Abstract
Hedgehog interacting protein (Hhip) is essential for islet formation and beta-cell proliferation during pancreatic development; abnormally elevated Hhip expression has been linked to human pancreatitis. Here, we investigate the role of Hhip in modulating insulin secretion in adult Hhip mice (Hhip +/− vs. Hhip+/+) fed high fat diets (HFD). Both sexes of HFD-Hhip +/+ mice developed impaired glucose intolerance, that was only ameliorated in male HFD-Hhip +/− mice that had high levels of circulating plasma insulin, but not in female HFD-Hhip +/− mice. HFD stimulated Hhip gene expression, mainly in beta cells. Male HFD-Hhip +/+ mice had more large islets in which insulin content was reduced; islet architecture was disordered; and markers of oxidative stress (8-OHdG and Nox 2) were increased. In contrast, male HFD-Hhip +/− mice had more small islets with increased beta cell proliferation, enhanced GSIS, less oxidative stress and preserved islet integrity. In vitro, recombinant Hhip increased Nox2 and NADPH activity and decreased insulin-positive beta cells. siRNA-Hhip increased GSIS and abolished the stimulation of sodium palmitate (PA)-BSA on Nox2 gene expression. We conclude that pancreatic Hhip gene inhibits insulin secretion by altering islet integrity and promoting Nox2 gene expression in beta cells in response to HDF-mediated beta cell dysfunction, a novel finding.
Similar content being viewed by others
Introduction
The incidence of type 2 diabetes (T2D) will account for approximately 85–90% of all diabetes cases by 2040 and constitutes a major health burden in developed countries1. T2D is a metabolic disorder characterized by hyperglycemia associated with obesity-induced peripheral insulin resistance and intrinsic pancreatic islet beta cell dysfunction2,3,4. Long-term complications of T2D may lead to multiple organ dysfunction affecting the heart, kidney, nervous system, eye, artery and blood vessels.
Pancreatic beta cells are primarily responsible for the production and secretion of insulin for the maintenance of metabolic homeostasis2,3,4. In T2D, chronic elevation of free fatty acids causes beta cell dysfunction, reflected by impaired insulin secretory responses to increased glucose levels. Consequently, accumulating free fatty acids and glucose levels in beta cells synergistically up-regulate endogenous triglyceride levels, leading to progressive cell toxicity and increased apoptosis, a metabolic phenomenon termed “gluco-lipotoxicity”2,3. A major challenge, given the epidemic of T2D, is to develop more effective preventive and therapeutic strategies based on better understanding of the underlying pathophysiology1,4,5.
Hedgehog (Hh) signaling is essential for the development and function of the endocrine and exocrine pancreas6,7,8,9. 3 secreted Hh ligands, i.e., Sonic (Shh), Indian (Ihh), and Desert (Dhh) bind to the Patched1 membrane receptor, releasing the tonic inhibition of Smoothened, which is responsible for the activation of transcriptional factors of cubitus interruptus homologs (Glis) – e.g., Gli1 and Gli2 (activators) and Gli3 (repressor) – resulting in the transcription of an array of target genes10,11,12,13,14,15. Pancreatic tissue responds to Hh signaling in a dose-dependent manner during pancreatic development7,9, and Hh signaling is also required for maintaining adult beta cell function7. An increase of Hh signaling in adult beta cells in vivo leads to either a loss of beta cell function resulting in decreased insulin production and impaired insulin response to a glucose challenge or even change differentiated beta cells into beta cell-derived undifferentiated tumor cells7.
Hedgehog interacting protein (Hhip) originally discovered as a putative antagonist of Hh ligands regulates cell function via either canonical- or non-canonical Hh pathways10,11,12,13,14,15,16,17. Hhip encodes a protein of 700 amino acids and is abundantly expressed in vascular endothelial cells-rich tissues, including the pancreas18,19. Hhip null mice (Hhip−/−) display markedly impaired pancreatic islet formation (45% reduction of islet mass with a decrease of beta cell proliferation by 40%)9, underscoring the importance of Hhip in normal pancreatic development, though Hhip−/− mice die shortly after birth mainly due to lung defects9,12. In addition, pancreatic islets are highly vascularized and contain a structurally and functionally unique cell composition (alpha-, beta-, and delta-cell)20. An earlier study showed that low-level Hhip expression could be detected in normal mature pancreas (i.e., in the cytoplasm of islet cells and in blood vessels), but abnormal elevated Hhip gene expression has been linked to human pancreatitis18, suggesting that tight regulation of pancreatic Hhip gene expression might be essential for maintaining normal pancreatic function. Moreover, a genome-wide diabetes profiling database (http://diabetes.wisc.edu) revealed that compared to lean animals, Hhip mRNA was markedly elevated in the islets of diabetic ob/ob mice (at the age of 4 and 10 weeks with both C57BL/6 and BTBR backgrounds), but not in other tissues such as liver, gastrocnemius and soleus muscles, adipose tissue and hypothalamus, underscoring the specificity of Hhip expression in murine T2D islets.
Less is known about Hhip expression pattern in mature islet cells and its function under normal and/or stressed conditions. In the present study, we systematically studied the role of pancreatic Hhip expression in response to high fat diet (HFD)-mediated beta cell dysfunction in vivo, ex vivo and in vitro. We showed that HFD-induced pancreatic Hhip gene expression targets beta cells and then inhibits glucose stimulated insulin secretion (GSIS). Mechanistically, Hhip activates oxidative stress-related NADPH oxidase 2 (Nox 2), one of the key factors implicated in beta cell dysfunction21,22, impairing insulin secretion and/or action.
Results
Metabolic characterization in vivo
Heterozygous Hhip (Hhip+/−) mice and control littermates (Hhip+/+) (Jackson Laboratories) were used [N.B., Adult Hhip+/– mice are phenotypically indistinguishable from control littermates (Hhip+/+), whereas Hhip−/− die after birth due to lung defects; thus, Hhip+/− mice were used in the current study9,12]. We compared the growth pattern and energy intake of both male and female animals fed either normal diet (ND) or HFD from the age of 6 until 14 weeks (Fig. 1a–c, male; Fig. 1d–f, female). As expected, HFD progressively increased body weight (BW) in both sexes of Hhip+/+ vs. Hhip+/− animals (Fig. 1a, male; Fig. 1d, female) with similar BW gain patterns (Fig. 1b, male; Fig. 1e, female). Notably, in both ND and HFD conditions, male Hhip+/− mice were heavier than male Hhip+/+ mice, while female animals (Hhip+/+ and Hhip+/−) had similar BW.
