Abstract
Cyclophilins (CYPs) are a group of ubiquitous prolyl cis/trans isomerases (PPIases). It was shown that plants possess the most diverse CYP families and that these are abundant in the phloem long-distance translocation stream. Since phloem exudate showed PPIase activity, three single-domain CYPs that occur in phloem samples from Brassica napus were characterised on functional and structural levels. It could be shown that they exhibit isomerase activity and that this activity is controlled by a redox regulation mechanism, which has been postulated for divergent CYPs. The structure determination by small-angle X-ray scattering experiments revealed a conserved globular shape. In addition, the high-resolution crystal structure of BnCYP19-1 was resolved and refined to 2.0 Å resolution, and the active sites of related CYPs as well as substrate binding were modelled. The obtained data and results support the hypothesis that single domain phloem CYPs are active phloem PPIases that may function as chaperones.
Similar content being viewed by others
Introduction
Cyclophilins (CYPs) are ubiquitous proteins involved in a number of fundamental cellular functions in a large number of organisms, such as animals, plants, fungi, bacteria and viruses1. Together with the structurally unrelated FK506-binding proteins (FKBPs), they belong to the superfamily of immunophilins that have originally been discovered as proteins binding to the immunosuppressant peptide drug cyclosporin A (CsA)2 or FK506/rapamycin3, respectively. Many proteins of both groups possess a peptidyl-prolyl cis/trans isomerase (PPIase) activity, implemented by the conserved FKBP- or CYP-like domain (CLD). By these isomerases the transition from cis to trans in an X-proline peptide bond, a rate-limiting step in protein folding4,5, is stabilised or accelerated. Furthermore, CYPs may also be involved in signalling6, pathogen response7, RNA processing8,9 gene repression10, as well as plant stress responses and development11,12.
Interestingly, plants possess the most diverse CYP families with rice (Oryza sativa) encoding 2713, Arabidopsis thaliana encoding 2914, soybean (Glycine max) encoding 6215, and oilseed rape (Brassica napus) encoding 9116 distinct CYP proteins. A. thaliana and B. napus, both belonging to the family of Brassicaceae, are of great importance as model organisms and in the case of B. napus also in agriculture. As has already been shown for various plant species, CYPs are abundant proteins in the phloem long-distance transport stream and it is assumed that they support protein refolding after trafficking into sieve elements17,18,19,20,21. With only few exceptions, functions of phloem CYPs are so far unknown. CYP1 from tomato (SlCYP1), however, has been suggested to be involved in long-distance signalling modulating auxin responses22.
Twenty distinct CYPs have been identified in the phloem of B. napus and all of them belong to the family of single-domain CYPs16. They are composed of the CLD with a common structure motif of an eight anti-parallel stranded right-handed β-barrel with two α-helices at the top and bottom23. Investigation of the most widely studied CYP, human CYPA (also known as hCYPA or HsCYPA), led to the identification of its CsA binding site24. Since the first structure of HsCYPA has been determined, four CYP structures from plants have been resolved (summarised in25). In contrast to the investigated CYPs from Citrus sinensis (CsCYP)26, Triticum aestivum (TaCYPA-1)27, and Catharanthus roseus (Cat r 1)28, which all constitute single-domain variants, A. thaliana AtCYP38 is a multi-domain protein consisting of the CLD plus a PsbQ-like helical bundle29. Yet, none of these proteins was assigned to the phloem. Since the structure of the tomato phloem CYP SlCYP1 has only been modelled22, experimental validation of a phloem mobile CYP structure is still missing.
The identification of CYPs in the phloem of B. napus under standard growth conditions supports the assumption that these proteins fulfil essential functions and may act as chaperones. In this context, the first question arising is whether CYPs can exercise their isomerase activity in the phloem. Therefore, we studied not only the PPIase activity of B. napus phloem exudate, but also of individual CYPs. The investigated candidate proteins BnCYP18-4, BnCYP18-5, and BnCYP19-1 were chosen because of their homology to already examined plant CYPs, either known to be phloem localised or from the close relative A. thaliana. Concerning AtCYPs, we included AtCYP19-3 into the study, which has not only been previously analysed in activity assays and by in silico modelling30, but also resembles the closest A. thaliana homolog to one of the investigated B. napus CYPs, BnCYP19-1. Small-angle X-ray scattering (SAXS) experiments of all four selected CYPs were performed to verify and compare their overall structure in solution. In addition, the high resolution structure of one phloem CYP, BnCYP19-1, was determined by X-ray crystallography. These data were further utilised to model active site residues of the other CYPs. The results show that the small specific activity differences observed cannot be explained by the conformation of the catalytic and CsA-binding residues alone.
Results and Discussion
B. napus phloem exudate has peptidyl-prolyl cis/trans isomerase activity
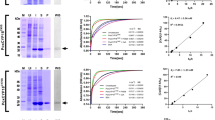
To support the hypothesis of CYPs being active PPIases in the phloem, the activity of freshly sampled phloem exudate was measured. A common assay to assess the isomerisation rate of PPIases has been first described by Fischer et al. in31. Mostly purified or recombinantly expressed proteins were investigated by this method, but it has also been applied to protein mixtures. In an attempt to answer the question whether PPIases are active in phloem exudate of Brassicaceae, we sampled B. napus phloem sap and added it directly to the assay mixture, what resulted in an enhanced isomerisation reaction (Fig. 1a). The observed rate constants showed a linear increase correlated with increasing amounts of phloem exudate (Fig. 1b). It is assumed that this activity results from a mixture of active CYPs, since 20 distinct CYPs have been identified in the phloem16. After the addition of CsA, a well-known cyclophilin inhibitor, the activity was reduced (Fig. 1c). In contrast, the addition of FK506, a FKBP inhibitor, did not result in any activity changes (Fig. 1d), demonstrating that the activity originates only from CYPs. Similar observations have been described for phloem exudate from Ricinus communis, a member of the family of Euphorbiaceae32.
