Abstract
Cholesterol oxidases are important enzymes with a wide range of applications from basic research to industry. In this study, we have discovered and described the first cell-associated cholesterol oxidase, ChoD, from Streptomyces lavendulae YAKB-15. This strain is a naturally high producer of ChoD, but only produces ChoD in a complex medium containing whole yeast cells. For characterization of ChoD, we acquired a draft genome sequence of S. lavendulae YAKB-15 and identified a gene product containing a flavin adenine dinucleotide binding motif, which could be responsible for the ChoD activity. The enzymatic activity was confirmed in vitro with histidine tagged ChoD produced in Escherichia coli TOP10, which lead to the determination of basic kinetic parameters with Km 15.9 µM and kcat 10.4/s. The optimum temperature and pH was 65 °C and 5, respectively. In order to increase the efficiency of production, we then expressed the cholesterol oxidase, choD, gene heterologously in Streptomyces lividans TK24 and Streptomyces albus J1074 using two different expression systems. In S. albus J1074, the ChoD activity was comparable to the wild type S. lavendulae YAKB-15, but importantly allowed production of ChoD without the presence of yeast cells.
Similar content being viewed by others
Introduction
Cholesterol oxidases (EC 1.1.3.6, 3β-hydroxysterol oxidase) are FAD-dependent (flavin adenine dinucleotide) enzymes, which oxidizes cholesterol to form cholest-4-en-3-one (cholestenone) and H2O2 (Fig. 1). Cholesterol oxidases have a broad range of applications including determination of food and serum cholesterol levels1, bioconversion of non-steroidal compounds2, allylic alcohols and sterols2, insecticidal activity3,4 and as a signal for the production of antifungal antibiotics5. Furthermore, cholesterol oxidases have been implicated in the manifestation of HIV, Alzheimer’s disease and tuberculosis6 and are needed for the biotransformation of cholesterol to cholestenone, which is an important precursor for the synthesis of hormones and steroidal drug intermediates7. More recently, cholesterol oxidases from Borodetella sp. have also been shown to promote cell apoptosis in lung adenocarcinoma8 and breast cancer9. Interestingly, enzymes extracted from Streptomyces sp. are typically preferred for industrial production as they are more stable than the ones isolated from Nocardia or Pseudomonas10.
Cholesterol oxidases in the context of basic biological function are utilized by various microorganisms to assimilate cholesterol as a carbon and energy source11. Two distinct classes of cholesterol oxidases have been found in microorganisms1. In many bacteria, including Streptomyces sp., cholesterol oxidases are intracellularly produced and secreted into the culture broth while in some others, e.g. Nocardia species, the enzyme is produced intrinsically membrane bound1,12. The intracellular proteins typically employ twin arginine transport (TAT) systems to export FAD-bound, fully folded proteins outside of cells, but the molecular details determining whether the proteins are bound to the extracellular matrix or secreted to the medium remain unresolved13,14,15. The strain S. lavendulae YAKB-15 has been noted to harbour significant cholesterol oxidase, ChoD, activity16, but appears to produce the enzyme only in a culture medium that contain whole yeast cells17. Originally this strain was cultivated with protein-vitamin concentrate as an organic nitrogen source, however, since it became unavailable other sources were tested (i.e. yeast extract, corn steep liquor, soy flour, peptone, casein hydrolysate, urea, beer yeast and baker’s yeast) and the use of baker’s yeast yielded the highest activity16. Interestingly, in contrast to several other strains of Streptomyces4,5,9,18,19, the activity is found associated with the fraction containing intact cells and only minor amounts can be detected from the supernatant.
In this study, we assembled a draft genome sequence of S. lavendulae YAKB-15 and identified a cholesterol oxidase, choD, gene in a putative operon (cho) containing two regulatory genes. Production of recombinant ChoD in Escherichia coli enabled the determination of kinetic parameters of the enzyme, whereas the overexpression in S. albus J1074 using a pSET-152-based vector20 led to comparative yields of ChoD production in comparison to S. lavendulae YAKB-15. Importantly, this allowed production of the ChoD in a simple medium without the presence of whole yeast cells.