Physiological parameters in both male (a-c, ND-Hhip+/+, n = 15; ND-Hhip+/−, n = 12; HFD-Hhip+/+, n = 18; and HFD-Hhip+/−, n = 14) and female (d-f, ND-Hhip+/+, n = 8; ND-Hhip+/−, n = 7; HFD-Hhip+/+, n = 12; and HFD-Hhip+/−, n = 12) mice from the age of 6 to 14 weeks. (a,d) body weight (BW, g); (b,e) body weight gain; (g,c,f) energy intake calculated based on food consumption per week (kCal/week). Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001 vs. ND-Hhip+/+; ǂp ≤ 0.05; ǂǂp ≤ 0.01; ǂǂǂp ≤ 0.001 vs. ND-Hhip+/−; †p ≤ 0.05; ††p ≤ 0.01, HFD-Hhip+/+ vs. HFD-Hhip+/−.
After 8 weeks of HFD, both sexes of 14 week-old HFD-Hhip+/+ mice developed a similar pattern of glucose intolerance (male, Fig. 2a; female, Fig. 2c), but hyperinsulinaemia was observed only in male HFD mice (Hhip+/+ and Hhip+/−) (Fig. 2b), not in female HFD mice (Hhip+/+ and Hhip+/−) (Fig. 2d), documented by plasma insulin levels measured during intraperitoneal glucose tolerance test (ipGTT). As compared to male HFD-Hhip+/+ mice, the impaired ipGTT was ameliorated in male HFD-Hhip+/− mice that had high levels of circulating plasma insulin (ng/ml) in total and/or in two insulin secretion phases (1st phase: 0–15 minutes; 2nd phase: 15–120 mins) during the ipGTT (Fig. 2b). In contrast, those changes did not occur in female HFD mice (Hhip+/+ vs. Hhip+/−) (Fig. 2d). Therefore, we focused on male mice for the rest of the studies.
Metabolic parameters in male (a,b) and female (c,d) mice at the age of 14 weeks. (a,c) ipGTT and ipGTT-area under the curve (AUC) quantification (0–120 mins). (a: ND-Hhip+/+, n = 17; ND-Hhip+/−, n = 15; HFD-Hhip+/+, n = 23; and HFD-Hhip+/−, n = 17); (c: ND-Hhip+/+, n = 8; ND-Hhip+/−, n = 7; HFD-Hhip+/+, n = 12; and HFD-Hhip+/−, n = 12); (b,d) plasma circulating insulin level (ng/ml) and its AUC quantifications (total, 0–120 mins; 1st phase, 0–15 mins; 2nd phase, 15–120 mins). (b: ND-Hhip+/+, n = 11; ND-Hhip+/−, n = 7; HFD-Hhip+/+, n = 13; and HFD-Hhip+/−, n = 13); (d: ND-Hhip+/+, n = 6; ND-Hhip+/−, n = 5; HFD-Hhip+/+, n = 11; and HFD-Hhip+/−, n = 10). Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001 vs. ND-Hhip+/+; ǂp ≤ 0.05; ǂǂp ≤ 0.01; ǂǂǂp ≤ 0.001 vs. ND-Hhip+/−; †p ≤ 0.05; ††p ≤ 0.01; †††p ≤ 0.001, HFD-Hhip+/+ vs. HFD-Hhip+/−; NS, non-significant.
Here, we did not observe any apparent changes in terms of intraperitoneal insulin sensitivity test (ipIST) (Fig. 3a) and longitudinal systolic blood pressure (SBP) measurement (Fig. 3b) among 4 subgroups of male mice (Hhip+/+ vs Hhip+/−; ND vs HFD). EchoMRI analysis revealed that both male Hhip+/+ and Hhip+/− mice had similar fat/lean mass at the age of 6 or 14 weeks in either ND or HFD conditions (Fig. 3c), even though male Hhip+/− mice are generally bigger and heavier through life as compared to male Hhip+/+ mice. Next, we analyzed several pancreatic parameters including pancreatic mass (ratio of pancreas weight to BW, Fig. 3d), beta cell mass (Fig. 3e) and Shh-Glis (Gli 1-Gli3) mRNA expression (Fig. S1), as well as total islets numbers (Fig. 3f) in the isolated islets among 4 subgroups of male mice (Hhip+/+ vs Hhip+/−; ND vs HFD) and we did not find any apparent changes, as well as no changes in mRNA expression levels of Irs 1, Irs 2, InsR and Glu2 genes in those isolated islets (Fig. 3g).
Metabolic and pancreatic parameters in male mice. (a) ipIST and ipIST- AUC quantification (0–90 mins) in mice at the age of 14 weeks (ND-Hhip+/+, n = 9; ND-Hhip+/−, n = 7; HFD-Hhip+/+, n = 14; and HFD-Hhip+/−, n = 7); (b) SBP measurement in mice from the age of 8 to 14 weeks (ND-Hhip+/+, n = 5; ND-Hhip+/−, n = 4; HFD-Hhip+/+, n = 9; and HFD-Hhip+/−, n = 6); (c) EchoMRI analysis in mice at the age of 6 weeks (ND-Hhip+/+, n = 21; ND-Hhip+/−, n = 13) and 14 weeks (ND-Hhip+/+, n = 9; ND-Hhip+/−, n = 7; HFD-Hhip+/+, n = 12; and HFD-Hhip+/−, n = 10); (d–f) pancreatic parameters in mice at the age of 14 weeks; (d) pancreatic mass (ratio of pancreas weight to BW (ND-Hhip+/+, n = 6; ND-Hhip+/−, n = 5; HFD-Hhip+/+, n = 5; and HFD-Hhip+/−, n = 7); (e) beta cell mass and (f) total islets numbers (ND-Hhip+/+, n = 3; ND-Hhip+/−, n = 3; HFD-Hhip+/+, n = 6; and HFD-Hhip+/−, n = 6); and (g) qPCR-mRNA expression of Irs1 (ND-Hhip+/+, n = 7; ND-Hhip+/−, n = 6; HFD-Hhip+/+, n = 5; and HFD-Hhip+/−, n = 6), Irs2 (ND-Hhip+/+, n = 7; ND-Hhip+/−, n = 6; HFD-Hhip+/+, n = 6; and HFD-Hhip+/−, n = 7), InsR (ND-Hhip+/+, n = 7; ND-Hhip+/−, n = 6; HFD-Hhip+/+, n = 6; and HFD-Hhip+/−, n = 7) and Glut2 genes (ND-Hhip+/+, n = 7; ND-Hhip+/−, n = 5; HFD-Hhip+/+, n = 6 and HFD-Hhip+/−, n = 7) (vs 18 S mRNA ratio), in isolated islets among 4 subgroups of male mice (Hhip+/+ vs Hhip+/−; ND vs HFD) at 14 week-old. Data shown as mean ± SEM; (a,b,d–f) 1 way-ANOVA followed by Bonferroni’s post hoc test; (c) unpaired student’s t-test. **p ≤ 0.01; ***p ≤ 0.001 vs. ND-Hhip+/+; ǂp ≤ 0.05; ǂǂǂp ≤ 0.001 vs. ND-Hhip+/−; NS, non-significant.
