Abstract
Store-operated Ca2+ entry (SOCE), the fundamental Ca2+ signaling mechanism in myogenesis, is mediated by stromal interaction molecule (STIM), which senses the depletion of endoplasmic reticulum Ca2+ stores and induces Ca2+ influx by activating Orai channels in the plasma membrane. Recently, STIM2β, an eight-residue-inserted splice variant of STIM2, was found to act as an inhibitor of SOCE. Although a previous study demonstrated an increase in STIM2β splicing during in vitro differentiation of skeletal muscle, the underlying mechanism and detailed function of STIM2β in myogenesis remain unclear. In this study, we investigated the function of STIM2β in myogenesis using the C2C12 cell line with RNA interference-mediated knockdown and CRISPR-Cas-mediated knockout approaches. Deletion of STIM2β delayed myogenic differentiation through the MEF2C and NFAT4 pathway in C2C12 cells. Further, loss of STIM2β increased cell proliferation by altering Ca2+ homeostasis and inhibited cell cycle arrest mediated by the cyclin D1-CDK4 degradation pathway. Thus, this study identified a previously unknown function of STIM2β in myogenesis and improves the understanding of how cells effectively regulate the development process via alternative splicing.
Similar content being viewed by others
Introduction
Myogenesis is the process by which somatic cells form mature muscle tissue through a series of complex regulated processes of proliferation, cell cycle arrest, and differentiation. Numerous signaling pathways help to precisely control these stages1,2. The mitogen-responsive signal then degrades cyclin D1 and its catalytic partner CDK4 to induce exit of the cell from the G1 phase to enter the G0 phase, a quiescent post-mitotic state3,4. Further, myogenin acts with the myocyte enhancer factor 2 (MEF2) transcription factor to regulate expression of the late myogenesis marker genes such as myosin heavy chain (MHC), which is usually expressed after the fusion of myoblasts into myotubes. In particular, MEF2C protein, which was first identified in the nuclei of myotube cells5, is a main regulator of muscle development6,7,8.
Store-operated Ca2+ entry (SOCE), orchestrated by the two key mediators stromal interaction molecule (STIM) and Orai, is involved in numerous cellular processes, including secretion, muscle contraction, cell growth, and muscle development9,10. STIM is a single-transmembrane Ca2+ sensor located in the endoplasmic reticulum (ER)11,12,13, whereas Orai is a plasma membrane-located Ca2+ channel with four transmembrane domains that harbor the pore subunit of SOCE14,15,16. Many studies have demonstrated that STIM1-dependent Ca2+ signaling is crucial for the onset of skeletal muscle development17,18. In addition, STIM2 regulates the intracellular Ca2+ distribution and myogenesis19,20.
SOCE modulates the activity of various Ca2+-dependent enzymes that regulate the myogenesis-related transcription factors, including nuclear factor of activated T cells (NFAT) that plays a regulatory role in skeletal muscle development21. In vivo, mice lacking NFAT4 showed impaired muscle development during embryogenesis22. In vitro, a calcineurin/NFAT4 pathway was found to regulate myogenin induction, and efficient induction and progression of myogenesis required the positive feedback between STIM1 and NFAT423,24.
There are many potential mechanisms by which SOCE is regulated, such as internalization and phosphorylation, and alternative splicing. The recently identified spliced isoform of STIM2, STIM2β (also known as STIM2.1), is highly conserved and acts as a potent inhibitor of SOCE25,26. STIM2β contains an additional eight amino acids in CRAC activation domains (CADs; also known as the STIM1 Orai1 activation region [SOAR] or CCb9)27,28,29,30 and forms a heterodimer with other STIM isoforms to inhibit Ca2+ release-activated Ca2+ (CRAC) channels through an allosteric mechanism31. Through this alternative splicing of STIM2, the cells can effectively control the Ca2+ homeostasis by preventing Orai1 channel cross-linking32 and regulate the balance between SOCE activators and inhibitors. However, the physiological role and detailed underlying mechanism of STIM2β remain unclear.
Accordingly, in this study we explored the detailed function of STIM2β in myogenesis using C2C12 cells with STIM2β knockdown or knockout approaches. We explored the effects of STIM2β loss or deficiency on the key regulators of myogenesis described above, including the NFAT4 and MEF2C signaling pathway, along with its effects on Ca2+ homeostasis and cell cycle arrest via analysis of changes in the expression of cyclin D1 and CDK4. Our subsequent molecular mechanistic study suggests that the STIM2β signaling pathway serves as a switch cell fate between proliferation and skeletal muscle differentiation. These results can provide new insight into the molecular interactions and roles of splicing events in the regulation of skeletal development with potential implications for regenerative medicine.
Results
Splicing form of STIM2β increased during myogenesis
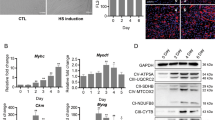
We first investigated the mRNA expression levels of STIM2α and STIM2β during C2C12 myoblast cells differentiation. C2C12 myoblasts were cultured in a differentiation medium and the mRNA expression level was assessed daily using quantitative PCR analysis. The STIM2β expression level showed an approximately 5-fold increase after induction of differentiation (Fig. 1A–C)26, whereas the mRNA levels of STIM2α and other SOCE components showed no dramatic change during myogenesis (Figs 1A and S1A). Therefore, we further explored the function of STIM2β during myogenesis.
