Abstract
Circulating microbial dysbiosis is associated with chronic liver disease including nonalcoholic steatohepatitis and alcoholic liver disease. In this study, we evaluated whether disease-specific alterations of circulating microbiome are present in patients with cirrhosis and hepatocellular carcinoma (HCC), and their potential as diagnostic biomarkers for HCC. We performed cross-sectional metagenomic analyses of serum samples from 79 patients with HCC, 83 with cirrhosis, and 201 matching healthy controls, and validated the results in the same number of subjects. Serum bacterial DNA was analyzed using high-throughput pyrosequencing after amplification of the V3–V4 hypervariable regions of 16S rDNA. Blood microbial diversity was significantly reduced in HCC, compared with cirrhosis and control. There were significant differences in the relative abundances of several bacterial taxa that correlate with the presence of HCC, thus defining a specific blood microbiome-derived metagenomic signature of HCC. We identified 5 microbial gene markers-based model which distinguished HCC from controls with an area under the receiver-operating curve (AUC) of 0.879 and a balanced accuracy of 81.6%. In the validation, this model accurately distinguished HCC with an AUC of 0.875 and an accuracy of 79.8%. In conclusion, circulating microbiome-based signatures may be potential biomarkers for the detection HCC.
Similar content being viewed by others
Introduction
Gut microbiota dysbiosis and increased bacterial translocation play an important role in the progression of chronic liver disease. Because liver receive most of the blood supply from intestine, it is exposed to gut microbiota, bacterial pathogen-associated molecular patterns and microbial metabolites which lead to chronic inflammation and progression of liver disease such as fibrosis and hepatocellular carcinoma (HCC)1,2.
Recent studies have reported the presence of circulating bacterial contents in healthy human blood by sequencing 16S ribosomal deoxyribonucleic acid (rDNA) genes3. Furthermore, the predictive roles of circulating microbiome on the onset of diabetes and cardiovascular events have been suggested in longitudinal studies4,5. In addition, changes in blood- and gut-microbiome signatures have been linked to liver fibrosis in obese patients6. Moreover, changes in blood-microbiome signatures associated with a shift in the metabolic functions has been reported in heavy alcohol drinkers without significant liver disease and those with alcoholic hepatitis7. Although it has not been known yet whether alteration in the blood microbiome is just a bystander of dysbiosis or a true player in the pathophysiology of disease, these findings collectively suggest that blood microbiota profiles might be used as a potential noninvasive biomarker. However, to our knowledge, the association of blood microbiota and HCC has not been studied, yet.
In this study, we evaluated whether disease-specific microbiome alterations are present in the blood microbiome from patients with cirrhosis and HCC, and their potential as diagnostic biomarkers for HCC.
Materials and Methods
Study population
The study population consisted of 158 patients with HCC and 166 with cirrhosis diagnosed between July 2013 and August 2017, at the Seoul National University Hospital (Seoul, Korea) and Ajou University Hospital (Suwon, Korea), and 402 healthy controls who underwent a medical checkup at the Seoul National University Boramae Medical Center (Seoul, Korea) and the Inje University Haeundae Paik Hospital (Busan, Korea) during the same period. Subjects within each hospital were randomly divided in a 1:1 ratio to a model development set and a test set (79 HCC, 83 cirrhosis and 201 controls for each set) to develop a model capable of distinguishing HCC from cirrhosis and controls.
The diagnosis of HCC was mainly based on the noninvasive criteria of the international guidelines8,9. Cirrhosis was diagnosed histologically, clinically, or by typical radiological findings10. The control group was defined by subjects with no clinical or imaging evidence of liver disease. Subjects with clinical symptoms and signs of infection were excluded from the study. Clinical data were collected using standard case report forms. Overnight fasting serum samples were obtained from all subjects before any treatment using standard protocols and stored at −80 °C until processing.
Ethics approval and consent to participate
This study was conducted in accordance with the Declaration of Helsinki principles. It was approved by the institutional review board (IRB) of Seoul National University Hospital (IRB No. 1704-021-843), and samples derived from the Korea Biobank Network were obtained with informed consent under IRB-approved protocols. The Reporting Recommendations for Tumor Marker Prognostic Studies criteria were followed throughout the study11.