Results and Discussion
Discovery of choD cholesterol oxidase from S. lavendulae YAKB-15
Streptomyces lavendulae YAKB-15 was found to produce ChoD under highly specific fermentation conditions15,16. The strain requires the presence of whole yeast cells, either live or autoclaved. No ChoD activity could be observed when yeast extract was used as a substitute in the production medium. In addition, the production appears to be strictly regulated in a temporal manner, with highest activity observed after 40 hours followed by rapid decline. Since these factors severely limit industrial strain development, we proceeded to obtain the draft genome sequence of S. lavendulae YAKB-15 and identify the gene responsible for ChoD production.
The sequencing data was acquired using MiSeq technology and resulted in 15,984,844 reads that were normalized, error corrected and trimmed down to 7,196,183 reads, which were then de novo assembled into 98 contigs. ABACAS ordered and aligned the contigs into 73 scaffolds with an N50 of 447,215 bp. The final genome assembly is 7.8 Mbp with a GC content of 72.2% and median coverage of 199x. The BUSCO analysis searched for 40 single-copy orthologs and found 36 (90%) were complete. Out of the 36 complete BUSCOs, 4 were found multiple times throughout the assembly and none were missing.
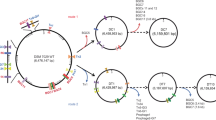
BLAST analysis was used to identify putative cholesterol oxidases and led to the discovery of a gene denoted choD that showed 82% identity to a cholesterol oxidase gene from Streptomyces sp. SA-COO (UniProt P1267621). The high sequence identity suggests that ChoD belongs to the glucose-methanol-choline oxidoreductase family22 and possesses the classical Rossman fold for dinucleotide binding, which is found in many flavin-dependent oxidases23. Further analysis of the gene product showed that all of the important catalytic residues were conserved (e.g. E389 and H484) and the N-terminal region contained a twin-arginine transport (TAT) signal21, which displayed 25/42 (60%) identity to P12676. The choD gene resides in a putative operon structure with six genes (Fig. 2). Two of which are regulatory genes of the LuxR24 and PadR25 families (Table 1). Related proteins in these families have been found to control many aspects of secondary metabolism in Streptomyces, including antibiotic production and resistance24,25,26,27, and therefore could be responsible for the transient transcription of choD. In addition, the cho operon contained three additional genes putatively encoding thioesterase, acyl-CoA dehydrogenase and methyltransferase (Table 1).
Enzyme kinetics of recombinant ChoD
In order to characterize ChoD, we ordered a synthetic gene codon optimized for expression in E. coli and cloned it in a modified pBAD vector. The N-terminally histidine-tagged ChoD was produced intracellularly in E. coli, possibly due to an impaired TAT-transport, and purified to near homogeneity by affinity chromatography (Figs 3a and S1). The enzyme activity was monitored spectrophotometrically by determining H2O2 concentration, which is formed during non-enzymatic oxidation of reduced flavin in the catalytic cycle, using a colour based reaction with ABTS (2,2′-azino-bis(3-ethylbenzthiazoline-6-sulfonic acid)) at 405 nm (Fig. 1). The progression curves displayed first-order kinetics leading to the determination of basic kinetic parameters for kcat (10.35 s−1) and Km (15.91 µM) (Fig. 3b). The affinity of ChoD towards cholesterol was higher than what has been reported for commercially available cholesterol oxidases from Brevibacterium (23 mM), Streptomyces (0.2 mM), Cellulomonas (84 µM) or Pseudomonas (61 µM)28. The Km of ChoD from Streptomyces lavendulae YAKB-15 resided between the values of the two enzymes from Streptomyces sp. SA-COO21 (3 µM) and B. sterolicum (>100 µM) for which crystal structures have been determined29. Furthermore, the substrate affinity of ChoD from Streptomyces lavendulae YAKB-15 is higher than that of the recently reported Streptomyces isolate, S. aegyptia NEAE 102 (152 µM)30. However, it should be noted that solubility issues and the use of detergents have been shown to have great influence on kinetic parameters of cholesterol oxidases, which makes comparing the properties of enzymes from various sources challenging31,32.
SDS-PAGE analysis of purified ChoD and spectrophotometric determination of enzyme kinetics for ChoD. (a) The SDS gel (cropped) was stained with Coomassie Blue and the original gel is presented in Supplementary Fig. S1. Lane MW: protein marker, Lane 1: purified ChoD. (b) Spectrophotometric assays were done in triplicate (grey squares) at seven different concentrations of cholesterol.