STIM2 splicing affects myogenesis. (A) Expression of STIM2 splicing variants during the differentiation of C2C12 myoblast cells (B) GAPDH-normalized expression levels of STIM2β during C2C12 myoblasts differentiation. (C) Histogram of the STIM2α/STIM2β ratio during C2C12 differentiation. (D) Polymerase chain reaction analysis of cDNA from Control (sh-Scr) and STIM2β knock-down (sh-STIM2β) cells 2 day after inducing differentiation. (E–H) GAPDH-normalized expression levels of STIM2α (E), STIM2β (F), myogenin (G) and MHC (H). (I,J) The fraction of myogenin positive cells from control cells (J, black bar) and STIM2β knock-down cells (J, white bar). (n > 80 cells for each group). (K,L) The fraction of MHC positive cells from control cells (L, black bar) and STIM2β knock-down cells (L, white bar). (n > 80 cells for each group). Error bars show means ± SEM. (All scale bars, 20 μm.) The results are representative of at least three independent experiments.
Knockdown of STIM2β delayed myogenesis
To identify the function of STIM2β in C2C12 myogenesis, we knocked down STIM2β using specific shRNA in C2C12 myoblasts. The specificity of the sh-STIM2β expression construct was confirmed, which effectively knocked down only STIM2β expression (>40%) but not STIM2α and other SOCE components expression in cells during differentiation (Figs 1D–F and S1A,B). Interestingly, STIM2β knockdown also caused significant suppression of the mRNA expression levels of the myogenesis marker genes myogenin and MHC by 40% and 50%, respectively, compared with control cells expressing sh-scramble after inducing differentiation for two days (Fig. 1D–H).
To further validate the function of STIM2β in myogenesis, we investigated the population of myogenin- and MHC-positive cells at day 2 after inducing differentiation following transfection of sh-Scr or sh-STIM2β. The C2C12 cells transiently expressing sh-STIM2β showed about 50% of reduced myogenesis compared to sh-Scr treated cells at differentiation day 2 (Fig. 1I–L). These results indicated that transient knockdown of STIM2β suppressed the expression of myogenic genes during C2C12 myogenesis.
Collectively, these results demonstrated that the splicing of STIM2 dramatically changed during myogenesis and that STIM2β regulates the expression of myogenic genes. However, the sh-RNA knockdown approach had a short lifespan to checked longer time period (Fig. S1C). Given the limitation of this approach for further exploration of C2C12 myogenesis, we further established STIM2β knockout cells using CRISPR-Cas9 technology.
STIM2β knockout cell line generated by CRISPR-Cas9
To generate the STIM2β knockout C2C12 myoblast cell line using the CRISPR-Cas9 gene editing system, we first designed sgRNA targeting the exon 9 region, which only includes STIM2β. After 4-day treatment of hygromycin for selecting the genome-edited cells, we observed an evident decrease in the length of the PCR product, and confirmed the 226-nucleotide deletion around the exon 9 region by sequencing (Fig. S1A). To test whether this CRISPR/Cas9-mediated deletion successfully knocked out the exon9 region which included only in STIM2β, we evaluated the expression level of the STIM2 splicing variants STIM2α, STIM2β and other SOCE components after inducing myogenesis. The C2C12 myoblasts showed up-regulation of STIM2β expression during myogenesis, whereas the expression of STIM2β was completely abolished in the CRISPR-Cas9-mediated exon 9-deleted cells. In contrast, the expression level of STIM2α and other SOCE components did not show a significant difference between wild-type and STIM2β-knockout cells (Fig. S1D–G), indicating that STIM2β-knockout cells were successfully generated by the CRISPR-Cas9 system targeting exon 9.
STIM2β knockout cells showed delayed expression of myogenic genes
Given that the results described above strongly implicated a role of STIM2β in C2C12 myogenesis, we next measured the expression of myogenic marker genes in both wild-type and STIM2β knockout cells after inducing myogenesis. The GAPDH-normalized mRNA expression levels of both myogenin and MHC were significantly decreased in STIM2β knockout cells compared to those of wild-type cells (Fig. 2A–C). Although the relative expression level of myosin increased during the 2-day differentiation period, reaching a level approximately one-fifth that of wild-type C2C12 myoblast cells (Fig. 2A,B), there was no such increase in MHC mRNA expression levels after differentiation induction, while the expression level in wild-type cells was significantly increased during differentiation (Fig. 2A,C).
STIM2β knockout delayed C2C12 myoblast differentiation. (A) Comparative myogenin and MHC mRNA expression by RT-PCR between wild-type and STIM2β knockout cells. (B,C) GAPDH-normalized expression levels of myogenin (B) and MHC (C) of wild-type cells (Black circle) and STIM2β knockout cells (White circle) during myogenesis. (D) Immunoblot analysis of wild-type cells and STIM2β knockout cells to measure the protein expression of myogenin and MHC during myogenesis. (E,F) Actin-normalized expression levels of Myogenin (E) and MHC (F) of wild-type cells (Black circle) and STIM2β knockout cells (White circle). (G) Immunofluorescence staining of myogenin protein during the differentiation of wild-type cells (top) and STIM2β knockout cells (bottom). (H) Myogenin positive nuclei population of wild-type cells (Black circle) and STIM2β knockout cells (White circle). (n > 80 cells for each group) (I) Immunofluorescence staining of MHC protein during the differentiation of wild-type cells (Top) and STIM2β knockout cells (Bottom). (J) The fraction of MHC positive cells of wild-type cell (Black circle) and STIM2β knockout cells (White circle). (n > 80 cells for each group) (K–N) Immunofluorescence staining of myogenin (K,L) and MHC (M,N) protein during the differentiation of wild-type cells (Top, black bar), STIM2β knockout cells (Middle, White bar) and transiently STIM2β expressed STIM2β knockout cells (Bottom, gray bar). (n > 80 cells for each group). Error bars show means ± SEM. (All scale bars: 20 μm.) The results are representative of at least three independent experiments.