Circulating cell-free DNA extraction from serum samples
Serum samples were centrifuged at 2,000 g for 15 min at 4 °C to remove cell debris, and the supernatant was collected and incubated with proteinase K at 56 °C for 30 minutes. Then, the samples were boiled for 40 min under 100 °C to extract DNA out of extracellular vesicles, and the supernatant was collated by centrifugation for 30 min at 10,000 g under 4 °C. Total DNA was extracted from the supernatant (1 mL) using a DNA isolation kit (DNeasy Blood & Tissue Kit, QIAGEN, Germany) and quantified by using the QIAxpert system (QIAGEN, Germany).
Metagenomic sequencing and annotation
Following DNA extraction, bacterial genomic DNA was amplified with 16S_V3_F (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′) and 16S_V4_R (5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′) primers, which are specific for V3-V4 hypervariable regions of 16S rDNA. The libraries were prepared using PCR products according to MiSeq System guide (Illumina, USA) and quantified using a QIAxpert (QIAGEN, Germany). Each amplicon is then quantified and sequenced on a MiSeq (Illumina, USA) according to the manufacturer’s recommendations.
Profiling
Raw sequencing reads obtained from the sequencer were filtered according to the barcode and primer sequences using MiSeq (Illumina, USA). Taxonomic assignment was performed by profiling program MDx-Pro ver.1. The high-quality sequencing reads were selected after checking the read length (≥300 bp) and the quality score (average Phred score ≥ 20). Operational Taxonomy Units (OTUs) were clustered by using sequence clustering algorithms CD-HIT. Subsequently, taxonomy assignment was carried out by using UCLUST and QIIME against the Greengenes reference database (gg_13_5_99)12,13. OTUs with a number of sequences <0.005% of the total were removed from the OTU table. After filtering, an average of 14,555 reads per sample was obtained (min: 19; max: 31139) and sample with low number of read counts (<2500) were filtered out for quality controls. The resulting OTU table was used for predictive functional analysis using the software Tax4Fun in the package metagenomics version 0.1.014.
Statistical analysis
All statistical analyses were performed with R version 3.4.4 on Windows 10 (Version 3.4.4, http://www.R-project.org). For significance tests for the difference among the three groups, one-way analysis of variance was used for continuous variables, and the chi-square test was used for categorical variables. α-diversity of microbiota for each sample was measured by Shannon index. To compare α-diversities between groups, the Wilcoxon rank-sum test was used for comparing two groups and the Kruskal-Wallis test was used for more than two groups. β-diversity of a pair of samples was measured by weighted and unweighted UniFrac distances15,16. Based on the UniFrac distances, principal coordinate analysis (PCoA) was performed and analysis of similarity (ANOSIM) permutation test was used to assess the statistical significance of the separation among groups17. The OTUs after removing the unassigned ones at phylum level were used to calculate the relative abundance for comparisons of groups and model construction.
To develop a model for discriminating HCC, we used the logistic regression model in which the response variable is a binary variable distinguishing HCC and control groups. We selected OTUs with significant effects in logistic regression model that consists of binary response variable and age/sex as adjustment variables.
We first randomly divided data into the model development and test sets. Data of the model development set were further randomly allocated into training and validation sets. As the rare OTUs differed significantly according to test methods and the relatively abundant OTUs were similar across the methods18, we selected candidate OTUs at genus level if their means of relative abundance were greater than 1% and if the p values were smaller than 0.05 in logistic regression models with age and sex as covariates. For abundance comparisons of OTUs between groups, p values are adjusted by Bonferroni correction based on the number of OTUs after filtering. Then all possible combinations of candidate OTUs were tested by repeating 10 times for two-fold cross validations to find the optimal model discriminating HCC. The final model was selected by the one with the lowest Akaike’s information criteria (AIC) from the model development set, and then its performance was assessed using the test set.
Results
Baseline characteristics
The baseline characteristics of subjects in the model development and test sets are described in Table 1. Age and gender were comparable between three groups in both sets. Most of the patients in HCC group had hepatitis virus-related compensated liver disease, whereas cirrhosis group showed relatively higher proportion of decompensated, non-viral etiology (mainly alcohol)-induced cirrhosis. In both sets, about 80% of the HCC cases belonged to stage I and II according to the 7th American Joint Committee on Cancer staging system.