B. napus phloem exudate has peptidyl-prolyl cis/trans isomerase activity. (a) Increasing amounts of phloem exudate show increasing catalytic activity. (b) The rate constants kobs demonstrate a linear behaviour. (c) The activity can be reduced by the addition of the cyclophilin inhibitor CsA. (d) The FKBP inhibitor FK506 does not influence the isomerisation rate.
Sequence similarities of plant single-domain CYPs
A challenge in studying highly similar proteins is to provide evidence that small differences in their sequence affect activities or structural arrangements. The three closely related BnCYPs examined in this study, BnCYP18-4, BnCYP18-5, and BnCYP19-1, are showing only minor variances in their amino acid composition (Supplementary Fig. S1), resulting in high sequence identities of 74 to 82%. In comparison with single-domain CYPs from other plant species, such as the A. thaliana homolog AtCYP19-3, the structurally resolved citrus CsCYP26 and wheat TaCYPA-126,27, the phloem CYPs SlCYP1 from tomato22, and RcCYP1 from castor bean32, the high sequence conservation even between different species becomes obvious. The closest B. napus homolog of SlCYP1 is BnCYP18-5 with a sequence identity of 83%, and BnCYP18-4 has a sequence identity of 85% with RcCYP1. To compare CYPs from the family of Brassicaceae, AtCYP19-3 has been chosen which shares 91% sequence identity with BnCYP19-1.
All of the amino acid residues that have been experimentally demonstrated to be important for PPIase activity, CsA binding or disulphide bridge formation26,33,34 are conserved in all comparatively analysed CYPs, except for Cys40, which is missing in HsCYPA. Furthermore, in contrast to HsCYPA all surveyed plant CYPs show an insertion which occurs in many plant14, nematode35, and human36 CYPs and is located within amino acid region 48 to 54 (Supplementary Fig. S1). These amino acids are positioned between α-helix-I and β-sheet-III (the so-called α-I/β-III junction14 or divergent loop37) and result in an extended loop region.
Activity of individual phloem CYPs
In order to find out whether the most abundant BnCYPs in the phloem, the 18–19 kDa representatives, contribute to its PPIase activity, we directly compared the catalytic activities of three abundant phloem single-domain CYPs from B. napus, BnCYP18-4 (18.6 kDa), BnCYP18-5 (18.7 kDa), BnCYP19-1 (19.9 kDa), and the A. thaliana AtCYP19-3 (19.2 kDa). The four proteins were expressed in E. coli and purified to homogeneity after cleaving off the His-tag.
While the PPIase assay raw data confirmed the intuitive concept that the more enzyme present, the higher the activity (Figs 2a, S2), plotting kobs-k0 versus the protein concentration showed a linear growth of the rate constant of the catalysed isomerisation reaction (Fig. 2b). The results showed that the surveyed CYPs were active PPIases with catalytic efficiencies in a similar range as reported for other plant CYPs (summarised in30). BnCYP18-4 had a catalytic activity kcat/Km of 9.02 ± 0.26 s−1 µM−1 and is therewith similarly active as BnCYP19-1 with 9.07 ± 0.17 s−1 µM−1. BnCYP18-5 had a slightly lower catalytic activity (5.30 ± 0.04 s−1 µM−1). The activity of AtCYP19-3 has been measured in a previous study with a kcat/Km value of 2.7 s−1 µM−1 30. In our study its activity was higher (4.91 ± 0.05 s−1 µM−1) what might be caused by optimisation of the assay set-up. For comparison, a catalytic activity of 1.4 s−1 µM−1 has been reported for HsCYPA38.
Determination of the enzymatic activity of recombinant Brassica napus CYPs and their inhibition by CsA. (a) Representative normalised raw data of BnCYP18-4 measured in a concentration range of 0.5 to 50 nM protein. (b) Influence of the protein concentration on the rate constant. (c) Representative normalised raw data of BnCYP18-4 inhibition by CsA with inhibitor concentrations of 0 to 1000 nM. (d) Dose-response curves of BnCYP18-4, BnCYP18-5, BnCYP19-1, and AtCYP19-3.
The addition of the inhibitor CsA resulted in reduced isomerisation rates (Fig. 2c) that can be visualised as a dose-response curve (Fig. 2d). All four CYPs bound CsA and concentrations above 500 nM CsA inhibited the reaction completely. The IC50 values of the inhibition kinetics showed that BnCYP18-4 (14.2 ± 2.5 nM), BnCYP18-5 (22.4 ± 2.1 nM), BnCYP19-1 (16.6 ± 2.0 nM), and AtCYP19-3 (20.4 ± 2.5 nM) bound CsA with only small variances in affinity.
Redox regulation of enzyme activity
Since all investigated CYPs belong to the divergent CYPs which were earlier proposed to be controlled by a redox regulated mechanism26 they were additionally subjected to activity assays under oxidative and reducing conditions. While BnCYP18-4, BnCYP18-5, BnCYP19-1, and AtCYP19-3 were highly active after reduction, all enzymes lost their activity after oxidative treatment (Fig. 3a). This is suggested to result from the formation of a disulphide bridge between Cys40 and Cys168, altering the enzyme structure allosterically via the divergent loop26. Supporting this hypothesis, a titration experiment of reducing versus oxidising agent revealed a redox state dependent migration pattern (Fig. 3b). Such changes in the migration behaviour are suggested to result mainly from the formation of disulphide bridges as it could be shown earlier for AtCYP20-334.
Redox regulation of PPIase activity via disulphide bond formation. (a) Reduced protein shows high activity in the PPIase assay, but its oxidation results in drastically reduced isomerisation rates. (b) Titrating reducing (0-50 mM DTT) versus oxidising (10 mM Cu2+) agent leads to an altered migration pattern on a non-reducing SDS-PAGE. Cropped gels are shown for clarity. Full size gels are presented in Supplementary Fig. S3. Oxidation by H2O2 showed similar results.