Western blot analysis confirmed these results at the protein level, in which STIM2β knockout cells showed significantly decreased protein expression levels of myogenesis marker genes. Myogenin was detected in the lysate of wild-type C2C12 myoblast cells as of 1 day after inducing myogenesis, but was only weakly detected in the lysate of STIM2β knockout cells with no tendency to increase during differentiation progression (Fig. 2D,E). Moreover, the MHC protein expression was strongly inhibited in STIM2β knockout cells, and was only weakly detected as of day 3 of differentiation, whereas MHC protein was clearly expressed as of 1 day after inducing myogenesis in wild-type cells and increased with the progress of differentiation (Fig. 2D,F). These results indicated that the expression of key myogenic genes was delayed in STIM2β knockout cells.
To further validate the delayed myogenesis of STIM2β knockout cells, we investigated the fraction of myogenic gene-positive nuclei among wild-type and STIM2β knockout cells. Immunofluorescence staining data showed that both myogenin (Fig. 2G,H) and MHC (Fig. 2I,J)-positive cells frequencies were decreased by about half in STIM2β knockout cells compared with those of the wild-type cells. These results further support the important role of STIM2β in regulating the expression of myogenic genes.
Moreover, after transiently expressing STIM2β in STIM2β knockout cells, immunofluorescence staining at 3 days after inducing differentiation showed that the fractions of myogenin-positive (Fig. 2K,L) cells and MHC-positive cells increased markedly (Fig. 2L,M). This observation indicated that the delayed expression of myogenic genes of STIM2β knockout cells was derived from the interference of STIM2β signaling.
STIM2β plays a role in NFAT4 and MEF2C-mediated myogenesis
We next compared the well-known Ca2+-related myogenesis regulator genes, NFAT4 and MEF2C, between wild-type and STIM2β knockout cells during myogenesis. First, to test whether STIM2β knockout could induce a downstream Ca2+ signaling, we introduced GFP-NFAT4 into wild-type and STIM2β knockout cells. Before inducing differentiation, wild-type and STIM2 knockout cells showed similar nuclear translocation of GFP-NFAT4 (Fig. S2). Unexpectedly, however, the mRNA expression level of NFAT4 rapidly decreased after inducing differentiation in STIM2β knockout cells but gradually increased in wild-type cells (Fig. 3A,B).
STIM2β knockout inhibited NFAT4 and MEF2c. (A) Comparative NFAT4 and MEF2c mRNA expression by RT-PCR between wild-type and STIM2β knockout cells. (B,C) GAPDH-normalized expression levels of NFAT4 (B) and MEF2c (C) of wild-type cells (Black circle) and STIM2β knockout cells (White circle) during myogenesis. (D) Immunoblot analysis of wild-type cells and STIM2β knockout cells to measure the protein expression of MEF2C during myogenesis. (E) Actin-normalized expression levels of MEF2C in wild-type cells (Black circle) and STIM2β knockout cells (White circle). (F) The fraction of MEF2c positive nuclei of wild-type cell (Black circle) and STIM2β over-expressed cells (White rectangular). (n > 60 cells for each group) (G) The fraction of MEF2c positive nuclei of wild-type cell (Black circle), STIM2β knockout cells (White circle) and transiently STIM2β expressed STIM2β knockout cells (Black rectangular). (n > 60 cells for each group). Error bars show means ± SEM. The results are representative of at least three independent experiments.
Notably, the mRNA expression of MEF2C in STIM2β knockout cells failed to increase even after inducing differentiation (Fig. 3A,C). We performed Western blot experiment of MEF2C. As expected, the expression pattern of MEF2C protein was different in both cells after differentiation. The wild type cells showed an increased expression level of MEF2C. However, the protein expression level of MEF2C was not changed in the STIM2β knockout cells after differentiation. It is consistent with the result of no change in the mRNA expression level of MEF2C in Stim2β knockout cells (Fig. 3D,E). Furthermore, 3 days after inducing differentiation, MEF2C-positive nuclei increased from 30% to 45% when transiently overexpressed STIM2β (Fig. 3F), and the significant decrease in MEF2C-positive cells from STIM2β knockout was rescued by transient expression of STIM2β (Fig. 3G). Taken together, these lines of evidence collectively suggest that STIM2β positively regulates myogenesis by activating the MEF2C and NFAT4 signaling pathway.
STIM2β regulates myoblast proliferation via Ca2+ homeostasis
Since myogenesis is accompanied by decreased cell proliferation, the transition from proliferation to differentiation is an important and irreversible checkpoint of myogenesis33,34. Therefore, we investigated the effects of STIM2β knockout on the proliferation of C2C12 myoblasts using an MTT assay. After culture in growth medium for 3 days, the growth of the wild-type cells increased by 4-fold based on the change in the MTT-based OD value, while the STIM2β knockout cells increased by 8-fold (Fig. 4A). To confirm this result, equal numbers of cells were seeded on the growth medium and counted 72 h later. Consistently, STIM2β knockout cells showed 3-fold higher cell numbers than those in wild-type cells (Fig. 4B), confirming that STIM2β knockout promotes the proliferation of C2C12 myoblasts.