Pancreatic Hhip expression and glucose stimulated insulin secretion (GSIS)
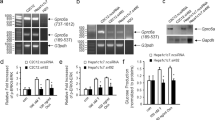
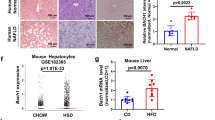
Although Hhip is abundantly expressed in the developing pancreas9 and in vascular endothelial cells19, less is known about its basal expression level in mature islet cells such as alpha and beta cells, and also, whether HFD could alter its expression profile in these islet cells and subsequently impact on insulin secretion/action. As compared to ND, HFD stimulated Hhip protein expression in freshly isolated islets of both Hhip+/+ and Hhip+/− mice, but, less in HFD- Hhip+/− mice vs HFD- Hhip+/+ mice by western blot (Fig. 4a). HFD increased Hhip-IHC expression in the islets of Hhip+/+ mice (Fig. 4b), predominantly evident in beta cells, not alpha cells (co-localized with insulin (Fig. 4c) and glucagon (Fig. 4d)). In addition, ex vivo studies revealed that the blunted GSIS in HFD mice reduced insulin secretion, ~7-fold less than in ND animals (Fig. 4e). Islets of Hhip heterozygous (Hhip+/−) mice had enhanced GSIS (ND, ~1.8 fold increase; HFD, ~1.6 fold increase) as compared to Hhip+/+ mice under ND and HFD conditions (Fig. 4e). In INS-1 832/13 cells in vitro, siRNA-Hhip (50 nM) further enhanced the stimulatory effects of 16.8 mM glucose on GSIS (Fig. 4f).
Pancreatic Hhip expression (a–d) and GSIS (e,f). (a–d) Hhip expression in the islets of male Hhip mice (Hhip+/+ vs. Hhip+/+; ND vs HFD) at 14 week-old; (a) Western blot; (b) Hhip-IHC staining (scale bar, 50 μm); (c) co-localization of IF-Hhip (red), IF-insulin (green) and DAPI (blue); (d) co-localization of IF-Hhip (red), IF-glucagon (green) and DAPI (blue); (e) GSIS in isolated islets from male Hhip+/+ vs Hhip+/− mice at 14 week-old under ND and HFD conditions (insulin secretion, % of total insulin content (ng/ml) measured in isolated islets cultured at 2.8 vs. 16.8 mM D-glucose medium). *p ≤ 0.05; ***p ≤ 0.001 vs. ND-Hhip+/+; ###p ≤ 0.001 vs. ND-Hhip+/−; ††p ≤ 0.01, HFD-Hhip+/+ vs. HFD-Hhip+/−; NS, non-significant. (f) siRNA-Hhip (50 nM) effect on GSIS in INS-1 832/13 cells (insulin secretion, % of total insulin content (ng/ml) measured in INS-1 832/13 cells cultured at 2.8 or 16.8 mM D-glucose medium). ***p ≤ 0.001 vs. INS-1 832/13 cells cultured in 2.8 mM D-glucose medium). NS, non-significant. Three to four separate experiments (ex vivo and in vitro); Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test.
Islet integrity and beta cell morphology
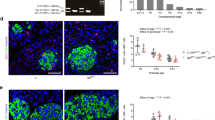
Next, we analyzed the frequency distribution profile of islet size by measuring islet surface areas from 0–500 µm2 to over 20000 µm2 in animals under both ND (Fig. 5a) and HFD (Fig. 5b) conditions. It appeared that ND-Hhip+/− and ND-Hhip+/+ mice displayed a similar islet-size distribution pattern (Fig. 5a). However, HFD-Hhip+/− had relatively more smaller-sized islets (<2000 µm2) as compared to HFD-Hhip+/+ mice, while HFD-Hhip+/+ mice had an increase in larger-sized islets (>2000 µm2) (Fig. 5b). Notably, morphological features including disoriented islet architecture with an invasion of α-cells into the central core of beta cells (Fig. 5c, insulin-IF staining and glucagon-IF staining) were observed in the pancreases of HFD-Hhip+/+ mice. More specifically, reduced insulin content (Fig. 6a, insulin-IHC staining) and decreased beta cell proliferation (Fig. 6a, Ki-67-IF staining) occurred significantly more often in the pancreases of HFD-Hhip+/+ mice. In contrast, HFD-Hhip+/− mice had a preponderance of smaller-sized islets (<2000 µm2) (Fig. 5b), in which islet integrity (Fig. 5c) and insulin content (Figs 5c and 6a) were preserved and protected, while increased beta cell proliferation (Fig. 6a) was detected as well.
Islet analysis of male mice at 14 week-old (Hhip+/+ vs Hhip+/−; ND vs HFD). (a,b) Distribution profile of islet size [a, ND (ND-Hhip+/+, n = 3; ND-Hhip+/−, n = 3); (b), HFD (HFD-Hhip+/+, n = 6; HFD-Hhip+/−, n = 6)]. Data shown as mean ± SEM; 2 way-ANOVA followed by Bonferroni’s post hoc test. †p ≤ 0.05; †††p ≤ 0.001, HFD-Hhip+/+ vs. HFD-Hhip+/−; (c) co-localization of IF-insulin (green) and –glucagon (red) staining (scale bar, 50 μm).
IHC/IF staining in vivo (a) and in vitro (b). (a) Insulin-IHC (ND-Hhip+/+, n = 3; ND-Hhip+/−, n = 3; HFD-Hhip+/+, n = 5; and HFD-Hhip+/−, n = 5); IF-Ki67 staining (Ki67, red; insulin, green; DAPI, blue; Ki-67 positive nuclei, pink arrows) (ND-Hhip+/+, n = 3; ND-Hhip+/−, n = 3; HFD-Hhip+/+, n = 5; and HFD-Hhip+/−, n = 5) and 8-OHdG-IHC staining (ND-Hhip+/+, n = 4; ND-Hhip+/−, n = 3; HFD-Hhip+/+, n = 6; and HFD-Hhip+/−, n = 5) in the islets among 4 subgroups of male mice (Hhip+/+ vs Hhip+/−; ND vs HFD) at 14 week-old (scale bar, 50 μm). Semi-quantification of staining; Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test. *p ≤ 0.05; ***p ≤ 0.001 vs. ND-Hhip+/+; ǂǂǂp ≤ 0.001 vs. ND-Hhip+/−; †p ≤ 0.05 HFD-Hhip+/+ vs. HFD-Hhip+/−; NS, non-significant. (b) IF-insulin staining (insulin, green; DAPI, blue) and DHE staining (DHE, red; DAPI, blue) (scale bar, 50 µm); Semi-quantification; Four separate experiments; Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test. **p ≤ 0.01; ***p ≤ 0.001; NS, non-significant vs INS-1 832/13 cells cultured in medium without rHhip (0 ng/ml) (100%).
