are particularly important.
A metabolic steady state is ideal for stable isotope tracing. Under conditions of a metabolic steady state, the intracellular metabolite concentrations and metabolic fluxes of a cell population are constant. Because progression of the cell cycle is coordinated with bioenergetic and biosynthetic demands4,5,6,7, differences in cell proliferation will affect substrate uptake and utilization, and thus metabolite labeling. The exponential phase of growth is commonly assumed to reflect a near steady state condition, if nutrient supply is maintained2,8. For controlling nutrient supply, a bioreactor (commonly called a nutrostrat) which adds fresh medium and removes waste products ensures that the extracellular environment is optimized for cell growth9,10. Often, experiments are performed at a metabolic pseudo-steady state, in which deviations in extracellular nutrient concentration and intracellular metabolite concentrations and fluxes are considered negligible over the course of the experiment. Many cell culture media compositions contain supraphysiologic substrate levels to prevent nutrient depletion over the course of culture. For example, incorporation of 13C from 13C6-glucose into glycolytic intermediates is expected to occur within minutes in most cell types, whereas tricarboxylic acid cycle (TCA) cycle intermediates may require a few hours for sufficient 13C accumulation. Some biosynthetic pathway-derived intermediates and end products require longer time frames for sufficient label incorporation (Fig. 1B). Thus, at physiological concentrations, glucose would not become limiting if only 13C enrichment in glycolytic intermediates are measured; however, higher concentrations of glucose may be required to prevent nutrient depletion for assessment of label incorporation into TCA cycle or biosynthetic pathway metabolites.
After provision of an isotopically labeled source, metabolite isotope incorporation will increase in a time-dependent manner. This phase of isotope labeling is referred to as dynamic labeling. After 13C incorporation in a metabolite pool is saturated, isotopic steady state labeling is achieved (Fig. 1C). Importantly, the time required to achieve isotopic steady state is both metabolite- and tracer-dependent and is assessed empirically. Dynamic and steady state labeling are discussed in detail below.
Stage 2 – Sample preparation, data acquisition, and data analysis
Proper sample preparation and data analysis are critical for reliable interpretations of SIRM experimental results. In this section, we discuss sample preparation, metabolite separation and detection by liquid chromatography mass spectrometry (LC-MS), and data processing and analysis.
Sample preparation
Rapid enzymatic inactivation and efficient metabolite extraction are crucial steps to provide a snap shot of isotopologue labeling. Collection vessels and solvents should be of the highest purity to prevent contamination, especially by typical metabolite contaminants such as lactate11 and palmitate12. The most effective solvents for quenching are cold (–20 °C) acetonitrile or ultracold (−80 °C) methanol13,14. While methanol is sufficient to quench and extract a broad range of polar and some nonpolar metabolites15, maximal extraction of polar metabolites—including central carbon intermediates, amino acids and dipeptides, nucleosides and nucleotides, short-chain fatty acids, and some mono- and di-glycerides—can be achieved using 50–60% acetonitrile (v/v; acetonitrile/water)13,14; for the empirical results presented below, a 60% acetonitrile solution was used. Extraction of more hydrophobic species and lipids may require solvents such as chloroform16 or methyl tert-butyl ether17.
After sample collection, the extracts are processed and centrifuged at ≤4 °C, and the supernatants may be stored at –80 °C. Frequently, the solvent volume is reduced or removed and replaced with a solvent suitable for metabolite detection. Volatile solvents (e.g., methanol, chloroform) are removed using a stream of nitrogen gas; other solvents such as acetonitrile/water can be removed with a freeze dryer. Vacuum concentrators and centrifugal dryers may contribute to sample loss, contamination, and metabolite degradation (due to overheating) and therefore are not recommended.
Platform considerations
While many platforms are suitable for SIRM, MS-based workflows are common in most laboratories due to their capacity for broad metabolite coverage, high sensitivity, and the availability of software packages for data analysis. Liquid chromatography prior to MS analysis reduces spectral complexity and minimizes spectral overlap between labeled species by separating some species of identical molecular mass (e.g., isobars and isomers)3. Pairing LC with high resolution, accurate mass (HRAM) spectrometers such as orbitraps or time-of-flight instruments empowers the LC-MS approach with the ability to perform SIRM in an untargeted manner. Knowledge of metabolite retention time, accurate m/z, MS/MS spectrum, and use of authentic standards improves peak assignment accuracy. Other techniques, such as gas chromatography mass spectrometry (GC-MS)18,19,20,21 and capillary electrophoresis MS (CE-MS)22, can be used in targeted and non-targeted applications, and unit resolution triple quadrupole-based LC-MS is effective for targeted metabolomics methods23,24.
Spectrum deconvolution and metabolite assignment
Mass spectra generated for stable isotope-labeled metabolites differ significantly from those of unlabeled samples. Incorporation of heavy atoms (e.g. 13C) from the stable isotope tracer increases the complexity of the mass spectrum and requires additional analytic steps. Several bioinformatics tools have been developed to analyze SIRM data, e.g., see25,26,27,28,29,30. Our group has used the following workflows for peak assignment: Premise31,32, MetSign33, and El-Maven/Polly PhiTM34,35; the last two of these software packages can facilitate large metabolomics databases or custom target lists. A key advantage of the El-Maven/Polly PhiTM software is that it automatically calculates all possible isotopologue m/z ratios for heavy atoms specified by the user (e.g. 13C, 15N, etc.) and performs peak matching and assignments based on its algorithm and a set of user modifiable criteria. Metabolite peaks are assigned and confirmed using accurate m/z, retention time, and the MS/MS spectrum performed on peaks from unlabeled sample controls. These unlabeled controls are biological equivalent samples generated in the same experimental settings but without incubation with the labeled tracer. Unlabeled samples are important because: (1) they help define the sample’s metabolome composition; (2) by absence of tracer atoms, they help validate isotopologue assignments; and (3) they can provide the relative concentration for a given metabolite.
Natural abundance correction
After spectral deconvolution and assignment, the contribution of naturally occurring heavy isotopes (e.g. 2H is 0.0115%, 13C is 1.07% and 15N is 0.36836) is removed, yielding corrected peak intensities. By removing the natural abundance from spectral data, overestimation of fractional enrichment can be avoided. Natural abundance correction algorithms are a part of several software pipelines. For example, we have used online platforms based on a script described by Moseley et al.37 as well as MetSign33. Here, we use the IsoCorrect module—part of a larger Elucidata PollyPhi™ Platform34. A more detailed discussion of natural abundance correction can be found in the following reference21.
Data analysis and verification
Proper vetting of the assignments is essential to exclude inaccuracies and avoid incorrect data interpretation. The steps involved for data inspection require knowledge of analytical chemistry and metabolism. Data validation starts by setting up a priori criteria to maintain coherence within the dataset and eliminate possible user bias. The granularity of these criteria varies between experiments, and commonly encountered problems are summarized in Table 2. In general, replicate data are manually inspected for assignment errors and omissions. This allows the user to identify and address software-related issues quickly. Nevertheless, reexamination of chromatograms and mass spectra may be necessary. This is especially true when there is high intra-replicate variability, which is commonly caused by poor detection of low abundance metabolites. For poorly detected metabolites, increasing the starting material may be sufficient to obtain interpretable results.
