Abstract
Iron is vital for nearly all living organisms, but during infection, not readily available to pathogens. Infectious bacteria therefore depend on specialized mechanisms to survive when iron is limited. These mechanisms make attractive targets for new drugs. Here, by genome-wide phenotypic profiling, we identify and categorize mycobacterial genes required for low iron fitness. Mycobacterium tuberculosis (Mtb), the causative agent of tuberculosis (TB), can scavenge host-sequestered iron by high-affinity iron chelators called siderophores. We take advantage of siderophore redundancy within the non-pathogenic mycobacterial model organism M. smegmatis (Msmeg), to identify genes required for siderophore dependent and independent fitness when iron is low. In addition to genes with a potential function in recognition, transport or utilization of mycobacterial siderophores, we identify novel putative low iron survival strategies that are separate from siderophore systems. We also identify the Msmeg in vitro essential gene set, and find that 96% of all growth-required Msmeg genes have a mutual ortholog in Mtb. Of these again, nearly 90% are defined as required for growth in Mtb as well. Finally, we show that a novel, putative ferric iron ABC transporter contributes to low iron fitness in Msmeg, in a siderophore independent manner.
Similar content being viewed by others
Introduction
Tuberculosis (TB), caused by Mycobacterium tuberculosis (Mtb), killed 1.6 million humans in 20181. The emerging threat of drug resistant Mtb adds burden to an already existing world health problem, and calls for urgent identification of new mycobacterial drug targets. Mtb depends on specialized mechanisms to access iron during infection2,3,4,5. These mechanisms make attractive targets for TB drug development6,7. Mycobacteria produce and secrete two types of high affinity iron chelators, siderophores, to scavenge ferric (Fe3+) iron; the mycobactins (here referring to both the insoluble membrane-embedded mycobactin and the soluble carboxymycobactin) and exochelin. Exochelin is produced only by rapidly growing, saprophytic mycobacteria like M. smegmatis (Msmeg) and M. neoaurum, while mycobactins are found in most mycobacterial species. The biosynthesis of siderophores and their transport across the inner membrane is well understood; however, transport across the outer membrane remains largely unknown. In addition to siderophores, mycobacteria can acquire iron by heme uptake (hemophores), sequestration of holo-transferrin and holo-lactoferrin, and through low-affinity porins (mycobacterial iron acquisition is thoroughly reviewed elsewhere7,8,9,10). Mtb is able to persist at low iron levels over a prolonged period of time, and launches a transcriptional response to adapt to iron availability11. The iron dependent repressor IdeR regulates one-third of the mycobacterial genes found to be repressed by iron, including genes for siderophore pathways12. Another protein, HupB, is shown to positively regulate Mtb siderophore biosynthesis in response to iron13. Furthermore, low environmental iron affects mycobacterial cell wall integrity (as well as membrane microvesicle production14), and the activity of diverse iron-containing enzymes7,15. Mechanisms not directly implicated in uptake of iron per se may, thus, play a pivotal role in mycobacterial fitness under iron-limited conditions.
Msmeg is widely used to study basic mycobacterial mechanisms, due to its non-pathogenic and fast-growing nature16. Pathways that promote low iron proliferation, separate from exochelin, are likely to be conserved within mycobacteria. In fact, we demonstrated the need of ESX-3, a type VII secretion system, for mycobactin-mediated iron uptake using Msmeg17,18, a finding later confirmed in Mtb19,20. Here, we take advantage of siderophore redundancy within Msmeg and define and categorize genes required for mycobactin, exochelin or siderophore independent modes of low iron growth by transposon insertion sequencing (Tnseq)21. (In fact, the power of Msmeg siderophore redundancy to identify genetic interaction partners to siderophore pathways was previously demonstrated by Judd el al, in a proof-of-principle synthetic genetic array22). To accomplish this, we developed protocols for efficient transposon mutagenesis in Msmeg23, which also allowed us to perform the first genome-wide identification of in vitro essential genes in this species. Interestingly, we found that the vast majority (90%) of Msmeg essential genes had a growth-required mutual ortholog in Mtb. Moreover, our low iron screens identified candidate genes for recognition, transport or utilization of siderophores, as well as new siderophore-independent low iron fitness mechanisms. For validation, we constructed a directed knockout of one of our hits, msmeg_3635 (a homologue of a ferric iron ABC permease), and found that this gene is indeed important during Msmeg iron starvation and appears to function independently of siderophores.
Results
Msmeg in vitro essential genes
Transposon mutagenesis, via mycobacteriophage ϕMycomarT7, has been less efficient in Msmeg than other mycobacterial species24. By systematically altering our transduction protocols we were able to obtain high-density transposon libraries also in Msmeg mc215523. The Himar1 transposon of ϕMycomarT7 inserts relatively randomly in the 77,755 TA dinucleotides present in the Msmeg genome. To confirm good representation, we selected a library of ~500,000 Msmeg mutants selected on Middlebrook 7H10 and performed Tnseq of three independent DNA libraries. We found an average of 1 345,678 unique template counts (transposon insertion counts per TA site), covering 78% of the TA sites in the genome, with an average count of 22 per TA (excluding TA’s with zero insertions). Transposon insertions were evenly distributed throughout the genome (Fig. 1a).
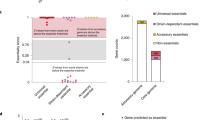
Msmeg in vitro growth requirement analysis. (a) Transposon insertion counts across the Msmeg genome (Middlebrook 7H10-selected library). The height of the black bars represents the number of insertion counts at the respective genome site. (b) Definition of the Msmeg wt in vitro requirement for growth (Middlebrook 7H10-selected library). 5815 genes were identified as non-essential (NE) for growth, 415 as causing growth advantage when disrupted (GA), 306 as essential (ES), and 97 as causing growth defect when disrupted (GD). 83 genes were not defined. (c) Venn diagram illustrating Msmeg required genes (ES and GD) and Mtb required genes (ES, GD and ESD, the latter genes with essential domains) as defined by DeJesus et al.30, relative to the entire pool of Msmeg (6716) and Mtb (4019) genes and their mutual orthologs (2547). Msmeg (403) and Mtb (625) required genes are shown within the blue and red circle, respectively. 343 genes are required in both species, 44 (Msmeg) and 247 (Mtb) required genes have a non-required mutual ortholog in the other species, and 16 (Msmeg) and 35 (Mtb) required genes do not have mutual orthologs in the other species (d) Venn diagram illustrating Msmeg and Mtb GA (causing growth advantage when disrupted) genes, as described for (c). Msmeg (415) and Mtb (283) GA genes are shown within the blue and red circle, respectively. 27 genes cause growth advantage in both species, 121 (Msmeg) and 123 (Mtb) GA genes have a non-GA mutual ortholog in the other species, and 267 (Msmeg) and 160 (Mtb) GA genes do not have mutual orthologs in the other species.