Oxidative stress
8-hydroxy-2-deoxyguanosine (8-OHdG) staining (Fig. 6a) revealed that oxidative stress was markedly elevated in the islets of HFD-Hhip+/+ mice vs. HFD-Hhip+/− mice, while no differences were observed between ND-Hhip+/+ mice and ND-Hhip+/− mice. In vitro, recombinant Hhip (rHhip) dose-dependently decreased and increased the numbers of insulin-positive cells (Fig. 6b) and dihydroethidium (DHE)-positive cells (Fig. 6b), respectively.
Then, we analyzed NADPH oxidase (Nox 1, 2 and 4 genes) mRNA expression by qPCR in isolated mouse islets and found that only Nox 2 mRNA was significantly increased in the islets of HFD-Hhip+/+ mice vs. HFD-Hhip+/− mice (Fig. 7a), while Nox 1 and Nox 4 mRNA were barely detectable in those islets (data not shown). Co-IF staining (Nox2 and insulin) further confirmed that the elevated Nox2-IF expression was far greater in the beta cells of HFD-Hhip+/+ mice vs. HFD-Hhip+/− mice (Fig. 7b). In INS-1 832/13 cells, we could detect both Nox 2 and Nox 4 mRNA expression, but not Nox 1 mRNA expression. As shown, rHhip dose-dependently increased Nox2 gene expression (mRNA, Fig. S2a; protein, Fig. 7c) and elevated NADPH activity (Fig. S2b), while Nox4 gene expression remained unchanged (mRNA, Fig. S2a; protein, Fig. 7c). In addition, 0.3 mM BSA-sodium palmitate (PA) stimulated Nox2 mRNA expression without affecting Nox4 mRNA expression (Fig. S2c). The immunoblotting data revealed that 0.3 mM BSA-PA stimulated both Hhip and Nox2 protein expression, which was increased by rHhip (Fig. 7d) or abolished by siRNA-Hhip (Fig. 7e).
Nox2 gene expression in vivo (a,b) and in vitro (c–f). (a) qPCR –Nox2 mRNA expression (ND-Hhip+/+, n = 7; ND-Hhip+/−, n = 6; HFD-Hhip+/+, n = 6; and HFD-Hhip+/−, n = 7); Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test. *p ≤ 0.05, HFD-Hhip+/+ vs. HFD-Hhip+/− male mice at 14 week-old; (b) IF-Nox2 staining (Nox2, green; insulin, red; DAPI, blue) in the islets among 4 subgroups of male mice (Hhip+/+ vs Hhip+/−; ND vs HFD) at 14 week-old (scale bar, 50 μm). (c) WB (Nox2 and Nox4 protein expression). The cells were treated by rHhip (0–5 ng/ml). (d,e) WB (Nox2 protein expression). The cells were treated by PA (0.3 mM) with rHhip (5 ng/ml) (d) or siRNA-Hhip (50 nM) (e). Three to four separate experiments; Data shown as mean ± SEM; 1 way-ANOVA followed by Bonferroni’s post hoc test; *p ≤ 0.05; ***p ≤ 0.001; NS, non-significant vs INS-1 832/13 cells cultured in medium with 0.5% BSA (100%).
Discussion
In the present study, we demonstrated that HFD stimulates Hhip gene expression in beta cells, resulting in disruption of islet integrity and decreased GSIS. Mechanistically, Hhip promotes oxidative stress in beta cells via the Nox 2 gene.
As expected, 8 weeks of HFD increased body weight over time in both sexes of HFD-Hhip+/+ and HFD-Hhip+/− mice. Despite the fact that female sex has been reported to be protective against HFD-induced metabolic disorders23,24,25, both sexes of HFD-Hhip+/+ mice displayed a similar ipGTT pattern in our study, and the impaired glucose intolerance was improved only in male HFD-Hhip+/− mice, but not in female HFD-Hhip+/− mice. To date, a few publications have explicitly looked at the role of sex in the response to HFD in mice23,24,26,27,28. Notably, HFD impact on metabolic phenotypes largely depends on sexually dimorphic variables, but strains, animal ages and environmental factors may be confounders23,24,26,27,28. Here, the sexual dimorphism might be explained by (1) the mixed genetic background in our JAX® Hhip mouse line. For instance, by using gold-standard methodologies, Fergusson et al. reported that C57BL/6J and C57BL/6N mice in response to glucose challenge to secrete insulin are completely different29. (2) the impact of body weight. As an example, Ingvorsen et al. studied the impact of diet and sex on 41 metabolic related variables from 1319 C57BL/6N mice (ND, N = 586 vs. HFD, N = 733) and found that for the majority of variables (79%), body weight on HFD-induced metabolic changes is a significant source of sexually dimorphic variation28. Nevertheless, seeing the differences between sexes, we then focused on male animals to elucidate the relevant mechanisms.
Glucose uptake and insulin secretion are the best known mechanisms that impact on systemic glucose tolerance. Relevant to the pancreas, insulin has been shown to be involved in glucose uptake from circulation into peripheral insulin-responsive tissues through glucose transporters, such as Glut2 in pancreatic islets30. Under basal conditions, Glut2 level is very low (sequestered internally), but in the presence of insulin (with the appropriate stimulus such as HFD), the expression of Glut2 might be altered to maintain glucose homeostasis30,31,32. Here, we did not observe any apparent changes of Glut2 mRNA in pancreatic islets, hinting that insulin-mediated glucose uptake in pancreatic islets might be normal. In line with others33,34,35, mouse islets express low levels of classical Shh signaling, underscoring the independence of Hhip action. Our data also revealed that the glucose response to exogenous insulin challenge (ipIST) and insulin signaling (Irs 1, Irs 2 and InsR mRNA) remained at similar levels among the 4 subgroups of mice fed either ND or HFD; and there were not any apparent changes in pancreatic mass, beta cell mass and total islet numbers in those experimental animals. Notably, depending on the composition and timing of diet, beta cells respond to HFD feeding in mice with two actions–insulin compensation (which results in hyperinsulinemia) and/or insulin resistance24,36,37,38,39,40,41. For example, within a 10-week period of HFD intake, beta cells are prone to secrete more insulin (hyperinsulinemia) to compensate the increasing obesity and glucose intolerance24,38, while long-term HFD consumption (more than 10 weeks) eventually culminates in beta cell failure and the change of those parameters is considered a major adaptation to insulin resistance36,37,38,39,41. Being neither insulin insensitivity nor insulin resistance, the current model with 8-week HFD might only induce beta cell proliferation as reported24,38.