Heterologous production of ChoD in S. lividans TK24 and S. albus J1074
In order to improve the production of ChoD, we opted to utilize two widely used and well-characterized Streptomyces hosts, S. lividans TK2420 and S. albus J107420, and two distinct vectors to drive the expression of the ChoD. For expression in S. lividans TK24, choD was cloned in the multi-copy plasmid pIJE48620 under the strong constitutive promoter ermEp by protoplast transformation. For expression in S. albus J1074, we elected to use the integrative single copy-number plasmid pS-GK, which is based on the pSET-15220 plasmid but contains a strong synthetic promoter SP4433, introduced into Streptomyces by intergeneric conjugation from E. coli ET12567/pUZ8002.
The native strain, S. lavendulae YAKB-15, produced ChoD rapidly, with the highest activity being 1.25 U/mL at 40 hours in Y medium containing whole yeast cells, whereas no production could be observed in YE medium (Fig. 4), where the whole yeast cells were replaced by yeast extract. S. albus J1074 produced ChoD in a similar fashion, but with lower activity (0.4 U/mL) in Y medium (Fig. 4a). In YE medium S. albus J1074 was the only strain able to produce significant amounts of ChoD, with the highest activity at 115 hours and a maximum of 0.78 U/mL (Fig. 4b). Curiously, S. lividans TK24 produced very little ChoD in both Y and YE medium (Fig. 4).
The production levels of the native strain, S. lavendulae YAKB-15, and the overexpression strain, S. albus J1074, are in line with previously reported levels from Streptomyces and Rhodococcus, which range from 0.2 U/mL to 9.75 U/mL18,34,35,36,37,38,39,40. However, it should be noted that in many previous reports the medium was extensively optimized to increase production levels. Furthermore, S. lavendulae YAKB-15 has the highest basal level of activity (1.25 U/mL) for a cell-associated cholesterol oxidase from Actinomycetales (Rhodococcus, 0.75 U/mL)37.
Properties of heterologously produced ChoD in S. albus J1074
The activity of cell-associated ChoD extract produced by the overexpression strain S. albus J1074/pS_ChoD was characterized using different temperatures and pH. The optimal temperature and pH were determined from a range of 25–75 °C and 4–9, respectively (Fig. 5). The optimal temperature was 65 °C and was dramatically lower (<60%) at any other tested temperature (Fig. 5a). The optimal pH was 5, although pH 6 and pH 7 both had 90% relative activity (Fig. 5b). Both the temperature and pH optima were in line with previously reported cholesterol oxidases as summarized by El-Naggar et al.30.
Concluding remarks
In this study we have identified and characterized a cell-associated cholesterol oxidase, ChoD, from S. lavendulae YAKB-15. To the best of our knowledge ChoD has the highest cholesterol affinity (Km 15.91 µM) and the highest basal activity (1.25 U/mL) of a cell-associated cholesterol oxidase. The optimum temperature and pH was 65 °C and 5, respectively. The presence of the TAT signal indicates that the protein is likely to be produced intracellularly in order to recruit FAD and is exported outside the cell membrane as a fully matured enzyme through the transport system. Since only minor enzymatic activity can be detected from fermentation broths, unlike in the case of other cholesterol oxidase proteins from Streptomyces, it is likely that the protein from S. lavendulae YAKB-15 becomes associated with components of the cell wall. Notably, after heterologous expression of the choD in S. lividans TK24 and S. albus J1074 the ChoD activity was still associated with the cell fraction and not the supernatant. The fact that S. lavendulae YAKB-15 only produces ChoD in the presence of whole-yeast raises the possibility that the strain utilizes ChoD as a signalling molecule to detect Streptomyces-fungal interactions. Such a role has been proposed for cholesterol oxidases residing in biosynthetic gene clusters responsible for production of several antifungal polyene macrolides5. However, no polyene gene clusters are found near choD in S. lavendulae YAKB-15. Future work currently in progress in our laboratory aims to uncover the role of ChoD in the biology of S. lavendulae YAKB-15.