STIM2β knockout promoted proliferation via Ca2+ homeostasis in C2C12 myoblast cells. (A) MTT Proliferation assay of wild-type cells (Black circle) and STIM2β knockout cells (White circle). (n = 4 respective experiment for each group) (B) Histogram of cell population of wild-type cells (Black bar) and STIM2β knockout cells (White bar). (C) Fura-2 Ca2+ measurements in 2 mM Ca2+ Tyrode’s solution. wild-type (black), STIM2β knockout cells (blue) and transiently STIM2β expressed in STIM2β knockout cells (green). (D) Histogram of basal calcium level of wild-type (black), STIM2β knockout cells (white) and transiently STIM2β expressed in STIM2β knockout cells (gray). (n > 40 cells for each group) (E) Fura-2 Ca2+ measurements in Tyrode’s solution. wild-type (black), STIM2β knockout cells (blue) in 2 mM extracellular Ca2+ Tyrode’ solution and wild-type (yellow), STIM2β knockout cells (green) in 10 mM extracellular Ca2+ Tyrode’ solution (F) Histogram of basal calcium level of wild-type (black), STIM2β knockout cells (white). (n > 30 cells for each group) (G,H) (Edu staining of C2C12 wild-type cells (G) and STIM2β knockout cells (H) in 2 mM and 10 mM Ca2+ medium. (I) Histogram of Edu positive nuclei of wildtype cells and STIM2β knockout cells cultured in 2 mM or 10 mM Ca2+ medium. (n > 80 cells for each group) Error bars show means ± SEM. (All scale bars: 20 μm.) The results are representative of at least three independent experiments.
We next asked whether the increased proliferation of STIM2β knockout cells occurred because of abnormal intracellular Ca2+ homeostasis in STIM2β knockout cells. We examined STIM2β-dependent Ca2+ homeostasis by measuring the basal intracellular Ca2+ level according to the ratio of the intensity of the fluorescent dye Fura2-AM detected under plain media and growth media. The results showed that a significant increase in the cytosolic Ca2+ level in STIM2β knockout cells, which was then reduced with transient expression of the STIM2β gene in the knockout cells. These results implied that STIM2 regulates basal Ca2+ homeostasis of C2C12 myoblast cells (Figs 4C,D and S3A,B).
Furthermore, we performed EdU staining in standard Ca2+ concentration (2 mM) and high Ca2+ concentration (10 mM) growth media to clarify the relationship between high basal calcium level and increased proliferation rate of STIM2β knockout cells. Interestingly, both wild-type and STIM2β knockout cells had a significantly increased basal Ca2+ level and a fraction of EdU positive cells when grown in high Ca2+ concentration (10 mM) compared to that under a standard Ca2+ concentration (2 mM). Moreover, the STIM2β knockout cells showed a tendency to be more sensitive to the extra cellular Ca2+ concentration (Figs 4E–I and S3C,D). Hence, the increase in the intracellular Ca2+ concentration in the absence of STIM2β likely contributed to the increased proliferation of STIM2β knockout cells.
STIM2β is involved in cyclin D1–CDK4-mediated cell-cycle arrest
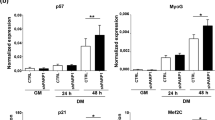
To gain insight into the regulatory mechanism of STIM2β in the proliferation and differentiation process, we investigated the S-phase cells population based on EdU staining patterns. Wild-type cells rapidly withdrew from the cell cycle after myogenesis was induced (Fig. 5A–E), whereas this did not occur in STIM2β knockout cells, rapidly leading to a significantly large S-phase population at 12 and 24 h after inducing myogenesis (Fig. 5B–E). These results suggest that STIM2β regulates cell-cycle arrest during myogenesis.
STIM2β knockout inhibited CyclinD1-CDK4 mediated cell cycle arrest. (A,B) Edu staining of C2C12 wild-type and STIM2β knockout cells. (C–E) Histogram of Edu positive nuclei of wild-type and STIM2β knockout cells in GM (C), 12 h in DM (D), and 24 h in DM (E). (n > 80 cells for each group) (F–H) Immunoblot analysis of wild-type cells and STIM2β knockout cells to measure the protein expression of CyclinD1 and CDK4 during myogenesis. Actin-normalized expression levels of CyclinD1 (G) and CDK4 (H) of wild-type cells (Black circle) and STIM2β knockout cells (White circle). (I,J) Immunofluorescence analysis of cyclin D1 expression in GM and DM 24 h (I) and relative quantification reporting the cyclin D1-positive nuclei percentage (J). (n > 130 cells for each group) Error bars show means ± SEM. (All scale bars: 20 μm.) The results are representative of at least three independent experiments.
Therefore, we next examined the expression of cell-cycle regulatory genes that control arrest (i.e., the G1–G0 transition). The mRNA expression levels of cyclin family genes showed similar patterns between wild-type and STIM2β knockout cells (Fig. S4). However, interestingly, when we checked the protein expression level of cyclin D1 and CDK4 that regulate cell-cycle arrest during myogenesis, wild-type cells showed decreased protein levels of cyclin D1 and CDK4 during myogenesis, while STIM2β knockout cells showed a 4–5 fold higher expression of cyclin D1 and 3–5 fold of CDK4 protein compared to wild-type cells (Fig. 5F–H), indicating that cell cycle arrest induced myogenesis might be delayed due to the altered the protein stability of cyclin D1. However, further study is needed how the protein stability of cyclin D1 is altered in STIM2β knockout cells. To further interrogate whether the stable protein expression of cyclin D1 in STIM2β knockout cells could induce the transcriptional activity of cyclin D1, we measured cyclin D1 positive nuclei portion. After inducing 1 day differentiation, only 25% of wild-type cells showed nuclear location of cyclin D1, compared with 97% of STIM2β knockout cells (Fig. 5I,J).
Taken together, these results implied that the STIM2β is required for the degradation of cyclinD1/CDK4 and cyclinD1/CDK4-mediated cell cycle arrest during skeletal muscle differentiation. Also, STIM2β dependent Ca2+ homeostasis regulates proliferation of myoblast cells, indicating that increased expression of STIM2β during myogenesis regulates transition of C2C12 cell-state from proliferation to differentiation.