In solution protein shape
Matching the PPIase assay data, the same four CYPs, BnCYP18-4, BnCYP18-5, BnCYP19-1, and AtCYP19-3 were structurally investigated by SAXS experiments. Beforehand, the samples were analysed by dynamic light scattering (DLS). This demonstrated that all four proteins were monomeric in solution (Supplementary Fig. S4). The hydrodynamic radii (Rh) are provided in Table 1. After this quality assessment monodisperse sample solutions were subjected to SAXS in order to compare the radii of gyration (Rg) and maximum diameters (Dmax) (Table 1), as well as to model the tertiary structure.
The SAXS data (Fig. 4a) strongly indicated a conserved nearly globular shape, which was also predicted by ab initio modelling (Fig. 4c). The CYP domain is consisting of a compact globular structure with an “opened” cleft, resembling the binding site of substrates and ligands like CsA. The dimensionless Kratky plots of all four CYPs are similar with a slight shift along the vertical axis for BnCYP19-1 and AtCYP19-3 at higher scattering angles (Fig. 4b). This observation might be related to elongated, flexible termini of the slightly larger CYPs. Supporting this observation, the ab initio models also strongly indicate a flexible C-terminus for all four CYPs. This flexibility is in agreement with the higher χ2 value of the BnCYP19-1 and AtCYP19-3 CRYSOL fit curves compared to these of BnCYP18-4 and BnCYP18-5. The addition of CsA resulted in no significant changes of the overall three-dimensional shape of the CYPs (data not shown), which is consistent with previous results for HsCYPA39.
Comparative SAXS data and solution structures. (a) X-ray scattering intensities depending on the momentum transfer s are shown. In addition, the theoretical scattering pattern calculated by CRYSOL based on the respective in silico model (depicted in c) is shown as well. The intensity distributions are displaced vertically for clarity. (b) Dimensionless Kratky plot. (c) Representation of a single ab initio shape of each CYP and its comparison to the in silico models. The putative substrate binding sites are indicated by black arrows.
High-resolution structure of a phloem CYP
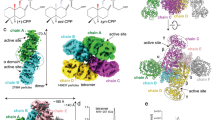
After verifying the monomeric state and overall protein shape of selected CYPs in solution, the crystal structure of BnCYP19-1 in complex with CsA (Fig. 5a,b) was resolved with one molecule per asymmetric unit and refined to a resolution of 2.0 Å. The tertiary structure is overall conserved compared to the plant homologues CsCYP and TaCYPA-1 (pdb id: 4JJM and 4E1Q), which share 67% and 66% sequence identity with BnCYP19-1, respectively. Also the homology models of BnCYP18-4, BnCYP18-5 and AtCYP19-3, which were calculated in silico and are based on the atom coordinates of BnCYP19-1, indicate a high degree of structural conservation compared to each other. The calculated respective RMS values for main chain atoms are around 1 Å, and the overall fold is highly similar compared to HsCYPA as well. BnCYP19-1 possesses a conserved β-barrel-like structure, consisting of eight β-sheets interconnected by three α-helices. Interestingly, the structure of BnCYP19-1 has a previously undescribed site for coordinative metal binding being occupied by Mg2+ (Fig. 5c). However, Mg2+ did not show any influence on the enzyme activity. Furthermore, the crystal structure revealed a malonate molecule forming non-bonded hydrophobic contacts with Ile46 and Gly47. This binding site might interact with an unknown small organic metabolite in vivo. While discussing the crystal structure it should also be noted that the C-terminus of BnCYP19-1 is not well-defined by the electron density map. BnCYP19-1 possesses additional disordered amino acids that are absent in other CYPs (Supplementary Fig. S1).
Crystal structure of BnCYP19-1. (a) Cartoon plot of BnCYP19-1. (b) Topology of the BnCYP19-1 secondary and tertiary structure as visualised by pdbsum (EMBL-EBI65). (c) Cation binding site of BnCYP19-1 superimposed with CsCYP. (d) Superimposition of the divergent loop residues 48-54. BnCYP19-1 is coloured green and the homologous loop of CsCYP in magenta for comparison. A cross-eye stereo image of a section of the BnCYP19-1 electron density map with a resolution of 1.98 Å is provided in Supplementary Fig. S6.
Structure and regulation of the active site
Most of the residues involved in substrate recognition and CsA binding are conserved with similar orientations (Fig. 6b,c). Therefore, it is consistent that the observed enzymatic activities and CsA binding kinetics are in a similar range for all CYPs examined.
Active site structure. (a) Active site of BnCYP19-1 with bound CsA. Serine 110 is highlighted. (b) Schematic representation of the BnCYP19-1 binding epitope according to pdbsum (EMBL-EBI65) with hydrogen bonds indicated by blue lines and hydrophobic interaction indicated by orange dashed lines. (c) Superimposition of active site residues involved in substrate and CsA interaction (crystal structure of BnCYP19-1 in cyan, BnCYP18-4 in purple, AtCYP19-3 in orange, BnCYP18-5 in beige, CsCYP in green and native TaCypA-1 without CsA (pdb ID: 4E1Q) in magenta). The active site geometry is widely conserved also in TaCYPA-1 in the absence of an active site compound.
Despite this fact, there are several structural differences. In contrast to most CYPs, including the two plant homologues from C. sinensis and T. aestivum, both BnCYP19-1 and AtCYP19-3 possess a serine at position 110 instead of an alanine. The high-resolution structure of BnCYP19-1 showed that the additional hydroxyl group is oriented towards a carbonyl oxygen (Aba2-O) of CsA, providing an additional hydrogen bond (2.9 Å, Fig. 6a). Thereby, binding to CsA is stabilised by a total of 6 hydrogen bonds involving 5 amino acids of BnCYP19-1 (Supplementary Table S3). Still, this additional bond does not result in an increased binding capacity.
In case of substrate binding, modelling approaches via superimposition based on the epitope of a Gly-Pro dipeptide binding to HsCYPA40 and an oligopeptide binding to a bacterial CYP41, showed that Ser110 may not interact with the proline peptide bond that is catalytically isomerised. But it could still alter the core substrate specificity.
