Abstract
Modeling in-vivo protein-DNA binding is not only fundamental for further understanding of the regulatory mechanisms, but also a challenging task in computational biology. Deep-learning based methods have succeed in modeling in-vivo protein-DNA binding, but they often (1) follow the fully supervised learning framework and overlook the weakly supervised information of genomic sequences that a bound DNA sequence may has multiple TFBS(s), and, (2) use one-hot encoding to encode DNA sequences and ignore the dependencies among nucleotides. In this paper, we propose a weakly supervised framework, which combines multiple-instance learning with a hybrid deep neural network and uses k-mer encoding to transform DNA sequences, for modeling in-vivo protein-DNA binding. Firstly, this framework segments sequences into multiple overlapping instances using a sliding window, and then encodes all instances into image-like inputs of high-order dependencies using k-mer encoding. Secondly, it separately computes a score for all instances in the same bag using a hybrid deep neural network that integrates convolutional and recurrent neural networks. Finally, it integrates the predicted values of all instances as the final prediction of this bag using the Noisy-and method. The experimental results on in-vivo datasets demonstrate the superior performance of the proposed framework. In addition, we also explore the performance of the proposed framework when using k-mer encoding, and demonstrate the performance of the Noisy-and method by comparing it with other fusion methods, and find that adding recurrent layers can improve the performance of the proposed framework.
Similar content being viewed by others
Introduction
Transcription factors can modulate gene expression by binding to specific DNA regions, which are known as transcription factor binding site (TFBS). Modeling in-vivo TF-DNA binding, also called motif discovery, is a fundamental yet challenging step towards deciphering transcriptional regulatory networks1,2.
In the past decades, the introduction of high-throughput sequencing technologies, especially ChIP-seq.3, dramatically increases the amount and spatial resolution of available data, which is helpful for the in-depth study of in-vivo protein-DNA binding. However, DNA sequences directly extracted from ChIP-seq cannot precisely represent TFBS since the outputs of such experiments contain a lot of noisy4. Thus lots of methods have been developed for precisely predicting protein-DNA binding sites, including conventional algorithms5,6,7,8,9 and deep-learning based methods10,11,12. Not surprisingly, deep-learning based methods are better than conventional algorithms at modeling protein-DNA binding. DeepBind10 and DeepSea11 were two famous deep-learning based methods, which used convolutional neural network (CNN) to model the binding preference of DNA-proteins with a superior performance over conventional methods. DanQ12 designed a hybrid deep neural network to quantify the function of DNA sequences, which first used a convolutional layer to detect regulatory motif features from DNA sequences, and subsequently employed a bi-directional recurrent layer to capture long-term dependencies between motif features. Soon after, a number of deep-learning based methods are proposed for modeling in-vivo protein-DNA binding13,14,15,16,17.
Although deep-learning based methods have achieved remarkable performance on modeling in-vivo protein-DNA binding, they usually overlook the weakly supervised information of genomic sequences that a bound DNA sequence may have multiple TFBS(s). In consideration of this information, Gao et al.18 developed a multiple-instance learning (MIL) based algorithm, which combines MIL with TeamD19, for modeling protein-DNA binding, and recently Zhang et al.20 also developed a weakly supervised convolutional neural network (WSCNN), which combines MIL with CNN, for modeling protein-DNA binding. Moreover, they are inclined to use one-hot encoding to encode DNA sequences, which means that it only considers the independent relationship among nucleotides. However, recent studies have shown that taking into consideration the high-order dependencies among nucleotides can improve the performance of modeling protein-DNA binding21,22,23. In consideration of this information, Zhou et al.24 evaluated DNA-binding specificities based on mononucleotide (1-mer), dinucleotide (2-mer), and trinucleotide (3-mer) identity, and stated that 2-mer and 3-mer may contain implicit DNA shape information and partially capture the effect of the DNA shape variation on binding. Zhang et al.25 proposed a high-order convolutional neural network, which first used k-mer encoding to transform DNA sequences into image-like inputs of high-order dependencies, and then applied CNN to extract motif features from these inputs.