Methods
Genomic DNA isolation and whole genome sequencing
S. lavendulae YAKB-15 was grown in 250 mL Erlenmeyer flasks containing 30 mL of GYM medium consisting of glucose 4 g/L, yeast extract 4 g/L, malt extract 10 g/L and 0.5% glycine. The pH of the medium was adjusted to 7.2 and the culture was incubated on a rotary shaker (300 rpm) at 30 °C for 96 h. Genomic DNA was extracted using the method from Nikodinovic et al. with slight modifications41. The DNA was sent to Eurofins Genomics (Ebersberg, Germany) for PCR-free shotgun library preparation (Illumina) and sequenced using MiSeq v3 producing 2 × 300 bp paired-end reads (Illumina).
The quality of the reads was manually checked before and after trimming and error correction using FASTQC (v0.11.2)42. The reads were normalized using BBNorm. Then the reads were error corrected and assembled using A5-miseq (v20150522)43, contiguated with ABACAS (v1.3.1)44 using Streptomyces albus NK660 (CP007574.1) as the reference, and the gaps were filled using IMAGE (v2.4.1)45. The final assembly was annotated using RAST46 and evaluated for completeness using BUSCO (v1.22)47. All programs were run with the default parameters on the CSC – IT Center for Science’s Taito super-cluster (Espoo, Finland). The final assembly was deposited in the National Center for Biotechnology Information (NCBI) database under the accession number SMSN00000000.
In silico analysis of cholesterol oxidase from S. lavendulae YAKB-15
The choD gene was identified in the assembly of S. lavendulae YAKB-15 using local Protein BLAST48 and UniProt sequence P12676 as a query. The sequence was further analysed by comparing important catalytic residues to the found gene31. This gene was targeted for cloning and recombinant expression in three different systems.
Expression systems
Three different expression systems were used to overproduce and quantify the production of ChoD via standard cloning methods20. First, an E. coli codon optimized synthetic choD gene was ordered (ThermoFisher Scientific) and cloned in a pBADΔHis49 expression plasmid using BglII/HindIII restriction sites, creating pBAD_ChoD, which was transformed into E. coli TOP10 (Invitrogen). Then the native choD gene was PCR amplified using the primer pair 5′-GCGTCTAGAGAAGCTCAGGAGCAACAGCG-3′ (XbaI site underlined) and 5′-CGAAGCTTGGATCCTCAGGAACCCGCGATGTCC-3′ (HindIII and BamHI sites underlined) from genomic DNA using Phusion high-fidelity DNA polymerase (ThermoFisher Scientific) and cloned in pUC18 (ThermoFisher Scientific) using XbaI/HindIII restriction sites and then transformed into E. coli TOP10 (Invitrogen). The DNA sequence of the cloned gene was confirmed by sequencing before subsequent subcloning. Second, the native choD gene was digested from the pUC18 cloning plasmid and ligated using the same restriction sites (XbaI/HindIII) in the expression plasmid pIJE48650, creating pIJ_ChoD, and then protoplast transformed into the expression host S. lividans TK24. Third, the native choD gene was digested from the pUC18 expression plasmid using XbaI/BamHI restriction sites and ligated in a modified pSET15251 expression vector using BcuI/BamHI restriction sites, creating pS_ChoD, and then transformed into E. coli ET12567/pUZ800220, which was then conjugated into the expression host S. albus J1074. The plasmid pS_ChoD contained choD and superfolder green fluorescence protein (sfGFP) genes with the corresponding ribosomal binding sites under the control of the strong synthetic promoter SP4433. To avoid promoter leakage due to the read-through from the upstream genes (i.e. bacteriophage phi31 integrase and apramycin resistance genes) two strong terminator, a synthetic T4 kurz52 and a natural terminator ECK12002960053 were placed upstream of the promoter.
Bacterial strains and culture conditions
S. lavendulae YAKB-15 was obtained from the Russian Collection of Agricultural Microorganisms (RCAM). S. lividans TK24 and S. albus J1074 originate from the John Innes Centre20. E. coli TOP10 (Invitrogen) was used for production of the histidine tagged ChoD.
S. lavendulae YAKB-15 and S. albus J1074/pS_ChoD were first grown on solid P medium containing 1 g/L peptone, 4.55 g/L glucose anhydrase, 0.4 g/L MgSO4 * 7H2O, 0.4 g/L K2HPO4, 22 g/L agar, and 100 g/L potato juice, until they sporulated. S. lividans TK24/ pIJ_ChoD was grown on ISP4 with 50 μg/mL thiostrepton until it sporulated. Spores were inoculated into 25 mL liquid medium, either Y medium or YE medium, in 250 mL Erlenmeyer flasks; S. lividans TK24 also contained 50 μg/mL thiostrepton. Y medium contains 9.1 g/L glucose anhydrase, 2 g/L NH4NO3, 2 g/L CaCO3, and 26 g/L common bakery yeast and YE medium is the same except yeast extract substituted common bakery yeast. Liquid pre-cultures were grown at 30 °C shaking at 300 rpm for 24 hours. These pre-cultures were used to inoculate main cultures in triplicates.
