Abstract
Mutations in isocitrate dehydrogenases 1 and 2 (IDHmut) are present in a variety of cancers, including glioma, acute myeloid leukemia (AML), melanoma, and cholangiocarcinoma. These mutations promote hypermethylation, yet it is only a favorable prognostic marker in glioma, for reasons that are unclear. We hypothesized that the patterns of DNA methylation, and transcriptome profiles, would vary among IDHmut cancers, especially gliomas. Using Illumina 450K and RNA-Seq data from The Cancer Genome Atlas, we show that of 365,092 analyzed CpG sites, 70,591 (19%) were hypermethylated in IDHmut gliomas compared to wild-type (IDHwt) gliomas, and only 3%, 2%, and 4% of CpG sites were hypermethylated in IDHmut AML, melanoma, and cholangiocarcinoma, relative to each of their IDHwt counterparts. Transcriptome differences showed pro-malignant genes that appear to be unique to IDHmut gliomas. However, genes involved in differentiation and immune response were suppressed in all IDHmut cancers. Additionally, IDHmut caused a greater degree of hypermethylation in undifferentiated neural progenitor cells than in mature astrocytes. These data suggest that the extent and targets of IDHmut-induced genomic hypermethylation vary greatly according to the cellular context and may help explain why IDHmut is only a favorable prognostic marker in gliomas.
Similar content being viewed by others
Introduction
Epigenetic modifications control gene expression via mechanisms that are highly coordinated throughout the life of a cell. In addition to regulating the development of distinct cell lineages, epigenetic modifications can also promote malignant transformation and cancer progression1. Methylation of a CpG site within or near a gene can change its expression, usually by suppressing it. If this happens to a tumor-suppressor gene, oncogenesis may occur2.
The mechanisms by which aberrant methylation occurs, and its consequences in cancer, are becoming better understood. One such mechanism involves point mutations in isocitrate dehydrogenases 1 and 2 (collectively “IDHmut”). These metabolic enzymes normally convert isocitrate into α-ketoglutarate, but mutations in key arginine residues that normally bind isocitrate substrate cause a radical change in enzymatic activity, wherein mutant enzyme converts α-ketoglutarate into D-2-hydroxyglutarate (D2HG)3. D2HG then acts as a competitive inhibitor of other enzymes that require α-ketoglutarate as a cofactor, including certain DNA demethylases, leading to genomic CpG hypermethylation and globally altered transcription4,5. IDHmut-induced hypermethylation may lead to suppression of cellular differentiation, thereby prolonging the window in which additional oncogenic “hits” can occur6.
IDHmut has been identified in a variety of malignancies, but the most common include infiltrative glioma7, acute myeloid leukemia (AML)8, melanoma9, cholangiocarcinoma10, and cartilaginous tumors11. Even though IDHmut has the same basic effects on the biochemistry and methylome of every cancer in which it has been detected, glioma is the only cancer in which IDHmut is a favorable prognostic marker6,12,13,14,15,16,17. The reasons for this are still unclear. In our recently published work, we found that the methylation-dependent suppression of Tissue Factor (TF) contributes to the reduced thrombogenicity and reduced malignancy of IDHmut gliomas, and that methylation and suppression of the gene encoding TF, F3, is far greater in IDHmut gliomas than in other IDHmut cancers18,19. This prompted a broader comparative analysis of the major IDHmut cancers in The Cancer Genome Atlas, demonstrating great diversity in DNA methylation patterns, resultant transcriptomic profiles, and specific genes and biological pathways altered by IDHmut depending on cellular context.
Results
Patient cohort characteristics
Whole-genome DNA methylation profiles of glioma, AML, cholangiocarcinoma, and melanoma from The Cancer Genome Atlas (TCGA) were grouped by isocitrate dehydrogenase 1 and 2 mutation status, referred to as IDHmut or IDHwt. The glioma cohort included 647 patients that consisted of World Health Organization (WHO) grade II-IV gliomas (427 IDHmut and 220 IDHwt). The AML cohort included 194 patients (15 IDHmut and 179 IDHwt), TCGA had 45 cholangiocarcinomas (7 IDHmut and 38 IDHwt), and melanomas consisted of 475 cases (23 IDHmut and 452 IDHwt). Chondrosarcomas are not represented in TCGA. Clinical data, including age, gender, race and ethnicity are summarized in Table 1. Survival plots are depicted in Fig. 1. IDHmut only had a significant favorable survival advantage in glioma patients (median 79.9 versus 13.3 months, HR = 0.18, 95% CI = 0.14–0.24, P < 0.0001), and was not associated with overall survival in the available AML, cholangiocarcinoma, or melanoma data (Fig. 1).
Overall survival of cancer patients stratified according to IDH mutation status. Kaplan-Meier curves of overall survival in IDHwt and IDHmut (a) glioma (IDHmut N = 151; IDHwt N = 302), (b) AML (IDHmut N = 34; IDHwt N = 137), (c) cholangiocarcinoma (IDHmut N = 7; IDHwt N = 29), (d) and melanoma (IDHmut N = 25; IDHwt N = 439).