Tnseq allows us to categorize genes by their requirement for growth25. Using a Hidden Markov Model incorporated into the TRANSIT platform26, we identified 306 Msmeg genes as essential (ES) for growth, 97 as causing growth defect (GD) and 415 as causing growth advantage (GA) when disrupted, and 5815 as non-essential (NE) (Fig. 1b, Dataset 1a). 83 genes were undefinable (N/A), due to insufficient number of TA sites present to robustly call their requirement status. As expected, we found insertions throughout two regions that are know to be copies of each other, ranging from msmeg_1002–1059 and msmeg_2282–233927,28. In addition, we discovered a novel region that is probably present in more than one copy, ranging from msmeg_4926–4946 (see Supplementary Fig. S1). Interestingly, this region contains genes encoding F1F0 ATP synthase and one of three native copies of rRNA operons (rrn) (the rrn copy number is relevant for maximum growth rate29). As expected, the duplicated regions were over-represented in GA genes (67% of total genes within the duplicated regions, 9 times more than would be expected by random). Because there are additional copies of each gene in these loci, we cannot draw any conclusions about their actual requirement.
Msmeg and Mtb in vitro essential genes compared
Differences and similarities in Msmeg and Mtb essential processes may illuminate our interpretation of data across species and influence our choice of study organism. Msmeg mc2155 shares 2547 mutual orthologous genes (meaning each gene in one species is the best match for the ortholog in the other species, with BLAST E-value < 10−10, Dataset 1b) with Mtb H37Rv. We found that nearly all (96%) of the Msmeg required (ES and GD) genes had a mutual ortholog in Mtb. 90% of these genes were required also for optimal Mtb in vitro growth (as defined by DeJesus et al.30, note that Dataset 1a also lists the genetic requirement of Mtb mutual orthologs as defined by Griffin et al. and Zhang et al.31,32) (Fig. 1c), indicating that, under the given conditions, Msmeg and Mtb largely depend on the same mechanisms for growth. On the contrary, only 6% of Msmeg GA genes had a GA ortholog in Mtb, and the majority (64%) of the Msmeg genes providing growth advantage when disrupted were species-specific (i.e. no mutual ortholog) (Fig. 1d).
Orthologous genes that differ in requirement between Msmeg and Mtb may represent differences in niche or suggest alternative pathways (Dataset 1a). For instance, the thiamin biosynthesis pathway is essential in Mtb, but dispensable in Msmeg. Furthermore, genes within the esx-3 gene cluster, required for mycobactin-mediated iron uptake, are essential in Mtb when cultured in standard (Middlebrook 7H10) medium but dispensable in Msmeg grown under similar conditions17,33. In Msmeg, a second iron acquisition system, exochelin, does not require ESX-317. Thus, functional redundancy between exochelin and mycobactin might allow ESX-3 components to be disrupted in Msmeg but not Mtb
Knockout mutants defective in siderophore-mediated iron acquisition
We have shown that Msmeg is suitable for genome wide phenotypic screening and uniquely apt to study mycobacterial low iron fitness mechanisms. We sought to identify and categorize Msmeg genes required for exochelin and/or mycobactin independent low iron survival by screening in strains deficient for siderophores (Fig. 2a). When one pathway is disrupted by a gene knockout, genes in that pathway are no longer likely to be essential, since the pathway itself no longer functions. However, genes in a compensatory or parallel pathway will become essential to rescue cells. This means that genes important for the mycobactin pathway are likely to be essential in low iron for cells unable to produce or utilize exochelin as the redundant siderophore is missing, and vice versa. Deletion of fxbA (msmeg_0014, a formyl transferase involved in synthesis of exochelin34) and mbtD (msmeg_4512, a polyketide synthase required for the synthesis of mycobactin35) previously abolished exochelin and mycobactin pathways, respectively17. Thus, we constructed Msmeg strains deficient in exochelin (∆fxbA), mycobactin (∆mbtD) and both siderophores (ΔmbtDΔfxbA). The genetic deletions had the predicted effect; the exochelin (∆fxbA) and mycobactin (∆mbtD) deficient single mutant grew on low iron, however, the ΔmbtDΔfxbA siderophore null mutant did not (Fig. 2b). This confirms that siderophores are fundamental for growth under the given condition. All strains grew in iron replete medium (Fig. 2b).
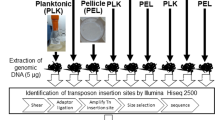
Schematic overview of screen for low iron fitness genes. (a) A transposon mutant library of Msmeg wt selected on high iron levels was compared to low iron-selected transposon libraries of Msmeg wt, ΔfxbA (exochelin knockout), ΔmbtD (mycobactin knockout) and ΔfxbAΔmbtD (double siderophore knockout) to identify and categorize genes involved in low iron growth. The amplitude of the blue vertical bars represents the hypothetical number of transposon insertions counted at the given TA dinucleotide site. (b) Msmeg wt, ΔfxbA, ΔmbtD and ΔfxbAΔmbtD mutants were prewashed in chelated Sauton’s before diluted and spotted on chelated Sauton-based agar plates with increasing concentrations of FeCl3. (c) Overview of library size, selective conditions, TA sites hit and total insertion count of the five sequenced libraries of the screen for low iron genes.
