Abstract
The Dallol geothermal area in the northern part of the Danakil Depression (up to 124–155 meter below sea level) is deemed one of the most extreme environments on Earth. The area is notable for being part of the Afar Depression, an incipient seafloor-spreading center located at the triple junction, between Nubian, Somali and Arabian plates, and for hosting environments at the very edge of natural physical-chemical extremities. The northern part of the Danakil Depression is dominated by the Assale salt plain (an accumulation of marine evaporite deposits) and hosts the Dallol volcano. Here, the interaction between the evaporitic deposit and the volcanisms have created the unique Dallol hot springs, which are highly acidic (pH ~ 0) and saline (saturation) with maximum temperatures ranging between 90 and 109 °C. Here we report for the first time evidence of life existing with these hot springs using a combination of morphological and molecular analyses. Ultra-small structures are shown to be entombed within mineral deposits, which are identified as members of the Order Nanohaloarchaea. The results from this study suggest the microorganisms can survive, and potential live, within this extreme environment, which has implications for understanding the limits of habitability on Earth and on (early) Mars.
Similar content being viewed by others
Introduction
The study of environmental limits of life provides useful information for assessing the limits of habitability both on Earth and elsewhere in the Solar System1,2. Understanding, and defining these limits, using extreme terrestrial environments and Earth analogues sites3,4 is therefore a crucial step in selecting sites for future life detection missions.
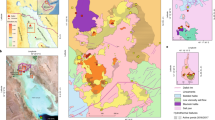
The Dallol geothermal area (14°14′21″N; 40°17′55″E) is located in the northern part of the Danakil Depression5. This is a narrow lowland salt plain (up to 124 m below sea level) running inland, quasi-parallel to the coast of the Red Sea (Fig. 1a), which formed when part of the Red Sea was isolated during the Pleistocene.
The area is located at the northern segment of the Afar triple junction and is characterized by an attenuated continental crust, which is less than 15 km thick with shallow (3–5 km deep) magma chambers beneath its axial zones6,7. The Dallol dome-shaped deposits formed as a result of the interaction between the evaporitic deposits and the shallow magma intrusions7,8.
The hydrothermalism of the Dallol area was possibly generated by a phreatic explosion in 1926. Since then volcanic episodes, seismicity and hydrothermal activity have been reported in the area5. As a consequence of geothermal activity7,9 several minerals, for example, pyrolusite, chlorargyrite, wurtzite and halite, precipitate forming colourful chimneys (Fig. 1b). It has been postulated that microbial activity does not play a major role in mineral precipitation and geochemical cycling within Dallol10.
At the surface, the water temperature at the source is above 100 °C and highly acidic (pH ~ 0). The resultant hot pools vary in color depending on the high metal concentration- (e.g., iron 35.6 g/L, copper 93 mg/l) (Table 1). A combination of these extreme chemical and physical parameters (e.g. temperature, pH, salinity and heavy metals) has resulted in a unique multi-extreme environment.
First reported thermoacidophilic prokaryote was isolated from coal refuse piles11 and the first eukaryote from upper oxygenic zones of acidic geothermally heated waters in solfataras12. In 1995 two species of a moderately thermophilic archaea living around pH 0 were reported13.
Here, we describe for the first time morphological and molecular evidence of thermohaloacidophilic nanomicroorganism existing in this novel multi-extreme environment.
Due to the unique geochemistry and volcanic activity of Dallol6,7 it is an ideal analogue site for studying Martian hydrothermal environments, such as Nili Patera caldera9, where hydrothermal sinter deposits are found in direct association with volcanic activity14, and Gusev crater15, the landing site of the Spirit MER rover.
Despite the Dallol hot springs being deemed one of the most extreme environments on Earth the possibility of microbial life existing within the springs has not been studied.
Results and Discussion
In this system many protuberances can be observed (Figs 1b and 6A–C), which are generated by the precipitation of minerals from the superheated underground solutions. The chimney selected for this study was Site D9 (Figs 1b and 6A) and composed of chlorargyrite (Fig. 2a), wurtzite (Fig. 2a), halite (Fig. 2b) and pyrolusite (Fig. 2c). Table 1 reports the physico-chemical parameters of Site D9 and the surrounding pool (Site D10) as shown in Fig. 6A. The pH and temperature between the two sites varied, for example at Site D9, the source of the hydrothermal fluid, the fluid was pH 0.25 and 86 °C compared to 2.42 and 47 °C at site D10.