Supplementary Fig. 1 illustrates the differences in chromatograms and raw mass spectra of citrate and α-ketoglutarate. High signal-to-noise chromatographic peaks were deconvoluted and assigned as citrate (Supplementary Fig. 1A). The corresponding mass spectrum (Supplementary Fig. 1C) revealed high abundance isotopologue peaks and undistorted labeling. In contrast, in this experiment, α-ketoglutarate signals were difficult to distinguish from instrument noise, increasing the likelihood for misassignment (Supplementary Fig. 1B). The MS spectrum of α-ketoglutarate showed that while the two most abundant isotopologues (m + 0 and m + 5) were present, others were undetected (Supplementary Fig. 1D). When peaks are below the software threshold of detection, metabolite peaks can be manually inspected, and assignments can be entered based on accurate retention times and masses. Removal of outliers is generally discouraged unless deemed statistically appropriate based on a priori criteria in the study design. For replicates with several poorly detected metabolites, the analyst may choose to reacquire the spectra or repeat the entire labeling experiment using more biological starting material.
Apart from analytical knowledge, expertise in metabolism, the integration of additional metabolic flux measurements, and the use of pharmacological controls may uncover additional data integrity issues. These frequently involve a mismatch in labeling patterns between a suspected metabolite and other pathway constituents. Such inconsistencies prompt additional steps to reexamine and evaluate assigned data. Furthermore, pharmacological controls that target critical steps in metabolic pathways can identify possible assignment issues, especially if the labeling patterns do not agree with textbook biochemistry. Finally, heavy isotope enrichment found in unlabeled samples that exceeds natural abundance suggests misassignments due to peak overlaps and requires intervention. Once assignments are corrected and possible outliers (e.g., undetected metabolites) are removed, they can be plotted and interpreted.
Stage 3 – Data interpretation
After metabolites and isotopologues are assigned and quantified, fractional enrichment data are plotted for interpretation. An example of a fractional enrichment plot is shown in Fig. 1D. Here, 13C enrichment into glutamate is shown for cells provided with 13C6-glucose. The degree of 13C incorporation is indicated by the fractional (isotopic) enrichment values for each isotopologue (m + x). As mentioned earlier, the m + 0 isotopologue indicates the fraction of unlabeled glutamate, whereas each successive isotopologue shows the relative abundance of 13C incorporated into 1, 2, 3, 4, or all 5 carbons of glutamate. For most LC-MS approaches, it is not possible to ascertain positional information of the labeled atom (i.e., distinguish isotopomers); however, strategic fragmentation and mathematical modeling strategies can provide position-specific enrichment information38.
Another way to examine metabolite labeling is by adding all isotopologues together, which provides the total isotopic enrichment (i.e. % metabolite enrichment) for a given metabolite pool. Importantly, when a time-course experiment is performed, plotting the isotopic enrichment is the only means by which dynamic and isotopic steady state may be distinguished. Shown in Supplementary Fig. 2 is a sampling of time-dependent isotopic enrichment in metabolites from several metabolic pathways from neonatal rat cardiomyocytes (NRCMs) incubated with 13C6-glucose. Differences in pathway flux are readily appreciated by comparing enrichment in metabolites derived from slower pathways, such as the pentose phosphate pathway, versus metabolites in faster pathways, such as glycolysis. For example, 3′AMP and glutathione are in the dynamic labeling phase throughout the time course, whereas the 13C label saturates the 3-phosphoglycerate (3PG) pool by at least 4 h. Metabolites associated with the Krebs cycle demonstrate intermediate labeling velocities, with the dynamic labeling phase occurring within 0–12 h of incubation with 13C6-glucose.
Relative flux differences in metabolic pathways may be inferred from incorporation of isotopic tracer into metabolites during the dynamic labeling phase. Resolving changes in relative metabolic pathway flux can be assessed by direct interpretation of stable isotope labeling patterns2, without the need for formal 13C flux analysis, which utilizes mathematical modeling39,40,41. For example, NRCMs incubated with 13C6-glucose show purine (3′AMP) labeling in the dynamic phase through 18 h. Therefore, a hypothetical decrease in m+0 fractional enrichment and an increase in m + 5 labeling in purines in an experimental group within this timeframe would suggest higher rates of ribose incorporation and increased pathway flux.
The contribution of a tracer to a metabolite pool should be assessed after isotopic enrichment has reached steady state. For example, 13C labeling in citrate saturates by 10–12 h of introducing 13C6-glucose to NRCMs. Enrichment approaches 90% by 18 h, suggesting that glucose-derived carbon is the predominant source of carbon for citrate synthesis (Supplementary Fig. 2).
Analysis of glycolysis: Although fractional enrichment plots of glycolytic intermediates can resolve changes in glycolysis, examining relative changes in glycolytic flux can be difficult. In NRCMs, glycolytic intermediates such as the glyceraldehyde 3-phosphate/dihydroxyacetone phosphate pool (GAP/DHAP), 3-phosphoglycerate (3PG), 2-phosphoglycerate (2PG), and phosphoenolpyruvate (PEP) show nearly complete saturation with label by 4 h (Fig. 2A–C). In another experiment, NRCMs were incubated with medium containing 13C6-glucose for 5 min (insets). Results from this experiment suggest that isotopic steady state is reached within minutes of 13C6-glucose provision. Such rapid attainment of isotopic steady state makes the assessment of glycolytic flux by isotopic labeling difficult. In such cases, glycolytic rates can be measured by assessing the levels of 13C3-lactate and 13C6-glucose in the incubation medium42 as well as by orthogonal methods, such as methods based on radiolabeled tracers ([5-3H]-glucose) or extracellular flux analysis.
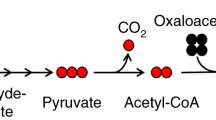
Time course of 13C enrichment in glycolytic intermediates. Neonatal rat cardiac myocytes (NRCMs) were cultured in medium containing 13C6-glucose for 5 min, 4 h, 8 h, 12 h, or 18 h, followed by quenching, metabolite extraction, and SIRM analyses: (A) Schematic of experimental protocol; (B) Condensed schematic of glycolysis; note that the metabolites in bold are shown in these analyses; and (C) Fractional enrichment of 13C into glycolytic intermediates. Insets are data derived from a separate experiment in which 13C6-glucose was provided for 5 min. n = 6 replicates per group, pooled from NRCMs isolated from three independent litters.
That 40–60% of the pyruvate and lactate pools were unlabeled suggests dilution by non-labeled substrates and is supported by the presence of unlabeled pyruvate in the medium. The presence of primarily the m + 3 isotopologue in these intermediates indicates that only 13C6-labeled glucose was used for glycolysis. Similarly, the hexose 6-phosphate pool consisted of only m + 6 labeling, which indicates low levels of fructose bisphosphatase activity in NRCMs. The presence of other isotopologues, e.g., m + 3 labeling, in hexose 6-phosphate in cells incubated with only 13C6-glucose could indicate gluconeogenic activity, but this isotopologue was not observed in these studies. Interestingly, in NRCMs, the hexose 6-phosphate pool had relatively modest labeling at the m + 6 isotopologue, which indicates unlabeled isomers of hexose 6-phosphate (Fig. 2C).
Analysis of Krebs cycle activity: As expected, Krebs cycle intermediates in NRCMs demonstrate time-dependent 13C labeling (Fig. 3), from which several conclusions may be drawn. The labeling patterns in Krebs cycle intermediates are suggestive of both pyruvate dehydrogenase-mediated and anaplerotic incorporation of the 13C atoms. Enrichment in m + 2 isotopologues indicate the first turn of the Krebs cycle (i.e. m + 2). The m + 4, m + 5, and m + 6 isotopologue labeling in citrate and aconitate may indicate the 2nd and 3rd turns of the Krebs cycle, with label saturation apparent in the m + 4 isotopologues by 12 h. Labeling the cells with 13C6-glucose for 5 min led to approximately 20% 13C enrichment (m + 2) in citrate and aconitate; however, little to no labeling was found in α-ketoglutarate (Fig. 3, insets) or any other Krebs cycle metabolite (data not shown). These data suggest that incorporation of label into citrate is faster than that through “downstream” metabolites, which is consistent with relatively higher metabolic needs for citrate due to fatty acid synthesis.