In order to identify the binding epitope of the activity assay peptide substrate, Suc-AAPF-pNA, at its BnCYP19-1 binding site, in silico docking was performed (Supplementary Fig. S5). Data obtained suggest a hydrogen bond via the hydroxyl group of Ser110 with the succinyl moiety and a hydrogen bond of Arg62 with the backbone carbonyl oxygen of the phenylalanine residue of the substrate among others within the elongated binding cleft. Given the conservation of many active site residues, the differences observed in catalytic activity of the substrate are rather explainable by non-conserved long range interactions and turnover limiting conformational changes of the whole protein as described for HsCYPA42,43 involving particularly residues 75–85, 101–110 and 147–155, which correspond to 82–92, 108–117 and 154–162 in BnCYP19-1. Further the relatively high B-values of Arg70, Lys154 and Arg159 of BnCYP19-1, which are not fully conserved, indicate an involvement in such a mechanism. Previous studies of HsCYPA applied NMR spectroscopy and MD simulations to probe tertiary structure conformational changes, which could further be studied in BnCYP19-1 using time-resolved X-ray crystallography similar to XFEL experiments performed with HsCYPA44.
Discussing the redox regulation model from Campos et al.26 in the context of our results from the activity assays and the structural analysis, it becomes obvious that the divergent BnCYPs and AtCYPs may be regulated by the same mechanism. For CsCYP it is reported that the formation of a disulphide bond between Cys40 and Cys168 induces a repositioning of α-helix I. Thereby hydrogen bonds of Glu83 with Lys48 and Ser49 are disrupted. In BnCYP19-1 (Fig. 5d), Glu83 forms hydrogen bonds with the backbone amide nitrogen of Gln48 (2.8 Å) and Ala49 (3.4 Å), respectively. Due to the alanine in position 49, the hydrogen bond of the Ser49 side chain reported for CsCYP cannot be formed. This leads to only two hydrogen bonds connecting the divergent loop to Glu83. The activity assays under oxidative treatment showed that this is still sufficient to provide the redox regulation mechanism via this interaction. According to the X-ray crystal structure of BnCYP19-1, all four Cys residues were clearly in a reduced state.
Conclusions
Specific CYPs are highly abundant in the phloem of B. napus and other plant species. In phloem exudate of Brassica napus, 18 and 19 kDa single-domain CYPs have been detected in 8 distinct spots on 2D-PAGE gels17 and 8 were detected by LC-MS/MS, with BnCYP18-5 being by far the most abundant CYP16.
In the present study we have compared three similar B. napus phloem CYPs plus a homologous A. thaliana CYP by enzyme activity assays and by structural analysis to understand if the minor differences in amino acid sequences alter structural or functional features.
The expressed and purified three phloem CYPs all showed PPIase activity with only minor differences in activity and inhibitor affinity. Furthermore, they may be regulated by a redox controlled mechanism. A crystal structure at 2 Å resolution enabled the analysis of the inhibitor binding cleft and suggested together with substrate docking no significant changes at the catalytic binding site, except for the non-conserved amino acid position 110.
In summary, our data confirm that the single domain phloem CYPs function as phloem PPIases and as protein chaperones, as it has been proposed earlier17,32. Because of the high number of CYPs in the phloem of B. napus and their high sequence similarity as well as functional and structural redundancy, it seems unlikely that these CYPs, in contrast to SlCYP1 in tomato22, function as long-distance signalling molecules. The data rather lead to the conclusion that the necessity of multiple enzymes may either result from their fundamental role which has to be maintained even under loss of individual CYP isoforms or from the modulation of substrate specificity by residues surrounding the catalytic cleft.
Methods
Multiple sequence alignment of CYP proteins
Sequences were aligned using ClustalOmega (http://www.ebi.ac.uk/Tools/msa/clustalo/)45 and displayed with Jalview 2.9.0b246. For comparison, RcCYP1 (UniProt KB ID: Q8VX73), SlCYP1 (P21568), HsCYPA (P62937), CsCYP (D0ELH5), and TaCYPA-1 (Q93W25) were aligned together with the examined oilseed rape and Arabidopsis cyclophilins.
Plant material, growth conditions and phloem exudate collection
Brassica napus cultivar ‘Drakkar’ plants were grown in 19 cm pots on soil (LAT-Terra Standard P, Industrie-Erdenwerk Archut, Germany) under controlled conditions in a glasshouse. The conditions applied were 70% humidity and a 16 h/8 h light/dark (day/night) with 22 °C/18 °C (day/night) cycle. Plants were watered once per day and fertilized with 2 g l−1 Osmocote Exact Standard High K (Scotts, the Netherlands). Phloem exudate was collected by exudation as described by Giavalisco et al.17 and used for the determination of its PPIase activity directly after collection.
Cloning, gene overexpression and purification
The open reading frames encoding BnCYP18-4 (BnaC03g60160D), BnCYP18-5 (BnaA09g08780D), BnCYP19-1 (BnaA09g35540D), and AtCYP19-3 (At3g56070) were amplified by PCR from genomic DNA of either Brassica napus cultivar ‘Drakkar’ or Arabidopsis thaliana ecotype Col-0 and cloned to pET28a+ with NdeI/XhoI (primers are listed in Supplementary Table S1).
Escherichia coli BL21 (DE3) were transformed with the pET28a+ constructs in order to produce His6-tagged BnCYP18-4, BnCYP18-5, BnCYP19-1, and AtCYP19-3 via recombinant gene overexpression. Liquid cultures in 2YT medium containing kanamycin (50 µg ml−1) were incubated at 37 °C and supplemented with IPTG at a final concentration of 1 mM after the optical density at 600 nm reached 1.0 AU. After 3 hours induction the cells were harvested by centrifugation and resuspended in lysis buffer (20 mM Tris/HCl pH 7.4, 300 mM NaCl, 2 mM imidazole) for cell disruption by lysozyme (f. c. 1 mg ml−1) and sonication. After centrifugation at 43000 x g the clarified supernatant was applied to Ni-NTA affinity chromatography and the His6-tagged proteins were eluted with 20 mM Tris/HCl pH 7.4, 300 mM NaCl, 200 mM imidazole. To remove imidazole and cleave the tag, thrombin digestion was combined with dialysis in 20 mM Tris/HCl pH 7.4, 300 mM NaCl. Subsequently, further purification was achieved by size exclusion chromatography. Finally, the proteins were either concentrated to the desired protein concentration and directly used or they were concentrated to 10 mg ml−1, supplied with 5% (v/v) glycerol and 0.1 mM AeBSF and stored at −80 °C. In order to check for purity, SDS-PAGE and MALDI-TOF mass spectrometry were carried out.