We found that the minimal amount of iron needed to rescue the siderophore null strain (ΔmbtDΔfxbA) was between 1 and 3 μM, where iron acquisition must occur independently of siderophores through porins, yet-to-be-discovered transporters, or alternative iron acquisition mechanisms (Fig. 2b). Therefore, genes that become essential in siderophore null cells grown with 3 μM iron may function in a novel iron uptake pathway, independently of siderophores, or be required for survival by other means. For the optimal outcome of our low iron screens we therefore selected wild type (wt), exochelin null (∆fxbA), and mycobactin null (∆mbtD) libraries in the absence of iron supplementation, and the siderophore null (∆mbtD∆fxbA) library in the presence of 3 μM FeCl3 supplement (Fig. 2c).
Differential growth analysis of high and low iron selected transposon libraries
We generated transposon libraries of Msmeg wt, mycobactin null (ΔmbtD), exochelin null (ΔfxbA) and siderophore null (ΔmbtDΔfxbA) mutants and selected on high or low iron media (Fig. 2c). Sequencing demonstrated that all libraries were adequately saturated with transposons (Fig. 3a, Dataset 1c). We expected that exochelin-related genes would become essential in the mycobactin null (∆mbtD) background in low iron. Indeed, genes in the exochelin pathway had few to no insertions in the ∆mbtD strain, while insertions in the same genes were readily apparent in ∆fxbA and ∆mbtD∆fxbA cells (Fig. 3b). Correspondingly, genes encoding enzymes required for mycobactin synthesis had lower counts in the exochelin null (∆fxbA) background than in ∆mbtD and ∆mbtD∆fxbA cells (Fig. 3c). Additionally, our previous finding that ESX-3 acts within the mycobactin utilization pathway was confirmed by the low insertion counts within the esx-3 genetic cluster in the ΔfxbA but not ΔmbtD and ∆mbtD∆fxbA low iron selected libraries (Fig. 3d)17.
Validation of screen for low iron fitness genes. Distribution of transposon insertion counts (blue vertical bars) for all sequenced libraries. (a) Whole genome. Insertion counts in log scale of 1–1500. (b) Exochelin gene cluster. Insertion counts in log scale of 0–50. The red triangles indicate the knocked out gene (msmeg_0014, fxbA). (c) Mycobactin gene cluster. Insertion counts in log scale of 0–50. The red triangles indicate the knocked out gene (msmeg_4512, mbtD). (d) ESX-3 gene cluster. Insertion counts in log scale of 0–25. Plots created using IGV - distributed by the Broad Institute (http://www.broadinstitute.org/igv/). Black dashed boxes show genes significantly under-represented in insertion counts in one or more of the low iron libraries compared to the high iron library.
To identify new genes required for low iron growth, we normalized the insertion counts between the libraries and compared the ratio of counts on a gene by gene basis from the wt high iron control library to the low iron selected experimental libraries (Fig. 4). Here, ‘under-represented’ and ‘over-represented’ refer to genes with significantly lower or higher insertion counts compared to wt high iron, respectively. We found that a large fraction of previously known iron uptake genes (Dataset 1d), are among those significantly under-represented in our screens, confirming the synthetic lethality between mycobactin and exochelin pathways (Fig. 4a–d). As expected, genes involved in siderophore mediated iron uptake were dispensable in the double siderophore null background (Fig. 4d).
Identification and categorization of low iron fitness genes. Transposon insertion counts presented relative to counts ratio (control/experimental) per gene between Msmeg wt high iron control library and (a) Msmeg wt, (b) ΔfxbA, (c) ΔmbtD or (d) ΔfxbAΔmbtD mutant low iron libraries (x-axis) and the corresponding P-values calculated by Mann Whitney U-test (y-axis). Genes previously known to be involved in mycobactin-mediated (green), exochelin-mediated (pink), or siderophore-independent (blue) iron uptake are color-coded. Red lines represent a cutoff of genes more than 5 fold under- (to the right) or over-represented (to the left) and with a P-value of less than 0.05. Genes knocked out are circled with pink (fxbA) or green (mbtD). In figure c is msmeg_0019 out of scale with P-value 1.5 × 10−9 and relative counts ratio 1.478.
Candidate genes required for low iron fitness
Comparing high and low iron-selected libraries allowed us to identify genes involved in iron acquisition as well as other processes directly or indirectly involved in low iron growth. For the wt library we found significant under-representation of 40 genes, of which 27.5% are previously known to be involved in siderophore-mediated iron uptake (Fig. 4a and Dataset 1e). We found several loci that had putative oxidation and reduction functions, all encoding proteins predicted to bind to metal co-factors. These enzymes might be required for basic cellular functions and bind iron even at very low iron concentrations. Other hits in our low iron screens included moxR (thought to chaperone insertion of metal cofactors into substrate molecules36), msmeg_3121 (encodes a protein with homology to a regulator of iron-sulfur cluster assembly, SufR37), and cysD (sulfate adenylyltransferase, involved in sulfur metabolism, important for iron-sulfur cluster formation38), all relevant to iron as a co-factor. We also found predicted iron-responsive candidate genes, with putative binding sites for the iron dependent regulator IdeR12,39,40,41,42, such as msmeg_6575 (lipE/rv3775), a putative β-lactamase/lipase.
Candidate genes required for mycobactin-mediated iron acquisition
We know of few proteins involved in the recognition and transport of mycobactins across the mycobacterial outer membrane. Nor is the precise mechanism of ESX-3 in utilization of mycobactin-bound iron fully understood. Using the exochelin null library, we specifically screened for genes required for mycobactin-mediated iron uptake. We identified 36 significantly under-represented (Dataset 1f) and 6 over-represented genes (Dataset 1g). Nearly one-third of the under-represented genes were previously found to be involved in iron uptake via the mycobactin pathway (Fig. 4b). 16 hits not previously related to iron acquisition have Mtb orthologs and are, therefore, candidates genes for mycobactin-mediated iron uptake. In addition, genes involved in ATP synthesis (atpA, atpB) were under-represented in the exochelin null screen, consistent with an energy requirement for iron uptake (e.g. for the ATP-dependent transport of mycobactin via ABC transporter IrtAB).