DNA was extracted from the salt precipitates at the hydrothermal fluid source (D9), 16S rDNA was amplified using Archaea primers (Fig. 4c) and phylogenetic analysis was performed. The OTU named Dallol 9 showed close similarity (95.8%) with the Nanohaloarchaea environmental clone ARDARCSS13 (GenBank accession number EU869371) (Fig. 3) which closely matched to uncultured sequences from Nanohaloarchaeales Order.
The clone ARDARCSS13 and Dallol 9 sequences are close to Candidatus Holaredivivus sp. G17 (Fig. 3) that has been described16 in a Bras of Port saltern pond (Alicante, Spain) as a photoheterotrophic microbe that present rhodopsin and photolyase. Candidatus Holaredivivus sp. G17 is capable to degrade polysaccharides, likewise presents typical genes of archaea. In relation to survival strategies in hypersaline environments, Haloredivivus sp. G17 is a salt-in strategist as the isoelectric point of his proteins shows16.
To confirm the presence of members of the Nanohaloarchaea group in the samples Fluorescence In Situ Hybridization (FISH) was carried out. Samples fixed in situ for FISH were sequentially filtered through 0.45, 0.22 and 0.1 µm filters to select for small (<0.1 µm) microorganisms. Inspection of the DNA stained material retained in the 0.2 µm filters with SybrGold suggested the presence of ultra-small microorganisms forming compact dense colonies, including mucilaginous films (Fig. 4a). Ultrasonication was further used to disintegrate the colonies prior to sequentially filtration. The ultra-small microorganisms passed through the 0.22 µm filter, but were again retained on the 0.1 µm filters (Fig. 4b). Figure 4c shows 16S rDNA amplification using universal archaea primers.
Living microorganisms at the Dallol hydrothermal outcrop. (a) SybrGold DNA staining of microorganisms retained in 0.2 µm filter before ultrasonication. (b) SybrGold DNA stain of microorganisms retained in 0.1 µm filter after ultrasonication. (c) Electrophoresis of archaeal amplified DNA from sample D9. (d–f) FISH images of Nanohaloarchaea microorganisms. In green, SybrGold DNA staining (d). In red, Narc1214 FISH probe signal (e). (f) d and e merge. Scale bars, 5 μm.
As it is shown in Fig. 4d–f, members of the Nanohaloarchaea group were detected with the Narc1214 FISH probe in the salt precipitates at the hydrothermal fluid source from D9, which corroborates the phylogenetic data.
The presence of ultra-small microorganisms was further investigated using Transmission Electron Microscopy (TEM) (Fig. 5), Scanning Electron Microscopy (SEM) and Scanning Transmission Electron Microscopy (STEM) (Fig. 6). Using these techniques, it was possible to identify ultra-small cellular morphologies (between 50 nm and 500 nm), which supports the molecular data (Figs 5 and 6). The Energy-dispersive X-ray (EDX) elemental microanalysis suggested that the cellular morphologies were unambiguously biological given their high carbon content (Fig. 5A–F). Whilst, SEM-EDX showed the presence of silica precipitates (Fig. 6G–M) surrounding and encrusting the ultra-small cellular morphologies (Fig. 5A–D), suggesting a hydrothermal origin with a high Fe content (Fig. 5E–F).
Figure 6 shows ultra-small microorganisms entombed within precipitated silica minerals (Fig. 6D–L). The cellular morphologies were associated with the needle-shaped crystals (Fig. 6M), which suggests a relationship between the microorganisms and the biomineralization process as has been previously reported in other acidic environments17.
The ultra-small microorganisms were entombed within the precipitated mineral structures (Fig. 6(D–F)). The mineral precipitation process gave rise to thin mineral layers (Fig. 6(D)). Figure 6(E,F) show an enlarged view of the thin mineral layers. Figure 6(G) shows ultra-small microorganisms, which were not covered by minerals, whilst Fig. 6(H,I) show ultra-small microorganisms partially covered with mineral precipitates. More biomineralized ultra-small microorganisms are visible in Fig. 6(J,K). Figure 6(M) shows STEM images with close views of ultra-small microorganisms surrounded by mineral needles and Fig. 6(F) shows mineral spheres resulting from the mineralization process. Needles with spike forms are observed in Fig. 6(J–L). Finally, Fig. 6(N,O) show ultra-small microorganisms surrounded by minerals in a close view using STEM.