Changes in the taxonomic signature of blood metagenomes according to liver disease
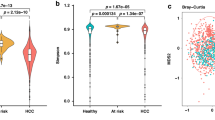
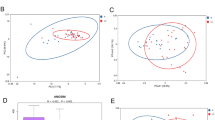
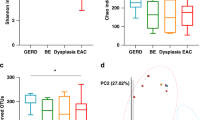
To investigate whether there are specific changes of blood metagenomes according to the liver disease, we assessed the relative abundance of taxa in the three group. We used the Shannon index at each level to compare α-diversity (within sample diversity). At the phylum level, α-diversity was significantly reduced in the HCC, compared with the cirrhosis and control groups (p = 0.006 and 4.7e-06, respectively), whereas there was no significant difference between the cirrhosis and control groups (p = 0.37; Fig. 1a). Similar trends were also observed at the genus level (Fig. 1b). To compare β-diversity (between sample diversity), we used the unweighted and weighted UniFrac distances. The PCoA plot based on the unweighted UniFrac distance showed the strong separation between control group and cirrhosis and HCC groups (Fig. 2a). The ANOSIM test confirmed the significance (control vs. cirrhosis: R = 0.45, p < 0.001; control vs. HCC: R = 0.49, p < 0.001; cirrhosis vs. HCC: R = 0.04, p < 0.001). While no clear separation was observed in the PCoA plot based on the weighted UniFrac distance (Fig. 2b), the ANOSIM test provided significant differences between groups (control vs. cirrhosis: R = 0.20, p < 0.001; control vs. HCC: R = 0.19, p < 0.001; cirrhosis vs. HCC: R = 0.04, p < 0.001). Compared to the unweighted UniFrac distance, the weighted UniFrac distance yielded much smaller R values. These results suggest that there are some microbiome factors that could distinguish the HCC group from the others.
Blood taxonomic signature characterizing patients with HCC in the development cohort
Next, we investigated abundant OTUs defined as mean relative abundances of OTUs >1% in total samples at phylum and genus levels. Relative abundances of OTUs are grouped for disease groups and plotted by the decreasing order of mean relative abundances of total samples in Fig. 3. The blood microbiomes in three groups were dominated by members of Firmicutes and Proteobacteria, followed by Actinobacteria and Bacteroidetes in much lower abundances. In addition, both Firmicutes and Proteobacteria were differentially abundant across the three groups (p < 0.05), with Firmicutes being highest in controls while Proteobacteria was highest in HCC group.
Relative abundances of abundant OTUs among the three groups. Boxplots of the relative abundances of abundant OTUs (a) at phylum and (b) at the genus levels in the three groups. Abundant OTUs are selected by relative abundance means of OTUs > 1% in total samples and listed by decreasing order of the mean values.
At the genus level, 7 bacterial taxa showed significantly different abundance between HCC and control groups (p < 0.05, multiple testing correction using Bonferroni, univariate test with adjustment of age and sex). Pseudomonas was the most abundant microbiome in the three groups, and significantly decreased in HCC, compared with control group. Of the remaining 6 taxa, Streptococcus and Bifidobacterium were significantly increased in controls, whereas 4 taxa including Staphylococcus, Acinetobacter, Klebsiella and Trabulsiella were significantly enriched in HCC-associated microbiomes. Remarkably, Staphylococcus showed the strongest association with HCC (p = 4.0e-08) and demonstrated a 4.3-fold increase in HCC, compared with controls.
We evaluated the effects of liver function and etiology of liver disease on the relative abundance of blood microbiome. When patients with cirrhosis and/or HCC were stratified into compensated and decompensated stages, there was no significant difference in α-diversity (supplementary Fig. S1A). However, the patterns of β-diversity provided some evidence of differences according to liver function (compensated vs. decompensated: weighted UniFrac ANOSIM, R = 0.06, p = 0.046; Unweighted UniFrac ANOSIM, R = 0.06, p = 0.06; supplementary Fig. S1B). On the contrary, there was no significant difference in α-diversity (supplementary Fig. S2A) and β-diversity according to the etiology of liver disease (viral vs. non-viral: weighted UniFrac ANOSIM, R = 0.02, p = 0.30; Unweighted UniFrac ANOSIM, R = 0, p = 0.49; supplementary Fig. S2B).