Inspired by the above observation, we extend our previous work WSCNN from three aspects in this paper. (1) WSCNN mainly employed CNN to learn motif features from DNA sequences, and did not take into consideration the long-term dependencies between motif features. In the weakly supervised framework, therefore we add a bi-directional recurrent layer after the convolutional layer to capture the forward and backward long-term dependencies between motif features. (2) WSCNN attempted to use four fusion methods to fuse the predicted values of all instances in a bag, and then selected the best one of them as the final prediction. However, it is inconvenient for user to decide which one is better, so they have to try the four fusion methods one by one. Therefore we offer a better and more robust fusion method Noisy-and26 to replace them. (3) WSCNN, like other deep-learning based methods, used one-hot encoding to transform DNA sequences into image-like inputs. However, the relationship between nucleotides is not independent in practice. Therefore we use k-mer encoding to transform DNA sequences into image-like inputs of high-order dependencies, and explore the performance of the proposed framework when using dinucleotide (2-mer) and trinucleotide (3-mer) as inputs in the weakly supervised framework. In summary, the proposed framework firstly use the concepts of MIL to segment DNA sequences into multiple instances, and adopt k-mer encoding to transform sequences into image-like inputs of high-order dependencies, and then design a hybrid neural network to compute a score for all instances, and finally employ the Noisy-and method to fuse the predicted values of all instances as the final prediction of a bag. We conducted a lot of comparative experiments on in-vivo datasets to show that our proposed framework outperforms other competing methods. Besides, we also show the performance gain of the proposed framework when using k-mer encoding, and compare the performance of the Noisy-and method with other fusion methods, and demonstrate the effectivness of adding recurrent layers.
The rest of the paper is organized as follows. We give a detailed description of the proposed framework, and introduce the fusion method Noisy-and in Section II. We give a detailed analysis of the experimental results, and discuss the hyper-parameter settings in Section III.
Methods
In this section, we give a detailed description of the proposed framework for modeling in-vivo protein-DNA binding. Actually, the task can be thought of as a binary classification problem that separates positive sequences (bound) from negative sequences (non-bound). The output of the network is a probability (a scalar in [0, 1]) distribution over two labels (1/0), since a binary classification problem can be addressed also through a binary output (1/0). This framework includes three stages in general: data processing, model designing, and results merging.
Data processing
Segmentation
Considering the weakly supervised information of DNA sequences, thus it is reasonable to use the concepts of MIL to deal with DNA sequences. Therefore we divided them into multiple overlapping instances following the works18,20, which ensures that (1) the weakly supervised information can be retained, and that (2) a large amount of instances containing TFBS are generated. This method is defined as a sliding window of length c, which divides DNA sequences of length l into multiple overlapping instances by a stride s. A bag is composed of all possible instances in the same sequence, and the number of instances in this bag is \(\lceil (l-c)/s\rceil +1\), where s and c are two hyper-parameters that need to be tuned by cross-validation. If (l − c) is not a multiple of s, we pad ‘0’ at the end of DNA sequences.
K-mer encoding
After segmenting DNA sequences, all instances should be transformed into image-like inputs that can be handled by CNN. One-hot encoding is a commonly-used method in deep-learning based methods, but it ignores high-order dependencies among nucleotides. In order to capture the dependencies, therefore we use the k-mer encoding method25 to transform all instances into image-like matrices of high-order dependencies. This method can be implemented according to (1):
where \(i\in [1,\,c-k+1]\), and c denotes the length of instances, and xi denotes a possible character from {A, C, G, T}, and Xi,j denotes a matrix generated by using k-mer encoding. According to the equation, we can find that one-hot encoding is a special case of k-mer encoding when k is set to 1. For example, 1-mer encoding: each nucleotide is mapped into a vector of size 4 (A → [1, 0, 0, 0]T, C → [0, 1, 0, 0]T, G → [0, 0, 1, 0]T, and T → [0, 0, 0, 1]T); 2-mer encoding: taking into consideration the dependencies between two adjacent nucleotides, and each dinucleotide is mapped into a vector of size 16 (AA → [1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0]T, ..., TT → [0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1]T); 3-mer encoding: taking into account the dependencies among three adjacent nucleotides, and each trinucleotide is mapped into a vector of size 64 (AAA → [1, 0, 0, 0, ..., 0, 0, 0, 0, 0], ...,TTT → [0, 0, 0, 0, ..., 0, 0, 0, 0, 0, 1]).
A graphical illustration of data processing when k = 1 is shown in Fig. 1, where l = 10, c = 8, s = 1, and the red dashed box denotes a sliding window of length c = 8. Through this stage, DNA sequences can be encoded into image-like inputs that can be easily handled by CNN.