For production of ChoD in E. coli TOP10/pBAD_ChoD, 2 × 500 mL of 2 x TY medium (tryptone 16 g/L, yeast extract 10 g/L, NaCl 5 g/L) or TB medium (tryptone 20 g/L, yeast extract 24 g/L, glycerol 4 mL/L, phosphate buffer (0.17 M KH2PO4 and 0.72 M K2HPO4) 100 mL/L) with 100 µg/mL of ampicillin were inoculated with 5 mL of pre-culture per flask. Cultures were cultivated at 37 °C for 3 hours with 250 rpm shaking and were induced with 0.02% L-(+)-Arabinose when the OD600 was 1.15. After induction, the cultivation was continued for 15.5 hours at 25 °C with 200 rpm shaking. The cells were harvested by centrifugation at 12,000 × g for 30 minutes at 4 °C resulting in wet cell weight of 4.3 g.
For characterization of ChoD properties S. albus J1074/pS_ChoD was pre-cultured in 15 mL of TSB medium containing 17 g/L tryptone, 3 g/L soy, 5 g/L NaCl, 2.5 g/L K2HPO4, and 2.5 g/L glucose inoculated from ISP4 spore plates and grown at 30 °C for 24 hours at 300 rpm. The main cultures were also grown in 15 mL of TSB, using 1 mL of pre-culture as inoculum, for at 30 °C for 40 hours at 300 rpm.
Purification of recombinant proteins
For purification of histidine tagged ChoD, the cells were suspended in 3 mL wash buffer (K2HPO4 50 mM, imidazole 5 mM, NaCl 50 mM, 10% glycerol) per gram of cells. The cells were sonicated with a cycle of 11 s of sonication and 40 s of rest on ice. The cycle was repeated 13 times (Sonicator MSE soniprep 150 with max amplitude). 1% of Triton X-100 was added after sonication to the supernatant. Samples were centrifuged 19,000 × g at 4 °C for 30 minutes and the supernatant was collected. The supernatant was mixed with 1 mL of TALON affinity resin (GE healthcare) and it was gently shaken for 60 minutes. The resin was washed with 5 mL of wash buffer and the protein was eluted with 2.5 mL of elution buffer (K2HPO4 50 mM, imidazole 250 mM, NaCl 50 mM, 10% glycerol). The buffer was changed to storage buffer (K2HPO4 50 mM, NaCl 50 mM, 10% glycerol) using a PD-10-column following manufacturer’s instructions. Finally, the glycerol concentration was increased to 40% and purified ChoD was stored at −20 °C. Purified ChoD was evaluated by SDS-PAGE 10%.
Analysis of cholesterol oxidase activity and enzyme kinetics
ChoD activity was measured spectrophotometrically by the modified method of Sasaki et al. The stoichiometric formation of H2O2 during the oxidation reaction of cholesterol was monitored with ABTS (2,2′-azino-bis(3-ethylbenzthiazoline-6-sulfonic acid)) at 405 nm. To determine the cell-bound ChoD, cultures were centrifuged at 15,000 × g for 10 min. The cell pellet was resuspended in extraction buffer (0.15% Tween 80 in 50 mM phosphate buffer solution) and mixed for 30 minutes at 24 °C. The suspension was centrifuged at 15,000 × g and ChoD activity was measured from the supernatant. The activity assay mixture contained 120 μL Triton X-100 (0.05%) in 50 mM sodium-potassium phosphate buffer (pH 7), 10 μL ABTS (9.1 mM in MQ H2O), 2.5 μL cholesterol in ethanol (1 mg/mL), 1.5 μL horseradish peroxidase solution (150 U/mL) and 20 μL of the extract preparation in a total volume of 154 μL. The spectrophotometric cholesterol activity assay was carried out in a 96-well plate. One unit of enzyme was defined as the amount of enzyme that forms 1 μmol of H2O2 per minute at pH 7.0 and 27 °C.