Patterns of genomic methylation vary across IDHmut cancers
To determine the methylation profile for each cohort, β-values for CpG sites that met mapping quality thresholds in the TCGA DNA Methylation Liftover Pipeline20 for all samples within a cancer type were analyzed, yielding 365,092 CpG sites for glioma, 393,152 CpG sites for AML, 379,101 CpG sites for cholangiocarcinoma, and 373,827 CpG sites for melanoma. A CpG site with a mean delta β-value > 0.15 (<0.15) for IDHmut relative to the matching IDHwt cancer, with FDR P < 0.05, was considered to be hypermethylated (hypomethylated). IDHmut was associated with genomic hypermethylation among all cancers, yet hypomethylation was also observed (Fig. 2). Gliomas showed extensive hypermethylation; of the 365,092 analyzed CpG sites, 70,591 (19%) were hypermethylated in IDHmut glioma compared to IDHwt gliomas, whereas the others ranged between 2–4% (Supplementary Table S1). To rule out the possibility that IDH1wt gliomas are just abnormally hypomethylated relative to other IDH1wt malignancies, a similar analysis was done directly comparing IDHmut gliomas to the other IDHmut cancers. In this setting, IDHmut gliomas showed much greater levels of hypermethylation when compared with other IDHmut cancers (Supplementary Table S2). The number of CpG sites hypermethylated in the other cancers were 13,128 (3%) in AML, 11,763 (3%) in cholangiocarcinoma, and 8,663 (2%) in melanoma relative to each of their IDHwt counterparts (Fig. 2b–d). While smaller sample sizes may have contributed to an overall lower frequency of statistically significant findings for AML, cholangiocarcinoma, and melanoma, differences in β-values for IDHmut compared to IDHwt tended to be more extreme across CpG sites for gliomas than for the other cancers, regardless of P-value. Differences in β-values approached 0.8 at the extremes for gliomas, while observed differences in β-values for the other cancers tended to be 0.5 or less (Fig. 2). Interestingly, IDHmut gliomas also showed the greatest degree of hypomethylation when compared with IDHmut AML, cholangiocarcinoma, and melanoma (Fig. 2). Overall, these results suggest that the methylation profiles associated with IDHmut vary greatly according to cellular and tissue contexts.
Methylation levels across IDHmut cancers. Volcano plots comparing CpG methylation levels of IDHmut versus IDHwt in (a) glioma, (b) AML, (c) cholangiocarcinoma, and (d) melanoma. Difference of mean methylation (x-axis) and significance of the difference (y-axis). Each point represents a unique CpG site. Hypermethylation is represented by a delta beta ≥0.15 and FDR-corrected P < 0.05 shown in red, and hypomethylation by delta beta ≤−0.15 and FDR-corrected P < 0.05.
To further investigate the unique methylation profiles observed among IDHmut TCGA cancers, methylation sites were grouped into six genomic regions relative to CpG islands: (1) north shelf, (2) north shore, (3) island, (4) south shore, (5) south shelf, (6) open sea21. ‘Shore’ regions immediately flank each CpG island up to 2 kilobases away, ‘shelf’ regions extend outwards from the ‘shore’ up to 2 kilobases away from the shelf, and ‘open sea’ indicates the rest of the genome. The ‘north’ region represents the 5′ end, and ‘south’ the 3′ end. Within each cancer type, IDHmut tumors consistently showed increased methylation across all genomic regions relative to their IDHwt counterparts (Fig. 3). However, the most striking IDHmut-IDHwt difference in methylation among cancer types was found in CpG islands, wherein IDHmut gliomas showed a greater increase in CpG island methylation, relative to IDHwt gliomas, than was found in other IDHmut cancers. IDHmut gliomas also showed increased methylation across all genomic regions when directly compared with other IDHmut cancers (Supplementary Fig. S2). This suggests that, while IDHmut is associated with increased CpG methylation across all major genomic regions, the greatest differences occur within CpG islands.
Distribution of mean site-specific CpG methylation levels by genomic region for CpG sites identified as hypermethylated in IDHmut versus IDHwt tumors. The density of probes at each CpG site (y-axis) is plotted versus methylation β-values (x-axis). Islands are genomic areas with relatively high CpG content flanked by shores (up to 2 kilobases) and shelves (2–4 kilobases from island). North shore and shelf represent 5′ end, and south shore and shelf represent 3′ end. The open sea represents the rest of the genome. Gold lines represent CpG methylation in IDHwt tumors and blue lines represent CpG methylation in IDHmut tumors.