Candidate genes required for exochelin-mediated iron acquisition
We found that 72 genes were uniquely required in the strain lacking mbtD (Dataset 1h), while 7 were over-represented (Dataset 1i, Fig. 4c). Surprisingly, genes encoding proteins involved in import (FxuA-D), but not synthesis and export of exochelin (FxbA-C and ExiT), seem to be essential for low iron growth in ∆mbtD cells. It is possible that the production and export of exochelin from neighboring colonies may rescue transposon mutants unable to produce or export the siderophore. A similar trans-complementation of transposon mutants was previously seen in genes encoding synthesis of the Yersinia pestis siderophore yersiniabactin43. Interestingly, we do not see the same division of essentiality in genes of the mycobactin pathway. It might be that mycobactin is not as accessible to adjacent colonies as exochelin under these conditions, particularly since mycobactin is produced in an insoluble and membrane anchored form in addition to the secreted soluble form.
Nevertheless, we found genes that may be specific for exochelin utilization. For example msmeg_4318 (encoding a hypothetical membrane transport protein), msmeg_6063 (encoding a predicted iron uptake membrane protein), and msmeg_6419 (encoding a conserved hypothetical predicted iron regulated protein12,39), are all required for Msmeg low iron growth in the absence of mycobactin synthesis and represent candidate exochelin uptake genes.
Candidate genes required for siderophore-independent low iron fitness
Using the strain lacking both siderophores (ΔmbtDΔfxbA) we found 57 significantly under-represented genes (Dataset 1j), and 42 over-represented genes (Dataset 1k, Fig. 4d). Genes involved in exochelin and mycobactin-mediated iron acquisition were not under-represented in this screen, which was expected, as the siderophore null (∆mbtD∆fxbA) mutant grows in a siderophore independent way. However, one gene known to be involved in heme uptake (mmpL11)44,45, and one involved in porin-mediated iron uptake (mspA)46, were found to be required for low iron growth in the absence of siderophores. This suggests that, while siderophores may be the predominant mechanism for Msmeg to access iron, under the given growth conditions, additional mechanisms can also take up the metal from the environment, albeit with lower efficiency. Novel genes of interest that may function in siderophore-independent iron uptake include msmeg_5420, which neighbors the predicted iron responsive iron permease msmeg_541839. Msmeg_5420 shows homology to the iron uptake gene ywbN of Bacillus subtilis47. Genes within a region spanning from msmeg_1701 to msmeg_1712 are under-represented in the ∆fxbA (msmeg_1701/deoD, msmeg_1703/amiA1), ΔmbtD (msmeg_1705) and ΔfxbAΔmbtD-screens (msmeg_1712). msmeg_1704-1712 encode a putative ABC transporter system, and this gene cluster might be important for low iron proliferation in a siderophore independent manner. Finally, two genes with homology to iron transport proteins, msmeg_3635 and msmeg_3636, were significantly under-represented in the siderophore null screen (as were they in the ΔmbtD-screen, Dataset 1h and 1j), and we further characterized the role of msmeg_3635 in mycobacterial low iron fitness below.
msmeg_3635 is required for Msmeg low iron fitness in the absence of siderophores
msmeg_3635 belongs to a three-gene operon consisting of msmeg_3633 (ATP binding protein), msmeg_3635 (permease) and msmeg_3636 (periplasmic protein), encoding the components of a putative ferric iron ABC transporter (Fig. 5c, Dataset 1a). To test whether this predicted transport system is indeed required for growth under low iron conditions, we disrupted it by constructing msmeg_3635 deletions in wt and siderophore mutant backgrounds. The Δmsmeg_3635 mutants showed impaired growth in low iron in the absence of both siderophores, but not in wt or single siderophore null backgrounds (Fig. 5a). All mutants grew at a rate similar to wt in high iron medium. Zinc, another ion required for optimal bacterial growth, is not able to rescue the double siderophore null mutant, suggesting the putative ABC transporter is iron-specific (Fig. 5b). Low iron growth of the ΔmbtDΔfxbAΔ3635 strain could be rescued by expression of the intact transport operon (Fig. 5d). Together, these results suggest that this putative transport system may translocate, or aid in translocation of, iron across the cell membrane in a manner independently of siderophore pathways.
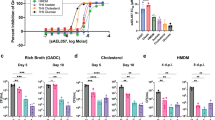
msmeg_3635 is important for Msmeg siderophore-independent low iron growth. (a) Growth of Msmeg strains in high (upper panel) or low iron (lower panel) monitored over time (x-axis) by OD600 (y-axis). Error bars represent standard error from the mean of three biological replicas. To the ΔfxbAΔmbtD and ΔfxbAΔmbtDΔ3635 curves, 3 μM FeCl3 was added in the low iron condition. To all other curves, 100 μM 2, 2′-bipyridine was add to low iron, and 150 μM FeCl3 was added to high iron condition. (b) Growth of ΔfxbAΔmbtD and ΔfxbAΔmbtDΔ3635 in low iron, low zinc (upper left), low iron, high zinc (lower left), high iron, low zinc (upper right), and high iron, high zinc (lower right). For all curves; low iron was supplemented with 3 μM FeCl3, low zinc with 0 μM ZnSO4, high iron with 150 μM FeCl3, and high zinc with 3.67 μM ZnSO4. (c) Distribution of transposon insertion counts (blue vertical bars) for all sequenced libraries in the msmeg_3630-3636 operon. Insertion counts in log scale of 0–75. Plot created using IGV - distributed by the Broad Institute (http://www.broadinstitute.org/igv/). The black dashed box shows genes significantly under-represented in insertion counts in one or more of the low iron libraries compared to the high iron library. (d) Complementation of the ΔfxbAΔmbtDΔ3635 low iron phenotype. Msmeg ΔmbtDΔfxbA, ΔmbtDΔfxbAΔ3635 and ΔmbtDΔfxbAΔ3635 compl. were grown in either high (150 μM FeCl3) or low (1.5 μM FeCl3) iron.