Non-reducing SDS-PAGE to visualise disulphide bond formation
Protein (25 µM) was pre-treated with oxidising agent (CuSO4, 10 mM) for 15 min at room temperature. The mixture was split and each aliquot was mixed with varying amounts of DTT (1-50 mM). After adding SDS sample buffer without reducing agent (60 mM Tris/HCl pH 6.8, 2% SDS, 10% glycerol, 0.02% bromophenol blue) 1 µg protein was directly loaded onto a 15% SDS-PAGE and stained with colloidal coomassie47.
Comparative determination of PPIase activity
Based on the protease-coupled PPIase assay described by Fischer et al.31, the PPIase activity was determined in 35 mM HEPES pH 8.0 as assay buffer and the whole approach was carried out at 4 °C. Either phloem exudate (1, 5 or 10 µl) or purified protein was added. Purified proteins were pre-diluted in assay buffer to stock solutions of 2.5 µM and 250 nM. From these different protein concentrations (0.5 to 50 nM) were prepared with assay buffer and pre-incubated for 5 min. 12 µM α-Chymotrypsin (from a stock solution of 2.4 mM in 1 mM HCl) as well as 80 µM substrate (from a stock solution of 8 mM) were added directly before the measurement. As a substrate N-Succinyl-Ala-Ala-Pro-Phe-p-Nitroanilide (short Suc-AAPF-pNA) was solubilised in 470 mM LiCl, 100% 2,2,2-Trifluorethanol (as described by Kofron et al.)48. After the addition, the total volume of 1 ml was mixed for 4 s and the absorption at 390 nm was recorded with 2 s intervals for 300 s. Three replicates were recorded for each protein concentration. The data were fitted with the first order reaction equation y = y0 + a e−kt (with k as the observed rate constant kobs). The rate constant of the isomerase was determined by subtracting k0 from kobs (with k0 as the spontaneous uncatalysed cis/trans isomerisation rate) and these values were plotted against the protein concentration. The data points could be fitted with a linear regression where the slope is kcat/Km.
To determine the activity after oxidative or reductive treatment, 25 µM protein was pre-treated with either 10 mM CuSO4 or 100 mM DTT for 15 min at room temperature and applied to the assay mixture as described above.
Inhibition of PPIase activity
To determine the inhibitory potential of CsA the PPIase assay was performed as described above except that the protein concentration was kept constant at 5 nM while the CsA concentration ranged from 0 to 1000 nM. To examine the inhibition of phloem exudate activity 10 µl phloem sap was applied and either 1000 nM CsA or FK506 were added.
CsA was solubilised in 100% DMSO to 10 mM and diluted to appropriate stock solutions with the same solvent. After adding the appropriate amount of CsA to the assay mixture, pure DMSO was added if necessary to keep the total amount of solvent constant at 0.1%. The assay mixture was incubated for 5 min at 4 °C before the reaction was started. The PPIase activity without any CsA was defined as maximal active (=kmax) so that the inhibition could be calculated by y = (kmax − (kobs-k0))/kmax. The obtained values were plotted against the CsA concentration. Changing the x-axis to a logarithmic scale resulted in a sigmoid shape of the data points which could be fitted with y = A1 + (A2 − A1)/(1 + (x/x0)p) to determine the IC50.
Dynamic light scattering (DLS)
Solution dispersity and oligomeric states of CYPs were determined using DLS. Protein solution aliquots in a quartz cuvette were exposed to a 660 nm laser applying a Spectrolight 300 instrument (Xtal Concepts, Germany). The time-dependent fluctuations of the scattering intensity at an angle of 90° were autocorrelated and evaluated using the CONTIN algorithm49. The determined diffusion coefficients were used to calculate the average hydrodynamic radii (Rh) distributions via the Stokes-Einstein-Equation.
Small-angle X-ray scattering (SAXS)
SAXS data were collected at the EMBL beamline P12 at PETRA III (DESY, Hamburg, Germany)50. A 2D photon-counting Pilatus 2 M pixel detector (Dectris) with a momentum transfer range 0.03 nm−1 < s < 4.8 nm−1 (\(s=4\pi \frac{sin\theta }{\lambda }\), where 2θ is the scattering angle) was applied. Data collection parameters are listed in Supplementary Table S2. The protein concentrations ranged from 1.4 to 7.7 mg ml−1 in 20 mM Tris pH 7.4, 300 mM NaCl, 1 mM DTT. Protein sample solutions and the corresponding buffer solutions were exposed to the X-rays in alternating order for twenty consecutive exposure frames of 45 ms each. Scattering intensities of individual exposures were averaged, and the latter scattering amplitudes were subtracted from the protein solution scattering amplitudes applying the automated data processing pipeline SASFLOW51. The resulting curves were further processed by PRIMUSqt52. Due to no significant changes in Rg, I(0) and MW with increasing protein concentrations, the subsequent analysis was carried out based on the highest measured concentration (3.1 mg ml−1 BnCYP18-4, 4.2 mg ml−1 BnCYP18-5, 7.1 mg ml−1 BnCYP19-1, 7.7 mg ml−1 AtCYP19-3). The Rg values and the maximum dimensions (Dmax) of native CYPs were obtained from AUTORG and the particle pair distance distribution functions P(r) were calculated via GNOM respectively52. DAMMIF was applied to generate 20 individual ab initio bead models for each protein sample53. The models are well superimposable as verified by their normalised spatial discrepancy (NSD). A bovine serum albumin reference solution (5 mg ml−1; 66.2 kDa) in 50 mM HEPES pH 7.5 was used to calibrate and verify the beamline operation.