ChoD optimal activity for various pH levels was determined as above with only changes in the buffer as needed for specific pH tests as follows: 50 mM citrate buffer (pH 4–5), 50 mM potassium phosphate buffer (pH 6–7), and 50 mM Tris-HCl buffer (pH 8–9). For optimal temperature activity, temperatures between 25 and 75 °C were obtained using heat blocks. Each condition was tested in triplicates.
For analysis of enzyme kinetics, 8.4 nM ChoD was utilized to probe reaction velocities with eight substrate concentrations ranging between 4–168 µM cholesterol in triplicates. The initial rate of the reaction was calculated from derivatives of progression curves (six initial measurement points over 25 s) and referenced to a H2O2 standard curve.
References
Pollegioni, L., Piubelli, L. & Molla, G. Cholesterol oxidase: biotechnological applications. FEBS J 276, 6857–6870 (2009).
Doukyu, N. Characteristics and biotechnological applications of microbial cholesterol oxidases. Appl. Microbiol. Biotechnol. 83, 825–837 (2009).
Bavari, S. et al. Lipid raft microdomains: a gateway for compartmentalized trafficking of ebola and marburg viruses. J. Exp. Med. 195, 593–602 (2002).
Purcell, J. P. et al. Cholesterol oxidase: a potent insecticidal protein active against boll weevil larvae. Biochem. Biophys. Res. Commun. 196, 1406–1413 (1993).
Mendes, M. V. et al. Cholesterol oxidases act as signaling proteins for the biosynthesis of the polyene macrolide pimaricin. Chem. Biol. 14, 279–290 (2007).
Kumari, L. & Kanwar, S. S. Cholesterol oxidase and its applications. Adv. Microbiol. 02, 49–65 (2012).
Ahmad, S. & Goswami, P. Application of chitosan beads immobilized Rhodococcus sp. NCIM 2891 cholesterol oxidase for cholestenone production. Process Biochem. 49, 2149–2157 (2014).
Liu, J. et al. Cholesterol oxidase from Bordetella species promotes irreversible cell apoptosis in lung adenocarcinoma by cholesterol oxidation. Cell Death Dis. 5, e1372–e1372 (2014).
El-Naggar, N. E.-A., Soliman, H. M. & El-Shweihy, N. M. Extracellular cholesterol oxidase production by Streptomyces aegyptia, in vitro anticancer activities against rhabdomyosarcoma, breast cancer cell-lines and in vivo apoptosis. Sci. Rep 8, 2706 (2018).
Lolekha, P. H. & Jantaveesirirat, Y. Streptomyces: a superior source for cholesterol oxidase used in serum cholesterol assay. J. Clin. Lab. Anal. 6, 405–9 (1992).
Stadtman, T., Cherkes, A. & Anfinsen, C. Studies on the microbiological degradation of cholesterol. J. Biol. Chem. 206, 511–23 (1954).
Kumari, L. & Shamsher, K. S. Cholesterol oxidase: role in biotransformation of cholesterol. J. Appl. Biol. Biotechnol 3, 53–65 (2015).
Barnett, J. P., Eijlander, R. T., Kuipers, O. P. & Robinson, C. A minimal Tat system from a Gram-positive organism. J. Biol. Chem. 283, 2534–2542 (2008).
Berks, B. C., Palmer, T. & Sargent, F. The Tat protein translocation pathway and its role in microbial physiology. Adv. Microb. Physiol 47, 187–254 (2003).
Widdick, D. A. et al. The twin-arginine translocation pathway is a major route of protein export in Streptomyces coelicolor. Proc. Natl. Acad. Sci. 103, 17927–17932 (2006).
Petrova, L. I., Podsukhina, G. M., Dikun, T. A. & Selezneva, A. A. Conditions for isolation of cholesterol oxidase from Actinomyces lavendulae mycelia. Prikl. Biokhim. Mikrobiol. 15, 250–3 (1979).
Buldakova, T. & Kolodyaznaya, V. Использование альтернативных источников органического азота при биосинтезе холестеролоксидаза Utilization of alternative organic nitrogen sources in cholesterol oxidase biosynthesis. In VII International Research Conference ‘Actual Scientific Achievements - 2012’ 35–38 (2012).
Varma, R. & Nene, S. Biosynthesis of cholesterol oxidase by Streptomyces lavendulae NCIM 2421. Enzyme Microb. Technol. 33, 286–291 (2003).