In the implementation code, each instance is firstly encoded into a tensor of shape 1 × 4k × 1 × (c − k + 1) (batchsize × channel × height × width), and then all instances of a bag can be concatenated along the height axis. Therefore a bag can be represented by a tensor of shape 1 × 4k × n × (c − k + 1), where n is the number of instances per bag (\(n=\lceil (l-c)/s\rceil +1\)). Details of implementation can refer to our open source code.
Model designing
Considering the spatial and sequential characteristics of DNA sequences, we design a hybrid deep neural network, which integrates convolutional and recurrent neural networks in this stage. Convolutional neural network (CNN) is a special version of article neural network (ANN)27,28,29, which adopts a weight-sharing strategy to capture local patterns in data such as DNA sequences. Recurrent neural network (RNN) is another variant of ANN where connections between neurons form a directed graph. Unlike CNN, RNN can use its internal state (memory) to exhibit dynamic temporal or spatial behavior. In the designed model, the convolution layer is used to capture motif features, while the recurrent layer is used to capture long-term dependencies between the motif features. The model is arranged in this order: a convolutional layer → a max-pooling layer → a dropout layer → a bi-directional recurrent layer → a dropout layer → a softmax layer.
Convolutional layer
This layer is used to capture motif features, which can be thought of as a motif scanner to compute a score for all potential motifs, and often followed by a rectified linear unit30 (ReLU) layer. The early work13 has explored the performance of using different number of convolutional kernels, and found that adding more kernels can significantly improve performance. Thus the number of kernels was set to a fixed value 16 in the proposed framework.
Max-pooling layer
Both DeepBind and DeepSea used a global max-pooling layer to pick out the maximum response of the whole sequence, while our deep model uses a max-pooling layer of a certain size (1, 8) to keep the local best values of the whole sequence.
Dropout layer
Dropout strategy31 is a widely-used regularization technique for reducing overfitting in deep neural networks by preventing complex co-adaptations on data, which randomly sets the outputs of the previous layer to zero with a dropout ratio. The dropout ratio is a hyper-parameter that was investigated by cross-validation in the experiments.
Recurrent layer
In order to capture the forward and backward long-term dependencies between the motif features, a bi-directional recurrent layer composed of long short-term memory (LSTM) units32 is used. A LSTM unit usually consists of a cell, an input gate, a forget gate and an output gate, where the cell remembers values over arbitrary time intervals and the three gates regulate information flows into and out of the cell. In this paper, we did not use a fully-connected layer to follow this layer, as this will result into worse performance. The number of neurons in this layer was set to 32, so the output size of this layer is 64.
Softmax layer
In order to get a probability distribution over two labels which separately represent bound or non-bound sequences, a softmax layer is used in this model. It is composed of two neurons, each of which is densely connected with the previous layer and computes a probability.
Results merging
MIL is commonly based on an assumption that a bag is labeled as positive if there is at least one instance that contains TFBS, and is labeled as negative if there are no any instances that contain TFBS. Therefore the Max function is frequently used as the fusion function in MIL. But Max only focuses on the most informative instance and overlooks other instances that may contain useful information. Therefore WSCNN used three additional fusion methods (Linear Regression, Average, and Top-Bottom Instances33) to utilize all instances that may contain useful information. However, both Average and Linear Regression take advantage of all information, inevitably containing useless information, and Top-Bottom Instances needs to manually determine the number of the highest and lowest scoring instances. Moreover, how to effectively take advantage of abundant positive instances is also a key point. To solve the above problems, we find a better and more elegant fusion method, named Noisy-and26, which is based on a different assumption that a bag is labeled as positive if the number of positive instances in the bag exceeds a threshold. This method is defined as follows:
where Pi,j denotes the score of the j-th instance at the i-th bag, and ni denotes the number of instances in the i-th bag, and \({\bar{P}}_{i}\) denotes the average score over n instances in the i-th bag. Noisy-and is designed to activate a bag level probability Pi once the mean of the instance level probabilities \({\bar{P}}_{i}\) exceeds a certain threshold. a is a fixed hyper-parameter that controls the slope of Noisy-and. bi represents an adaptable soft threshold for each class i and needs to be learned during training. σ(a(1 − bi)) and σ(−abi) are included to normalized Pi to [0, 1] for bi in [0, 1] and a > 0.
Through this stage, the predicted values of all instances in a bag are fused to yield a final prediction (probability) over ‘bound’ and ‘non-bound’ labels.