Discussion
Iron is an essential nutrient for bacteria, and is especially critical during host infection where iron itself is limited. Mtb depends on iron acquisition genes for virulence in animal models2,3,4,5, however, it is worth noting that Tufariello et al. found that the essentiality of Mtb mycobactin during mouse infection depended on host genotype19. Even so, many aspects of mycobacterial growth under iron limited conditions remain unknown, and the presence of a second high affinity iron uptake pathway in Msmeg allowed us to define a network of factors critical for mycobacterial low iron fitness. We identified novel genes potentially required for recognition, uptake or utilization of mycobactin and exochelin. Over all, nearly half (46%) of the low iron genes we identified had an ortholog in Mtb. However, when we look at the genes we found specifically required for the exochelin and mycobactin pathways, 32% and 72%, respectively, had Mtb orthologs. This reflects that the mycobactin pathway, but not the exochelin pathway, is present in Mtb. We also identified new genetic clusters and operons possibly involved in low iron growth independently of siderophores. To validate our screen, we investigated one such operon, msmeg_3633-36, encoding a putative ferric iron ABC transporter (with orthologs in M. gilvum, M. vanbaalenii, M. thermoresistible, and a multitude of other mycobacterial species, but not in Mtb). By disrupting the transporter we confirmed its involvement in siderophore independent low iron fitness. MSMEG_3635, encoded by the gene we knocked out, shares homology with the ferric iron transport system permease protein SfuB. In Serratia marcescens, SfuB was shown to be part of a putative transporter that enabled a siderophore-deficient strain of Escherichia coli K-12 to grow in iron-limited medium48, suggesting a conserved role for this protein in bacterial iron uptake.
Along with genes required for low iron growth, we found a surprising number of genes that became dispensable in low iron, as evidenced by over-representation in transposon libraries. This was particularly true when both siderophore pathways were deleted (Fig. 4), and might therefore be an effect of higher selective pressure in extremely iron-restricted environments. Some of the over-represented genes encode transcriptional repressors, and their absence might increase the expression of beneficial proteins. Interestingly, mbtT and fxuB (involved in mycobactin synthesis and exochelin import, respectively34,35) were over-represented in the ΔmbtD and/or ΔmbtDΔfxbA screen(s). Both these genes had very few insertion counts in the wt high iron control library but not the respective low iron libraries. Their disruption might be toxic to the cells when iron is plentiful, perhaps by creating harmful intermediates or causing toxic siderophore accumulation, as previously seen for mycobactin49.
Msmeg has previously been central for our understanding of conserved mycobacterial functions (exemplified in50,51,52), though, as an environmental organism, it likely must adapt to a broader range of environmental stresses than the pathogen Mtb. It was therefore surprising to find that, under the given conditions, Msmeg and Mtb (as defined by DeJesus et al.30) largely depended on the same genes for optimal growth. In fact, only 4% of the Msmeg required genes did not have an Mtb mutual ortholog, a very small fraction considering the Msmeg genome contains 4169 (62%) genes with no Mtb mutual ortholog (albeit, some of the ‘non-mutual-orthologs’ will have partial or ambiguous Mtb orthologs). This large overlap (96%) in essential genes between the two species might reflect the reliance on a smaller set of conserved house-keeping genes required in a nutrient-rich, “non-stress” environment, like our test condition. Perhaps, if we subjected Msmeg to other growth conditions, more closely resembling its natural niches, we would find a larger fraction of species-specific genes required for growth. On the contrary, the majority (64%) of Msmeg GA genes (prompting growth advantage when disrupted) did not have a mutual orthologs in Mtb, in fact, only 6% of Msmeg GA genes had a GA Mtb mutual ortholog. Among the 6% mutual GA genes were several putative transcriptional regulators; their disruption might increase expression of genes benefiting growth under the given conditions. Many of the Msmeg GA genes were found within duplicated genome regions, partially explaining the difference in growth advantage seen between the two species. Also, the Mtb screen by DeJesus et al. was based on libraries selected in the presence of oleic acid supplement30, while our Msmeg library was not, potentially causing differences in genetic requirement between the two species.
Among genes that we found non-essential in Msmeg, but required in Mtb, were genes encoding thiamine and ATP biosynthesis (we have already mentioned the ESX-3 secretion system). Interestingly, Mtb does not appear to have a thiamine salvage and transport/uptake system53, while Msmeg encodes a putative thiaminase II (msmeg_3478), potentially involved in salvage of thiamine intermediates and thus rescue of de novo thiamine synthesis mutants54. A region we discovered to be present in more than one copy (msmeg_4926-4946) encodes F1F0 ATP synthase, probably explaining the non-essential nature of ATP synthesis in Msmeg. Actually, F1F0 ATP synthase was previously found essential in Msmeg, however, that was after knocking out both copies of atpD55. Taken together, the remarkable overlap in Msmeg and Mtb in vitro required genes, as well as the possibility to dissect differentially essential pathways using the non-pathogenic fast-growing strain, in our opinion, strengthen Msmeg as a model organism to study basic mycobacteriology and anti-mycobacterial drug discovery.
In summary; iron plays a central role in bacterial metabolism. Because of its scarcity, bacteria require highly avid molecules to adapt to iron limitation. Paradoxically, too much iron is toxic. Thus, these organisms have developed complex mechanisms to take up iron, regulate its uptake and maintain growth in iron-scarce host environments. Our work suggests that, in mycobacteria, several proteins have evolved to play critical roles in sustaining fitness when iron is low, and may propose attractive targets for new drugs.
Methods
Strains and growth conditions
M. smegmatis mc215556,57 was cultured in Middlebrook 7H9 (BD Difco) medium supplemented with 0.2% glycerol, 0.05% Tween 80, and 10% albumin-dextrose-catalase (ADC) (5% [wt/vol] bovine serum albumin fraction V, 2% [wt/vol] dextrose, 145.5 mM NaCl, 0.003% [wt/vol] catalase), unless otherwise stated. E. coli DH5α was used for cloning and grown in LB medium. Kanamycin was added to 20 μg/ml or 50 μg/ml, and zeocin to 50 μg/ml or 25 μg/ml, in mycobacteria and E. coli, respectively. Hygromycin and gentamicin were added to 50 μg/ml and 7.5 μg/ml in mycobacteria, respectively. Liquid medium for Msmeg growth curves was prepared by adding 100 μM 2, 2′-bipyridine (Alfa Aesar) or 150 μM FeCl3 to chelated Sauton’s (prepared as previously described, using Chelex 100 resin from Bio-Rad17) for low or high iron, respectively. Intermediate iron medium (chelated Sauton’s with 1.5 or 3 μM FeCl3) did not contain 2, 2′-bipyridine. Msmeg transposon mutant library for in vitro gene requirement analysis (essential genes) was selected on Middlebrook 7H10 (BD Difco) agar medium supplemented with 0.5% glycerol, 0.1% Tween 80, and 10% ADC. Msmeg transposon mutant libraries for low iron gene requirement analysis were selected on agar medium prepare with chelated Sauton’s mixed with bacto agar (BD Difco) (see Supplementary SI Methods). All low iron media were prepared using plasticware.