Ishizaki, T., Hirayama, N., Shinkawa, H., Nimi, O. & Murooka, Y. Nucleotide sequence of the gene for cholesterol oxidase from a Streptomyces sp. J. Bacteriol. 171, 596–601 (1989).
Kieser, T., Bibb, M., Buttner, M., Chater, K. & Hopwood, D. Practical Streptomyces Genetics. (The John Innes Foundation, 2000).
Yue, Q. K., Kass, I. J., Sampson, N. S. & Vrielink, A. Crystal structure determination of cholesterol oxidase from Streptomyces and structural characterization of key active site mutants. Biochemistry 38, 4277–4286 (1999).
Cavener, D. R. GMC oxidoreductases. A newly defined family of homologous proteins with diverse catalytic activities. J. Mol. Biol. 223, 811–4 (1992).
Wierenga, R. K., Drenth, J., Schulz, G. E. & Huber, R. Comparison of the three-dimensional protein and nucleotide structure of the FAD-binding domain of p-hydroxybenzoate hydroxylase with the FAD- as well as NADPH-binding domains of glutathione reductase. J. Mol. Biol. 167, 725–739 (1983).
Chen, J. & Xie, J. Role and regulation of bacterial LuxR-like regulators. J. Cell. Biochem. 112, 2694–2702 (2011).
Isom, C. E. et al. Crystal structure and DNA binding activity of a PadR family transcription regulator from hypervirulent Clostridium difficile R20291. BMC Microbiol. 16, 231 (2016).
Anton, N., Mendes, M. V., Martin, J. F. & Aparicio, J. F. Identification of PimR as a positive regulator of pimaricin biosynthesis in Streptomyces natalensis. J. Bacteriol. 186, 2567–2575 (2004).
Vicente, C. M. et al. PAS-LuxR transcriptional control of filipin biosynthesis in S. avermitilis. Appl. Microbiol. Biotechnol. 98, 9311–9324 (2014).
Srisawasdi, P., Jearanaikoon, P., Kroll, M. H. & Lolekha, P. H. Performance characteristics of cholesterol oxidase for kinetic determination of total cholesterol. J. Clin. Lab. Anal. 19, 247–252 (2005).
Vrielink, A., Lloyd, L. F. & Blow, D. M. Crystal structure of cholesterol oxidase from Brevibacterium sterolicum refined at 1.8 Å resolution. J. Mol. Biol. 219, 533–554 (1991).
El-Naggar, N. E. A., Deraz, S. F., Soliman, H. M., El-Deeb, N. M. & El-Shweihy, N. M. Purification, characterization and amino acid content of cholesterol oxidase produced by Streptomyces aegyptia NEAE 102. BMC Microbiol. 17, 1–12 (2017).
Vrielink, A. & Ghisla, S. Cholesterol oxidase: Biochemistry and structural features. FEBS J 276, 6826–6843 (2009).
Reiss, R., Faccio, G., Thöny-Meyer, L. & Richter, M. Cloning, expression and biochemical characterization of the cholesterol oxidase CgChoA from Chryseobacterium gleum. BMC Biotechnol. 14 (2014).
Bai, C. et al. Exploiting a precise design of universal synthetic modular regulatory elements to unlock the microbial natural products in Streptomyces. Proc. Natl. Acad. Sci. 112, 12181–12186 (2015).
Pathak, L. et al. Artificial intelligence versus statistical modeling and optimization of cholesterol oxidase production by using Streptomyces sp. PLoS One 10, e0137268 (2015).
Niwas, R., Singh, V., Singh, R., Tripathi, D. & Tripathi, C. K. M. Production, purification and characterization of cholesterol oxidase from a newly isolated Streptomyces sp. World J. Microbiol. Biotechnol. 29, 2077–2085 (2013).
Fernández de las Heras, L. et al. ChoG is the main inducible extracellular cholesterol oxidase of Rhodococcus sp. strain CECT3014. Microbiol. Res. 166, 403–418 (2011).
Ahmad, S. & Goswami, P. Enhanced production of cell-bound cholesterol oxidase from Rhodococcus sp. NCIM 2891 by the statistical method. Ann. Microbiol. 63, 199–205 (2013).