Methylome and transcriptome changes in IDHmut gliomas are distinct from other IDHmut cancers
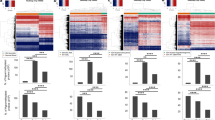
Hierarchical cluster analysis of differences in β-values for genes that were hypermethylated in IDHmut compared to IDHwt cancers, in at least one cancer type, highlighted the distinct differential methylation profile for CpG sites between IDHmut and IDHwt subsets across tumor types. In particular, many CpG sites with substantial hypermethylation in gliomas (dark blue) consistently showed far more modest differences in methylation for IDHmut versus IDHwt (yellow) in the other tumors, regardless of statistical significance (Fig. 4A). Of these CpG sites meeting formal criteria for differential methylation, 56,801 were unique to glioma, and were not identified as differentially methylated in any other IDHmut-IDHwt tumor pairings. Notably, despite smaller sample sizes for AML, cholangiocarcinoma, and melanoma, statistical analyses did identify unique sites of differential methylation in those tumors (6,271 AML; 4,808 cholangiocarcinoma; 3,519 melanoma). Since those CpG sites were not significantly methylated in IDHmut gliomas, even though the glioma cohort was much larger, these results indicate that a great deal of IDHmut-related CpG site methylation is tissue-specific. Only 217 similar CpG sites were hypermethylated across all IDHmut cancers, compared to all IDHwt cancers (Fig. 4B). Transcriptome comparisons showed similar results, with IDHmut gliomas showing unique alteration of more mRNA transcripts (4,214) than any other tumor type (Fig. 4C,D). Stratifying IDHmut gliomas into oligodendroglioma and astrocytoma revealed a large amount of similar CpG sites hypermethylated when compared to IDHwt gliomas (Supplemental Fig. S1a,b). There were some variances in differential gene expression between IDHmut oligodendrogliomas and astrocytomas. Oligodendrogliomas showed unique down regulation of genes linked to angiogenesis, cell proliferation, and integrin binding (Supplemental Fig. S1c). Hierarchical clustering of methylation sites again demonstrated that, whereas the methylation patterns of IDHmut AML, cholangiocarcinomas, and melanomas were relatively similar, IDHmut gliomas were quite distinct (Fig. 4A). Glioma showed the greatest IDHmut-IDHwt differential gene expression (4,214), compared to AML (159), cholangiocarcinoma (139), and melanoma (416) (Fig. 4D). IDHmut gliomas also had the most concordance of genes that were both hypermethylated and differentially expressed when compared with other cancers (Supplementary Table S3).
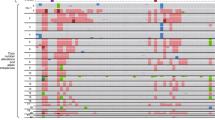
Differential methylation and transcription signatures between IDHmut and IDHwt groups within each cancer type. (A) Methylation heatmap of differences in CpG site beta values between IDHmut and IDHwt tumors for all genes demonstrating statistically significant hypermethylation in at least one tumor type. (B) Venn diagram representing the overlap of hypermethylated genes of IDHmut cancers compared with IDHwt control. (C) Gene expression heatmap of log fold change (FC) gene expression between IDHmut and IDHwt tumors for all genes demonstrating statistically significant differential expression in at least one tumor type. (D) Venn diagram representing the overlap of differentially expressed genes of IDHmut cancers compared with IDHwt control.
To better understand how these methylation and transcription differences among IDHmut cancers might impact cell biology, we performed gene set enrichment analyses (GSEA) of genes that were differentially expressed within each pair of IDHmut and IDHwt cancers. Using an FDR-corrected P < 0.05, we found the following differentially expressed genes: 4,691 (glioma); 282 (AML); 246 (cholangiocarcinoma); 758 (melanoma). Compared to IDHwt gliomas, IDHmut gliomas showed increased expression of 1,629 genes and decreased expression of 3,063 genes (Supplementary Spreadsheet). Glioma GSEA of IDHmut and IDHwt tumors revealed a down-regulation of multiple biological processes; most notable were tissue development (GO:0009888), immune response (GO:0006955), angiogenesis (GO:0001525), and cell proliferation (GO:0008283) (Fig. 5a). GSEA of AML, melanoma, and cholangiocarcinoma showed enrichment of fewer pathways (Fig. 5b–d), which is likely related to smaller sample sizes and lower statistical power. However, all IDHmut cancers showed suppression of GSEA pathways involved in tissue development (GO:0009888) and immune response (GO:0006955).
Gene Set Enrichment Analysis (GSEA) showing gene sets in an interaction network for (a) glioma, (b) AML, (c) cholangiocarcinoma, and (d) melanoma. Each circular node represents enriched gene sets, where node color intensity corresponds with statistical significance (P-value). Blue nodes represent negative enrichment and red nodes represent positive enrichment. Edges (grey connection lines) represent overlap between gene sets with line thickness correlating to the degree of overlap. Networks of nodes that reflect generic function were circled and assigned group labels.