Concluding Remarks
We demonstrated the presence of living ultra-small microorganisms in a multi-extreme environment with adverse conditions for life: extreme low pH (0.25), temperature (90 °C), redox potential, salinity and heavy metals content. Molecular studies, electron microscopies observation and phylogenetic analyses of amplified rDNA sequences showed the presence of ultra-small microorganisms related to the Order Nanohaloarchaea. The ultra-small bacteria observed in the samples were morphologically spherical and entombed in the mineral layers that form the small chimneys in Dallol under high acidity and high temperature conditions. The saturation of salts and minerals in the superheated water, which results in precipitation and the formation of the chimneys may be influenced by entombed microorganisms. We described the presence of ultra-small microorganisms in a natural environment which is an Earth analogue of some regions of Mars such as Nili Patera Caldera9,10. The presence of life in the Dallol hot springs expands our understanding of the limits of habitability on Earth and beyond. However, future work is needed to understand how these nanobacteria survive in such an extreme environment and whether they play a role in geochemical cycling.
Materials and Methods
Sampling
The sampling campaign was carried out in January 2017 and samples were collected from a fumarole’s wall (14°14′21″N; 40°17′55″E) located at the main Dallol hydrothermal outcrop in the Danakil depression and a blue pool surrounding the small fumarole. Samples were collected aseptically, using sterilized spatulas and plastic aseptic materials in 12 mL sterile glass vials. The vials were completely filled to prevent head space. To ensure that the samples did not oxidize during transportation and storage, the vials were sealed with a septum tap, covered with parafilm tape and kept under anaerobic conditions. Samples were transported at room temperature.
“In Situ” physico-chemical parameters measurement
Physico-chemical parameters (T: °C, Eh: mV; conductivity: mS/cm2) were measured “in situ” using a multi-parametric probe, YSI 556 MPS. Gas composition by gas chromatography and elemental composition using ICP-MS.
Electron microscopy
Samples were fixed in 4% paraformaldehyde and 2% glutaraldehyde, in 0.1 M phosphate buffer (pH 7.2), for 2 h at room temperature. Fixed samples were washed three times in the same buffer and post-fixed with 1% OsO4 in water for 60 min at room temperature in the dark. After three washes in distillate water, samples were incubated with 2% aqueous uranil acetate for 1 h at room temperature, washed again, and dehydrated in increasing concentrations of ethanol 30, 50, and 70% 20 min each, 90% 2 × 20 min, and 100% 2 × 30 min at room temperature. Dehydration was completed with a mixture of ethanol/propylene oxide (1:1) for 10 min and pure propylene oxide 3 × 10 min. Infiltration of the resin was accomplished with propylene oxide/Epon (1:1) for 45 min and pure LR White resin (London Resin Company limited, England), overnight at room temperature. Polymerization of infiltrated samples was done at 60 °C for 2 days. Ultrathin sections of the samples were stained with uranyl acetate and lead citrate by standard procedures.
Four types of electronic microscopy techniques were used for this study: Scanning Electron Microscope (SEM) (JEOL-5600 coupled to an EDX, INCA) with an Energy Dispersive X-Ray Analyzer (EDX) and SEM-FI (Philips XL30-FEG); Scanning Electron Microscopy-Field Emission Gun (SEM-FEG) (Philips XL30-FEG); Transmission Electronic Microscope (TEM) (JEM-1010); TEM/STEM electronic microscope (JEOL 2100 K) with FEG. Samples were mounted on conductive graphite stubs and sputter and gold-coated in a Quorum, Q150T-S apparatus to ensure electrical conductivity and prevent charging under electron beams.