Construction of mutant strains
Msmeg mutant strains were constructed by recombineering, replacing the gene in question with an antibiotic resistance marker, using strains expressing the mycobacteriophage recombinases gp60 and gp61 on a nitril-inducible, counter-selectable plasmid or on the acetamide-inducible plasmid pJV5358. Details for construct preparations and selection processes are contained in Supplementary SI Methods59,60.
Generation of Msmeg transposon mutant library
Msmeg transposon mutant libraries were prepared using the ϕMycomarT7 phagemid as recently described23.
Preparation of libraries for Illumina sequencing
Total DNA of the transposon mutant libraries was purified using Masterpure DNA Purification kit (Epicentre). DNA fragmentation, end repair, A-tailing and adapter ligation was performed as previously described32. Transposon junctions were amplified by PCR and 200–400 bp fragments were isolated from gel and sequenced by Illumina GAII instrument.
Genetic growth requirement analysis
The gene requirement calls were made using a Hidden Markov Model25, as implemented in TRANSIT26. The raw read counts mapping to each TA site in the Msmeg mc2155 genome (Msmeg mc2155_tamu) were reduced to unique templates counts by grouping based on nucleotide barcodes embedded in read 261. Template counts at each TA site were exponentially scaled (f(t) = t1.5) so that the histogram of template counts better matches a Geometric distribution based on a Q-Q (quantile-quantile) plot, which is an assumption built-in to the HMM. The HMM uses 4-states, with labels ES (essential), GD (growth-defect), NE (non-essential), and GA (growth-advantage), and the likelihood that a TA site will be labeled with each state is based on differences between local read-count and the global average. The HMM was run on the Middlebrook 7H10-selected wt transposon insertion sequencing dataset, and the state labels of each site were post-processed to determine the majority label for each gene (ES, GD, NE, or GA), excluding TA sites in the N-terminal 10% and C-terminal 10% of the ORF.
Differential growth analysis
The control library (wt selected on high iron) was compared to wt, ΔmbtD and ΔfxbA and ΔmbtDΔfxbA libraries selected on low iron levels by determining the ratio of transposon insertion counts for the middle 94% of each gene. P-values were calculated by Mann-Whitney U test, treating the insertion counts as non-parametric distributions. A null hypothesis was used assuming the distribution of transposon insertion counts were similar for the respective genes between the two compared conditions. Promising candidates involved in iron acquisition were searched for within a list of genes with a P-value less than 0.05 (two tailed test) and a counts ratio of more than 5.
Growth curve experiments
For Msmeg, cultures were grown in Middlebrook 7H9 for 2–3 days before washed once in chelated Sauton’s and diluted to OD600 0.01 or 0.05 in the appropriate medium. Growth was monitored as 200 μl cultures (in triplicates) in microplate honeycomb wells using a Bioscreen growth curve reader (Oy Growth Curves Ab Ltd.).
Data Availability
All data generated during this study are included in this published article (and its Supplementary Information files).
References
World Health Organization. Global Tuberculosis Report Executive Summary 2018., https://www.who.int/tb/publications/global_report/en/ (2019).
Rodriguez, G. M. & Smith, I. Identification of an ABC Transporter Required for Iron Acquisition and Virulence in Mycobacterium tuberculosis. J. Bacteriol. 188, 424–430 (2005).
Wells, R. M. et al. Discovery of a Siderophore Export System Essential for Virulence of Mycobacterium tuberculosis. PLoS Pathog. 9, e1003120, https://doi.org/10.1371/journal.ppat.1003120.s024 (2013).
Reddy, P. V. et al. Disruption of Mycobactin Biosynthesis Leads to Attenuation of Mycobacterium tuberculosis for Growth and Virulence. The Journal of infectious diseases 208, 1255–1265 (2013).
Madigan, C. A. et al. Lipidomic analysis links mycobactin synthase K to iron uptake and virulence in M. tuberculosis. PLoS Pathog. 11, e1004792, https://doi.org/10.1371/journal.ppat.1004792 (2015).
Hameed, S., Pal, R. & Fatima, Z. Iron Acquisition Mechanisms: Promising Target Against Mycobacterium tuberculosis. Open Microbiol J 9, 91–97 (2015).
Ratledge, C. In Iron Acquisition by the Genus Mycobacterium History, Mechanisms, Role of Siderocalin, Anti-Tuberculosis Drug Development SpringerBriefs in Molecular Science (ed. Byers, B. R.) Ch. 2, 3–39 (Springer, 2013).
Banerjee, S., Farhana, A., Ehtesham, N. Z. & Hasnain, S. E. Iron acquisition, assimilation and regulation in mycobacteria. Infect. Genet. Evol. 11, 825–838 (2011).
Fang, Z., Sampson, S. L., Warren, R. M., Gey van Pittius, N. C. & Newton-Foot, M. Iron acquisition strategies in mycobacteria. Tuberculosis 95, 123–130 (2015).
Chao, A., Sieminski, P. J., Owens, C. P. & Goulding, C. W. Iron Acquisition in Mycobacterium tuberculosis. Chem. Rev. 119, 1193–1220 (2019).
Kurthkoti, K. et al. The Capacity of Mycobacterium tuberculosis To Survive Iron Starvation Might Enable It To Persist in Iron-Deprived Microenvironments of Human Granulomas. mBio 8, e01092–17, https://doi.org/10.1128/mBio.01092-17 (2017).
Rodriguez, G. M., Voskuil, M. I., Gold, B., Schoolnik, G. K. & Smith, I. ideR, an Essential Gene in Mycobacterium tuberculosis: Role of IdeR in Iron-Dependent Gene Expression, Iron Metabolism, and Oxidative Stress Response. Infect. Immun. 70, 3371–3381 (2002).