Thus, we speculate that the impaired ipGTT observed in HFD-Hhip+/+ mice might be as a result of pancreatic beta cell dysfunction on GSIS. Indeed, we here found that Hhip expression was significantly increased in the islets of HFD-Hhip+/+ mice (>HFD- Hhip+/− mice), mainly co-localized in beta cells and none in alpha cells, pin-pointing beta cells as a main target of HFD-induced Hhip expression. The islets isolated from Hhip+/− mice under both ND and HFD diet had enhanced GSIS responses as compared to those from Hhip+/+ mice. Similarly, siRNA-Hhip in INS-1 832/13 cells resulted in an increase of GSIS in response to glucose challenge from 2.8 mM to 16.8 mM. Together, these data suggest that Hhip deficiency might directly preserve the capacity and ability of GSIS in insulin production and lower Hhip gene expression could enhance GSIS, and that might also explain the phenotype of HFD- Hhip+/− mice showing an improved ipGTT with higher circulating plasma insulin levels in two insulin secretion phases. We are unable to capture the obvious expected 1st phase of insulin secretion (data not shown), that result might be due to the mixed genetic background of our mice. As noted by others42,43,44, glucose metabolism on insulin secretory function indeed varies in a strain-dependent manner.
Islet size, insulin content and the amount of insulin released are known to be correlated in proportional population existing in animals and humans, evidenced by small islets that are comprised of more beta cells with higher insulin content than large islets45,46. When there is increased insulin demand as in diabetes and obesity, islet cell composition and size distribution pattern appear to be altered, and some islets exhibit abnormal architecture with intermingled alpha and beta cells, particularly large islets and distribution with a disproportionate number of larger size islets populations45,47. As noted, HFD-Hhip+/+ mice phenotypically had larger-sized islets with alpha cell invasion, lower insulin content and less beta cell proliferation. In contrast, Hhip deficiency protected islet integrity, in response to HFD challenge over time; and there were increased numbers smaller islets and beta cell proliferation in HFD-Hhip+/− mice, thereby improving GSIS.
Mechanistically, evidence indicates that under oxidative stress, Hhip gene expression may redirect cells toward apoptosis, fibrosis, angiogenesis and/or tumorigenesis19,48,49,50. Previously, we demonstrated that Hhip could directly elevate ROS generation; interact with NOX isoform(s) (i.e., Nox 4 in the kidney), subsequently activating TGFβ1-signaling to result in renal cell fibrosis/apoptosis, irrespective of diabetes50,51. Here, oxidative stress in the islets and beta cells was markedly elevated in HFD-Hhip+/+ mice vs. HFD-Hhip+/− mice. In vitro, rHhip dose-dependently decreased insulin-positive beta cells number; increased the number of DHE-positive cells and NADPH activity. Together, these data suggest that HFD-increased Hhip gene expression might elevate oxidative stress in beta cells, thereby impairing GSIS on insulin secretion.
In the pancreas, the NOX family represents one potential source of ROS in insulin-secreting cells22. Among NOX isoforms, we detected Nox2 gene expression (islets and INS-1 832/13 cells); Nox4 (INS-1 832/13 cells); but not Nox1 (neither islets nor INS-1 832/13 cells), in line with others21,22. Also, only Nox2 (but not Nox4) gene expression was elevated in the islets of HFD-Hhip+/+ mice, and that elevated Nox2 expression was ameliorated in the islets of HFD-Hhip+/− mice. Currently, it is still unclear whether Nox2 has an impact on GSIS or not, since both positive21 and negative52 results have been reported in Nox2 KO mice (C57BL/6) (Jackson Laboratory, Bar Harbor, ME) as well. Nevertheless, we further validated Hhip impact on Nox2 gene expression in vitro. We found that rHhip directly stimulated Nox2 expression (mRNA and protein) in a dose-dependent manner; 0.3 mM BSA-PA stimulated both Hhip and Nox2 protein expression, that could be either aggravated or abolished by rHhip or siRNA-Hhip, respectively. Taken together, not only did we confirm the notion of Nox2 acting as the predominant isoform and therefore Nox2 elevates ROS production to antagonize GSIS in the regulation of insulin secretion21, but we also established that a lower Hhip gene expression might protect islet integrity against HFD-mediated beta cell dysfunction; improve GSIS on maintaining sufficient levels of insulin secretion via ameliorating ROS-Nox2 gene expression. Clearly, beta cell-specific gain- and/or loss-of-Hhip function/expression models would be merited to circumvent the potential pitfall of the current whole body Hhip-deficient model in the future.
In conclusion, pancreatic Hhip gene regulates insulin secretion by altering islet integrity and promoting Nox2 gene expression in beta cells in response to HFD-mediated beta cell dysfunction. As a perspective, exploring a method to decrease/lower Hhip expression may provide a new therapeutic strategy in the diagnosis, prevention and treatment of T2D.
Materials and Methods
Animal models
Heterozygous Hhip (Hhip+/−) mice and control littermates (Hhip+/+) (Jackson Laboratories, Hhiptm1Amc/J; mixed background of C57BL/6, Swiss-Webster, 129) were used (N.B., Adult Hhip+/– mice are phenotypically indistinguishable from control littermates (Hhip+/+), whereas Hhip−/− die after birth due to lung defects; thus, Hhip+/− mice were used in the current study9,12). In brief, both sexes of Hhip mice (Hhip+/− vs. Hhip+/+) at the age of 6 weeks were fed with normal chow (ND) (18% protein with 6.2% fat, calories from protein 24%, fat 18% and carbohydrate 58%) (Harlan Teklad, Montreal, Canada) or HFD (20.5% protein with 36% fat, calories from protein 14%, fat 60% and carbohydrate 26%) (Bio-Serv, Flemington, NJ) until 14 weeks of age, as reported previously53. After euthanasia (75 mg/kg sodium pentobarbital i.p.), pancreata were rapidly processed and harvested for either islets isolation or fixation for immunohistochemistry (IHC) and/or immunofluorescence (IF).
All animal protocols were carried out in strict accordance with the recommendations in the NIH Guide for the Care and Use of Laboratory Animals and followed the Principles of Laboratory Animal Care [National Institutes of Health (NIH) publication no. 85–23, revised 1985: http://grants1.nih.gov/grants/olaw/references/phspol.htm]. Animal care and experimental procedures were approved by the Animal Care Committee from the Centre de recherche du centre hospitalier de l’Université de Montréal (CRCHUM). Animals were housed in ventilated cages in SPF conditions under a 12 hours light-dark cycle with free access to chow and water at the animal facility of the CRCHUM. Breeding was carried out in pairs (Hhip+/+ with Hhip+/− mating) under the same housing conditions.
Biological parameters measurement
Body weight (BW, g) and energy intake (kCal/week) were monitored weekly through the experimental course. With the aid of the Rodent Cardiovascular Phenotyping Core Facility (CRCHUM), fat and lean mass as a percentage of body weight for mice were assessed using an EchoMRI-700 (EchoMRI™, Houston, TX); and plasma insulin was measured by mouse ultrasensitive insulin ELISA jumbo kit (Alpco Diagnostics, Salem, NH). Longitudinal systolic blood pressure (SBP) (from the age of 8 until 14 weeks following one week of pre-training) was monitored by the tail-cuff method with a BP-2000 Blood Pressure Analysis System (Visitech Systems Inc., Apex, NC), as reported elsewhere50,53. Intraperitoneal glucose tolerance test (ipGTT) and insulin sensitivity test (ipIST) were performed according to a standard protocol, with 6 and 4 hours fasting periods before sacrifice at the age of 14 weeks, respectively53. Blood glucose was quantified with an Accu-Chek Performa glucose meter (Roche Diagnostics, Laval, QC, Canada)50,53.