For SEM analysis, samples were also analyzed with EDX to obtain semiquantitative chemical data. A STEM unit was coupled to the microscope with acquisition of contrast images z (HAADF). The qualitative element composition of samples was determined using an INCA-X-SIGHT with a Si-Li detector (Oxford, England) with a detection limit of 10% for the main element. The operating energy was 200 kv. For TEM analyses, the instrument was operated at 200 kv (Catalysis Institute and PETROQUIMICA-CSIC), equipped with an energy dispersive X-RAY microanalysis instrument INCA (Oxford, England). The TEM/STEM was operated at 200 kv with an EDX, which was coupled to a STEM unit, with acquisition of contrast images z (HAADF). The same grids were used for both TEM and TEM-STEM.
Fluorescence microscopy
Samples were fixed in 4% formaldehyde in phosphate-buffered saline (PBS) for 2 h and stored at 4 °C until further processing. Fixed samples were dispersed by 3 cycles of sonication of 30 sec, with a 30 sec break in between with 1 pulse per second (intensity of 20%). The samples were filtered sequentially through a 0.4 μm pore size polycarbonate filters, a 0.2 μm pore size filter and a 0.1 μm pore size filter (Millipore, Germany). Filters were pretreatment as previously described18. FISH was performed as described by Glöckner, et al.19 using Cy3 single-labeled Narc1214 probe20 (Biomers, Ulm, Germany). Stringencies were regulated adjusting formamide and NaCl concentration in hybridization and washing buffer at 30% (vol/vol) and 0.112 M respectively. Filters were counterstaining by incubation with SYBR® Gold (Molecular Probes, Eugene, OR, USA) 1X diluted in milliQ water for 15 min, before being mounted on glass a slide using Vectashield (Vector Laboratories, Burlingame, CA, USA): Citifluor (Citifluor, London, United Kingdom) (1:4). Samples were imaged using a confocal laser scanning microscope LSM710 (Carl Zeiss, Jena, Germany) equipped with diodo (405 nm), argon (458/488/514 nm) and helium and neon (543 and 633 nm) lasers. Fiji software21 was used to process images.
DNA extraction and amplification of ribosomal genes
Samples were filtered through polycarbonate membranes (0.45, 0.2 and 0.1 µm diameter; Millipore, USA). Total genomic DNA was extracted from those membranes using the DNeasy PowerSoil Kit (Qiagen, Germany) as described in the manufacture’s instructions, and concentrated using a SpeedVac Concentrator. The quantity of the extracted DNA was analysed by fluorimetry using a Qubit 2.0 fluorometer (Thermo Fhiser Scientific, USA). Total genomic DNA were stored at −20 °C for sequencing.
Ribosomal genes were amplified using a primers set specific to amplify V2-V3 region of 16S rRNA in Archaea domain (Arch1F 5′-CGGRAAACTGGGGATAAT-3′ and Arch1R 5′-TRTTACCGCGGCGGCTGBCA-3′). PCR reactions were performed as described22,23. Gel electrophoreses (1% agarose Conda, Spain in 0.5X TBE buffer, 90 mV during 30 min and staining with GreenSafe Premium NZYTech) were carried out to check the size and quality of PCR products.
Ribosomal genes library preparation and sequencing
Library preparation and 2 × 300 pair-end sequencing by Illumina MiSeq were made by Genomic Unit in Parque Científico de Madrid Foundation/FPCM (Madrid, Spain).
Total genomic DNAs was quantified by Picogreen. Then, an input of 16 pg of DNA and 27 cycles were used in a first PCR with Q5® Hot Start High-Fidelity DNA Polymerase (New England Biolabs, USA) in the presence of 100 nM primers for 16S amplification (5′-ACACTGACGACATGGTTCTACACCTACGGGNGGCWGCAG-3′ and 5′- TACGGTAGCAGAGACTTGGTCTGACTACHVGGGTATCTAATCC-3′, these primers amplify the V3-V4 region of 16S), 200 nM primers for Archaea amplification (5′-ACACTGACGACATGGTTCTACACGGRAAACTGGGGATAAT-3′ and 5′-TACGGTAGCAGAGACTTGGTCTTRTTACCGCGGCGGCTGBCA-3′),
After the first PCR, a second PCR of 15 cycles was perfomed with Q5® Hot Start High-Fidelity DNA Polymerase (New England Biolabs, USA) in the presence of 400 nM of primers (5′-AATGATACGGCGACCACCGAGATCTACACTGACGACATGGTTCTACA-3′ and 5′-CAAGCAGAAGACGGCATACGAGAT-[10 nucleotides barcode]-TACGGTAGCAGAGACTTGGTCT-3′) of the Access Array Barcode Library for Illumina Sequencers (Fluidigm, USA).