Pandey, S. D. et al. Iron-Regulated Protein HupB of Mycobacterium tuberculosis Positively Regulates Siderophore Biosynthesis and Is Essential for Growth in Macrophages. J. Bacteriol. 196, 1853–1865 (2014).
Prados-Rosales, R. et al. Role for Mycobacterium tuberculosis membrane vesicles in iron acquisition. J. Bacteriol. 196, 1250–1256 (2014).
Pal, R., Hameed, S. & Fatima, Z. Iron Deprivation Affects Drug Susceptibilities of Mycobacteria Targeting Membrane Integrity. J Pathog 2015, 938523 (2015).
Shiloh, M. U. & Champion, P. A. To catch a killer. What can mycobacterial models teach us about Mycobacterium tuberculosis pathogenesis? Curr. Opin. Microbiol. 13, 86–92 (2010).
Siegrist, M. S. et al. Mycobacterial Esx-3 is required for mycobactin-mediated iron acquisition. Proc. Natl. Acad. Sci. USA 106, 18792–18797 (2009).
Siegrist, M. S. et al. Mycobacterial Esx-3 requires multiple components for iron acquisition. mBio 5, e01073–01014, https://doi.org/10.1128/mBio.01073-14 (2014).
Tufariello, J. M. et al. Separable roles for Mycobacterium tuberculosis ESX-3 effectors in iron acquisition and virulence. Proc. Natl. Acad. Sci. USA 113, E348–357 (2016).
Serafini, A., Pisu, D., Palu, G., Rodriguez, G. M. & Manganelli, R. The ESX-3 Secretion System Is Necessary for Iron and Zinc Homeostasis in Mycobacterium tuberculosis. PLoS One 8, e78351, https://doi.org/10.1371/journal.pone.0078351 (2013).
van Opijnen, T. & Camilli, A. Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms. Nat Rev Microbiol 11, 435–442 (2013).
Judd, J., Boucher, N., Van Roey, E., Gray, T. A. & Derbyshire, K. M. Application of Distributive Conjugal DNA Transfer in Mycobacterium smegmatis To Establish a Genome-Wide Synthetic Genetic Array. J. Bacteriol. 199 (2017).
Majumdar, G. et al. Genome-Wide Transposon Mutagenesis in Mycobacterium tuberculosis and Mycobacterium smegmatis. Methods Mol. Biol. 1498, 321–335 (2017).
Sassetti, C. M., Boyd, D. H. & Rubin, E. J. Comprehensive identification of conditionally essential genes in mycobacteria. Proc. Natl. Acad. Sci. USA 98, 12712–12717 (2001).
DeJesus, M. A. & Ioerger, T. R. A Hidden Markov Model for identifying essential and growth-defect regions in bacterial genomes from transposon insertion sequencing data. BMC Bioinformatics 14, 303 (2013).
DeJesus, M. A., Ambadipudi, C., Baker, R., Sassetti, C. & Ioerger, T. R. TRANSIT - A Software Tool for Himar1 TnSeq Analysis. PLoS Comput. Biol. 11, e1004401, https://doi.org/10.1371/journal.pcbi.1004401 (2015).
Galamba, A. et al. Disruption of adhC reveals a large duplication in the Mycobacterium smegmatis mc(2)155. genome. Microbiol-Sgm 147, 3281–3294 (2001).
Warner, D. F. et al. A derivative of Mycobacterium smegmatis mc2155 that lacks the duplicated chromosomal region. Tuberculosis 86, 438–444 (2006).
Roller, B. R., Stoddard, S. F. & Schmidt, T. M. Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat Microbiol 1, 16160 (2016).
DeJesus, M. A. et al. Comprehensive Essentiality Analysis of the Mycobacterium tuberculosis Genome via Saturating Transposon Mutagenesis. mBio 8, 02133–16, https://doi.org/10.1128/mBio.02133-16 (2017).
Griffin, J. E. et al. High-Resolution Phenotypic Profiling Defines Genes Essential for Mycobacterial Growth and Cholesterol Catabolism. PLoS Pathog. 7, e1002251, https://doi.org/10.1371/journal.ppat.1002251.g004 (2011).
Zhang, Y. J. et al. Global Assessment of Genomic Regions Required for Growth in Mycobacterium tuberculosis. PLoS Pathog. 8, e1002946, https://doi.org/10.1371/journal.ppat.1002946.g005 (2012).
Serafini, A., Boldrin, F., Palu, G. & Manganelli, R. Characterization of a Mycobacterium tuberculosis ESX-3 conditional mutant: essentiality and rescue by iron and zinc. J. Bacteriol. 191, 6340–6344 (2009).
Fiss, E. H., Yu, S. & Jacobs, W. R. Jr. Identification of genes involved in the sequestration of iron in mycobacteria: the ferric exochelin biosynthetic and uptake pathways. Mol. Microbiol. 14, 557–569 (1994).
Quadri, L. E., Sello, J., Keating, T. A., Weinreb, P. H. & Walsh, C. T. Identification of a Mycobacterium tuberculosis gene cluster encoding the biosynthetic enzymes for assembly of the virulence-conferring siderophore mycobactin. Chem. Biol. 5, 631–645 (1998).
Snider, J. & Houry, W. A. MoxR AAA+ ATPases: A novel family of molecular chaperones? J. Struct. Biol. 156, 200–209 (2006).
Huet, G., Daffe, M. & Saves, I. Identification of the Mycobacterium tuberculosis SUF Machinery as the Exclusive Mycobacterial System of [Fe-S] Cluster Assembly: Evidence for Its Implication in the Pathogen’s Survival. J. Bacteriol. 187, 6137–6146 (2005).
Pinto, R., Tang, Q. X., Britton, W. J., Leyh, T. S. & Triccas, J. A. The Mycobacterium tuberculosis cysD and cysNC genes form a stress-induced operon that encodes a tri-functional sulfate-activating complex. Microbiology (Reading, England) 150, 1681–1686 (2004).
Yellaboina, S., Ranjan, S., Vindal, V. & Ranjan, A. Comparative analysis of iron regulated genes in mycobacteria. FEBS Lett. 580, 2567–2576 (2006).
Prakash, P., Yellaboina, S., Ranjan, A. & Hasnain, S. E. Computational prediction and experimental verification of novel IdeR binding sites in the upstream sequences of Mycobacterium tuberculosis open reading frames. Bioinformatics 21, 2161–2166 (2005).