The effect of IDHmut on CpG methylation varies according to cellular differentiation
A causal link between IDHmut and genomic hypermethylation was originally established by inserting IDHmut into IDHwt cells and observing its effects on methylation5,22. However, the choice of IDHwt cell models varies greatly among laboratories, even among those focusing on the same kind of IDHmut tumor. For example, in gliomas, some use immortalized differentiated normal human astrocytes (NHAs), whereas others use immature neural progenitor cells (NPCs). To compare the effects of IDHmut-induced methylation between different experimental models from the same type of tissue, two publicly available datasets were analyzed: (i) Infinium 450K methylation data from NHAs expressing ectopic R132H IDH1 or IDH1wt for 15 consecutive passages (GSE30339)5; (ii) Infinium 450K methylation data from NPCs, also expressing ectopic R132H IDH1 or IDH1wt for 15 consecutive passages (GSE94962)22. R132H IDH1 had a far greater effect on the methylation of undifferentiated NPCs than of mature NHAs, resulting in 87,541 versus 15,976 hypermethylated CpG sites, respectively (Fig. 6a,b). Basal methylation levels of IDHwt NPCs compared with IDHwt astrocytes showed extensive hypomethylation, suggesting that they may be more amenable to IDHmut (Supplementary Fig. S6). NPCs with IDHmut versus IDHwt showed a much larger shift towards hypermethylation at the ‘island’, ‘north shore’, and ‘south shore’ genomic regions when compared to NHAs (Fig. 6c,d). Since these studies were analyzed retrospectively and performed using separate experimental conditions, caution must be used when interpreting the results. However, these data may suggest that the effect of IDHmut on the methylome not only depends on cellular lineage, but also on the differentiation state and preexisting DNA methylation of the cell.
Differences in plasticity of DNA methylation in IDHmut astrocytes and neural progenitor cells. Volcano plots comparing CpG methylation levels of IDHmut versus IDHwt in (a) astrocytes and (b) neural progenitor cells. Difference of mean methylation (x-axis) and significance of the difference (y-axis). Each point represents a unique CpG site. Hypermethylation is represented by a delta beta ≥0.15 and FDR-corrected P < 0.05 shown in red, and hypomethylation by delta beta ≤−0.15 and FDR-corrected P < 0.05. Distribution of mean site-specific CpG methylation across genomic regions for IDHwt and IDHmut (c) astrocytes and (d) neural progenitor cells.
Discussion
The first published case of an IDHmut cancer was in 2006, when a study of over 1,600 breast and colorectal cancers found a single colorectal adenocarcinoma with the R132C variant of IDH123. In 2008–2009, a pair of studies reported a surprisingly high proportion of infiltrative gliomas with IDHmut, most commonly R132H IDH1; such tumors tended to be much less aggressive than their IDHwt counterparts7,24. Follow-up studies showed that, while IDHmut can rarely be found in a variety of cancers, it is most common in glioma, acute myeloid leukemia, cholangiocarcinoma, melanoma, and cartilaginous tumors7,8,9,10,11,24. This was puzzling, insofar as none of the cancers, or the tissues from which they arise, have any obvious connections with each other. Although IDHmut produces D2HG and causes genomic hypermethylation in each type of cancer, only in gliomas does IDHmut show a consistent, powerful association with prognosis (Fig. 1). Guilhamon et al. previously reported on shared signaling pathway alterations in IDHmut cancers25, yet to the best of our knowledge, no study has focused on what distinguishes IDHmut gliomas from the others. Our TCGA-based data strongly suggests that IDHmut has a unique effect on the methylome and transcriptome of gliomas relative to other IDHmut cancers, and that this could help explain why IDHmut is only a favorable prognostic marker in gliomas.
Assuming that mutations in dividing cells occur in a more or less stochastic fashion, the study of mutation patterns among cancers raises provocative questions regarding the conditions under which specific alterations exert a positive (or negative) evolutionary advantage. In contrast to TP53, which is frequently mutated in a wide range of cancers, IDHmut is only enriched in a relatively small number of malignancies, none of which bear any clear resemblance to each other. This would seem to indicate that IDHmut, and its D2HG product, are beneficial to cell growth and oncogenesis in only a few settings. Perhaps only a few tissues express wild-type IDH1 and IDH2 in sufficient amounts, or perhaps carbon flux through these enzymes is only strong enough in particular microenvironments, for IDHmut to have a significant effect on metabolism. Wild-type IDH1 and IDH2 normally produce α-ketoglutarate (α-KG) and reduce nicotinamide adenine dinucleotide phosphate (NADPH); the latter then serves as an important antioxidant in the cell6. Because IDHmut consumes α-KG and NADPH, some cell types may be better able to compensate for the resultant metabolic deficiencies than others. Since D2HG acts as a competitive inhibitor of certain demethylating dioxygenases that require α-ketoglutarate as a cofactor6, the pro-methylating (and pro-oncogenic) effects of IDHmut may depend on the degree to which specific cells express those dioxygenases. And, since a consistent effect of IDHmut is increased genomic methylation, its ability to promote cellular transformation may depend on the patterns of euchromatin and heterochromatin in the cell, which in turn would dictate which regions of the genome are amenable to methylation.
Even among IDHmut cancers, there exists a remarkable preference for certain IDHmut variants according to cancer type. For example, AML has a high proportion of mutations in IDH2, whereas the others are mostly IDH1 (Supplementary Figs S3 and S4). This variant still produces D2HG, but IDH2 is localized to the mitochondrial matrix, while IDH1 is in the cytosol and peroxisomes6. IDHmut glioma is distinct in being the only cancer in the entire body to show enrichment for R132H IDH1 (Supplementary Fig. S4). Indeed, R132H IDH1 is so prevalent in gliomas, compared to other variants that are far more common in other IDHmut cancers, it has become standard practice to routinely screen gliomas with an R132H-specific antibody6. Yet R132H also appears to be the least efficient producer of D2HG, compared to all other IDH1 and IDH2 variants26. This could mean that, in the unique microenvironment of the brain, less D2HG is required for genomic hypermethylation and neoplastic transformation, and that the capacity of glial cells (or glial progenitor cells) to tolerate very high D2HG may be lower than cells elsewhere in the body.