Islet isolation and glucose stimulated insulin secretion
Islet isolation was performed following a previously reported protocol54. In brief, mouse islets were isolated by collagenase P digestion and purified on a bilayer HBSS/Histopaque 1091 and 1077 (Sigma-Aldrich Canada, Oakville, ON, Canada) gradient centrifugation. Freshly isolated islets were used for Western Blot (WB) and qPCR analysis. For glucose-stimulated insulin secretion (GSIS) experiments, isolated islets were kept in culture at 37 °C in 2.8 mM glucose RPMI complete medium supplemented with 10% fetal bovine serum (FBS) for overnight recovery. GSIS and insulin content in both islets and culture media were assessed in 1-hour static incubation in Krebs-Ringer bicarbonate buffer containing 10 mM HEPES (pH 7.4, KRBH), 0.5% defatted BSA (d-BSA) at 2.8 mM glucose and 16.8 mM glucose, respectively (CRCHUM Cellular Physiology Core Facility).
Islets numbers, size and beta cell mass
With the aid of the CRCHUM Cellular Physiology Core Facility, we measured several islet parameters including total islet number, size and beta cell mass, as reported29,55. In brief, whole pancreata were removed, weighed, fixed and embedded in paraffin blocks. Longitudinal pancreatic cross sections (5 μm thick) were collected at 30 μm intervals. Insulin-IHC (anti-guinea pig insulin antibody, Agilent-DAKO, Santa Clara, CA) with counterstaining with hematoxylin was performed according to a standard protocol. At least 4–6 slides from each pancreas were processed for islet and beta cell mass measurements. First, slides were scanned at 20X magnification by using an Aperio ScanScope model CS slide scanner (Leica Biosystems Inc., Concord, ON, Canada) to assess islet/beta cell area and whole pancreas area via the Aperio Pixel count algorithm v9 (ImageScope v12.3.2.5030, Leica Biosystems Inc.), followed by calculation of the ratio of beta cell area to whole pancreas area. Then, beta cell mass was calculated by multiplying the ratio of beta cell area to whole pancreas area with whole pancreatic mass (mg) measured before fixation. Finally, morphometric measurements were performed by identifying manually regions of interest (ROIs) around insulin-IHC positive islets. The surface of all islets from at least 4 sections (>400 islets) were calculated for each ROI (ImageScope) and used to generate the size frequency distribution (surface) profile.
Cell culture
We employed a pancreatic beta cell line, rat INS-1 832/1356,57 kindly provided by Dr. Marc Prentki (CRCHUM, Montreal, QC, Canada) for our in vitro study. In brief, INS-1 832/13 cells (passage 51–53) have robust glucose responsiveness over the physiological range of glucose concentrations (i.e., 2.8–16.8 mM glucose) and were cultured in RPMI 1640 medium supplemented with 10% FBS, 10 mM HEPES, 2 mM L-glutamine, 1 mM sodium pyruvate, and 50 µM β-mercaptoethanol.
Oxidative stress and real-time quantitative PCR (qPCR)
Oxidative stress was determined by 8-hydroxy-2-deoxyguanosine (8-OHdG) staining (a biomarker for oxidative stress or a marker for reactive oxygen species (ROS) damage)58; dihydroethidium (DHE) staining (a cell-permeable fluorescent dye, redox indicator), NADPH oxidases (Nox 1, 2 and 4) gene expression and NADPH activity50,51,59,60. qPCR (Fast SYBR green master mix kit and 7500 Fast real-time PCR system; Applied Biosystems, Foster City, CA, USA) was performed as reported previously50,51. Primer sequences for qPCR were listed in Table 1.
Reagents and chemicals
Antibodies (method with dilution) used included anti-Hhip (5D11) (WB, 1:2000; IF/IHC, 1:300), glucagon (IF, 1:200), β-actin (WB, 1:100000) from Sigma-Aldrich Canada (Oakville, ON, Canada); anti-Ki 67 (IF, 1: 50) from BD Bioscience (San Jose, CA, USA); anti-insulin (H-86) (IF/IHC, 1:500) and gp91-phox antibody (K-15) (IF, 1:200) from Santa Cruz Biotechnology (Santa Cruz, CA, USA); anti-mouse 8-OHdG (IHC, 1: 50) from Abcam (Toronto, ON, Canada). Mouse anti-gp91phox-Cter for WB (1:10000) was obtained from Dr. Nathalie Grandvaux (CRCHUM) as reported elsewhere61. Chemical reagents included small interfering RNA (siRNA) of Hhip from Santa Cruz Biotechnology (Santa Cruz, CA, USA), which pools three target-specific 19–25 nucleotide sequences (Fig. S2e); recombinant Hhip (rHhip) from R&D Systems, Inc. (Burlington, ON, Canada); BSA (fatty acid free) and sodium palmitate (PA) were procured from Sigma-Aldrich Canada. The preparation of 0.3 mM BSA-PA (vs. the control, 0.5%BSA) was done as reported elsewhere53,62. Semi-quantitation of relative staining values was performed by NIH Image J software (Bethesda, MD). The images (N = 8~15 per animal) were analyzed and quantitated in a randomized and blinded fashion.
Statistical analysis
For animal studies, groups of 8 to 23 mice were studied (N.B. The precise number of animals used for each specific experiment is either labeled in the figures or the individual data points are shown on a column scatter graph). Ex vivo and in vitro, three to four separate experiments were performed for each protocol. All values represent mean ± SEM. Statistical significance between the experimental groups was analyzed by Student’s t-test, 1-way ANOVA and/or 2-way ANOVA (islet size frequency distribution) followed by the Bonferroni test using Prism 5.0 software (GraphPad, San Diego, CA). A probability level of p ≤ 0.05 was considered to be statistically significant50,53.
References
Ogurtsova, K. et al. IDF Diabetes Atlas: Global estimates for the prevalence of diabetes for 2015 and 2040. Diabetes Res Clin Pract 128, 40–50 (2017).
Prentki, M., Joly, E., El-Assaad, W. & Roduit, R. Malonyl-CoA signaling, lipid partitioning, and glucolipotoxicity: role in beta-cell adaptation and failure in the etiology of diabetes. Diabetes 51(Suppl 3), S405–413 (2002).
Prentki, M. & Madiraju, S. R. Glycerolipid/free fatty acid cycle and islet beta-cell function in health, obesity and diabetes. Molecular and cellular endocrinology 353, 88–100 (2012).
Nolan, C. J., Ruderman, N. B., Kahn, S. E., Pedersen, O. & Prentki, M. Insulin resistance as a physiological defense against metabolic stress: implications for the management of subsets of type 2 diabetes. Diabetes 64, 673–686 (2015).