Chauhan, A. K., Survase, S. A., Kishenkumar, J. & Annapure, U. S. Medium optimization by orthogonal array and response surface methodology for cholesterol oxidase production by Streptomyces lavendulae NCIM 2499. J. Gen. Appl. Microbiol 55, 171–80 (2009).
Moradpour, Z., Ghasemian, A., Safari, A., Mohkam, M. & Ghasemi, Y. Isolation, molecular identification and statistical optimization of culture condition for a new extracellular cholesterol oxidase-producing strain using response surface methodology. Ann. Microbiol. 63, 941–950 (2013).
El-Naggar, N. E.-A., El-Shweihy, N. M. & El-Ewasy, S. M. Identification and statistical optimization of fermentation conditions for a newly isolated extracellular cholesterol oxidase-producing Streptomyces cavourensis strain NEAE-42. BMC Microbiol. 16, 217 (2016).
Nikodinovic, J., Barrow, K. D. & Chuck, J. A. High yield preparation of genomic DNA from Streptomyces. Biotechniques 35, 932–936 (2003).
Andrews, S. FASTQC a quality control tool for high throughput sequence data. Babraham Institute (2015). Available at, http://www.bioinformatics.babraham.ac.uk/projects/fastqc/Help/3 Analysis Modules/. (Accessed: 9th February 2016).
Coil, D., Jospin, G. & Darling, A. E. A5-miseq: an updated pipeline to assemble microbial genomes from Illumina MiSeq data. Bioinformatics 31, 587–589 (2015).
Assefa, S., Keane, T. M., Otto, T. D., Newbold, C. & Berriman, M. ABACAS: algorithm-based automatic contiguation of assembled sequences. Bioinformatics 25, 1968–1969 (2009).
Tsai, I. J., Otto, T. D. & Berriman, M. Improving draft assemblies by iterative mapping and assembly of short reads to eliminate gaps. Genome Biol. 11, R41 (2010).
Brettin, T. et al. RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci. Rep 5, 8365 (2015).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Kallio, P., Sultana, A., Niemi, J., Mäntsälä, P. & Schneider, G. Crystal structure of the polyketide cyclase AknH with bound substrate and product analogue: implications for catalytic mechanism and product stereoselectivity. J. Mol. Biol. 357, 210–220 (2006).
Ylihonko, K., Tuikkanen, J., Jussila, S., Cong, L. & Mäntsälä, P. A gene cluster involved in nogalamycin biosynthesis from Streptomyces nogalater: sequence analysis and complementation of early-block mutations in the anthracycline pathway. MGG Mol. Gen. Genet. 251, 113–120 (1996).
Flett, F., Mersinias, V. & Smith, C. P. High efficiency intergeneric conjugal transfer of plasmid DNA from Escherichia coli to methyl DNA-restricting streptomycetes. FEMS Microbiol. Lett 155, 223–229 (2006).
Horbal, L., Siegl, T. & Luzhetskyy, A. A set of synthetic versatile genetic control elements for the efficient expression of genes in Actinobacteria. Sci. Rep 8, 491 (2018).
Chen, Y.-J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Acknowledgements
The authors wish to acknowledge CSC – IT Center for Science, Finland, for computational resources. As well as the Jane and Aatos Erkko Foundation, Finland, and the Saint Petersburg State Chemical Pharmaceutical University, Russia, for kindly providing funds for materials and sequencing. Furthermore, the deposition and long-term storage of the strain, S. lavendulae YAKB-15, in the RCAM collection was supported by the Program for the Development and Inventory of Bioresource Collections, Russia.
Author information
Authors and Affiliations
Contributions
K.Y. was responsible for experiment planning and execution, data analysis and writing the manuscript. A.K. was responsible for cloning, enzymatic work and data analysis. M.L. was responsible for cloning. N.O. was responsible for the enzymatic work. A.A. was involved in the cloning and experimental design. V.S. and T.B. were responsible for DNA work and experimental design. E.Y. and V.K. were responsible for S. lavendulae YAKB-15 strain development. M.M.K. was involved in the experimental design, data analysis, manuscript writing and funding.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamada, K., Koroleva, A., Laughlin, M. et al. Characterization and overproduction of cell-associated cholesterol oxidase ChoD from Streptomyces lavendulae YAKB-15. Sci Rep 9, 11850 (2019). https://doi.org/10.1038/s41598-019-48132-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-48132-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.