These data also raise the question as to why there is such heterogeneity among CpG methylation sites, particularly in gliomas relative to the other cancers. Even in IDHmut glioma, which shows the greatest extent of CpG hypermethylation compared to its IDHwt counterpart (Fig. 2), not every CpG site is methylated, and not every gene amenable to methylation-induced silencing is suppressed (Supplementary Table S3), or IDHmut tumors would probably never form. To reiterate, the D2HG product of IDHmut only inhibits demethylases, and does not directly cause methylation. In addition to tissue- and cell lineage-specific differences in baseline patterns of euchromatin and heterochromatin, there are other downstream mechanisms that determine which CpG sites are methylated and which ones are not. Such mechanisms include targeting of specific CpG islands, as those regions show the greatest cancer-related variations in IDHmut versus IDHwt methylation (Fig. 3). Prominent among possible mechanisms is cell-specific regulation of DNA methyltransferases, through altered expression, degradation, and complex-mediated targeting of specific CpG sites, which would greatly influence the extent and targets of IDHmut-induced methylation27. This could help explain some interesting features of IDHmut gliomas. For example, it is well known that EGFR mutation and/or amplification, which occurs in nearly 50% of IDHwt gliomas, is nearly mutually exclusive with IDHmut 28. Our data indicate that EGFR is strongly and uniquely hypermethylated in IDHmut gliomas (Supplementary Spreadsheet) (Supplementary Fig. S5a); such silencing would likely diminish any selection pressure toward activating EGFR alterations29,30. Conversely, IDHmut gliomas can have PDGFRA amplification31, and the PDGFRA gene is mostly hypomethylated in this subset of gliomas (Supplementary Fig. S5a)32. Similarly, PROM1 is markedly hypermethylated and downregulated only in IDHmut gliomas, not other IDHmut cancers (Supplementary Fig. S5). PROM1 encodes CD133, a well-known marker of tumor-self-renewal and tumor malignancy33. One of the phenotypic hallmarks of high CD133-expressing tumors is the ability to grow as patient-derived xenografts in immunocompromised mice—an ability that IDHmut gliomas, but not other IDHmut cancers, typically lack6.
Analysis of tumor-specific patterns in IDHmut-associated methylation and mRNA transcription yielded other interesting differences, as well as some profound similarities, among IDHmut malignancies (Figs 4 and 5). All four IDHmut cancers showed relative suppression of genes involved in tissue development (Fig. 5). This has been found to be a consistent effect of IDHmut by multiple laboratories22,34,35,36,37, and helps explain why IDHmut is associated with neoplasia. While neither IDH1 nor IDH2 are classic oncogenes or tumor suppressor genes, and IDHmut is not enough to cause cancer by itself, it may facilitate oncogenesis by extending the window of vulnerability in which additional pro-oncogenic mutations can arise, such as those involving TP53 in IDHmut astrocytomas6. Another common feature of IDHmut cancers is the downregulation of genes associated with immune response (Fig. 5). This is increasingly becoming a recognized feature of IDHmut gliomas38,39,40,41; our data show that this applies to all IDHmut malignancies, with implications for the efficacy of immune-based therapies in this subset of cancers.
Studies that have demonstrated a direct causal link between IDHmut, its D2HG product, and genomic hypermethylation have often done so by inserting one of the IDHmut variants into IDHwt cells, then passaging those cells multiple times and assessing their methylation via the Infinium 450K array. However, the choice of cell model has varied greatly from group to group. Data from two of those models, one expressing R132H IDH1 in differentiated, immortalized human astrocytes5, and the other expressing the same mutation in less differentiated neural progenitor cells22, for the same number of passages apiece, demonstrate that the ability of R132H IDH1 to promote methylation is much greater in less differentiated cells (Fig. 6). This is consistent with the prevailing thinking, based on patient data, that IDHmut occurs at a very early step in oncogenesis, in cells that have not yet fully differentiated42. Whether this window of sensitivity to IDHmut is due to greater amounts of open euchromatin in earlier stages of differentiation, and/or other downstream modulators as discussed above, is not yet clear. But these data do suggest that the choice of cell differentiation state, as well as cell lineage, could greatly impact the results generated in experimental IDHmut research. It may also help explain conflicting results in the literature, including why some investigators have found that IDHmut impairs cancer cell malignancy, whereas others have reported no difference, or even that IDHmut enhances malignant behavior. The same holds for preclinical studies of IDHmut inhibitors, in which some have reported responses to targeted inhibition, whereas others have not.
Finally, these data raise the question as to why certain settings, in which IDHmut showed the greatest amount of hypermethylation, also showed increased hypomethylation relative to matched IDHwt tumors (Figs 2a and 6b). To date, the IDHmut literature has focused on hypermethylation, but these data suggest that there may be another aspect to the effects of IDHmut on the genome, perhaps in the form of increased methylation turnover, or alteration of chromatin structure via IDHmut-induced histone methylation, leading to altered accessibility of DNA for methylation.