Crystallisation
Cyclosporin A (Sigma-Aldrich) was dissolved in pure DMSO and slowly added to the protein after tag cleavage at a molar ratio of 2:1 (CsA:protein). The CYP:CsA complex was mixed with an equal volume of reservoir solution. Initially, 800 different crystallization conditions were screened using the vapour diffusion and microbatch technique, respectively. X-ray suitable crystals of BnCYP19-1 in complex with CsA were obtained from sitting drop vapour diffusion crystallisation trials applying a protein concentration of 10 mg ml−1. The reservoir contained 2.4 M sodium malonate, pH 7.0. Bipyramidal crystals grew to a maximum size of approximately 200 × 200 × 200 µm3 within 3 weeks and were cryo-protected prior to X-ray data collection by slowly supplementing the drop with glycerol to a final concentration of 12% (v/v) on a micromesh.
Diffraction data collection and processing
Diffraction data of BnCYP19-1 in complex with CsA were collected at the EMBL beamline P14 (PETRA III, DESY, Germany) applying an X-ray wavelength of 1.0332 Å and using one loop-mounted and flash-cooled crystal at 100 K. A single crystal was exposed to the beam at a rotation increment of 0.1°. Indexing was performed by IMOSFLM54. The space group was assigned to be I4122. Unit cell and refinement parameters are summarized in Table 2.
Structure refinement
Diffraction data obtained were reduced and scaled applying SCALA, part of the CCP4 suite55, and converted via FREEFLAG56. The phase problem was solved by molecular replacement using MOLREP57 and coordinates of pdb entry 4JJM. For iterative refinement the program REFMAC58 was applied and manual refinement was performed using COOT59. The quality of refinement and coordinates was verified using PROCHECK60 and the REDO server61.
Homology modelling
The tertiary structure of B. napus and A. thaliana CYPs in the absence of an active site ligand based on the obtained high-resolution structure of BnCYP19-1 was modelled using SWISS MODEL62,63. For comparison I-TASSER64 was applied and these models were also fitted into ab initio shapes.
In silico ligand docking
The Flexidock™ subprogram implemented in Sybyl-X 1.2 (Tripos International) was applied for docking studies in order to determine the epitope of a substrate peptide at the active site of BnCYP19-1. In preparation, hydrogen atoms were added to the protein and Gasteiger-Hückel charges were calculated and assigned. The free energy of the protein was minimised over 500 cycles and 50 cycles for preparation of the ligand. The dielectric constant of the Tripos force field was set to 20.
Data Availability
The resolved macromolecular structure of BnCYP19-1 is deposited in the protein data bank via pdb ID 6HMZ.
References
Galat, A. Peptidylprolyl cis/trans isomerases (immunophilins): biological diversity-targets-functions. Curr. Top. Med. Chem. 3, 1315–1347 (2003).
Handschumacher, R. E., Harding, M. W., Rice, J., Drugge, R. J. & Speicher, D. W. Cyclophilin: a specific cytosolic binding protein for cyclosporin A. Science 226, 544–547 (1984).
Harding, M. W., Galat, A., Uehling, D. E. & Schreiber, S. L. A receptor for the immunosuppressant FK506 is a cis-trans peptidyl-prolyl isomerase. Nature 341, 758–760, https://doi.org/10.1038/341758a0 (1989).
Kiefhaber, T., Quaas, R., Hahn, U. & Schmid, F. X. Folding of ribonuclease T1. 2. Kinetic models for the folding and unfolding reactions. Biochemistry 29, 3061–3070 (1990).
Jager, M. & Pluckthun, A. The rate-limiting steps for the folding of an antibody scFv fragment. FEBS Lett. 418, 106–110 (1997).
Yurchenko, V. et al. Active site residues of cyclophilin A are crucial for its signaling activity via CD147. J. Biol. Chem. 277, 22959–22965, https://doi.org/10.1074/jbc.M201593200 (2002).
Pogorelko, G. V. et al. Characterization of three Arabidopsis thaliana immunophilin genes involved in the plant defense response against Pseudomonas syringae. Gene 538, 12–22, https://doi.org/10.1016/j.gene.2014.01.029 (2014).
Krzywicka, A. et al. KIN241: a gene involved in cell morphogenesis in Paramecium tetraurelia reveals a novel protein family of cyclophilin-RNA interacting proteins (CRIPs) conserved from fission yeast to man. Mol. Microbiol. 42, 257–267 (2001).
Thapar, R. Roles of prolyl isomerases in RNA-mediated gene expression. Biomolecules 5, 974–999, https://doi.org/10.3390/biom5020974 (2015).
Li, H. et al. A WD40 domain cyclophilin interacts with histone H3 and functions in gene repression and organogenesis in Arabidopsis. Plant Cell 19, 2403–2416, https://doi.org/10.1105/tpc.107.053579 (2007).
Park, S. W. et al. Cyclophilin 20-3 relays a 12-oxo-phytodienoic acid signal during stress responsive regulation of cellular redox homeostasis. Proc. Natl. Acad. Sci. USA 110, 9559–9564, https://doi.org/10.1073/pnas.1218872110 (2013).
Geisler, M., Bailly, A. & Ivanchenko, M. Master and servant: Regulation of auxin transporters by FKBPs and cyclophilins. Plant Sci 245, 1–10, https://doi.org/10.1016/j.plantsci.2015.12.004 (2016).
Ahn, J. C. et al. Classification of rice (Oryza sativa L. Japonica nipponbare) immunophilins (FKBPs, CYPs) and expression patterns under water stress. BMC Plant Biol. 10, 253, https://doi.org/10.1186/1471-2229-10-253 (2010).
Romano, P. G., Horton, P. & Gray, J. E. The Arabidopsis cyclophilin gene family. Plant Physiol. 134, 1268–1282, https://doi.org/10.1104/pp.103.022160 (2004).