Discussion
STIM2β is considered to be a unique inhibitor of SOCE among all known STIM isoforms. A small increase of STIM2β can inhibit Orai function by forming heterodimer with STIM. Despite the fact that this feature was predicted to have an important effect on STIM2β during the development process, the exact underlying molecular mechanism is still unclear. In the present study, we found that the STIM2β signaling pathway plays a critical role in myogenesis. Our results demonstrate that STIM2β mRNA expression level was dramatically increased during early myogenesis. Knockout of STIM2β in C2C12 myoblast cells using the CRISPR/Cas9 technique show delayed myogenesis and increased proliferation process. Most importantly, we found that STIM2β regulates myogenesis by activating the MEF2C and NFAT4 signaling pathway, and promotes cell cycle arrest by decreasing cyclin D1 and its catalytic partner CDK4. Further, STIM2β reached the highest mRNA expression level at 6 days after inducing differentiation. Also, STIM2β knockout cells exhibited delayed myogenesis even at 7 days after inducing differentiation and showed impaired multinucleated muscle fiber formation, indicating that STIM2β might be required for not only early myogenesis but also later myogenesis (Fig. S5).
The effect of Ca2+ signaling on cell proliferation has been well documented35. Particularly, SOCE is the most extensively studied Ca2+ ignaling involved in cell proliferation. NFAT and CaM stimulation by SOCE is involved in the proliferation of various cell types and G1/S transition36,37,38,39,40,41. A previous study suggested that SOCE was relevant only in the first 12 h after the onset of C2C12 myogenesis and became less important in subsequent stages of the transition to mature muscle tissue differentiation17. These findings agree with those of other studies suggesting that myogenesis is accompanied by a transition between the proliferation and differentiation processes33,34. However, other studies have revealed that STIM and Orai are both strongly expressed in mature skeletal muscle cells and remain fully functional42,43,44. In this discrepancy, our results suggested that STIM2β is involved in SOCE through Orai1 channel during myogenesis (Fig. S6A). And confirmed by puncta formation of STIM2β with STIM1 and weakly recruited by Orai1 (Fig. S6 C). Also, STIM2β knockout cells showed increased SOCE amplitude and sustained SOCE peak compared with wild-type cells even after inducing differentiation (Fig. S6A,B). These results suggest that myoblast cells efficiently regulate SOCE via STIM2 alternative splicing during myogenesis.
In this study, we found that STIM2β is essential for intracellular Ca2+ signaling in myogenesis. This finding is inconsistent with a previous study suggesting that STIM2β did not have a significant effect on basal [Ca2+]i25. This discrepancy might suggest that around 40% expression level of STIM2β in naïve CD4 + T cells enough to regulate the basal intracellular Ca2+. Consistently, our results showed low expression of STIM2β in C2C12 myoblasts was still enough to regulate the basal intracellular Ca2+ level. Interestingly, the proliferation of C2C12 myoblasts was susceptible to changes in Ca2+ levels, and this phenomenon was more pronounced in STIM2β knockout cells. Moreover, the increased basal Ca2+ level of STIM2β knockout cells and requirement for STIM2ββ-dependent Ca2+ signaling for fine-tuning the proliferation of C2C12 myoblasts suggests that STIM2β regulates basal Ca2+ homeostasis in C2C12 myoblast cells. In this study, we suggest that STIM2β appeared to function as the key regulator of Ca2+ homeostasis in C2C12 myoblasts, and increased STIM2β expression induced a transition from proliferation to differentiation during myogenesis.
Given its high evolutionary conservation and widespread expression, the importance of STIM2β in various physiological functions is increasingly becoming recognized26. Moreover, the generation of STIM2β through alternative splicing is a particularly useful mechanism for modulating SOCE. Thus, the present insight of the role of STIM2β in myogenesis further highlights alternative splicing as an effective way to modulate the SOCE during development. Based on our findings, further investigation will be necessary to clarify the importance of STIM2β not only in skeletal muscle development but also in the developmental processes of other tissues and other physiological functions.
In summary, the present study demonstrates a previously unrecognized mechanism by which the STIM2β signaling pathway controls myogenesis. Under normal skeletal muscle development condition, the STIM2β positively regulates myogenesis by controlling cell-cycle arrest through regulating the degradation of cyclin D1-CDK4. Increased STIM2β will induce inhibition of SOCE during myogenesis to block its action on cell proliferation, which efficiently controls the transition of cell fate from proliferation to differentiation. At the same time, We further highlight that STIM2β activates myogenic factors through the MEF2C and NFAT4 pathway. Our findings contribute important insight toward gaining a better understanding of how myoblasts effectively regulate various signaling pathways during myogenesis processes through the alternative splicing of STIM2.
Material and Methods
Cell culture and transfection
C2C12 cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) at 37 °C in 5% CO2. For the muscle cell differentiation, C2C12 myoblast cells were cultured to 70–80% confluency and then change cultured media to DMEM containing 2% horse serum. We changed the medium every day. For transient transfection, the cells were transfected at 70% confluency with 0.3–3 μg DNA using Jet Prime (PolyPlus) according to the manufacturer’s instructions.