In summary, the proposed framework is arranged in this order: data processing (segmentation + k-mer encoding) → a convolutional layer → a max-pooling layer → a dropout layer → a bi-directional recurrent layer → a dropout layer → a softmax layer → a fusion layer. A graphical illustration of the proposed framework is shown in Fig. 2.
Results
For brevity, the proposed framework is named as WSCNNLSTM. In this section, the performance of WSCNNLSTM is systematically evaluated by comparing it with other deep-learning based algorithms. We carried out a series of experiments on in-vivo ChIP-Seq datasets to show that the overall performance of WSCNNLSTM is superior to the competing methods.
Experimental setup
Data preparation
We collected 50 public ChIP-seq datasets from the HaibTfbs group, which stems from three different cell lines (Gm12878, H1hesc, and K562). For each public dataset, an average number of ~15000 top ranking sequences were chosen as the positive data where each sequence is composed of 200 bps, and the corresponding negative data was generated by matching the repeat fraction, length and GC content of the positive ones following the work9, and the number of the negative data is 1~3 times more than the positive data. Moreover, 1/8 of the training data were randomly sampled as the validation data during training.
Competing methods
To better evaluate the performance of WSCNNLSTM, we constructed three deep-learning based models, which are similar to DeepBind10, DanQ12, and WSCNN20, respectively.
Model 1: This model is a single-instance learning (SIL) based method, and has the similar architecture to DeepBind. It is arranged in this order: data processing (one-hot encoding) → a convolutional layer → a global max-pooling layer → a fully connected layer → a dropout layer → a softmax layer.
Model 2: This model is a single-instance learning (SIL) based method, and has the similar architecture to DanQ. It is arranged in this order: data processing (one-hot encoding) → a convolutional layer → a max-pooling layer → a dropout layer → a bi-directional recurrent layer → a dropout layer → a softmax layer.
Model 3: This model is a multiple-instance learning (MIL) based method, and has the similar architecture to WSCNN. It is arranged in this order: data processing (segmentation + one-hot encoding) → a convolutional layer → a global max-pooling layer → a fully connected layer → a dropout layer → a softmax layer → a fusion layer.
Evaluation metrics
To comprehensively assess the performance of WSCNNLSTM, we adopted three standard evaluation metrics in this paper, including area under receiver operating characteristic curve (ROC AUC), area under precision-recall curve (PR AUC), and F1-score, which are widely used in machine learning and motif discovery34,35,36,37,38,39,40,41,42,43,44,45,46.
ROC AUC47 and PR AUC two commonly-used metrics in which PR AUC is often used under the situation of imbalanced data. Since PR AUC needs not to consider the number of true negative samples, thus it is less prone to influenced by the class imbalance than the ROC AUC metric is12,48.
F1-score is a solid metric to measure the classification performance of the classifier, which simultaneously takes into consideration the precision and the recall when computing the score of a test49.
Hyper-parameter settings
We implemented WSCNNLSTM and the competing methods by Keras with the tensorflow backend, which are freely available at: https://github.com/turningpoint1988/WSCNNLSTM. The parameters in the deep-learning based methods were initialized by Glorot uniform initializer50, and optimized by AdaDelta algorithm51 with a mini-batchsize of 300. For some sensitive hyper-parameters (i.e., dropout ratio, Momentum in AdaDelta, Delta in AdaDelta), we selected the best configuration using a grid-search strategy. Epochs of training were set to 60, where after each epoch of training, the accuracy of the validation set was assessed and monitored, and the model with the best accuracy in the validation set was saved. The instance length c and segmentation stride s were set to 120 and 10, since WSCNN has stated that the two hyper-parameters has little effect on the main conclusion. The hyper-parameter a in Noisy-and was set to 7.5 following the work26. The hyper-parameter settings of all the deep-learning based methods in this paper are detailed in Table 1.
Performance comparison on in-vivo data
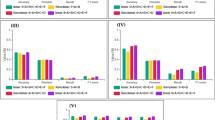
A comparison of WSCNNLSTM and the competing methods on 50 in-vivo ChIP-seq datasets is shown in Fig. 3 and Supplementary Tables 1, 2, 3. Evaluation is done with three-fold cross validation, and prediction performance is measured by the PR AUC, ROC AUC and F1-score metrics.
A comparison of WSCNNLSTM and the competing methods on in-vivo data, where the first column corresponds to a comparison of WSCNNLSTM and DeepBind under the ROC AUC, PR AUC and F1-score metrics, and the second column corresponds to a comparison of WSCNNLSTM and DanQ, and the third column corresponds to a comparison of WSCNNLSTM and WSCNN.