Boyle, J. P., Thompson, T. J., Gregg, E. W., Barker, L. E. & Williamson, D. F. Projection of the year 2050 burden of diabetes in the US adult population: dynamic modeling of incidence, mortality, and prediabetes prevalence. Popul Health Metr 8, 29 (2010).
Hebrok, M. Hedgehog signaling in pancreas development. Mech Dev 120, 45–57 (2003).
Landsman, L., Parent, A. & Hebrok, M. Elevated Hedgehog/Gli signaling causes beta-cell dedifferentiation in mice. Proc Natl Acad Sci USA 108, 17010–17015 (2011).
Thomas, M. K., Lee, J. H., Rastalsky, N. & Habener, J. F. Hedgehog signaling regulation of homeodomain protein islet duodenum homeobox-1 expression in pancreatic beta-cells. Endocrinology 142, 1033–1040 (2001).
Kawahira, H. et al. Combined activities of hedgehog signaling inhibitors regulate pancreas development. Development 130, 4871–4879 (2003).
Bishop, B. et al. Structural insights into hedgehog ligand sequestration by the human hedgehog-interacting protein HHIP. Nat. Struct. Mol. Biol 16, 698–703 (2009).
Bosanac, I. et al. The structure of SHH in complex with HHIP reveals a recognition role for the Shh pseudo active site in signaling. Nat. Struct. Mol. Biol 16, 691–697 (2009).
Chuang, P. T. & McMahon, A. P. Vertebrate Hedgehog signalling modulated by induction of a Hedgehog-binding protein. Nature 397, 617–621 (1999).
Coulombe, J., Traiffort, E., Loulier, K., Faure, H. & Ruat, M. Hedgehog interacting protein in the mature brain: membrane-associated and soluble forms. Mol. Cell Neurosci 25, 323–333 (2004).
Holtz, A. M. et al. Secreted HHIP1 interacts with heparan sulfate and regulates Hedgehog ligand localization and function. The Journal of cell biology 209, 739–757 (2015).
Kwong, L., Bijlsma, M. F. & Roelink, H. Shh-mediated degradation of Hhip allows cell autonomous and non-cell autonomous Shh signalling. Nat. Commun 5, 4849 (2014).
Chuang, P. T., Kawcak, T. & McMahon, A. P. Feedback control of mammalian Hedgehog signaling by the Hedgehog-binding protein, Hip1, modulates Fgf signaling during branching morphogenesis of the lung. Genes Dev 17, 342–347 (2003).
Lao, T. et al. Hhip haploinsufficiency sensitizes mice to age-related emphysema. Proc Natl Acad Sci USA 113, E4681–4687 (2016).
Kayed, H. et al. Localization of the human hedgehog-interacting protein (Hip) in the normal and diseased pancreas. Mol. Carcinog 42, 183–192 (2005).
Olsen, C. L., Hsu, P. P., Glienke, J., Rubanyi, G. M. & Brooks, A. R. Hedgehog-interacting protein is highly expressed in endothelial cells but down-regulated during angiogenesis and in several human tumors. BMC cancer 4, 43 (2004).
Brissova, M. et al. Islet microenvironment, modulated by vascular endothelial growth factor-A signaling, promotes beta cell regeneration. Cell metabolism 19, 498–511 (2014).
Li, N. et al. NADPH oxidase NOX2 defines a new antagonistic role for reactive oxygen species and cAMP/PKA in the regulation of insulin secretion. Diabetes 61, 2842–2850 (2012).
Newsholme, P. et al. Insights into the critical role of NADPH oxidase(s) in the normal and dysregulated pancreatic beta cell. Diabetologia 52, 2489–2498 (2009).
Pettersson, U. S., Walden, T. B., Carlsson, P. O., Jansson, L. & Phillipson, M. Female mice are protected against high-fat diet induced metabolic syndrome and increase the regulatory T cell population in adipose tissue. PLoS One 7, e46057 (2012).
Oliveira, R. B., Maschio, D. A., Carvalho, C. P. & Collares-Buzato, C. B. Influence of gender and time diet exposure on endocrine pancreas remodeling in response to high fat diet-induced metabolic disturbances in mice. Annals of anatomy = Anatomischer Anzeiger: official organ of the Anatomische Gesellschaft 200, 88–97 (2015).
Gannon, M., Kulkarni, R. N., Tse, H. M. & Mauvais-Jarvis, F. Sex differences underlying pancreatic islet biology and its dysfunction. Molecular metabolism 15, 82–91 (2018).
El Akoum, S., Lamontagne, V., Cloutier, I. & Tanguay, J. F. Nature of fatty acids in high fat diets differentially delineates obesity-linked metabolic syndrome components in male and female C57BL/6J mice. Diabetology & metabolic syndrome 3, 34 (2011).
Nishikawa, S., Yasoshima, A., Doi, K., Nakayama, H. & Uetsuka, K. Involvement of sex, strain and age factors in high fat diet-induced obesity in C57BL/6J and BALB/cA mice. Experimental animals 56, 263–272 (2007).
Ingvorsen, C., Karp, N. A. & Lelliott, C. J. The role of sex and body weight on the metabolic effects of high-fat diet in C57BL/6N mice. Nutrition & diabetes 7, e261 (2017).
Fergusson, G. et al. Defective insulin secretory response to intravenous glucose in C57Bl/6J compared to C57Bl/6N mice. Molecular metabolism 3, 848–854 (2014).
Thorens, B. GLUT2, glucose sensing and glucose homeostasis. Diabetologia 58, 221–232 (2015).
Lu, B. et al. Improved glucose-stimulated insulin secretion by intra-islet inhibition of protein-tyrosine phosphatase 1B expression in rats fed a high-fat diet. Journal of endocrinological investigation 35, 63–70 (2012).
Salimi, M., Zardooz, H., Khodagholi, F., Rostamkhani, F. & Shaerzadeh, F. High-fat diet with stress impaired islets’ insulin secretion by reducing plasma estradiol and pancreatic GLUT2 protein levels in rats’ proestrus phase. Journal of physiology and pharmacology: an official journal of the Polish Physiological Society 67, 653–666 (2016).
Lau, J. & Hebrok, M. Hedgehog signaling in pancreas epithelium regulates embryonic organ formation and adult beta-cell function. Diabetes 59, 1211–1221 (2010).
Lau, J., Kawahira, H. & Hebrok, M. Hedgehog signaling in pancreas development and disease. Cell Mol Life Sci 63, 642–652 (2006).
Lee, J. J. et al. Stromal response to Hedgehog signaling restrains pancreatic cancer progression. Proc Natl Acad Sci USA 111, E3091–3100 (2014).
Ahren, J., Ahren, B. & Wierup, N. Increased beta-cell volume in mice fed a high-fat diet: a dynamic study over 12 months. Islets 2, 353–356 (2010).