Time course of 13C enrichment in Krebs cycle intermediates. Neonatal rat cardiac myocytes (NRCMs) were cultured in medium containing 13C6-glucose for 5 min, 4 h, 8 h, 12 h, or 18 h, followed by quenching, metabolite extraction, and SIRM analyses: (A) Fractional enrichment of 13C into Krebs cycle intermediates. Insets are data derived from a separate experiment in which 13C6-glucose was provided for 5 min. n = 4–6 replicates per group, pooled from NRCMs isolated from three independent litters. PDH, pyruvate dehydrogenase; ME, malic enzyme; PC, pyruvate carboxylase; ADC, aspartate decarboxylase.
Product-precursor relationships are also important to consider. For example, by 4 h, >70% of the citrate pool was labeled (at m + 1, m + 2, m + 3, m + 4, or m + 5), yet pyruvate (see Fig. 2C) showed only 40% labeling (at m + 3). This could indicate the presence of more than one pool of pyruvate, with the labeled pyruvate pool contributing more to the Krebs cycle than the unlabeled pool; however, the Krebs cycle carries large flux, and given that the levels of unlabeled pyruvate in the medium would progressively decrease over time, earlier times of labeling may be informative. Indeed, a 5 min labeling study revealed lower fractional enrichment in citrate than in pyruvate (Supplementary Fig. 3), which refutes the idea that two pools of pyruvate contribute to citrate formation in NRCMs. These data exemplify the importance of time course data for delineating product-precursor relationships and for interpreting metabolic pathway activities.
The fact that glutamate and glutathione (GSH) were labeled suggests that NRCMs use glucose as a carbon source for glutathione synthesis. Glutamine was not labeled, which is expected given that unlabeled glutamine is provided in relatively high amounts in the culture medium (Fig. 3). The m + 3 labeling, e.g., in 13C3-malate and -fumarate, may also indicate anaplerotic activity43,44, likely through either pyruvate carboxylase44 or malic enzyme45. The kinetics of labeling in these isotopologues, when compared to that of the PDH-supported labeling (m + 2), indicate that these carboxylation reactions are generally slower than pyruvate decarboxylation reactions in NRCMs. Indeed, 5 min of culture of NRCMs with 13C6-glucose led to labeling of citrate at only m + 2 and did not label malate (Supplementary Fig. 3C).
While the labeling patterns of Krebs cycle intermediates and associated metabolites (e.g., aspartate, glutamate) are generally consistent, questions arise with regard to succinate and alanine. Succinate was almost completely devoid of 13C incorporation derived from PDH-mediated Krebs cycle activity. Nevertheless, the fact that independent studies [e.g.44,] show relatively low levels of 13C incorporation into succinate compared with other Krebs cycle intermediates suggest that these data may be valid; they might be explained by different intracellular pools of succinate, in which a relatively minor succinate pool is used for fumarate production. Surprisingly, alanine labeling is most similar to that of malate and aspartate (Fig. 3) and does not follow that of pyruvate (see Fig. 2C). This would suggest that, in NRCMs, alanine does not arise by alanine transamination of pyruvate but rather via aspartate decarboxylase. Conclusive evidence for this mechanism could be provided by interventions that target aspartate decarboxylase directly.
Importance of complementary metabolic assays and pharmacological controls: The preceding data illustrate how isotopologue pattern matching and time course information can impart confidence or skepticism in stable isotope metabolomics data as well as give rise to new, interesting questions. While technical controls—including proper retention time and mass values according to metabolite standards—provide confidence in metabolite assignment and isotopologue quantification, the metabolome consists of many similar metabolites that may have nearly identical masses and may be difficult to resolve by chromatography. While some newer technologies (e.g., differential mobility spectrometry) can address some of these challenges38,46, additional methods to assess the veracity of stable isotope metabolomics data and to standardize workflows within and between laboratories could increase the reproducibility, certainty, and usefulness of SIRM results.
We gain additional confidence in SIRM data by including pharmacological controls in experimental design. By predicting 13C labeling based on the metabolic steps and pathways known to be affected by pharmacological compounds, these experimental groups could be used as positive or negative controls. To examine this, we treated NRCMs with koningic acid (KA)—a known glyceraldehyde 3-phosphate dehydrogenase inhibitor; rotenone—a commonly used inhibitor of mitochondrial Complex I; oligomycin—a commonly used inhibitor of Complex V; or FCCP—an uncoupler that stimulates mitochondrial oxygen consumption (Fig. 4A). We measured mitochondria and glycolytic activity with acute (17 or 100 min: Protocol I) and chronic (12 h: Protocol II, III) treatments with each compound (Fig. 4B,C). As expected, both acute and chronic treatment with FCCP increased oxygen consumption rate (OCR); rotenone decreased OCR both acutely and chronically (Fig. 4D,F). Oligomycin initially decreased OCR by 50%, which is consistent with our previous studies (Fig. 4D)42,47,48; however, upon sustained (12 h) exposure, OCR rebounded to baseline levels (Fig. 4F). This is expected given the propensity of oligomycin to increase Δp and thereby promote proton leakage or electron slippage49. While KA had no effect on OCR (Fig. 4D,F), KA acutely decreased the extracellular acidification rate (ECAR), whereas oligomycin and rotenone increased ECAR (Fig. 4E). Radiolabeled tracing experiments using [5-3H]-glucose (Protocol III; Fig. 4C) provided absolute flux measurements and demonstrated sustained changes in glucose utilization caused by each pharmacological compound (Fig. 4H). Importantly, the pharmacological agents did not affect protein content significantly (Supplementary Fig. 4), which suggests minimal changes in cell viability.
Mitochondrial and glycolytic activity under controlled metabolic states. Extracellular flux analyses and radiolabeled glucose utilization assays in NRCMs treated with metabolic inhibitors or activators: (A) Schematic illustrating the inhibitors used: KA, koningic acid (10 µM); Rot, rotenone (1 µM); Oligo, oligomycin (1 µM); FCCP, carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone (1 µM); (B) Schematic of the protocol for measuring acute changes in mitochondrial and glycolytic activity with each pharmacological compound; (C) Schematic of the protocol for measuring chronic changes in mitochondrial and glycolytic activity with each pharmacological compound; (D) Oxygen consumption rate (OCR) after incubation with each compound for 17 and 100 min; (E) Extracellular acidification rate (ECAR) after incubation with each compound for 17 and 100 min; (F) OCR after incubation with each compound for 12 h; (G) ECAR after incubation with each compound for 12 h; (H) Glucose utilization with each compound, measured using the [5-3H]-glucose assay (see Protocol III, panel C). n = 4 independent isolations per group. Data from XF analyses were log-transformed prior to statistical analysis. *p < 0.05.
We next examined 13C metabolite labeling in NRCMs cultivated with 13C6-glucose for 12 h in the absence or presence of each pharmacological compound (Fig. 5A). As shown in Fig. 5B, compounds that increased glycolytic activity (i.e., FCCP, oligomycin, and rotenone) robustly increased the fractional enrichment of 13C-glucose-derived carbon into hexose 6-phosphate. That KA also augmented 13C enrichment into the hexose phosphate pool and that all glycolytic intermediates indicate nearly maximal 13C enrichment appear at first inconsistent; however, analysis of relative pool size, knowledge of biochemistry, and additional experiments can provide an explanation. Because KA inhibits GAPDH, crossover theorem50,51,52,53 predicts that the concentrations of metabolites should increase before the inhibition site and decrease after the inhibition site. Indeed, hexose 6-phosphate and GAP/DHAP levels were remarkably elevated, and levels of metabolites after the GAPDH reaction (e.g., 3PG) were diminished (Fig. 5C). Moreover, because high levels of G6P promote glycogen synthesis54 and inhibit glycogen breakdown55, the high level of 13C enrichment that occurs in the hexose 6-phosphate pool with KA treatment is likely due to diminished G6P dilution by the unlabeled glycogen pool.