The amplicons were validated and quantified by a Bioanalyzer. An equimolecular pool was purified by gen extraction and titrated by quantitative PCR using the “Kapa-SYBR FAST qPCR kit for LightCycler480” with a reference standard for quantification. The pooled amplicons were denatured and added to the flowcell at a density of 9 pM. Clusters formed, which were sequenced using a “MiSeq Reagent Kit v3”, in a 2 × 300 pair-end sequencing run on an Illumina MiSeq sequencer.
Detection of DPANN OTUs by microbiome analyse
Quality of reads was evaluated by means of FastQC software. PANDAseq Assembler was using for assembling forward and reverse reads and convert in a fasta file24. Sequencing data were processed using Qiime software package version 1.9.025. High quality contigs were clustered into OTUs based on 94% sequence similarity with UCLUST. The first sequence for each OTU as the representative OTU, which were aligned using PYNAST26. The taxonomic identity of each phylotype was determined using the SILVA_132_QIIME database (https://www.arb-silva.de/download/archive/qiime/)27. Since Ultra small microorganisms are the target of this study, their representative sequences were filtered from biom table in order to analysis only these taxa using filter_taxa_from_otu_table.py and filter_fasta.py scripts in Qiime. Finally, the ARB software package28 was used to reconstruct the phylogenetic tree. The neighbour-joining with Felsenstein correction included in the ARB package was used for phylogenetic inference. The robustness of the reconstructed trees was evaluated by bootstrap analysis of 1000 resampled datasets.
Change history
26 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Horneck, G. et al. AstRoMap European Astrobiology Roadmap. Astrobiology. 16(3), 201–243, https://doi.org/10.1089/ast.2015.1441 (2016).
Stetter, K. O. Extremophiles and their adaptation to hot environments. FEBS Letters. 452, 1–2, https://doi.org/10.1016/S0014-5793(99)00663-8 (2003).
Fairén, A. G. et al. Astrobiology through the Ages of Mars: The Study of Terrestrial Analogues to Understand the Habitability of Mars. Astrobiology. 10(8), 821–843, https://doi.org/10.1089/ast.2009.0440 (2010).
Rothschild, L. J. Earth analogs for Martian life. Microbes in evaporites, a new model system for life on Mars. Icarus. 88-1, 246–260, https://doi.org/10.1016/0019-1035(90)90188-F (1990).
Miruts, H., Bheemalingeswara, K. & Jemal, A. A preliminary geological and generalized stratigraphy of western margin of northerm Afar depression, Dallol area, northern Ethiopia. Momona Ethiopian Journal of Science. 8(1), 1–22 (2016).
Pagli, C. et al. Shallow axial magmachamber at the slow spreading Erta Ale Ridge. Nat Geosci 5, 284–288 (2012).
Nobile, A. et al. Dyke-fault interaction during the 2004 Dallol intrusion at the northern edge of the Erta AleRidge (Afar, Ethiopia). Geophys Res Lett 39, L19305 (2012).
Cavalazzi, B. et al. The Dallol Geothermal Area, Northern Afar (Ethiopia)—An Exceptional Planetary Field Analog on Earth. Astrobiology Published Online: 17 Jan, https://doi.org/10.1089/ast.2018.1926 (2019).
Fawdon, P. et al. The geological history of Nili Patera, Mars. JGR. 120(5), 951–977, https://doi.org/10.1002/2015JE004795 (2015).
Kotopoulou, E. et al. A Polyextreme hydrothermal system controlled by iron: the case of Dallol at the Afar triangle. ACS Earth Space Chem. 3(1), 90–99, https://doi.org/10.1021/acsearthspacechem.8b00141 (2019).
Darland, G., Brock, T. D., Samsonoff, W. & Conti, S. F. A thermophilic acidophilic Mycoplasma isolated from a coal refuse pile. Science 170, 1416–1418 (1970).
Brock, T. D. Thermophilic microorganisms and life at high temperatures. Springer-Verlag, New York (1978).