Intracellular Ca2+ imaging
Cells were loaded with 5 μM Fura-2/AM in DMEM at 37 °C for 30 min. Ratiometric Ca2+ imaging was performed at 340 and 380 nm in 2 mM Ca2+ Tyrode’s solution (129 mM NaCl, 5 mM KCl, 2 mM CaCl2, 1 mM MgCl2, 30 mM glucose, and 25 mM Hepes, pH 7.4) or cultued medium with a IDX81 microscope (Olympus) equipped with an Olympus x40 oil (NA 1.30) objective equipped with a fluorescent arc lamp (LAMDA LS), excitation filter wheel (SUTTER, LAMBDA 10–2), stage controller (ASI, MS-2000) and a CCD camera (HAMAMATSU, C10600) at room temperature. Images were processed with Metamorph and analyzed with Igor Pro.
Reagents
Fura-2/AM was obtained from Invitrogen. Jet Prime was from Poly Plus. Thapsigargin (TG) was from Santa Cruz. Antibodies of myogenin(F5D, DSHB), MHC(MF-20, DSHB), CyclinD1(A-12, Santa Cruz), CDK4(B-10, Santa Cruz), β-actin(C4, Santa Cruz), MEF2C(MAB6786, R&D Systems) were purchased from the indicated vendor.
mRNA expression level check
Total RNA was extracted from cells using RiboEx (GeneAll) following the manufacturer’s protocol. cDNA was made from 2~3 μg of RNA reverse transcribed using oligo (dT) primers and First Strand cDNA Synthesis Kit (TOYOBO). The PCR amplification was done using a C1000 Touch thermal Cycler (Bio-Rad). Amplification started with initial denaturation at 95 °C for 3 min and then 30–40 cycles of denaturation at 95 °C for 30 sec, annealing at 55 °C for 30 sec, and extension at 72 °C for 1 min. Gel electrophoresis was used to identify the PCR products in a 1.5% agarose gel using ethidium bromide staining. For real-time quantitative PCR analysis, synthesized cDNA and SYBR green Master Mix (Roche) were run on a LightCycler480 II (Roche). Relative expression levels of mRNA were calculated using the 2 − Ct method45.
Western blotting
Cell lysates in lysis buffer (100 mM Tris-HCl, pH 8, 150 mM NaCl, 1% Triton X-100) were subjected to SDS-PAGE and electro-transferred onto PVDF membranes. The PVDF membrane was blocked with 7% Skim milk dissolved in Tris-buffered saline containing 0/1% Tween 20 for two h at room temperature with gentle shaking. After blocking, PVDF membrane was probed overnight at 4 °C, with specific primary antibodies in 3% BSA solution. The membrane was incubated for 30 min at RT with a horseradish peroxidase-conjugated anti-mouse or rabbit IgG in TBST. Detection was performed with the enhanced chemiluminescence reagent (ECL). Quantification of bands was performed by using the ImageJ software.
Generation of knockout cell line with CRISPR/Cas9
Guide RNA sequences for mouse STIM2β (5′-CCTGCAGGTTAGTAGTTACTAGA-3′) are inserted in pRGEN vector and pRGEN-reporter (ToolGen). C2C12 cells were transfected with pRGEN-mSTIM2β, pRGEN-Cas9, and pRGEN-reporter using Lipofectamine 2000, according to the manufacture’s instructions. Two days after transfection, cells were selected with 1000 μg/ml of hygromycin for four days. After one week, colonies were an isolated and genomic deletion, and mRNA depletion was analyzed with sequencing and RT-PCR assay.
Immunocytochemistry
C2C12 cells were seed on Collagen coated glass and induce differentiation. Every indicated timepoint, the cells were fixed with 4% paraformaldehyde for 10 min and treated with 0.1% Triton X-100 in PBS for 10 min at room temperature, and then block with 3% BSA in PBS for 2 hr. Following incubation with the indicated antibody over night at 4 °C, cells were incubated with a conjugated secondary antibody at 37 °C for 1 hr. Finally, the nuclei were stained with Hoechest (Molecular Probes) for 5 min.
EdU staining
Cell-cycle analysis was performed by assaying the incorporation of 5-ethynyl-2′-deoxyuridine (EdU) according to the manufacturer’s instruction (Invitrogen). Briefly, cells were cultured in medium containing 1X EdU component for an additional time, as indicated. Following incubation, the cells were rinsed with PBS and fixed with 4% PFA for 15 mins at RT.
MTT assay
C2C12 cells were seeded at a density of 7500 cells/well in 96-well plates. Next day, cells were incubated with medium containing 1 mg/ml of MTT. After 4 hr incubation at 37 °C, The medium was then aspirated and add DMSO and gently shake 15 min for formazan solubilization. The absorbance was measured at a wavelength of 562 nm by using a fluorescence plate reader (Spectra MAX Pro 5, Molecular Devices).
Statistics
All statistical analysis was performed in Prism 6 (GraphPad Software). All error bars represent SEM. All pairwise differences were tested for significance using a two-tailed t test. At least three independent experiments were performed. P-values < 0.05 were considered statistically significant and indicated as following: *P < 0.05; **P < 0.01; ***P < 0.001.
Data Availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files), are available from the corresponding author on reasonable request.
References
Sassoon, D. et al. Expression of two myogenic regulatory factors myogenin and MyoDl during mouse embryogenesis. Nature 341, 303–307, https://doi.org/10.1038/341303a0 (1989).
Wright, W. E., Sassoon, D. A. & Lin, V. K. Myogenin, a factor regulating myogenesis, has a domain homologous to MyoD. Cell 56, 607–617, https://doi.org/10.1016/0092-8674(89)90583-7 (1989).
Guo, K. & Walsh, K. Inhibition of Myogenesis by Multiple Cyclin-Cdk Complexes: Coordinate Regulation Of Myogenesis And Cell Cycle Activity At The Level Of E2f. 272, 791–797, https://doi.org/10.1074/jbc.272.2.791 (1997).