13C enrichment in glycolytic intermediates under controlled metabolic states. (A) NRCMs were treated with the indicated inhibitor (as in Fig. 4) for 12 h in medium containing 25 mM 13C6-glucose. Intracellular metabolites were then extracted and subjected to LC/MS for SIRM analysis; (B) Fractional enrichment plots of glycolytic intermediates; and (C) Relative pool size for indicated glycolytic intermediates. n = 3–6 replicates per group, pooled from NRCMs isolated from three independent litters. **p < 0.01, ****p < 0.0001 vs. Ctrl.
In separate experiments, we also evaluated 3H-2-deoxyglucose (2DG) uptake, which provides a measure of glucose uptake, in NRCMs treated with KA and FCCP; treatment of NRCMs with unlabeled 2DG (100 mM) was used as a control. As predicted from the labeling patterns of hexose 6-phosphate in Fig. 5, FCCP increased glucose uptake in NRCMs; however, KA did not affect glucose uptake (Supplementary Fig. 5), which suggests that in KA-treated cells glucose-derived intermediates upstream of the GAPDH reaction are likely shunted to ancillary pathways. Such complementary assays help to better define changes in metabolism, especially when the metabolites of interest are assessed at the isotopic steady state phase of labeling.
Analysis of 13C incorporation into Krebs cycle metabolites provides information on mitochondrial activity. As expected from the mitochondrial OCR measurements (see Fig. 4), FCCP led to higher 13C incorporation into Krebs cycle intermediates, with the exception of α-ketoglutarate, which showed less 13C incorporation with FCCP (Fig. 6A). Reanalysis of the α-ketoglutarate spectral data indicated a poor signal-to-noise ratio (Supplementary Fig. 1), which diminishes confidence in its adequate detection in this set of experiments. The markedly higher fractional enrichment of the m + 6 isotopologues of citrate and aconitate and the m + 4 isotopologues of succinate, fumarate, and malate could indicate more turns of the Krebs cycle; however, it is important to consider other explanations as well. Longer periods of labeling can increase the possibility of tracer scrambling, which can occur after the tracer has been extensively recycled, or promote more isotope labeling due to less dilution by the unlabeled pool of metabolite precursors. Although increased m + 6 and m + 4 labeling could suggest more turns of the Krebs cycle, it is possible that the same number of turns are occurring with less input from other carbon-donating sources or that there is more input from 13C-labeled metabolic products (e.g., 13C3-lactate or labeled non-essential amino acids); however, integration of pharmacological controls in labeling studies and OCR measurements help resolve these issues. For example, in FCCP-treated NRCMs, complementary data showing higher 13C enrichment (Fig. 6) and lower pool size of several Krebs cycle intermediates (Supplementary Fig. 6) as well as higher OCR (Fig. 4F) confirms higher Krebs cycle metabolite turnover and flux.
13C enrichment in Krebs intermediates under controlled metabolic states. Fractional enrichment plots of Krebs cycle intermediates: NRCMs were treated with the indicated inhibitor (as in Fig. 4) for 12 h in medium containing 25 mM 13C6-glucose. Intracellular metabolites were then extracted and subjected to LC/MS for SIRM analysis. n = 4–6 replicates per group, pooled from NRCMs isolated from three independent litters. PDH, pyruvate dehydrogenase; ME, malic enzyme; PC, pyruvate carboxylase; ADC, aspartate decarboxylase.
FCCP also increased 13C incorporation into glutathione (GSH) and alanine, indicating that mitochondrial uncoupling also increases the biosynthetic rates of GSH and alanine. The observation that chronic oligomycin treatment did not markedly affect 13C labeling of Krebs cycle intermediates (Fig. 6) is not surprising, as OCR was unaffected after 12 h of incubation. As expected based on the OCR data, the Complex I inhibitor rotenone diminished fractional enrichment in all Krebs cycle intermediates as well as in GSH and alanine (Fig. 6A). Collectively, these data illustrate the fractional enrichment patterns that occur in uncoupled mitochondria and when complex I is inhibited.
The glycolysis inhibitor, KA, remarkably diminished 13C incorporation into Krebs cycle intermediates, indicating that it decreases glucose oxidation in NRCMs (Fig. 6A). That 13C6-glucose-derived carbon labeled approximately 80% of the Krebs cycle intermediate pool (e.g., see citrate in Supplementary Fig. 2) indicates that glucose is a primary nutrient for mitochondrial oxidation in cultured NRCMs; that KA generally diminished the levels of Krebs cycle intermediates (see Supplementary Fig. 6) further supports the concept that cultured NRCMs rely on glucose for mitochondrial respiration. Interestingly, mitochondrial respiration data indicate that KA does not markedly inhibit basal OCR (Fig. 4), which suggests inherent flexibility in recruiting other nutrients (e.g., unlabeled pyruvate) to support mitochondrial respiration when glucose is limiting or glucose catabolism is inhibited.
Biosynthetic pathway activity: In addition to catabolic pathways, SIRM provides information of how nucleotides, UDP-HexNAc (comprising UDP-GlcNAc and UDP-GalNAc), amino acids, and glycerophospholipids are built, which delineates anabolic pathway activity in cells or tissues31,42,56. As an example, we show in Fig. 7 fractional enrichment data in the pyrimidine, uridine monophosphate (UMP), and in the endproduct of the hexosamine biosynthetic pathway (HBP), UDP-HexNAc. The former is synthesized via the pentose phosphate pathway and from aspartate-derived carbons (Fig. 7A), and the latter is synthesized from carbons derived from not only F6P, but from acetyl CoA and UDP as well (Fig. 7D). In NRCMs, both UMP and UDP-HexNAc appear to remain in the dynamic phases of labeling through 12 h of 13C6-glucose provision (Fig. 7B,E).
13C enrichment in biosynthetic pathway metabolites. Time course and pharmacological control 13C enrichment data: (A) Atom resolved map of pyrimidine (UMP) biosynthesis; (B) NRCMs were cultured in medium containing 25 mM 13C6-glucose for 4 h, 8 h, 12 h, or 18 h, followed by quenching, metabolite extraction, and SIRM analyses. Shown is the fractional enrichment plot of UMP; (C) NRCMs were incubated in 25 mM 13C6-glucose for 12 h with the indicated inhibitor (as in Figs 4–6), followed by SIRM analysis. Shown is the fractional enrichment plot of UMP; (D) Illustration of UDP-GlcNAc synthesis; (E) NRCMs were cultured in medium containing 25 mM 13C6-glucose for 4 h, 8 h, 12 h, or 18 h, followed by quenching, metabolite extraction, and SIRM analyses. Shown is the fractional enrichment plot of UDP-HexNAc (composed of both UDP-GlcNAc and UDP-GalNAc); and (F) NRCMs were incubated in 25 mM 13C6-glucose and the indicated inhibitor (as in Figs 4–6), followed by SIRM analysis. Shown is the fractional enrichment plot of UDP-HexNAc. n = 6 replicates per group, pooled from NRCMs isolated from three independent litters.
The UMP labeling profile indicates that all inhibitors diminish incorporation of glucose-derived carbon into pyrimidines, with FCCP inhibiting glucose carbon incorporation into the ribose ring (m + 5). The labeling patterns of the m + 6, m + 7, and m + 8 isotopologues of UMP in FCCP-treated cells are consistent with that of the m + 2, m + 3, and m + 4 isotopologues of asparate, respectively (see Fig. 6). The pool size of UMP was modestly lower in FCCP-treated NRCMs (Supplementary Fig. 7).