Manabe, Y. C., Saviola, B. J., Sun, L., Murphy, J. R. & Bishai, W. R. Attenuation of virulence in Mycobacterium tuberculosis expressing a constitutively active iron repressor. Proc. Natl. Acad. Sci. USA 96, 12844–12848 (1999).
Gold, B., Rodriguez, G. M., Marras, S. A., Pentecost, M. & Smith, I. The Mycobacterium tuberculosis IdeR is a dual functional regulator that controls transcription of genes involved in iron acquisition, iron storage and survival in macrophages. Mol. Microbiol. 42, 851–865 (2001).
Palace, S. G., Proulx, M. K., Lu, S., Baker, R. E. & Goguen, J. D. Genome-wide mutant fitness profiling identifies nutritional requirements for optimal growth of Yersinia pestis in deep tissue. mBio 5, e01385–14, https://doi.org/10.1128/mBio.01385-14 (2014).
Owens, C. P. et al. The Mycobacterium tuberculosis secreted protein Rv0203 transfers heme to membrane proteins MmpL3 and MmpL11. J. Biol. Chem. 288, 21714–21728 (2013).
Tullius, M. V. et al. Discovery and characterization of a unique mycobacterial heme acquisition system. Proc. Natl. Acad. Sci. USA 108, 5051–5056 (2011).
Jones, C. M. & Niederweis, M. Role of porins in iron uptake by Mycobacterium smegmatis. J. Bacteriol. 192, 6411–6417 (2010).
van der Ploeg, R. et al. Environmental salinity determines the specificity and need for Tat-dependent secretion of the YwbN protein in Bacillus subtilis. PLoS One 6, e18140, https://doi.org/10.1371/journal.pone.0018140 (2011).
Angerer, A., Gaisser, S. & Braun, V. Nucleotide sequences of the sfuA, sfuB, and sfuC genes of Serratia marcescens suggest a periplasmic-binding-protein-dependent iron transport mechanism. J. Bacteriol. 172, 572–578 (1990).
Jones, C. M. et al. Self-poisoning of Mycobacterium tuberculosis by interrupting siderophore recycling. Proc. Natl. Acad. Sci. USA 111, 1945–1950 (2014).
Aldridge, B. B. et al. Asymmetry and aging of mycobacterial cells lead to variable growth and antibiotic susceptibility. Science 335, 100–104 (2012).
Kieser, K. J. et al. Phosphorylation of the Peptidoglycan Synthase PonA1 Governs the Rate of Polar Elongation in Mycobacteria. PLoS Pathog. 11, e1005010, https://doi.org/10.1371/journal.ppat.1005010 (2015).
Gray, T. A. et al. Intercellular communication and conjugation are mediated by ESX secretion systems in mycobacteria. Science 354, 347–350 (2016).
Du, Q., Wang, H. & Xie, J. Thiamin (vitamin B1) biosynthesis and regulation: a rich source of antimicrobial drug targets? Int. J. Biol. Sci. 7, 41–52 (2011).
Jenkins, A. H., Schyns, G., Potot, S., Sun, G. X. & Begley, T. P. A new thiamin salvage pathway. Nat. Chem. Biol. 3, 492–497 (2007).
Tran, S. L. & Cook, G. M. The F1F0-ATP synthase of Mycobacterium smegmatis is essential for growth. J. Bacteriol. 187, 5023–5028 (2005).
Panas, M. W. et al. Noncanonical SMC protein in Mycobacterium smegmatis restricts maintenance of Mycobacterium fortuitum plasmids. Proc. Natl. Acad. Sci. USA 111, 13264–13271 (2014).
Snapper, S. B., Melton, R. E., Mustafa, S., Kieser, T. & Jacobs, W. R. Jr. Isolation and characterization of efficient plasmid transformation mutants of Mycobacterium smegmatis. Mol. Microbiol. 4, 1911–1919 (1990).
van Kessel, J. C. & Hatfull, G. F. Recombineering in Mycobacterium tuberculosis. Nat Methods 4, 147–152 (2007).
Dragset, M. S. et al. Benzoic Acid-Inducible Gene Expression in Mycobacteria. PLoS One 10, e0134544, https://doi.org/10.1371/journal.pone.0134544 (2015).
Dragset, M. S. et al. A Novel Antimycobacterial Compound Acts as an Intracellular Iron Chelator. Antimicrob. Agents Chemother. 59, 2256–2264 (2015).
Long, J. E. et al. In Gene Essentiality: Methods and Protocols (ed. Lu, Long Jason) 79–95 (Springer New York, 2015).
Acknowledgements
We gratefully acknowledge Dr. Chris B. Ford, Mark Ragheb and Brian Shuster for contributing in preparation of transposon library samples for sequencing. We also thank Dr. Manuel Linares for guidance in large dataset handling, Dr. Michael Chao for proofreading and scientific discussion and Dr. Fabian Lorenzo-Diaz for providing office space during parts of the writing process. This work was partly supported by the Research Council of Norway through its Centres of Excellence funding scheme project number 223255, Research Council of Norway grants 220836/H10, 246944/F10, 249901 and Central Norway Regional Health Authority.
Author information
Authors and Affiliations
Contributions
M.S.D.: conception, design of the work, acquisition, interpretation, and analysis of data, drafted and revised the manuscript. T.R.I.: design of work, acquisition, interpretation, and analysis of data, drafted and revised the manuscript. Y.J.Z.: acquisition, interpretation, and analysis of data. M.M.: acquisition, interpretation, and analysis of data. Z.G.: acquisition, interpretation, and analysis of data. J.C.S.: acquisition of data. T.H.F.: acquisition, interpretation, and analysis of data, revised the manuscript. E.J.R.: conception, design of the work, revised the manuscript. M.S.: conception, design of the work, acquisition, interpretation, and analysis of data, revised the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dragset, M.S., Ioerger, T.R., Zhang, Y.J. et al. Genome-wide Phenotypic Profiling Identifies and Categorizes Genes Required for Mycobacterial Low Iron Fitness. Sci Rep 9, 11394 (2019). https://doi.org/10.1038/s41598-019-47905-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47905-y
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.