Schleper, C., Pühler, G., Kühlmorgen, B. & Zillig, W. Life at extremely low pH. Nature 375, 741–742 (1995).
Skok, J. R., Mustard, J. F., Ehlmann, B. L., Milliken, R. E. & Murchie, S. L. Silica deposits in the Nili Patera caldera on the Syrtis Major volcanic complex on Mars. Nature Geoscience. 3, 838–841, https://doi.org/10.1038/ngeo990 (2010).
Ruff, S. W. & Farmer, J. D. Silica deposits on Mars with features resembling hot spring biosignatures at El Tatio in Chile. Nature communications. 7, 13554, https://doi.org/10.1038/ncomms13554 (2016).
Ghai, R. et al. New abundant microbial groups in aquatic hypersaline environments. Scientific reports 1, 135 (2011).
Oggerin, M., Tornos, F., Rodriguez, N., Pascual, L. & Amils, R. Fungal Iron biomineralization in Río Tinto. Minerals 6, 37, https://doi.org/10.3390/min6020037 (2016).
Ishii, K., Mussmann, M., Macbregor, B. J. & Amann, R. An improved fluorescence in situ hybridization protocol for the identification of bacteria and archaea in marine sediments. FEMS Microbiol Ecol. 50, 203–13 (2004).
Glöckner, F. O. et al. An In Situ Hybridization Protocol for Detection and Identification of Planktonic Bacteria. Systematic and Applied Microbiology 19, 403–406, https://doi.org/10.1016/S0723-2020(96)80069-5 (1996).
Narasingarao, P. et al. De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. The ISME journal 6(1), 81 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 9(7), 676–82, https://doi.org/10.1038/nmeth.2019 (2012).
Cruaud, P. et al. Influence of DNA extraction method, 16S rRNA targeted hypervariable regions, and sample origin on microbial diversity detected by 454 pyrosequencing in marine chemosynthetic ecosystems. Applied and Environmental Microbiology. 80(15), 4626–4639 (2014).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic acids research. 41(1), 1–11 (2013).
Bartram, A. K., Lynch, M. D., Stearns, J. C., Moreno-Hagelsieb, G. & Neufeld, J. D. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end Illumina reads. Applied and environmental microbiology. 77(11), 3846–3852 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods. 7(5), 335–336 (2010).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics. 26(2), 266–267 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic acids research. 41(D1), D590–D596 (2012).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic acids research 32(4), 1363–1371 (2004).
Acknowledgements
Europlanet 2020 RI received funding from the European Union’s Horizon 2020 research and innovation program under Grant Agreement No 654208.
Author information
Authors and Affiliations
Contributions
Felipe Gómez led the work, did the sampling and the experimentation and wrote the article, including Figures 1–3. Barbara Cavalazzi contributed to design the field campaign, to the description of the geological setting and to the corrections of the article. Nuria Rodríguez contributed in the sampling, experimental and electromicroscopy of the work. She did the Figures 4 and 5 and Table 1. Ricardo Amils advised on the microbiological part of the work and contributed to the correction of the manuscript. Gian Gabriele Ori contributed to design the field campaign and advised on the geological part of the work. Karen Olsson-Francis contributed to the microbiological work and to the correction of the article. Cristina Escudero contributed to the hybridization and staining experiments. Jose M. Martínez contributed with the metagenomic study of the work. Hagos Miruts contributed to the sampling campaign and advised on the contributed to the geological interpretation of the geological Dallol model.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gómez, F., Cavalazzi, B., Rodríguez, N. et al. Ultra-small microorganisms in the polyextreme conditions of the Dallol volcano, Northern Afar, Ethiopia. Sci Rep 9, 7907 (2019). https://doi.org/10.1038/s41598-019-44440-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44440-8
This article is cited by
-
Temperature and elemental sulfur shape microbial communities in two extremely acidic aquatic volcanic environments
Extremophiles (2021)
-
In silico determination of nitrogen metabolism in microbes from extreme conditions using metagenomics
Archives of Microbiology (2021)
-
Geoheritage and Resilience of Dallol and the Northern Danakil Depression in Ethiopia
Geoheritage (2020)
-
Microbial unknowns at the saline limits for life
Nature Ecology & Evolution (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.