Zhang, J. M., Wei, Q., Zhao, X. & Paterson, B. M. Coupling of the cell cycle and myogenesis through the cyclin D1‐dependent interaction of MyoD with cdk4. 18, 926–933, 10.1093/emboj/18.4.926 The EMBO Journal (1999).
Gossett, L. A., Kelvin, D. J., Sternberg, E. A. & Olson, E. N. A new myocyte-specific enhancer-binding factor that recognizes a conserved element associated with multiple muscle-specific genes. Molecular and Cellular Biology 9, 5022–5033, https://doi.org/10.1128/MCB.9.11.5022 (1989).
Edmondson, D. G., Cheng, T. C., Cserjesi, P., Chakraborty, T. & Olson, E. N. Analysis of the myogenin promoter reveals an indirect pathway for positive autoregulation mediated by the muscle-specific enhancer factor MEF-2. Molecular and Cellular Biology 12, 3665–3677, https://doi.org/10.1128/MCB.12.9.3665 (1992).
Edmondson, D. G., Lyons, G. E., Martin, J. F. & Olson, E. N. Mef2 gene expression marks the cardiac and skeletal muscle lineages during mouse embryogenesis. 120, 1251–1263 (1994).
Molkentin, J. D., Black, B. L., Martin, J. F. & Olson, E. N. Mutational analysis of the DNA binding, dimerization, and transcriptional activation domains of MEF2C. Molecular and Cellular Biology 16, 2627–2636, https://doi.org/10.1128/MCB.16.6.2627 (1996).
Clapham, D. E. Calcium Signaling. Cell 131, 1047–1058, https://doi.org/10.1016/j.cell.2007.11.028 (2007).
Li, T. et al. STIM1-Ca2+ Signaling Is Required for the Hypertrophic Growth of Skeletal Muscle in Mice. Molecular and Cellular Biology 32, 3009–3017, https://doi.org/10.1128/MCB.06599-11 (2012).
Liou, J. et al. STIM Is a Ca2+ Sensor Essential for Ca2+-Store-Depletion-Triggered Ca2+ Influx. Current Biology 15, 1235–1241, https://doi.org/10.1016/j.cub.2005.05.055 (2005).
Roos, J. et al. STIM1, an essential and conserved component of store-operated Ca2+ channel function. The Journal of Cell Biology 169, 435–445, https://doi.org/10.1083/jcb.200502019 (2005).
Zhang, S. L. et al. STIM1 is a Ca2+ sensor that activates CRAC channels and migrates from the Ca2+ store to the plasma membrane. Nature 437, 902, https://doi.org/10.1038/nature04147 (2005).
Prakriya, M. et al. Orai1 is an essential pore subunit of the CRAC channel. Nature 443, 230, https://doi.org/10.1038/nature05122 (2006).
Vig, M. et al. CRACM1 Multimers Form the Ion-Selective Pore of the CRAC Channel. Current Biology 16, 2073–2079, https://doi.org/10.1016/j.cub.2006.08.085 (2006).
Yeromin, A. V. et al. Molecular identification of the CRAC channel by altered ion selectivity in a mutant of Orai. Nature 443, 226, https://doi.org/10.1038/nature05108 (2006).
Darbellay, B. et al. STIM1- and Orai1-dependent Store-operated Calcium Entry Regulates Human Myoblast. Differentiation. 284, 5370–5380, https://doi.org/10.1074/jbc.M806726200 (2009).
Darbellay, B. et al. Human Muscle Economy Myoblast Differentiation and Excitation-Contraction Coupling Use the Same Molecular Partners. STIM1 and STIM2. 285, 22437–22447, https://doi.org/10.1074/jbc.M110.118984 (2010).
Phuong, T. T. T. & Kang, T. M. Stromal interaction molecule 2 regulates C2C12 myoblast differentiation. Integrative Medicine. Research 4, 242–248, https://doi.org/10.1016/j.imr.2015.09.001 (2015).
Oh, M. R. et al. STIM2 regulates both intracellular Ca2+ distribution and Ca2+ movement in skeletal myotubes. Scientific Reports 7, 17936, https://doi.org/10.1038/s41598-017-18256-3 (2017).
Armand, A.-S. et al. Cooperative Synergy between NFAT and MyoD Regulates Myogenin Expression and Myogenesis. 283, 29004–29010, https://doi.org/10.1074/jbc.M801297200 (2008).
Kegley, K. M., Gephart, J., Warren, G. L. & Pavlath, G. K. Altered Primary Myogenesis in NFATC3−/− Mice Leads to Decreased Muscle Size in the Adult. Developmental Biology 232, 115–126, https://doi.org/10.1006/dbio.2001.0179 (2001).
Delling, U. et al. A Calcineurin-NFATc3-Dependent Pathway Regulates Skeletal Muscle Differentiation and Slow Myosin Heavy-Chain Expression. Molecular and Cellular Biology 20, 6600–6611, https://doi.org/10.1128/MCB.20.17.6600-6611.2000 (2000).
Phuong, T. T. T., Yun, Y.-H., Kim, S. J. & Kang, T. M. Positive feedback control between STIM1 and NFATc3 is required for C2C12 myoblast differentiation. Biochemical and Biophysical Research Communications 430, 722–728, https://doi.org/10.1016/j.bbrc.2012.11.082 (2013).
Miederer, A.-M. et al. A STIM2 splice variant negatively regulates store-operated calcium entry. Nature Communications 6, 6899, https://doi.org/10.1038/ncomms7899 (2015).
Rana, A. et al. Alternative splicing converts STIM2 from an activator to an inhibitor of store-operated calcium channels. The Journal of Cell Biology 209, 653–670, https://doi.org/10.1083/jcb.201412060 (2015).