Gupta, D. et al. Temporal characterization of beta cell-adaptive and -maladaptive mechanisms during chronic high-fat feeding in C57BL/6NTac mice. J Biol Chem 292, 12449–12459 (2017).
Mosser, R. E. et al. High-fat diet-induced beta-cell proliferation occurs prior to insulin resistance in C57Bl/6J male mice. Am J Physiol Endocrinol Metab 308, E573–582 (2015).
Peyot, M. L. et al. Beta-cell failure in diet-induced obese mice stratified according to body weight gain: secretory dysfunction and altered islet lipid metabolism without steatosis or reduced beta-cell mass. Diabetes 59, 2178–2187 (2010).
Prentki, M. & Nolan, C. J. Islet beta cell failure in type 2 diabetes. J Clin Invest 116, 1802–1812 (2006).
Saisho, Y. et al. beta-cell mass and turnover in humans: effects of obesity and aging. Diabetes Care 36, 111–117 (2013).
Andrikopoulos, S. et al. Differential effect of inbred mouse strain (C57BL/6, DBA/2, 129T2) on insulin secretory function in response to a high fat diet. The Journal of endocrinology 187, 45–53 (2005).
Berglund, E. D. et al. Glucose metabolism in vivo in four commonly used inbred mouse strains. Diabetes 57, 1790–1799 (2008).
Kaku, K., Fiedorek, F. T. Jr., Province, M. & Permutt, M. A. Genetic analysis of glucose tolerance in inbred mouse strains. Evidence for polygenic control. Diabetes 37, 707–713 (1988).
Farhat, B. et al. Small human islets comprised of more beta-cells with higher insulin content than large islets. Islets 5, 87–94 (2013).
Aizawa, T. et al. Size-related and size-unrelated functional heterogeneity among pancreatic islets. Life sciences 69, 2627–2639 (2001).
Kilimnik, G. et al. Altered islet composition and disproportionate loss of large islets in patients with type 2 diabetes. PLoS One 6, e27445 (2011).
Nie, D. M. et al. Endothelial microparticles carrying hedgehog-interacting protein induce continuous endothelial damage in the pathogenesis of acute graft-versus-host disease. American journal of physiology. Cell physiology 310, C821–835 (2016).
Zeng, X. et al. A freely diffusible form of Sonic hedgehog mediates long-range signalling. Nature 411, 716–720 (2001).
Zhao, X. P. et al. Hedgehog Interacting Protein Promotes Fibrosis and Apoptosis in Glomerular Endothelial Cells in Murine Diabetes. Sci Rep 8, 5958 (2018).
Liao, M. C. et al. AT2 R deficiency mediated podocyte loss via activation of ectopic hedgehog interacting protein (Hhip) gene expression. The Journal of pathology 243, 279–293 (2017).
de Souza, A. H. et al. NADPH oxidase-2 does not contribute to beta-cell glucotoxicity in cultured pancreatic islets from C57BL/6J mice. Molecular and cellular endocrinology 439, 354–362 (2017).
Aliou, Y. et al. Post-weaning high-fat diet accelerates kidney injury, but not hypertension programmed by maternal diabetes. Pediatric research 79, 416–424 (2016).
Jacqueminet, S., Briaud, I., Rouault, C., Reach, G. & Poitout, V. Inhibition of insulin gene expression by long-term exposure of pancreatic beta cells to palmitate is dependent on the presence of a stimulatory glucose concentration. Metabolism: clinical and experimental 49, 532–536 (2000).
Jetton, T. L. et al. Mechanisms of compensatory beta-cell growth in insulin-resistant rats: roles of Akt kinase. Diabetes 54, 2294–2304 (2005).
Hectors, T. L., Vanparys, C., Pereira-Fernandes, A., Martens, G. A. & Blust, R. Evaluation of the INS-1 832/13 cell line as a beta-cell based screening system to assess pollutant effects on beta-cell function. PLoS One 8, e60030 (2013).
Hohmeier, H. E. et al. Isolation of INS-1-derived cell lines with robust ATP-sensitive K+ channel-dependent and -independent glucose-stimulated insulin secretion. Diabetes 49, 424–430 (2000).
Zong, C. et al. The stress-response molecule NR4A1 resists ROS-induced pancreatic beta-cells apoptosis via WT1. Cellular signalling 35, 129–139 (2017).
Godin, N. et al. Catalase overexpression prevents hypertension and tubular apoptosis in angiotensinogen transgenic mice. Kidney Int 77, 1086–1097 (2010).
Shi, Y. et al. Angiotensin-(1-7) prevents systemic hypertension, attenuates oxidative stress and tubulointerstitial fibrosis, and normalizes renal angiotensin-converting enzyme 2 and Mas receptor expression in diabetic mice. Clinical science (London, England: 1979) 128, 649–663 (2015).
Soucy-Faulkner, A. et al. Requirement of NOX2 and reactive oxygen species for efficient RIG-I-mediated antiviral response through regulation of MAVS expression. PLoS pathogens 6, e1000930 (2010).
Roduit, R. et al. A role for hormone-sensitive lipase in glucose-stimulated insulin secretion: a study in hormone-sensitive lipase-deficient mice. Diabetes 50, 1970–1975 (2001).
Acknowledgements
We specially thank Dr. Marc Prentki (CRCHUM) for his valuable suggestions and discussions; Dr. Erik Joly (CRCHUM) for his technical advice on the analysis of islets number, size and beta cell mass. This project was supported by Natural Sciences and Engineering Research Council of Canada (NSERC) grant (RGPIN-2017-05615) to S-L Zhang. Editorial assistance was provided by the CRCHUM’s Research Support Office.
Author information
Authors and Affiliations
Contributions
S.L.Z. is the guarantor of this work, had full access to all study data, and takes responsibility for data integrity and the accuracy of data analysis. J.S.D.C. and S.L.Z. were principal investigators and were responsible for the study conception and design. H.N. wrote the first draft of the manuscript, and S.L.Z. drafted/reviewed/edited the manuscript. H.N., M.C.L., X.P.Z., S.Y.C., C.S.L., I.C. and S.L.Z. contributed to the experiments and collection of data. J.R.I. contributed to data discussion, and reviewed/edited the manuscript. All authors were involved in the analysis and interpretation of data and contributed to the critical revision of the manuscript. All authors provided final approval of the version to be published.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nchienzia, H., Liao, MC., Zhao, XP. et al. Hedgehog Interacting Protein (Hhip) Regulates Insulin Secretion in Mice Fed High Fat Diets. Sci Rep 9, 11183 (2019). https://doi.org/10.1038/s41598-019-47633-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47633-3
This article is cited by
-
Hedgehog interacting protein activates sodium–glucose cotransporter 2 expression and promotes renal tubular epithelial cell senescence in a mouse model of type 1 diabetes
Diabetologia (2023)
-
AT2R deficiency in mice accelerates podocyte dysfunction in diabetic progeny in a sex-dependent manner
Diabetologia (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.