Figure 3a,d,g show a superior performance of WSCNNLSTM over DeepBind under the ROC AUC, PR AUC, and F1-score metrics. As WSCNNLSTM combines MIL for learning the weakly supervised information of sequences with RNN for capturing the forward and backward long-term dependencies between the motif features, so it outperforms DeepBind by a large margin. Figure 3b,e,h show a superior performance of WSCNNLSTM over DanQ under the ROC AUC, PR AUC, and F1-score metrics, demonstrating the advantages of allowing for the weakly supervised information of sequences. Figure 3c,e,i show a superior performance of WSCNNLSTM over WSCNN under the ROC AUC, PR AUC, and F1-score metrics, demonstrating the benefits of allowing for the forward and backward long-term dependencies between the motif features. Figure 4 records the average values on ROC AUC, PR AUC, and F1-score, which also shows the consistent conclusion that WSCNNLSTM outperforms the three competing methods. In summary, the above results show that the overall performance of WSCNNLSTM is better than DeepBind, DanQ, and WSCNN.
The k-mer encoding method can significantly improve the performance of modeling in-vivo protein-DNA binding
Unlike one-hot encoding, the k-mer encoding method can take into consideration the high-order dependencies among nucleotides, which may improve the performance of modeling in-vivo protein-DNA binding. As stated by Zhou et al.24, 2-mer (dinucleotide) and 3-mer (trinucleotide) may contain implicit DNA shape information and partially capture the effect of the DNA shape variation on binding. Moreover, Zhang et al.25 has stated that the number of learnable parameters will grow exponentially with the increase of k. Therefore, in order to make a trade-off between performance and computational complexity, we transformed DNA sequences into matrices that consist of 2-mer or 3-mer by setting k to 2 or 3 in k-mer encoding.
To test the performance of k-mer encoding in the weakly supervised framework, we carried out some comparative experiments on 23 ChIP-seq datasets from the Gm12878 cell line, and the detailed results are shown in Supplementary Tables 4, 5.
Figure 5 shows a comparison of one-hot encoding and k-mer encoding in WSCNN under the ROC AUC, PR AUC, and F1-score metrics. We can find that the performance of 2-mer and 3-mer encoding is much better than that of one-hot encoding, demonstrating the effectiveness of k-mer encoding for modeling in-vivo protein-DNA binding. Figure 6 shows a comparison of one-hot encoding and k-mer encoding in WSCNNLSTM under the ROC AUC, PR AUC, and F1-score metrics. We can also find the same trend that the performance of 2-mer and 3-mer encoding is much better. Figure 7 records the average values on ROC AUC, PR AUC, and F1-score, which concludes that k-mer encoding is superior to one-hot encoding. Moreover, we find that the performance of WSCNN and WSCNNLSTM is improved with the increase of k. We think that the reason of the good performance may lie in: it explicitly considers the high-order dependencies among nucleotides (which contains implicit DNA shape information).
A comparison of WSCNN when using one-hot, 2-mer, and 3-mer encoding, where the first row corresponds to a comparison of WSCNN when using one-hot and 2-mer encoding under the ROC AUC, PR AUC and F1-score metrics, and the second row corresponds to a comparison of WSCNN when using one-hot and 3-mer encoding.
A comparison of WSCNNLSTM when using one-hot, 2-mer, and 3-mer encoding, where the first row corresponds to a comparison of WSCNNLSTM when using one-hot and 2-mer encoding under the ROC AUC, PR AUC and F1-score metrics, and the second row corresponds to a comparison of WSCNNLSTM when using one-hot and 3-mer encoding.
The Noisy-and function is a better fusion method in the weakly supervised framework
WSCNN employed four fusion methods (Max, Linear Regression, Average and Top-Bottom Instances) to fuse the predicted values of all instances in a bag, and then selected the best one as the final prediction. However, Max only focuses on the most informative instance and overlooks other instances that may contain useful information, and both Average and Linear Regression take advantage of all information, inevitably containing useless information, and Top-Bottom Instances needs to manually determine the number of the highest and lowest scoring instances. Moreover, how to effectively take advantage of abundant positive instances is also a key point. Thus we adopt a better and more elegant fusion method, named Noisy-and, in this paper.