HCC discrimination with the 5-genera microbiome signature
To evaluate the potential of blood taxonomic signature to discriminate HCC from controls, all combination models of candidate OTUs, which are selected at the genus level (supplementary Table 1), were tested with adjustment for age and sex. Table 2 showed performance of top models for each number of microbiome markers, and the model composed of 5 HCC-associated genera (i.e., Pseudomonas, Streptococcus, Staphylococcus, Bifidobacterium, and Trabulsiella) was finally selected. The model based on these five OTUs showed an area under the receiver-operating curve (AUC) value of 0.879 (sensitivity, 0.729; specificity, 0.850; accuracy, 0.816) in the model development set. Subsequently, we evaluated its performance in the test set. The AUC was 0.875 (sensitivity, 0.756; specificity, 0.797; accuracy, 0.798; Fig. 4), suggesting a potential for the blood microbiome-based signature to accurately discriminate HCC from controls. When the model was applied to three groups, the probability of disease was significantly increased in the HCC group versus the control group, and the cirrhosis group was intermediate level between the HCC and control group both in the model development set and test set (supplementary Fig. S3). In addition, these trends significantly maintained regardless of underlying liver function status, suggesting that the changes from healthy control to cirrhosis and further to HCC might be correlated with the disease progression status, rather than liver function status. In addition, we performed additional 5-fold and 10-fold cross validations and compared their performances. We found that high number of folds increases model performances (i.e. sensitivity, specificity, accuracy and AUC) significantly which might be due to a small sample size (data not shown). To provide a more conservative (i.e. smaller AUC values) results, we decided to present the 2-fold cross validation results.
Discussion
In this study, we investigated the relationship between circulating microbiota and HCC for the first time. The main result of this study was that HCC was associated with altered composition of circulating microbiota, as well as significant lower level of diversity. In addition, we validated the diagnostic accuracy of the blood microbiome-derived metagenomic signature to detect HCC, and suggested their potential as diagnostic markers for HCC.
A recent study has reported that the blood 16S rDNA concentration was shown to be increased in obese patients with hepatic fibrosis, whereas the bacterial diversity was decreased6. Another study has shown that there were multiple alterations in the circulating microbiome in heavy drinkers and patients with alcoholic hepatitis, and these alterations were associated with changes in the metabolic functions such as activation of the type III secretion system associated with gram-negative bacteria, and increased isoprenoid synthesis and anthranilate degradation which are well-known modulators of biofilm formation and gram-positive bacterial growth7. In our study, we showed changes in the composition of the circulating microbiota that correlate with the presence of HCC. These results indicate that specific blood dysbiosis is associated with chronic liver disease including nonalcoholic steatohepatitis, alcoholic liver disease and HCC.
Emerging studies have shown that gut microbiome can affect the pathophysiology of HCC. Gut microbiome has been known to activate lipopolysaccharide (LPS)-toll-like receptor 4 (TLR4) pathway, leading to promotion of HCC growth in mice2,19. Furthermore, gut microbiome–mediated bile acid metabolism modulates antitumor surveillance against HCC via hepatic natural killer T cells20. In addition, a recent study has reported characteristic alterations of gut microbiome in patients with early HCC, and suggested OTUs markers as a diagnostic tool for HCC21. These results indicate a global shift in gut microbiome in HCC, and the altered microbial community might play an important role in the development and progression of HCC. Our study showed that circulating microbiota as well as gut microbiota also presented a moderate dysbiosis in patients with HCC, and this signature might be a potential noninvasive biomarker for detecting HCC.
In this study, the relative abundance of Bifidobacterium genus was notably decreased in HCC. Bifidobacterium has been known to reinforce gut barrier function and have protective effects on liver injury22. In addition, it can promote anticancer immunity and enhance the efficacy of immunotherapy23. Moreover, probiotics has been shown to inhibit the development and growth of HCC in animal model by changing the composition of gut microbiota and recovering intestinal permeability24,25. Akkermansia, a well-known intestinal commensal promoting intestinal integrity and ameliorating liver injury26, also showed decreasing tendency in HCC compared with controls, even though the difference was not statistically significant. On the contrary, the relative abundance of potentially pathogenic gram-negative bacteria producing LPS, such as Klebsiella and Acinetobacter, were increased in HCC. High levels of LPS activate NF-κB pathway and enhance inflammatory damage in liver, thereby promoting the development of HCC27. Furthermore, LPS-TLR4 pathway promotes epithelial-mesenchymal transition, invasion and metastasis of HCC2,28. Collectively, the decrease of potentially beneficial bacteria protecting intestinal barrier, and the increase of potentially pathogenic bacteria might affect intestinal and hepatic inflammation, promoting the development and progression of HCC.