A similar picture emerges with UDP-HexNAc. FCCP decreased glucose carbon incorporation into the m + 5, m + 6, m + 11, and m + 13 isotopologues of UDP-HexNAc, which largely represent carbons derived from UDP (m + 5), glycolysis-derived F6P (m + 6), UDP + F6P (m + 11), and UDP + F6P + glucose oxidation-derived acetyl CoA (m + 13) (Fig. 7F). These data indicate that mitochondrial uncoupling by FCCP diminishes pyrimidine synthesis by decreasing pentose phosphate pathway flux and that it may decrease UDP-HexNAc synthesis by diminishing carbon incorporation from UDP, F6P, and acetyl CoA.
Exposure of NRCMs to KA markedly diminished glucose-derived carbon incorporation into UMP (Fig. 7B), and it diminished UMP pool size (Supplementary Fig. 7A). Although KA did not affect ribose incorporation into UMP (m + 5), it markedly diminished incorporation from aspartate (see m + 6, m + 7, m + 8 isotopologues). Because KA markedly inhibited glucose oxidation and diminished the overall Krebs cycle intermediate pool size (Supplementary Fig. 6), these data indicate that KA inhibited pyrimidine biosynthesis by decreasing glucose oxidation and aspartate synthesis.
The 13C labeling patterns and pool size in UDP-HexNAc from KA-exposed NRCMs suggest that although inhibition of GAPDH drives F6P-derived carbon into the HBP, UDP-HexNAc biosynthesis is limited by availability of Krebs cycle-derived products. This conclusion is supported by the fact that KA increased 13C enrichment in the m + 6 isotopologue, yet diminished 13C incorporation into the ribose of UDP (m + 5 isotopologue) and into those isotopologues built from Krebs cycle-derived products (m + 7–m + 16 isotopologues). Furthermore, KA remarkably decreased UDP-HexNAc pool size. Oligomycin and rotenone treatment did not have large effects on UMP or UDP-HexNAc pool size or 13C enrichment. Collectively, these findings illustrate critical relationships between catabolic and anabolic pathways and show that mitochondrial uncoupling and glucose oxidation affect the rates of pyrimidine and UDP-HexNAc biosynthesis.
Summary: The findings of this study illustrate how to integrate SIRM with the use of pharmacological controls and complementary flux analyses, which collectively provide a more comprehensive view of metabolism. The pharmacological controls used here provide the experimenter the ability to appraise confidence in fractional enrichment data. An example shown here is α-ketoglutarate, the enrichment pattern of which led to its spectral reanalysis and exclusion from interpretation. The combined use of time course data and metabolic inhibitors and activators can be used to assess the relative difference in pathway activity. We provide evidence that, in NRCMs, glucose oxidation occurs at a higher rate than anaplerotic activity from pyruvate carboxylase or malic enzyme. Moreover, we provide tangible examples of how labeling at isotopic steady state does not provide an index of flux. Additionally, we found that, in NRCMs, alanine may be synthesized via an aspartate decarboxylase. The data also provide unique insights into how glycolysis, glucose oxidation, and mitochondrial uncoupling regulate ancillary biosynthetic pathway activity. Although these findings may be specific to the cultured NRCM system used here, they provide evidence that may extend to other in vitro or to in vivo systems.
Materials and Methods
Materials
DMEM growth medium containing l-glutamine, D-glucose, and sodium pyruvate was purchased from Sigma-Aldrich (St. Louis, MO, USA). Koningic acid was from Cayman Chemical (Ann Arbor, MI, USA). All other reagents were from Sigma-Aldrich Corp. (St. Louis, MO, USA), unless indicated otherwise.
Rodent models & primary cell isolation
All procedures were approved by the University of Louisville Institutional Animal Care and Use Committee and were in accordance with NIH guidelines. The euthanasia procedures were consistent with the AVMA Guidelines for the Euthanasia. Neonatal rat cardiomyocytes (NRCMs) were isolated from 1- to 2-day-old Sprague-Dawley rats as previously described57,58,59. Briefly, neonatal rat hearts were excised, rinsed, and minced in calcium- and bicarbonate-free Hank’s balanced salt solution containing HEPES (pH 7.0). Cells were dissociated by stepwise trypsin digestion, after which the trypsin in the dissociated cell mixture was quenched with FBS and centrifuged at 180 g for 5 min. The pellet was then resuspended in warm DMEM (25 mM glucose, 4 mM glutamine, 1 mM sodium pyruvate) supplemented with 5% FBS, 1% penicillin/streptomycin, 0.1 mM BrdU, and 2 µg/mL vitamin B12 and preplated in 10 cm dishes for 1 h to allow for fibroblast adherence and removal. The non-adherent cardiomyocytes were then collected and plated at the desired cell density, as outlined below.
Cell culture
NRCMs were plated into tissue culture treated 6-well dishes (1.7 × 106 cells/well) or XF24 cell culture microplates (7.5 × 104 cells/well) in DMEM (25 mM glucose, 4 mM glutamine, 1 mM sodium pyruvate) supplemented with 5% FBS, 1% penicillin/streptomycin, 0.1 mM BrdU, and 2 µg/mL vitamin B12. The following day (day 1), media was replaced with the same plating media. On day 4, culture media was changed to media lacking BrdU. On day 5, media was replaced to serum-free media for at least 18 h prior to experimental treatments. Following serum starvation, cells were treated with pharmacological and chemical reagents: FCCP − carbonyl cyanide 4-(trifluoromethoxy) phenylhydrazone (1 µM); KA – koningic acid (10 µM); Oligo – oligomycin (1 µM); Rot – rotenone (1 µM); or 2DG – 2-deoxyglucose (100 mM).
Cell viability before and following treatments were assessed via visual inspection under microscopy; there were no indications (vesicle formation, cell debris, etc.) of a change in cell viability following any the treatments. Assessment of total protein was used as an additional surrogate for changes in cell viability.
Stable isotope tracing
NRCMs were incubated in 6-well plates for 5 min or 4–18 h in DMEM media (US Biological; Swampscott, MA, USA) containing 1 mM pyruvate, 4 mM glutamine, and 25 mM [13C6]-glucose (99% purity, microbiological and pyrogen tested; Cambridge Isotope Laboratories, Inc., Tewksbury, MA). Additionally, for compound/inhibitor experiments, FCCP, KA, Oligo, Rot, or 2DG were added to this [13C6]-glucose containing media. After the specified isotope labeling time point, cell reactions were quenched in cold acetonitrile, and extracted in acetonitrile:water:chloroform (v/v/v, 2:1.5:1), as described previously31,32,60,61, to obtain the polar, nonpolar, and insoluble proteinaceous fractions. The nonpolar (lipid) layer was collected, dried under a stream of nitrogen gas, and reconstituted in 0.1 ml of chloroform:methanol:butylated hydroxytoluene (2:1 + 1 mM) mixture and stored at −80 °C for future analysis. The polar fraction was lyophilized using a Freezone 2.5 L −84 °C benchtop freeze dryer (Labconco, Kansas City, CO, USA). The dried sample was reconstituted in 100 μl 20% acetonitrile and used for LC-MS analysis.