To test the performance of Noisy-and in the weakly supervised framework, we carried out some comparative experiments on 23 ChIP-seq datasets from the Gm12878 cell line, and the detailed results are shown in Supplementary Tables 6, 7.
Figure 8 shows a comparison of Noisy-and and Max, Average in WSCNN under the ROC AUC, PR AUC, and F1-score metrics. We can find that the performance of Noisy-and is much better than that of Max and Average. Figure 9 shows a comparison of Noisy-and and Max, Average in WSCNNLSTM under the ROC AUC, PR AUC, and F1-score metrics. We find that the performance of Noisy-and is also much better than that of Max and Average. Figure 10 records the average values on ROC AUC, PR AUC, and F1-score, which concludes the same conclusion. We think that the reason of good performance may result from: the weakly supervised framework (WSCNN, WSCNNLSTM) segments DNA sequences into multiple overlapping instances, producing enough positive instances, which is in accordance with the assumption of Noisy-and that a bag is labeled as positive if the number of positive instances in the bag exceeds a threshold.
A comparison of WSCNNLSTM when using Max, Average, and Noisy-and functions, where the first row corresponds to a comparison of WSCNNLSTM when using Max and Noisy-and under the ROC AUC, PR AUC and F1-score metrics, and the second row corresponds to a comparison of WSCNNLSTM when using Average and Noisy-and.
Adding recurrent layers can improve the performance of the proposed framework
Convolutional layers are often used as motif scanners to capture motifs in the task of modeling in-vivo protein-DNA binding, but they ignores the long-term dependencies between motifs. Therefore we add a bi-directional recurrent layer after the convolutional layer in the weakly supervised framework, just like DanQ does, to capture the forward and backward long-term dependencies between motifs.
Figure 11 shows a comparison of the methods with RNN (WSCNNLSTM, DanQ) and the ones without RNN (WSCNN, DeepBind) under the ROC AUC, PR AUC and F1-score metrics. The above results show that the models with RNN outperform the ones without RNN by a health margin, demonstrating the effectiveness of allowing for the forward and backward long-term relationship between motif features.
Conclusions
In this paper, we propose a weakly supervised framework, which combines multiple-instance-learning with a hybrid deep neural network, for modeling in-vivo protein-DNA binding. The proposed framework contains three stages: data processing, model designing, and results merging, where the first stage contains the segmentation process and k-mer encoding, and the second stage contains a hybrid deep neural network, and the last stage contains the Noisy-and fusion method. The experimental results on in-vivo ChIP-seq datasets show that our proposed framework performs better than the competing methods. In addition, we also explore the performance of the proposed framework when using k-mer encoding and show that the k-mer encoding can significantly improve the performance of modeling in-vivo protein-DNA binding, and demonstrate that the Noisy-and function is a better fusion method in the weakly supervised framework.
From the above results, we find that the performance of WSCNN and WSCNNLSTM is improved with the increase of k. However, the big k will bring out exponentially growing learnable parameters, and the performance of models may be degenerated when k reaches a certain value. Therefore, we will explore the performance of k-mer encoding with a big k value, and propose a corresponding solution to it in the future works.
Data Availability
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
References
Elnitski, L., Jin, V. X., Farnham, P. J. & Jones, S. J. M. Locating mammalian transcription factor binding sites: a survey of computational and experimental techniques. Genome Research 16, 1455–1464 (2006).
Orenstein, Y. & Shamir, R. A comparative analysis of transcription factor binding models learned from PBM, HT-SELEX and ChIP data. Nucleic acids research 42, e63–e63 (2014).
Furey, T. S. ChIP–seq and beyond: new and improved methodologies to detect and characterize protein–DNA interactions. Nature Reviews Genetics 13, 840–852 (2012).
Jothi, R., Cuddapah, S., Barski, A., Cui, K. & Zhao, K. Genome-wide identification of in vivo protein–DNA binding sites from ChIP-Seq data. Nucleic acids research 36, 5221–5231 (2008).
Stormo, G. D. Consensus patterns in DNA. Methods in enzymology 183, 211–221 (1990).
Stormo, G. D. DNA binding sites: representation and discovery. Bioinformatics 16, 16–23 (2000).
Zhao, X., Huang, H. & Speed, T. P. Finding short DNA motifs using permuted Markov models. Journal of Computational Biology 12, 894–906 (2005).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Ghandi, M. et al. gkmSVM: an R package for gapped-kmer SVM. Bioinformatics 32, 2205–2207 (2016).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA-and RNA-binding proteins by deep learning. Nature biotechnology 33, 831–838 (2015).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nature methods 12, 931–934 (2015).