Mainali, H. R., Chapman, P. & Dhaubhadel, S. Genome-wide analysis of cyclophilin gene family in soybean (Glycine max). BMC Plant Biol. 14, 282, https://doi.org/10.1186/s12870-014-0282-7 (2014).
Hanhart, P. et al. Bioinformatic and expression analysis of the Brassica napus L. cyclophilins. Sci. Rep. 7, 1514, https://doi.org/10.1038/s41598-017-01596-5 (2017).
Giavalisco, P., Kapitza, K., Kolasa, A., Buhtz, A. & Kehr, J. Towards the proteome of Brassica napus phloem sap. Proteomics 6, 896–909, https://doi.org/10.1002/pmic.200500155 (2006).
Barnes, A. et al. Determining protein identity from sieve element sap in Ricinus communis L. by quadrupole time of flight (Q-TOF) mass spectrometry. J. Exp. Bot. 55, 1473–1481, https://doi.org/10.1093/jxb/erh161 (2004).
Lin, M. K., Lee, Y. J., Lough, T. J., Phinney, B. S. & Lucas, W. J. Analysis of the pumpkin phloem proteome provides insights into angiosperm sieve tube function. Mol. Cell. Proteomics 8, 343–356, https://doi.org/10.1074/mcp.M800420-MCP200 (2009).
Rodriguez-Medina, C., Atkins, C. A., Mann, A. J., Jordan, M. E. & Smith, P. M. Macromolecular composition of phloem exudate from white lupin (Lupinus albus L.). BMC Plant Biol. 11, 36, https://doi.org/10.1186/1471-2229-11-36 (2011).
Schobert, C. et al. Identification of immunologically related proteins in sieve-tube exudate collected from monocotyledonous and dicotyledonous plants. Planta 206, 245–252 (1998).
Spiegelman, Z., Omer, S., Mansfeld, B. N. & Wolf, S. Function of Cyclophilin1 as a long-distance signal molecule in the phloem of tomato plants. J. Exp. Bot. 68, 953–964, https://doi.org/10.1093/jxb/erw487 (2017).
Ke, H. M., Zydowsky, L. D., Liu, J. & Walsh, C. T. Crystal structure of recombinant human T-cell cyclophilin A at 2.5 A resolution. Proc. Natl. Acad. Sci. USA 88, 9483–9487 (1991).
Pflugl, G. M., Kallen, J., Jansonius, J. N. & Walkinshaw, M. D. The molecular replacement solution and X-ray refinement to 2.8 A of a decameric complex of human cyclophilin A with the immunosuppressive drug cyclosporin A. J. Mol. Biol. 244, 385–409, https://doi.org/10.1006/jmbi.1994.1738 (1994).
Vasudevan, D. et al. Plant immunophilins: a review of their structure-function relationship. Biochim. Biophys. Acta, https://doi.org/10.1016/j.bbagen.2014.12.017 (2015).
Campos, B. M. et al. A redox 2-Cys mechanism regulates the catalytic activity of divergent cyclophilins. Plant. Physiol. 162, 1311–1323, https://doi.org/10.1104/pp.113.218339 (2013).
Sekhon, S. S. et al. Structural and biochemical characterization of the cytosolic wheat cyclophilin TaCypA-1. Acta Cryst. D 69, 555–563, https://doi.org/10.1107/S0907444912051529 (2013).
Ghosh, D. et al. Primary identification, biochemical characterization, and immunologic properties of the allergenic pollen cyclophilin Cat r 1. J. Biol. Chem. 289, 21374–21385, https://doi.org/10.1074/jbc.M114.559971 (2014).
Vasudevan, D., Fu, A., Luan, S. & Swaminathan, K. Crystal structure of Arabidopsis cyclophilin38 reveals a previously uncharacterized immunophilin fold and a possible autoinhibitory mechanism. Plant Cell 24, 2666–2674, https://doi.org/10.1105/tpc.111.093781 (2012).
Kaur, G. et al. Characterization of peptidyl-prolyl cis-trans isomerase- and calmodulin-binding activity of a cytosolic Arabidopsis thaliana cyclophilin AtCyp19-3. PloS one 10, e0136692, https://doi.org/10.1371/journal.pone.0136692 (2015).
Fischer, G., Bang, H. & Mech, C. Determination of enzymatic catalysis for the cis-trans-isomerization of peptide binding in proline-containing peptides. Biomed. Biochim. Acta 43, 1101–1111 (1984).
Gottschalk, M. et al. Ricinus communis cyclophilin: functional characterisation of a sieve tube protein involved in protein folding. Planta 228, 687–700, https://doi.org/10.1007/s00425-008-0771-8 (2008).
Zydowsky, L. D. et al. Active site mutants of human cyclophilin A separate peptidyl-prolyl isomerase activity from cyclosporin A binding and calcineurin inhibition. Protein Sci. 1, 1092–1099, https://doi.org/10.1002/pro.5560010903 (1992).
Laxa, M., Konig, J., Dietz, K. J. & Kandlbinder, A. Role of the cysteine residues in Arabidopsis thaliana cyclophilin CYP20-3 in peptidyl-prolyl cis-trans isomerase and redox-related functions. Biochem. J. 401, 287–297, https://doi.org/10.1042/BJ20061092 (2007).
Page, A. P., MacNiven, K. & Hengartner, M. O. Cloning and biochemical characterization of the cyclophilin homologues from the free-living nematode Caenorhabditis elegans. Biochem. J. 317, 179–185 (1996).
Davis, T. L. et al. Structural and biochemical characterization of the human cyclophilin family of peptidyl-prolyl isomerases. PLoS Biol. 8, e1000439, https://doi.org/10.1371/journal.pbio.1000439 (2010).
Dornan, J. et al. Biochemical and structural characterization of a divergent loop cyclophilin from Caenorhabditis elegans. J. Biol. Chem. 274, 34877–34883 (1999).
Daum, S. et al. Isoform-specific inhibition of cyclophilins. Biochemistry 48, 6268–6277, https://doi.org/10.1021/bi9007287 (2009).