There are some limitations to this study, including the comparatively fewer numbers of non-glial cancers with IDHmut. However, our analyses showed CpG sites and mRNA transcripts that were specifically altered in non-glioma IDHmut cancers, but not in gliomas (Figs 4 and 5). Furthermore, well-described aspects of IDHmut biology, including suppression of differentiation and immunity, were recapitulated in all four IDHmut cancers, thus supporting measured interpretation of intergroup comparisons. The data are also not merely a consequence of IDHwt gliomas being hypomethylated relative to the other IDHwt cancers, because a direct comparison between IDHmut gliomas and other IDHmut malignancies demonstrate that IDHmut gliomas methylate a greater proportion of their genome (Supplementary Tables S1, S2, and Fig. S2). The focus of this project was on TCGA cancers, which does not include chondrosarcomas, another cancer in which IDHmut is frequently seen11. Finally, histone methylation data is not present in TCGA, which means that it is not yet clear whether there are also tissue-specific effects of IDHmut on histone modifications.
These data represent a novel comparative analysis of IDHmut cancers, yielding several key insights. While IDHmut does have similar biochemical effects in all tissue types, there is a great deal of heterogeneity regarding the results of that altered biochemistry on the cellular genome and transcriptome. This likely plays a major role in determining whether, and how, IDHmut affects cancer behavior, specifically concerning prognosis and response to IDHmut inhibitors. Furthermore, research involving IDHmut must take into careful consideration the choice of cell model, including the differentiation state of the cell. This study provides a compelling rationale for new lines of investigation into the biology of IDHmut, including studies that simultaneously compare several types of cancer, and explore the impact of cellular context.
Methods
Analysis of DNA methylation
DNA methylation data from Illumina Human Methylation 450K arrays were downloaded from the Cancer Genome Atlas using the TCGAWorkflow and TCGAbiolinks R packages43,44. Legacy data with methylation levels expressed as β-values were downloaded during October 201820. Primary solid tumor data were downloaded for glioma and cholangiocarcinoma. Primary solid tumor and metastatic tumor data were downloaded for melanoma.
CpG genomic region methylation analysis
CpG sites were mapped to regions (North Shelf, North Shore, Island, South Shore, South Shelf, Open Sea) using the IlluminaHumanMethylation450kanno.ilmn12.hg19 R package.
Analysis of gene expression
Transcriptome profiles from the Illumina HiSeq platform were downloaded from The Cancer Genome Atlas using the TCGAWorkflow and TCGAbiolinks R packages43,44. Harmonized data with gene expression levels quantified as counts were downloaded during October 2018. Data were normalized according to the ‘gcContent’ method45. Primary solid tumor data were downloaded for glioma and cholangiocarcinoma. Primary solid tumor and metastatic tumor data were downloaded for melanoma.
Gene ontology analysis
Gene Ontology (GO) analyses were conducted using differentially expressed genes comparing IDHmut versus IDHwt within each cancer type. The cutoff value for the differentially expressed genes was set P < 0.05. The web-based application Metascape (http://www.metascape.org) was used to determine significantly enriched GO biological processes46. The Metascape analysis was performed using the default settings.
Gene set enrichment analysis (GSEA) and network construction
Mean differential gene expression analysis was conducted on IDHmut and IDHwt cancers using the gene set enrichment analysis (GSEA) Desktop v3.0 software (http://www.software.broadinstitute.org/gsea/index.jsp). The number of gene permutations was set to 1,000. The gene-sets were derived from the Broad institute’s Molecular Signatures Database (MSigDB)47. Network data visualization and analysis were performed with Cytoscape v3.5.1 software (http://www.cytoscape.org) and with the EnrichmentMap plugin application48,49. EnrichmentMap threshold settings: false-discovery rate (FDR) q-value < 0.05; overlap coefficient cutoff = 0.5; P-value < 0.05. The functional networks of nodes that represent generic function were determined using the Cytoscape AutoAnnotate application50.
Meta-analysis of astrocyte and neural progenitor cell methylation
For the analysis comparing genome methylation in IDHmut astrocytes and neural progenitor cells data were downloaded from the Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo/) using data sets GSE30339 and GSE949625,22. In brief, immortalized human astrocytes were modified to express the IDH1 (R132H) mutation, cultured for 15 passages, and DNA extracted for methylation analysis. Illumina Methylation 450 K data from these data sets were then analyzed using the same approach described in the analysis of DNA methylation methods section.
Statistical analysis
Differential methylation was evaluated according to Wilcoxon rank sum tests with false discovery rate (FDR) correction. Differential gene expression was evaluated using the edgeR package with FDR correction51. Venn diagrams and heatmaps were used to visualize pairwise comparisons of methylation and gene expression.
Data Availability
The results shown here are in part based upon data generated by the TCGA Research Network (http://cancergenome.nih.gov/). Methylation profiles of astrocytes and neural progenitor cells was downloaded from the GEO (http://www.ncbi.nlm.nih.gov/geo/) using data set GSE30339 and GSE94962.