Quang, D. & Xie, X. DanQ: a hybrid convolutional and recurrent deep neural network for quantifying the function of DNA sequences. Nucleic acids research 44, e107–e107 (2016).
Zeng, H., Edwards, M. D., Liu, G. & Gifford, D. K. Convolutional neural network architectures for predicting DNA–protein binding. Bioinformatics 32, i121–i127 (2016).
Kelley, D. R., Snoek, J. & Rinn, J. L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome research 26, 990–999 (2016).
Hassanzadeh, H. R. & Wang, M. D. DeeperBind: Enhancing prediction of sequence specificities of DNA binding proteins. In IEEE International Conference on Bioinformatics and Biomedicine. 178–183 (2017).
Shrikumar, A., Greenside, P. & Kundaje, A. Reverse-complement parameter sharing improves deep learning models for genomics. bioRxiv, 103663 (2017).
Bosco, G. L. & Gangi, M. A. D. Deep Learning Architectures for DNA Sequence Classification. International Workshop on Fuzzy Logic and Applications, 162–171 (2016).
Gao, Z. & Ruan, J. Computational modeling of in vivo and in vitro protein-DNA interactions by multiple instance learning. Bioinformatics 33(14), 2097–2105 (2017).
Annala, M., Laurila, K., Lähdesmäki, H. & Nykter, M. A linear model for transcription factor binding affinity prediction in protein binding microarrays. PloS one 6, e20059 (2011).
Zhang, Q., Zhu, L., Bao, W. & Huang, D. S. Weakly supervised Convolutional Neural Network Architecture for Predicting Protein-DNA Binding. IEEE/ACM Transactions on Computational Biology and Bioinformatics PP, 1–1 (2018).
Keilwagen, J. & Grau, J. Varying levels of complexity in transcription factor binding motifs. Nucleic Acids Research 43, e119 (2015).
Siebert, M. & Söding, J. Bayesian Markov models consistently outperform PWMs at predicting motifs in nucleotide sequences. Nucleic Acids Research 44, 6055–6069 (2016).
Eggeling, R., Roos, T., Myllymäki, P. & Grosse, I. Inferring intra-motif dependencies of DNA binding sites from ChIP-seq data. Bmc Bioinformatics 16, 1–15 (2015).
Zhou, T. et al. Quantitative modeling of transcription factor binding specificities using DNA shape. Proceedings of the National Academy of Sciences 112(15), 4654–4659 (2015).
Zhang, Q., Zhu, L. & Huang, D. S. High-Order Convolutional Neural Network Architecture for Predicting DNA-Protein Binding Sites. IEEE/ACM Transactions on Computational Biology and Bioinformatics 1, 1–1 (2018).
Kraus, O. Z., Ba, J. L. & Frey, B. J. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 32, i52–i59 (2016).
Huang, D. S. Systematic theory of neural networks for pattern recognition. Publishing House of Electronic Industry of China, Beijing 201 (1996).
Huang, D. S. Radial basis probabilistic neural networks: model and application. International Journal of Pattern Recognition and Artificial Intelligence 13, 1083–1101 (1999).
Huang, D. S. & Du, J. X. A Constructive Hybrid Structure Optimization Methodology for Radial Basis Probabilistic Neural Networks. IEEE Transactions on Neural Networks 19, 2099–2115 (2008).
Glorot, X., Bordes, A. & Bengio, Y. Deep sparse rectifier neural networks. In Proceedings of the Fourteenth International Conference on Artificial Intelligence and Statistics, 315–323 (2011).
Srivastava, N., Hinton, G. E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. Journal of machine learning research 15, 1929–1958 (2014).
Hochreiter, S. & Schmidhuber, J. Long Short-Term Memory. Neural Computation 9, 1735–1780 (1997).
Durand, T., Thome, N. & Cord, M. Weldon: Weakly supervised learning of deep convolutional neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 4743–4752 (2016).
Deng, S. P., Zhu, L. & Huang, D. S. Predicting hub genes associated with cervical cancer through gene co-expression networks. (IEEE Computer Society Press, 2016).
Weirauch, M. T. et al. Evaluation of methods for modeling transcription factor sequence specificity. Nature biotechnology 31, 126 (2013).