Kawasaki, T., Lange, I. & Feske, S. A minimal regulatory domain in the C terminus of STIM1 binds to and activates ORAI1 CRAC channels. Biochemical and Biophysical Research Communications 385, 49–54, https://doi.org/10.1016/j.bbrc.2009.05.020 (2009).
Muik, M. et al. A Cytosolic Homomerization and a Modulatory Domain within STIM1 C Terminus Determine Coupling to ORAI1 Channels. 284, 8421–8426, https://doi.org/10.1074/jbc.C800229200 (2009).
Park, C. Y. et al. STIM1 Clusters and Activates CRAC Channels via Direct Binding of a Cytosolic Domain to Orai1. Cell 136, 876–890, https://doi.org/10.1016/j.cell.2009.02.014 (2009).
Yuan, J. P. et al. SOAR and the polybasic STIM1 domains gate and regulate Orai channels. Nature Cell Biology 11, 337, https://doi.org/10.1038/ncb1842 (2009).
Chung, S., Zhang, M. & Stathopulos, P. B. The 2β Splice Variation Alters the Structure and Function of the Stromal Interaction Molecule Coiled-Coil Domains. 19, 3316 (2018).
Zhou, Y. et al. Cross-linking of Orai1 channels by STIM proteins. Proceedings of the National Academy of Sciences 115, E3398–E3407, https://doi.org/10.1073/pnas.1720810115 (2018).
Shi, X. & Garry, D. J. Muscle stem cells in development. regeneration, and disease. 20, 1692–1708, https://doi.org/10.1101/gad.1419406 (2006).
Buckingham, M. & Rigby Peter, W. J. Gene Regulatory Networks and Transcriptional Mechanisms that Control Myogenesis. Developmental Cell 28, 225–238, https://doi.org/10.1016/j.devcel.2013.12.020 (2014).
Thierry, C. The Need for Calcium Channels in Cell Proliferation. Recent Patents on Anti-Cancer Drug Discovery 8, 4–17, https://doi.org/10.2174/1574892811308010004 (2013).
Hidaka, H. et al. N-(6-aminohexyl)-5-chloro-1-naphthalenesulfonamide, a calmodulin antagonist, inhibits cell proliferation. Proceedings of the National Academy of Sciences of the United States of America 78, 4354–4357 (1981).
Rasmussen, C. D. & Means, A. R. Calmodulin is required for cell-cycle progression during G1 and mitosis. The EMBO journal 8, 73–82 (1989).
Kahl, C. R. & Means, A. R. Regulation of Cell Cycle Progression by Calcium/Calmodulin-Dependent Pathways. Endocrine Reviews 24, 719–736, https://doi.org/10.1210/er.2003-0008 Endocrine Reviews (2003).
Hu, F. et al. Elevation of Extracellular Ca2+ Induces Store-Operated Calcium Entry via Calcium-Sensing Receptors: A Pathway Contributes to the Proliferation of Osteoblasts. PLOS ONE 9, e107217, https://doi.org/10.1371/journal.pone.0107217 (2014).
Somasundaram, A. et al. Store-Operated CRAC Channels Regulate Gene Expression and Proliferation in Neural Progenitor Cells. The Journal of Neuroscience 34, 9107–9123, https://doi.org/10.1523/JNEUROSCI.0263-14.2014 (2014).
Chen, Y.-W., Chen, Y.-F., Chen, Y.-T., Chiu, W.-T. & Shen, M.-R. The STIM1-Orai1 pathway of store-operated Ca2+ entry controls the checkpoint in cell cycle G1/S transition. Scientific Reports 6, 22142, https://doi.org/10.1038/srep22142 https://www.nature.com/articles/srep22142#supplementary-information (2016).
Stiber, J. et al. STIM1 signalling controls store-operated calcium entry required for development and contractile function in skeletal muscle. Nature Cell Biology 10, 688, https://doi.org/10.1038/ncb1731 (2008).
Edwards, J. N., Blackmore, D. G., Gilbert, D. F., Murphy, R. M. & Launikonis, B. S. Store-operated calcium entry remains fully functional in aged mouse skeletal muscle despite a decline in STIM1 protein expression. 10, 675–685, https://doi.org/10.1111/j.1474-9726.2011.00706.x (2011).
Tu, M. K., Levin, J. B., Hamilton, A. M. & Borodinsky, L. N. Calcium signaling in skeletal muscle development, maintenance and regeneration. Cell Calcium 59, 91–97, https://doi.org/10.1016/j.ceca.2016.02.005 (2016).
Choe, J. H. et al. The relation between glycogen, lactate content and muscle fiber type composition, and their influence on postmortem glycolytic rate and pork quality. Meat Science 80, 355–362, https://doi.org/10.1016/j.meatsci.2007.12.019 (2008).
Acknowledgements
Authors thank S.K.L., Y.C.K., A.R.L., J.S.J., Y.Y.L., K.H.J. and S.G.M. for the reading and comments for clarifying the manuscript. This work was supported by the National Research Foundation of Korea (NRF) Grant funded by the Korean Government (MEST) NRF-2018R1D1A1B07051128, 2018R1A5A1024340, 2019R1A2C2002235 to C.Y.P.
Author information
Authors and Affiliations
Contributions
K.M.K., A.R., C.Y.P. designed and performed the experiments. And K.M.K., A.R., C.Y.P. wrote the manuscript. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, K.M., Rana, A. & Park, C.Y. Orai1 inhibitor STIM2β regulates myogenesis by controlling SOCE dependent transcriptional factors. Sci Rep 9, 10794 (2019). https://doi.org/10.1038/s41598-019-47259-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47259-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.