References
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692, https://doi.org/10.1016/j.cell.2007.01.029 (2007).
Pfeifer, G. P. Defining Driver DNA Methylation Changes in Human Cancer. International journal of molecular sciences 19, https://doi.org/10.3390/ijms19041166 (2018).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744, https://doi.org/10.1038/nature08617 (2009).
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer cell 17, 510–522, https://doi.org/10.1016/j.ccr.2010.03.017 (2010).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483, https://doi.org/10.1038/nature10866 (2012).
Horbinski, C. What do we know about IDH1/2 mutations so far, and how do we use it? Acta Neuropathol 125, 621–636, https://doi.org/10.1007/s00401-013-1106-9 (2013).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med 360, 765–773, https://doi.org/10.1056/NEJMoa0808710 (2009).
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N Engl J Med 361, 1058–1066, https://doi.org/10.1056/NEJMoa0903840 (2009).
Lopez, G. Y. et al. IDH1(R132) mutation identified in one human melanoma metastasis, but not correlated with metastases to the brain. Biochemical and biophysical research communications 398, 585–587, https://doi.org/10.1016/j.bbrc.2010.06.125 (2010).
Borger, D. R. et al. Frequent mutation of isocitrate dehydrogenase (IDH)1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. The oncologist 17, 72–79, https://doi.org/10.1634/theoncologist.2011-0386 (2012).
Amary, M. F. et al. IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. The Journal of pathology 224, 334–343, https://doi.org/10.1002/path.2913 (2011).
Feng, J. H. et al. Prognostic significance of IDH1 mutations in acute myeloid leukemia: a meta-analysis. American journal of blood research 2, 254–264 (2012).
Lugowska, I. et al. IDH1/2 Mutations Predict Shorter Survival in Chondrosarcoma. Journal of Cancer 9, 998–1005, https://doi.org/10.7150/jca.22915 (2018).
Shibata, T., Kokubu, A., Miyamoto, M., Sasajima, Y. & Yamazaki, N. Mutant IDH1 confers an in vivo growth in a melanoma cell line with BRAF mutation. The American journal of pathology 178, 1395–1402, https://doi.org/10.1016/j.ajpath.2010.12.011 (2011).
Wang, P. et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene, 315 (2012).
Goyal, L. et al. Prognosis and Clinicopathologic Features of Patients With Advanced Stage Isocitrate Dehydrogenase (IDH) Mutant and IDH Wild-Type Intrahepatic Cholangiocarcinoma. Oncologist. 20, 1019–1027, 1010.1634/theoncologist.2015-0210. Epub 2015 Aug 1015 (2015).
Chou, W. C. et al. Distinct clinical and biologic characteristics in adult acute myeloid leukemia bearing the isocitrate dehydrogenase 1 mutation. Blood 115, 2749–2754, https://doi.org/10.1182/blood-2009-11-253070 (2010).
Unruh, D. et al. Mutant IDH1 and thrombosis in gliomas. Acta Neuropathol, https://doi.org/10.1007/s00401-016-1620-7 (2016).
Unruh, D. et al. Methylation-dependent Tissue Factor suppression contributes to the reduced malignancy of IDH1 mutant gliomas. Clin Cancer Res 28, 1078–0432 (2018).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic acids research 45, e22, https://doi.org/10.1093/nar/gkw967 (2017).
Bibikova, M. et al. High density DNA methylation array with single CpG site resolution. Genomics 98, 288–295, https://doi.org/10.1016/j.ygeno.2011.07.007 (2011).
Modrek, A. S. et al. Low-Grade Astrocytoma Mutations in IDH1, P53, and ATRX Cooperate to Block Differentiation of Human Neural Stem Cells via Repression of SOX2. Cell reports 21, 1267–1280, https://doi.org/10.1016/j.celrep.2017.10.009 (2017).
Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274, https://doi.org/10.1126/science.1133427 (2006).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812, https://doi.org/10.1126/science.1164382 (2008).
Guilhamon, P. et al. Meta-analysis of IDH-mutant cancers identifies EBF1 as an interaction partner for TET2. Nature communications 4, 2166, https://doi.org/10.1038/ncomms3166 (2013).
Jin, G. et al. 2-hydroxyglutarate production, but not dominant negative function, is conferred by glioma-derived NADP-dependent isocitrate dehydrogenase mutations. PloS one 6, e16812, https://doi.org/10.1371/journal.pone.0016812 (2011).
Jeltsch, A. & Jurkowska, R. Z. Allosteric control of mammalian DNA methyltransferases - a new regulatory paradigm. Nucleic acids research 44, 8556–8575, https://doi.org/10.1093/nar/gkw723 (2016).
Louis, D. N. et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131, 803–820, https://doi.org/10.1007/s00401-016-1545-1 (2016).
Alberti, S., Nutini, M. & Herzenberg, L. A. DNA methylation prevents the amplification of TROP1, a tumor-associated cell surface antigen gene. Proc Natl Acad Sci USA 91, 5833–5837 (1994).