Huang, D. S. & Jiang, W. A general CPL-AdS methodology for fixing dynamic parameters in dual environments. IEEE Transactions on Systems Man & Cybernetics Part B 42, 1489–1500 (2012).
Yu, H.-J. & Huang, D. S. Normalized feature vectors: a novel alignment-free sequence comparison method based on the numbers of adjacent amino acids. IEEE/ACM Transactions on Computational Biology and Bioinformatics (TCBB) 10, 457–467 (2013).
Zhu, L., You, Z. H., Huang, D. S. & Wang, B. t-LSE: A Novel Robust Geometric Approach for Modeling Protein-Protein Interaction Networks. Plos One 8, e58368 (2013).
Huang, D. S. et al. Prediction of protein-protein interactions based on protein-protein correlation using least squares regression. Curr Protein Pept Sci 15, 553–560 (2014).
Zhu, L., Deng, S.-P. & Huang, D. S. A Two-Stage Geometric Method for Pruning Unreliable Links in Protein-Protein. Networks. NanoBioscience, IEEE Transactions on 14, 528–534 (2015).
Zhu, L., Guo, W. L., Deng, S. P. & Huang, D. S. ChIP-PIT: Enhancing the Analysis of ChIP-Seq Data Using Convex-Relaxed Pair-Wise Interaction Tensor Decomposition. IEEE/ACM Transactions on Computational Biology and Bioinformatics 13, 55–63 (2016).
Zheng, C. H., Huang, D. S., Zhang, L. & Kong, X. Z. Tumor clustering using nonnegative matrix factorization with gene selection. IEEE Transactions on Information Technology in Biomedicine A Publication of the IEEE Engineering in Medicine & Biology Society 13, 599–607 (2009).
Huang, D. S. & Zheng, C. H. Independent component analysis-based penalized discriminant method for tumor classification using gene expression data. Bioinformatics 22, 1855–1862 (2006).
Deng, S. P. & Huang, D. S. In IEEE International Conference on Bioinformatics and Biomedicine. 29–34.
Zheng, C.-H., Zhang, L., Ng, V. T.-Y., Shiu, S. C.-K. & Huang, D. S. Molecular pattern discovery based on penalized matrix decomposition. Computational Biology and Bioinformatics, IEEE/ACM Transactions on 8, 1592–1603 (2011).
Deng, S. P., Zhu, L. & Huang, D. S. Mining the bladder cancer-associated genes by an integrated strategy for the construction and analysis of differential co-expression networks. Bmc Genomics 16, S4 (2015).
Fawcett, T. An introduction to ROC analysis. Pattern recognition letters 27, 861–874 (2006).
Davis, J. & Goadrich, M. The relationship between Precision-Recall and ROC curves. In ICML ‘06: Proceedings of the International Conference on Machine Learning, New York, Ny, Usa, 233–240 (2006).
Sasaki, Y. The truth of the F-measure. Teach Tutor mater 1(5), 1–5 (2007).
Glorot, X. & Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. Journal of Machine Learning Research 9, 249–256 (2010).
Zeiler, M. D. ADADELTA: An Adaptive Learning Rate Method. Computer Science (2012).
Acknowledgements
This work was supported by the grants of the National Science Foundation of China, Nos 61672382, 61732012, 61772370, 61520106006, 61772357, 31571364, 61532008, U1611265, 61702371, and 61572447, China Postdoctoral Science Foundation Grant, Nos 2017M611619 and 2016M601646, and supported by “BAGUI Scholar” Program of Guangxi Province of China.
Author information
Authors and Affiliations
Contributions
Q.H.Z. and Z.S. designed the method. Q.H.Z. conducted the experiments and wrote the main manuscript text. D.S.H. supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Q., Shen, Z. & Huang, DS. Modeling in-vivo protein-DNA binding by combining multiple-instance learning with a hybrid deep neural network. Sci Rep 9, 8484 (2019). https://doi.org/10.1038/s41598-019-44966-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44966-x
This article is cited by
-
KDeep: a new memory-efficient data extraction method for accurately predicting DNA/RNA transcription factor binding sites
Journal of Translational Medicine (2023)
-
Human DNA/RNA motif mining using deep-learning methods: a scoping review
Network Modeling Analysis in Health Informatics and Bioinformatics (2023)
-
DNA sequence classification based on MLP with PILAE algorithm
Soft Computing (2021)
-
Prediction Model of Organic Molecular Absorption Energies based on Deep Learning trained by Chaos-enhanced Accelerated Evolutionary algorithm
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.