There are several limitations to the present study. First, although there was no subject presenting symptoms and signs of infection at the time of blood sampling, medication history was not available in the healthy control group. Therefore, those taking antibiotics or pro/prebiotics could be included and affect the results. However, the key features of the changes across groups in the development set were maintained in the test set. In addition, a previous study showed that even though antibiotic exposure altered blood microbiome, it did not affect the principal differences between alcoholic hepatitis and control7. Second, this study could not reveal whether blood dysbiosis is only a bystander or a true player in the pathogenesis of HCC. Further studies are warranted to assess the functional and metabolic potential of the circulating microbial communities. Third, we could not analyse gut microbiome. Blood microbiome derives probably mostly from gut microbiome; however, both differ substantially from each other which suggest that gut barrier, host immune, and liver may act as modifiers6. Future mechanistic studies may elucidate the cross-talk between gut and blood microbiome and their functional roles in HCC.
In conclusion, this study revealed compositional dysbiosis in circulating microbiome of patients with HCC, and their potential as diagnostic biomarkers for HCC. Further independent and larger cohort studies and functional mechanistic studies are warranted to validate the identified HCC microbial signature.
References
Tilg, H., Cani, P. D. & Mayer, E. A. Gut microbiome and liver diseases. Gut 65, 2035–2044 (2016).
Dapito, D. H. et al. Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 21, 504–516, https://doi.org/10.1016/j.ccr.2012.02.007 (2012).
Paisse, S. et al. Comprehensive description of blood microbiome from healthy donors assessed by 16S targeted metagenomic sequencing. Transfusion 56, 1138–1147, https://doi.org/10.1111/trf.13477 (2016).
Amar, J. et al. Involvement of tissue bacteria in the onset of diabetes in humans: evidence for a concept. Diabetologia 54, 3055–3061, https://doi.org/10.1007/s00125-011-2329-8 (2011).
Amar, J. et al. Blood microbiota dysbiosis is associated with the onset of cardiovascular events in a large general population: the D.E.S.I.R. study. PLoS One 8, e54461, https://doi.org/10.1371/journal.pone.0054461 (2013).
Lelouvier, B. et al. Changes in blood microbiota profiles associated with liver fibrosis in obese patients: A pilot analysis. Hepatology 64, 2015–2027, https://doi.org/10.1002/hep.28829 (2016).
Puri, P. et al. The circulating microbiome signature and inferred functional metagenomics in alcoholic hepatitis. Hepatology 67, 1284–1302, https://doi.org/10.1002/hep.29623 (2018).
European Association for the Study of the Liver. Electronic address, e. e. e. & European Association for the Study of the, L. EASL Clinical Practice Guidelines: Management of hepatocellular carcinoma. J Hepatol 69, 182–236 https://doi.org/10.1016/j.jhep.2018.03.019 (2018).
Heimbach, J. K. et al. AASLD guidelines for the treatment of hepatocellular carcinoma. Hepatology 67, 358–380, https://doi.org/10.1002/hep.29086 (2018).
Nishida, T. et al. Oral glucose tolerance test predicts prognosis of patients with liver cirrhosis. Am J Gastroenterol 101, 70–75, https://doi.org/10.1111/j.1572-0241.2005.00307.x (2006).
McShane, L. M. et al. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst 97, 1180–1184, https://doi.org/10.1093/jnci/dji237 (2005).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461, https://doi.org/10.1093/bioinformatics/btq461 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Asshauer, K. P., Wemheuer, B., Daniel, R. & Meinicke, P. Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 31, 2882–2884, https://doi.org/10.1093/bioinformatics/btv287 (2015).
Lozupone, C. A., Hamady, M., Kelley, S. T. & Knight, R. Quantitative and qualitative beta diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73, 1576–1585, https://doi.org/10.1128/AEM.01996-06 (2007).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71, 8228–8235, https://doi.org/10.1128/AEM.71.12.8228-8235.2005 (2005).
Clarke, K. R. Non‐parametric multivariate analyses of changes in community structure. Australian journal of ecology 18, 117–143 (1993).