Sun, W. et al. The association between copy number aberration, DNA methylation and gene expression in tumor samples. Nucleic acids research 46, 3009–3018, https://doi.org/10.1093/nar/gky131 (2018).
Phillips, J. J. et al. PDGFRA amplification is common in pediatric and adult high-grade astrocytomas and identifies a poor prognostic group in IDH1 mutant glioblastoma. Brain pathology (Zurich, Switzerland) 23, 565–573, https://doi.org/10.1111/bpa.12043 (2013).
Alvarez, H. et al. Widespread hypomethylation occurs early and synergizes with gene amplification during esophageal carcinogenesis. PLoS genetics 7, e1001356, https://doi.org/10.1371/journal.pgen.1001356 (2011).
Visvader, J. E. & Lindeman, G. J. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nature reviews. Cancer 8, 755–768, https://doi.org/10.1038/nrc2499 (2008).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478, https://doi.org/10.1038/nature10860 (2012).
Sasaki, M. et al. IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature 488, 656–659, https://doi.org/10.1038/nature11323 (2012).
Saha, S. K. et al. Mutant IDH inhibits HNF-4alpha to block hepatocyte differentiation and promote biliary cancer. Nature 513, 110–114, https://doi.org/10.1038/nature13441 (2014).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer cell 18, 553–567, https://doi.org/10.1016/j.ccr.2010.11.015 (2010).
Berghoff, A. S. et al. Correlation of immune phenotype with IDH mutation in diffuse glioma. Neuro-oncology 19, 1460–1468, https://doi.org/10.1093/neuonc/nox054 (2017).
Amankulor, N. M. et al. Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes & development 31, 774–786, https://doi.org/10.1101/gad.294991.116 (2017).
Kohanbash, G. et al. Isocitrate dehydrogenase mutations suppress STAT1 and CD8+ T cell accumulation in gliomas. The Journal of clinical investigation 127, 1425–1437, https://doi.org/10.1172/jci90644 (2017).
Zhang, X. et al. IDH mutant gliomas escape natural killer cell immune surveillance by downregulation of NKG2D ligand expression. Neuro-oncology 18, 1402–1412, https://doi.org/10.1093/neuonc/now061 (2016).
Watanabe, T., Nobusawa, S., Kleihues, P. & Ohgaki, H. IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. The American journal of pathology 174, 1149–1153, https://doi.org/10.2353/ajpath.2009.080958 (2009).
Silva, T. C. et al. TCGA Workflow: Analyze cancer genomics and epigenomics data using Bioconductor packages. F1000 Research 5, 1542, https://doi.org/10.12688/f1000research.8923.2 (2016).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic acids research 44, e71, https://doi.org/10.1093/nar/gkv1507 (2016).
Benjamini, Y. & Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic acids research 40, e72, https://doi.org/10.1093/nar/gks001 (2012).
Tripathi, S. et al. Meta- and Orthogonal Integration of Influenza “OMICs” Data Defines a Role for UBR4 in Virus Budding. Cell host & microbe 18, 723–735, https://doi.org/10.1016/j.chom.2015.11.002 (2015).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics (Oxford, England) 27, 1739–1740, https://doi.org/10.1093/bioinformatics/btr260 (2011).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome research 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PloS one 5, e13984, https://doi.org/10.1371/journal.pone.0013984 (2010).
Kucera, M., Isserlin, R., Arkhangorodsky, A. & Bader, G. D. AutoAnnotate: A Cytoscape app for summarizing networks with semantic annotations. F1000 Research 5, 1717, https://doi.org/10.12688/f1000research.9090.1 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics (Oxford, England) 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
Acknowledgements
C.H. was supported by NIH grants K08CA155764 and R01NS102669. D.U. was supported by F32CA216996. This work was also supported by the Northwestern University SPORE in Brain Cancer P50CA221747 (C.H. and D.M.S.).
Author information
Authors and Affiliations
Contributions
D.U., D.M.S. and C.H. conceptualized and designed the study. M.Z., A.B., M.R.D., A.N.T. helped with bioinformatic analyses. D.U., D.M.S., and C.H. wrote and edited the manuscript. D.M.S. and C.H. supervised all aspects of the study.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Unruh, D., Zewde, M., Buss, A. et al. Methylation and transcription patterns are distinct in IDH mutant gliomas compared to other IDH mutant cancers. Sci Rep 9, 8946 (2019). https://doi.org/10.1038/s41598-019-45346-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45346-1
This article is cited by
-
Translocator protein (18kDA) (TSPO) marks mesenchymal glioblastoma cell populations characterized by elevated numbers of tumor-associated macrophages
Acta Neuropathologica Communications (2023)
-
Meta-analysis of whole-genome gene expression datasets assessing the effects of IDH1 and IDH2 mutations in isogenic disease models
Scientific Reports (2022)
-
Major Features of the 2021 WHO Classification of CNS Tumors
Neurotherapeutics (2022)
-
DNA methylation-based classification of sinonasal tumors
Nature Communications (2022)
-
The implications of IDH mutations for cancer development and therapy
Nature Reviews Clinical Oncology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.