LC-MS analysis
All samples were analyzed on a Thermo Q Exactive HF Hybrid Quadrupole-Orbitrap Mass Spectrometer with a Thermo DIONEX UltiMate 3000 HPLC system (Thermo Fisher Scientific, Inc., Germany). Separation was performed in parallel mode on a reversed phase (RP) Waters ACQUITY UPLC HSS T3 column (150 × 2.1 mm i.d., 1.8 µm, part number: 186003540) and a hydrophilic interaction chromatography (HILIC) Thermo Accucore HILIC column (100 × 3 mm i.d., 2.6 µm, part number: 17526–103030). The temperature of both columns was set to 40 °C. The HILIC column was operated as follows: mobile phase A was 10 mM ammonium acetate (pH adjusted to 3.25 with acetate) and mobile phase B was acetonitrile. The gradient was: 0 min, 100% B; 0 to 5 min, 100% B to 35% B; 5 to 12.7 min, 35% B; 12.7 to 12.8 min, 35% B to 95% B; 12.8 to 14.3 min, 95% B. The flow rate was set 0.3 mL/min. For the RP column, the mobile phase A was water with 0.1% formic acid and mobile phase B was 100% acetonitrile. The gradient was as follows: 0 min, 0% B; 0 to 5 min, 0% B; 5 to 6.1 min, 0 to 15% B; 6.1 to 10 min, 15 to 60% B; 10 to 12 min, 60% B; 12 to 14 min, 60% to 100% B; 14 to 14.1 min, 100% to 5% B; 14.1 to 16 min, 5% B. The flow rate was 0.4 mL/min.
The electrospray ionization probe was fixed at level C. The parameters for the probe were set as follows: sheath gas = 55 arbitrary units, auxiliary gas = 15 arbitrary units, sweep gas = 3 arbitrary units, spray voltage = 3.5 kV, capillary temperature = 320 °C, S-lens RF level = 65.0, auxiliary gas heater temperature = 450 °C. The method of mass spectrometer was set as follows: full scan range = 50 to 750 (m/z); resolving power R = 30,000 at m/z = 200, 10% valley; maximum injection time = 50 ms; automatic gain control (AGC) = 106 ions for both positive and negative modes.
Unlabeled samples were analyzed in full MS scan and data-dependent MS/MS modes, while the labeled specimens were only analyzed in full MS mode. The LC methods and electrospray ionization conditions were set as follows: for full MS scan, scan range = 50–750 (m/z), R = 30,000 at m/z = 200, 10% valley, maximum injection time = 50 ms, automatic gain control (AGC) = 106 ions; for data dependent MS/MS scan, R = 15,000, maximum injection time = 100 ms, automatic gain control (AGC) = 5 × 104, loop count = 6, isolation window = 1.3 m/z, dynamic exclusion time = 1.2 s, the collision energy was set 10, 20, 40, 60 and 150 eV, respectively.
Stable isotope data analyses
Full MS.raw files were first converted to.mzML format with msConvert tool, a part of an open-source ProteoWizard suite, described in detail by Adusumilli and Mallick62. Isotopologue peak deconvolution and assignments were performed using El-MAVEN. Peaks were assigned using a metabolite library first generated and verified using full scan MS and MS/MS spectra of unlabeled samples, as described previously33,61. The library contained metabolite names and corresponding molecular formulae used for generation of theoretical m/z values for all possible isotopologues, and retention times for each entry. The El-MAVEN parameters for compound library matching were as follows: EIC Extraction Window ± 7 ppm; Match Retention Time ± 0.60 min. For 13C isotopologue peak detection, the software criteria were set as follows: Minimum Isotope-parent correlation 0.5; Isotope is within 7 scans of parent; Abundance threshold 1.0; Maximum Error To Natural Abundance 100%. All assignments were visually inspected and compared to unlabeled samples for reference. Any peak groups assigned in error, e.g. not present or having different retention time than in the unlabeled samples, were deleted and correct peak assignments were added manually. Finally, the peak list with corresponding abundances was exported to a comma-separated (CSV) file and uploaded to the Polly workflow to perform natural abundance correction and to calculate total pool size for each metabolite (by summing peak areas of each detected isotopologue) using Polly Isocorrect. Finally, the data were analyzed and plotted with GraphPad Prism 8.0 (GraphPad Software, San Diego, Ca, USA).
Radiolabeled glucose utilization assay
Glucose utilization was determined using [5-3H]-glucose63,64,65. Briefly, NRCMs were cultured in 6-well dishes and culture media was replaced with 2 ml of fresh, serum-free culture media with the addition of 2 µCi/ml [5-3H]-glucose (Moravek Biochemicals, Brea, CA, USA). Following incubation for 3 h, 100 µl of media was collected and added to 100 µl of 0.2 M HCl in a microcentrifuge tube. This tube, with the tube cap removed, was placed in a scintillation vial containing 500 µl of dH2O to allow for evaporation diffusion of [3H]2O in the microcentrifuge tubes into the scintillation vials. To account for incomplete equilibration of [3H]2O and background, known amounts (µCi) of [5-3H]-glucose and [3H]2O (Moravek Biochemicals) were placed into microcentrifuge tubes, and these were placed into separate scintillation vials containing 500 µl dH2O. After incubation at 37 °C for 72 h, the microcentrifuge tube was removed from the vial, 10 ml of scintillation fluid was added, and scintillation counting was performed using a Tri-Carb 2900TR Liquid Scintillation Analyzer (Packard Bioscience Company, Meriden, CT, USA). Glucose utilization was then calculated as reported by Ashcroft et al.63, with considerations for the specific activity of [5-3H]-glucose, incomplete equilibration and background, dilution of [5-3H]- to unlabeled-glucose, and scintillation counter efficiency. To normalize glucose utilization to total protein, the cells were washed with PBS (to remove adherent radioactivity) and then lysed in common lysis buffer. Protein concentration was measured using the Lowry assay kit (Bio-Rad Laboratories).
Glucose uptake measurements
Glucose uptake was estimated by uptake of 3H-2-deoxyglucose (2DG)59. For this, NRCMs in 6-well plates were exposed to 2 µCi/ml 3H-2-DG in the presence or absence of 100 mM unlabeled 2-DG, 10 µM KA, or 1 µM FCCP for 3 h. The cells were then washed 6 times with ice-cold phosphate-buffered saline to remove adherent radioactivity, and lysed in 100 µl of lysis buffer (20 mM Hepes, pH 7.0, 1 mM EDTA, 1% Nonidet P40 and 0.1% SDS). After centrifugation at 13,000 g for 10 min, radioactivity in the supernatant was measured by scintillation counting. The radioactivity (in counts per minute; CPM) was normalized to total protein content.
Extracellular flux analysis of cellular energetics
To measure cellular energetics in intact cardiomyocytes, the Seahorse Bioscience XF24 extracellular flux analyzer was used as previously described47. For this, 75,000 NRCMs were plated into each well and cultured as described above. An hour before each experiment, the media was replaced with 675 µl of assay media, i.e., unbuffered DMEM containing glucose (25 mM), glutamine (4 mM), and pyruvate (1 mM), pH 7.4. Following 1 h in a 37 °C, CO2-free incubator, the cells were placed in the instrument for analysis. Basal oxygen consumption rates (OCR) and extracellular acidification rates (ECAR) were measured using a programmed protocol: 3 cycles of 2 min mix, 2 min wait, and 3 min measure. The OCR and ECAR values were normalized to the total amount of protein in each well.
Statistical analysis
All values are mean ± S.D. Comparisons were made to the relevant control group, and statistical tests were performed using one-way analysis of variance (ANOVA) with Dunnett’s correction for multiple comparisons. The null hypothesis was rejected when p < 0.05.
Data Availability
Raw data are available upon request.
References
Lehninger, A. L., Nelson, D. L. & Cox, M. M. Lehninger principles of biochemistry. 3rd edn, (Worth Publishers 2000).
Buescher, J. M. et al. A roadmap for interpreting (13)C metabolite labeling patterns from cells. Curr Opin Biotechnol 34, 189–201, https://doi.org/10.1016/j.copbio.2015.02.003 (2015).