Hamilton, G. S. & Steiner, J. P. Immunophilins: beyond immunosuppression. J. Med. Chem. 41, 5119–5143, https://doi.org/10.1021/jm980307x (1998).
Zhao, Y. & Ke, H. Mechanistic implication of crystal structures of the cyclophilin-dipeptide complexes. Biochemistry 35, 7362–7368, https://doi.org/10.1021/bi960278x (1996).
Christoforides, E., Dimou, M., Katinakis, P., Bethanis, K. & Karpusas, M. Structure of a bacterial cytoplasmic cyclophilin A in complex with a tetrapeptide. Acta Cryst. F 68, 259–264, https://doi.org/10.1107/S1744309112000188 (2012).
McGowan, L. C. & Hamelberg, D. Conformational plasticity of an enzyme during catalysis: intricate coupling between cyclophilin A dynamics and substrate turnover. Biophys. J. 104, 216–226, https://doi.org/10.1016/j.bpj.2012.11.3815 (2013).
van den Bedem, H., Bhabha, G., Yang, K., Wright, P. E. & Fraser, J. S. Automated identification of functional dynamic contact networks from X-ray crystallography. Nat. Methods 10, 896–902, https://doi.org/10.1038/nmeth.2592 (2013).
Keedy, D. A. et al. Mapping the conformational landscape of a dynamic enzyme by multitemperature and XFEL crystallography. eLife 4, https://doi.org/10.7554/eLife.07574 (2015).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539, https://doi.org/10.1038/msb.2011.75 (2011).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191, https://doi.org/10.1093/bioinformatics/btp033 (2009).
Kang, D. H., Gho, Y. S., Suh, M. K. & Kang, C. H. Highly sensitive and fast protein detection with coomassie brilliant blue in sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Bull. Korean Chem. Soc. 23, 1511–1512 (2002).
Kofron, J. L., Kuzmic, P., Kishore, V., Colon-Bonilla, E. & Rich, D. H. Determination of kinetic constants for peptidyl prolyl cis-trans isomerases by an improved spectrophotometric assay. Biochemistry 30, 6127–6134 (1991).
Provencher, S. W. CONTIN: a general purpose constrained regularization program for inverting noisy linear algebraic and integral equations. Comput. Phys. Commun. 27, 229–242 (1982).
Blanchet, C. E. et al. Versatile sample environments and automation for biological solution X-ray scattering experiments at the P12 beamline (PETRA III, DESY). J. Appl. Crystallogr. 48, 431–443, https://doi.org/10.1107/S160057671500254X (2015).
Franke, D., Kikhney, A. G. & Svergun, D. I. Automated acquisition and analysis of small angle X-ray scattering data. Nucl. Instrum. Methods Phys. Res. 689, 52–59, https://doi.org/10.1016/j.nima.2012.06.008 (2012).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225, https://doi.org/10.1107/S1600576717007786 (2017).
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J. Appl. Crystallogr. 42, 342–346, https://doi.org/10.1107/S0021889809000338 (2009).
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Cryst. D 67, 271–281, https://doi.org/10.1107/S0907444910048675 (2011).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Cryst. D 67, 235–242, https://doi.org/10.1107/S0907444910045749 (2011).
Brunger, A. T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025, https://doi.org/10.1107/S0021889897006766 (1997).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Cryst. D 53, 240–255, https://doi.org/10.1107/S0907444996012255 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Cryst. D 60, 2126–2132, https://doi.org/10.1107/S0907444904019158 (2004).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291, https://doi.org/10.1107/S0021889892009944 (1993).
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220, https://doi.org/10.1107/S2052252514009324 (2014).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–258, https://doi.org/10.1093/nar/gku340 (2014).
Bienert, S. et al. The SWISS-MODEL repository-new features and functionality. Nucleic Acids Res. 45, D313–D319, https://doi.org/10.1093/nar/gkw1132 (2017).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8, https://doi.org/10.1038/nmeth.3213 (2015).
Laskowski, R. A., Jablonska, J., Pravda, L., Varekova, R. S. & Thornton, J. M. PDBsum: Structural summaries of PDB entries. Protein Sci. 27, 129–134, https://doi.org/10.1002/pro.3289 (2018).
Acknowledgements
The authors acknowledge excellent support by the EMBL beamline scientists at PETRA III beamline P14 and P12 (DESY, Hamburg, Germany) from the groups of Thomas Schneider and Dmitri Svergun. S.F. is supported by ‘The Hamburg Centre for Ultrafast Imaging’ excellence cluster of the Deutsche Forschungsgemeinschaft (DFG). This work was further financially supported by a Career Integration Grant (CIG; PCIG14-GA-2013-63 0734) by the European Commission within the 7th framework program, the grant LFF-GK06 ‘DELIGRAH’ (Landesforschungsförderung Hamburg), and a DFG grant (DFG KE 856_6-1) all awarded to J.K.
Author information
Authors and Affiliations
Contributions
P.H. and J.K. designed the study. P.H. and M.G. cloned the constructs and purified the proteins. P.H. performed enzyme assays with phloem exudate and untreated purified proteins, DLS, SAXS measurements and the respective data analysis. P.H. and V.R. investigated the purified proteins under oxidative and reducing conditions. M.G. and S.F. set up the crystallisation. S.F. did diffraction data collection and processing, docking and supported SAXS experiments. P.H., S.F. and M.T. wrote the manuscript. C.B. and J.K. corrected the manuscript. J.K. supervised the study. All authors reviewed the manuscript and approved the final article.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hanhart, P., Falke, S., Garbe, M. et al. Enzyme activity and structural features of three single-domain phloem cyclophilins from Brassica napus. Sci Rep 9, 9368 (2019). https://doi.org/10.1038/s41598-019-45856-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45856-y
This article is cited by
-
Characterization and regulation of salt upregulated cyclophilin from a halotolerant strain of Penicillium oxalicum
Scientific Reports (2023)
-
Cyclophilin anaCyp40 regulates photosystem assembly and phycobilisome association in a cyanobacterium
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.