Lee, C., Lee, S. & Park, T. Statistical methods for metagenomics data analysis. International Journal of Data Mining and Bioinformatics 19, 366–385 (2017).
Schnabl, B. & Brenner, D. A. Interactions between the intestinal microbiome and liver diseases. Gastroenterology 146, 1513–1524, https://doi.org/10.1053/j.gastro.2014.01.020 (2014).
Ma, C. et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, https://doi.org/10.1126/science.aan5931 (2018).
Ren, Z. et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut. https://doi.org/10.1136/gutjnl-2017-315084 (2018).
Fang, D. et al. Bifidobacterium pseudocatenulatum LI09 and Bifidobacterium catenulatum LI10 attenuate D-galactosamine-induced liver injury by modifying the gut microbiota. Sci Rep 7, 8770, https://doi.org/10.1038/s41598-017-09395-8 (2017).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089, https://doi.org/10.1126/science.aac4255 (2015).
Zhang, H. L. et al. Profound impact of gut homeostasis on chemically-induced pro-tumorigenic inflammation and hepatocarcinogenesis in rats. J Hepatol 57, 803–812, https://doi.org/10.1016/j.jhep.2012.06.011 (2012).
Li, J. et al. Probiotics modulated gut microbiota suppresses hepatocellular carcinoma growth in mice. Proc Natl Acad Sci USA 113, E1306–1315, https://doi.org/10.1073/pnas.1518189113 (2016).
Grander, C. et al. Recovery of ethanol-induced Akkermansia muciniphila depletion ameliorates alcoholic liver disease. Gut 67, 891–901, https://doi.org/10.1136/gutjnl-2016-313432 (2018).
Yu, L. X. et al. Endotoxin accumulation prevents carcinogen-induced apoptosis and promotes liver tumorigenesis in rodents. Hepatology 52, 1322–1333, https://doi.org/10.1002/hep.23845 (2010).
Jing, Y. Y. et al. Toll-like receptor 4 signaling promotes epithelial-mesenchymal transition in human hepatocellular carcinoma induced by lipopolysaccharide. BMC Med 10, 98, https://doi.org/10.1186/1741-7015-10-98 (2012).
Acknowledgements
The biospecimens for this study were provided by the Ajou University Human Bio-Resource Bank (AHBB) and Seoul National University Hospital-Human Biobank, members of Korea Biobank Network, which are supported by the Ministry of Health and Welfare, Republic of Korea. This work was supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute funded by the Ministry of Health & Welfare (No. HI16C2037), the Bio-Synergy Research Project (2013M3A9C4078158) of the Ministry of Science, ICT and Future Planning through the National Research Foundation (NRF), Basic Science Research Program through the NRF funded by the Ministry of Education (2017R1D1A1B03034639), grants from the SNUH Research Fund (No. 04-2017-0640) and Liver Research Foundation of Korea.
Author information
Authors and Affiliations
Contributions
E.J.C. and S.L. designed the experiments, analyzed data and prepared the manuscript. S.A.K. analyzed data. J.Y., Y.B.L., S.S.K., J.Y.C., S.W.C., J.W.K. and S.-M.K. helped with acquisition and interpretation of data. J.-H.Y. and T.P. designed and were the overseers of the entire study. All authors read and approved the final manuscript for publication.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cho, E.J., Leem, S., Kim, S.A. et al. Circulating Microbiota-Based Metagenomic Signature for Detection of Hepatocellular Carcinoma. Sci Rep 9, 7536 (2019). https://doi.org/10.1038/s41598-019-44012-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44012-w
This article is cited by
-
Urinary microbiome-based metagenomic signature for the noninvasive diagnosis of hepatocellular carcinoma
British Journal of Cancer (2024)
-
Leveraging circulating microbiome signatures to predict tumor immune microenvironment and prognosis of patients with non-small cell lung cancer
Journal of Translational Medicine (2023)
-
A new horizon of precision medicine: combination of the microbiome and extracellular vesicles
Experimental & Molecular Medicine (2022)
-
Dysbiotic change in gastric microbiome and its functional implication in gastric carcinogenesis
Scientific Reports (2022)
-
Machine-learning algorithms for asthma, COPD, and lung cancer risk assessment using circulating microbial extracellular vesicle data and their application to assess dietary effects
Experimental & Molecular Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.