Jang, C., Chen, L. & Rabinowitz, J. D. Metabolomics and Isotope Tracing. Cell 173, 822–837, https://doi.org/10.1016/j.cell.2018.03.055 (2018).
Robbins, E. & Morrill, G. A. Oxygen uptake during the HeLa cell life cycle and its correlation with macromolecular synthesis. J Cell Biol 43, 629–633 (1969).
Cai, L. & Tu, B. P. Driving the cell cycle through metabolism. Annual review of cell and developmental biology 28, 59–87, https://doi.org/10.1146/annurev-cellbio-092910-154010 (2012).
Moncada, S., Higgs, E. A. & Colombo, S. L. Fulfilling the metabolic requirements for cell proliferation. Biochem J 446, 1–7, https://doi.org/10.1042/BJ20120427 (2012).
Jorgensen, P. & Tyers, M. How cells coordinate growth and division. Curr Biol 14, R1014–1027, https://doi.org/10.1016/j.cub.2004.11.027 (2004).
Ahn, W. S. & Antoniewicz, M. R. Metabolic flux analysis of CHO cells at growth and non-growth phases using isotopic tracers and mass spectrometry. Metab Eng 13, 598–609, https://doi.org/10.1016/j.ymben.2011.07.002 (2011).
Birsoy, K. et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508, 108–112, https://doi.org/10.1038/nature13110 (2014).
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc Natl Acad Sci USA 104, 19345–19350, https://doi.org/10.1073/pnas.0709747104 (2007).
Trotzmuller, M., Guo, X., Fauland, A., Kofeler, H. & Lankmayr, E. Characteristics and origins of common chemical noise ions in negative ESI LC-MS. J Mass Spectrom 46, 553–560, https://doi.org/10.1002/jms.1924 (2011).
Yao, C. H., Liu, G. Y., Yang, K., Gross, R. W. & Patti, G. J. Inaccurate quantitation of palmitate in metabolomics and isotope tracer studies due to plastics. Metabolomics 12, doi:ARTN 14310.1007/s11306-016-1081-y (2016).
Fan, T. W.-M., Lane, A. N. & Higashi, R. M. The handbook of metabolomics. (Springer 2012).
Dietmair, S., Timmins, N. E., Gray, P. P., Nielsen, L. K. & Kromer, J. O. Towards quantitative metabolomics of mammalian cells: development of a metabolite extraction protocol. Anal Biochem 404, 155–164, https://doi.org/10.1016/j.ab.2010.04.031 (2010).
Ser, Z., Liu, X., Tang, N. N. & Locasale, J. W. Extraction parameters for metabolomics from cultured cells. Anal Biochem 475, 22–28, https://doi.org/10.1016/j.ab.2015.01.003 (2015).
Fan, T. W.-M. In The Handbook of Metabolomics (eds Teresa Whei-Mei Fan, Andrew N. Lane, & Richard M. Higashi) 7–27 (Humana Press 2012).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J Lipid Res 49, 1137–1146, https://doi.org/10.1194/jlr.D700041-JLR200 (2008).
Fan, T. W. M., Bandura, L. L., Higashi, R. M. & Lane, A. N. Metabolomics-edited transcriptomics analysis of Se anticancer action in human lung cancer cells. Metabolomics 1, 325–339, https://doi.org/10.1007/s11306-005-0012-0 (2005).
Wittmann, C. Fluxome analysis using GC-MS. Microb Cell Fact 6, 6, https://doi.org/10.1186/1475-2859-6-6 (2007).
Mairinger, T., Sanderson, J. & Hann, S. GC-QTOFMS with a low-energy electron ionization source for advancing isotopologue analysis in (13)C-based metabolic flux analysis. Anal Bioanal Chem, https://doi.org/10.1007/s00216-019-01590-y (2019).
Midani, F. S., Wynn, M. L. & Schnell, S. The importance of accurately correcting for the natural abundance of stable isotopes. Anal Biochem 520, 27–43, https://doi.org/10.1016/j.ab.2016.12.011 (2017).
Zeng, J. et al. Comprehensive Profiling by Non-targeted Stable Isotope Tracing Capillary Electrophoresis-Mass Spectrometry - A Novel Tool Complementing Metabolomics Analyses of Polar Metabolites. 0, https://doi.org/10.1002/chem.201900539.
Cocuron, J. C., Tsogtbaatar, E. & Alonso, A. P. High-throughput quantification of the levels and labeling abundance of free amino acids by liquid chromatography tandem mass spectrometry. J Chromatogr A 1490, 148–155, https://doi.org/10.1016/j.chroma.2017.02.028 (2017).
Cocuron, J.-C. & Alonso, A. P. In Plant Metabolic Flux Analysis 131–142 (Springer 2014).
Creek, D. J. et al. Stable isotope-assisted metabolomics for network-wide metabolic pathway elucidation. Analytical chemistry 84, 8442–8447, https://doi.org/10.1021/ac3018795 (2012).
Huang, X. et al. X13CMS: global tracking of isotopic labels in untargeted metabolomics. Analytical chemistry 86, 1632–1639, https://doi.org/10.1021/ac403384n (2014).
Poskar, C. H. et al. iMS2Flux – a high–throughput processing tool for stable isotope labeled mass spectrometric data used for metabolic flux analysis. 13, 295, https://doi.org/10.1186/1471-2105-13-295 (2012).
Capellades, J. et al. geoRge: A Computational Tool To Detect the Presence of Stable Isotope Labeling in LC/MS-Based Untargeted Metabolomics. Analytical Chemistry 88, 621–628, https://doi.org/10.1021/acs.analchem.5b03628 (2016).
Chokkathukalam, A. et al. mzMatch-ISO: an R tool for the annotation and relative quantification of isotope-labelled mass spectrometry data. Bioinformatics 29, 281–283, https://doi.org/10.1093/bioinformatics/bts674 (2013).
Mashego, M. R. et al. MIRACLE: mass isotopomer ratio analysis of U-C-13-labeled extracts. A new method for accurate quantification of changes in concentrations of intracellular metabolites. Biotechnology and Bioengineering 85, 620–628, https://doi.org/10.1002/bit.10907 (2004).
Lorkiewicz, P., Higashi, R. M., Lane, A. N. & Fan, T. W. High information throughput analysis of nucleotides and their isotopically enriched isotopologues by direct-infusion FTICR-MS. Metabolomics 8, 930–939, https://doi.org/10.1007/s11306-011-0388-y (2012).
Lane, A. N., Fan, T. W., Xie, Z., Moseley, H. N. & Higashi, R. M. Isotopomer analysis of lipid biosynthesis by high resolution mass spectrometry and NMR. Analytica chimica acta 651, 201–208, https://doi.org/10.1016/j.aca.2009.08.032 (2009).
Wei, X. et al. Analysis of Stable Isotope Assisted Metabolomics Data Acquired by High Resolution Mass Spectrometry. Anal Methods 9, 2275–2283, https://doi.org/10.1039/C7AY00291B (2017).
Elucidata. http://www.elucidata.io/polly (2019).
Agrawal, S. et al. In High-Throughput Metabolomics: Methods and Protocols Vol. 1978 (ed. D’Alessandro, A.) (Springer New York 2019).
Berglund, M. & Wieser, M. E. Isotopic compositions of the elements 2009 (IUPAC Technical Report). Pure and Applied Chemistry 83, 397–410, https://doi.org/10.1351/Pac-Rep-10-06-02 (2011).
Moseley, H. N. Correcting for the effects of natural abundance in stable isotope resolved metabolomics experiments involving ultra-high resolution mass spectrometry. BMC Bioinformatics 11, 139, https://doi.org/10.1186/1471-2105-11-139 (2010).
Alves, T. C. et al. Integrated, Step-Wise, Mass-Isotopomeric Flux Analysis of the TCA Cycle. Cell Metab 22, 936–947, https://doi.org/10.1016/j.cmet.2015.08.021 (2015).
Sauer, U. Metabolic networks in motion: 13C-based flux analysis. Mol Syst Biol 2, 62, https://doi.org/10.1038/msb4100109 (2006).
Antoniewicz, M. R. 13C metabolic flux analysis: optimal design of isotopic labeling experiments. Curr Opin Biotechnol 24, 1116–1121, https://doi.org/10.1016/j.copbio.2013.02.003 (2013).
Crown, S. B. & Antoniewicz, M. R. Parallel labeling experiments and metabolic flux analysis: Past, present and future methodologies. Metab Eng 16, 21–32, https://doi.org/10.1016/j.ymben.2012.11.010 (2013).
Gibb, A. A. et al. Integration of flux measurements to resolve changes in anabolic and catabolic metabolism in cardiac myocytes. Biochem J 474, 2785–2801, https://doi.org/10.1042/BCJ20170474 (2017).
Chen, P. H. et al. Metabolic diversity in human non-small cell lung cancer cells. bioRxiv, https://doi.org/10.1101/561688 (2019).
Sellers, K. et al. Pyruvate carboxylase is critical for non-small-cell lung cancer proliferation. J Clin Invest 125, 687–698, https://doi.org/10.1172/JCI72873 (2015).
Lahey, R. et al. Enhanced Redox State and Efficiency of Glucose Oxidation With miR Based Suppression of Maladaptive NADPH-Dependent Malic Enzyme 1 Expression in Hypertrophied Hearts. Circ Res 122, 836–845, https://doi.org/10.1161/CIRCRESAHA.118.312660 (2018).
Campbell, J. L., Le Blanc, J. C. & Kibbey, R. G. Differential mobility spectrometry: a valuable technology for analyzing challenging biological samples. Bioanalysis 7, 853–856, https://doi.org/10.4155/bio.15.14 (2015).
Hill, B. G., Dranka, B. P., Zou, L., Chatham, J. C. & Darley-Usmar, V. M. Importance of the bioenergetic reserve capacity in response to cardiomyocyte stress induced by 4-hydroxynonenal. Biochem J 424, 99–107, https://doi.org/10.1042/BJ20090934 (2009).
Sansbury, B. E., Jones, S. P., Riggs, D. W., Darley-Usmar, V. M. & Hill, B. G. Bioenergetic function in cardiovascular cells: the importance of the reserve capacity and its biological regulation. Chemico-biological interactions 191, 288–295, https://doi.org/10.1016/j.cbi.2010.12.002 (2011).
Jastroch, M., Divakaruni, A. S., Mookerjee, S., Treberg, J. R. & Brand, M. D. Mitochondrial proton and electron leaks. Essays Biochem 47, 53–67, https://doi.org/10.1042/bse0470053 (2010).
Chance, B. & Williams, G. R. Respiratory enzymes in oxidative phosphorylation. III. The steady state. J Biol Chem 217, 409–427 (1955).
Chance, B., Williams, G. R., Holmes, W. F. & Higgins, J. Respiratory enzymes in oxidative phosphorylation. V. A mechanism for oxidative phosphorylation. J Biol Chem 217, 439–451 (1955).
Chance, B. & Williams, G. R. Respiratory enzymes in oxidative phosphorylation. VI. The effects of adenosine diphosphate on azide-treated mitochondria. J Biol Chem 221, 477–489 (1956).
Heinrich, R. & Rapoport, T. A. A linear steady-state treatment of enzymatic chains. Critique of the crossover theorem and a general procedure to identify interaction sites with an effector. Eur J Biochem 42, 97–105 (1974).
Bouskila, M. et al. Allosteric regulation of glycogen synthase controls glycogen synthesis in muscle. Cell Metab 12, 456–466, https://doi.org/10.1016/j.cmet.2010.10.006 (2010).
Aiston, S., Andersen, B. & Agius, L. Glucose 6-phosphate regulates hepatic glycogenolysis through inactivation of phosphorylase. Diabetes 52, 1333–1339 (2003).
Moseley, H. N., Lane, A. N., Belshoff, A. C., Higashi, R. M. & Fan, T. W. A novel deconvolution method for modeling UDP-N-acetyl-D-glucosamine biosynthetic pathways based on (13)C mass isotopologue profiles under non-steady-state conditions. BMC Biol 9, 37, https://doi.org/10.1186/1741-7007-9-37 (2011).
Jones, S. P. et al. Cardioprotection by N-acetylglucosamine linkage to cellular proteins. Circulation 117, 1172–1182, https://doi.org/10.1161/CIRCULATIONAHA.107.730515 (2008).
Ngoh, G. A. et al. Unique hexosaminidase reduces metabolic survival signal and sensitizes cardiac myocytes to hypoxia/reoxygenation injury. Circ Res 104, 41–49, https://doi.org/10.1161/circresaha.108.189431 (2009).
Sansbury, B. E. et al. Responses of hypertrophied myocytes to reactive species: implications for glycolysis and electrophile metabolism. Biochemical Journal 435, 519–528 (2011).
Le, A. et al. Glucose-independent glutamine metabolism via TCA cycling for proliferation and survival in B cells. Cell Metab 15, 110–121, https://doi.org/10.1016/j.cmet.2011.12.009 (2012).
Salabei, J. K. et al. Type 2 Diabetes Dysregulates Glucose Metabolism in Cardiac Progenitor Cells. J Biol Chem 291, 13634–13648, https://doi.org/10.1074/jbc.M116.722496 (2016).
Adusumilli, R. & Mallick, P. Data Conversion with ProteoWizard msConvert. Methods Mol Biol 1550, 339–368, https://doi.org/10.1007/978-1-4939-6747-6_23 (2017).
Ashcroft, S. J., Weerasinghe, L. C., Bassett, J. M. & Randle, P. J. The pentose cycle and insulin release in mouse pancreatic islets. Biochem J 126, 525–532 (1972).
Radde, B. N. et al. Bioenergetic differences between MCF-7 and T47D breast cancer cells and their regulation by oestradiol and tamoxifen. Biochem J 465, 49–61, https://doi.org/10.1042/bj20131608 (2015).
Salabei, J. K. et al. Glutamine Regulates Cardiac Progenitor Cell Metabolism and Proliferation. Stem Cells 33, 2613–2627, https://doi.org/10.1002/stem.2047 (2015).
Acknowledgements
The authors acknowledge funding support from the National Institutes of Health (NIH) [BGH: HL130174, HL147844, ES028268, HL078825, GM127607; XZ: S10OD020106 and P20GM113226), an AHA predoctoral fellowship (AAG: 16PRE31010022), and the American Diabetes Association Pathway to Stop Diabetes Grant (BGH: ADA 1-16-JDF-041.)].
Author information
Authors and Affiliations
Contributions
P.K.L., project planning, execution of experiments, data analysis and interpretation, writing of manuscript; A.A.G., project planning, execution of experiments, data analysis and interpretation, writing of manuscript; B.R.R., execution of experiments, data analysis; L.H., execution of experiments; Y.Z., execution of experiments; B.F.C., data analysis and interpretation, writing of manuscript; X.Z., execution of experiments; B.G.H., project planning, data analysis and interpretation, writing of manuscript, financial support.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lorkiewicz, P.K., Gibb, A.A., Rood, B.R. et al. Integration of flux measurements and pharmacological controls to optimize stable isotope-resolved metabolomics workflows and interpretation. Sci Rep 9, 13705 (2019). https://doi.org/10.1038/s41598-019-50183-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50183-3
This article is cited by
-
Palbociclib treatment alters nucleotide biosynthesis and glutamine dependency in A549 cells
Cancer Cell International (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.