Abstract
Newcastle disease virus (NDV) is a threat to the global poultry industry, but particularly for smallholder farmers in low- and middle-income countries. Previous reports suggest that some breeds of chickens are less susceptible to NDV infection, however, the mechanisms contributing to this are unknown. We here examined the comparative transcriptional responses of innate immune genes to NDV infection in inbred sublines of the Fayoumi and Leghorn breeds known to differ in their relative susceptibility to infection as well as at the microchromosome bearing the major histocompatability complex (MHC) locus. The analysis identified a set of five core genes, Mx1, IRF1, IRF7, STAT1, and SOCS1, that are up-regulated regardless of subline. Several genes were differentially expressed in a breed- or subline-dependent manner. The breed-dependent response involved TLR3, NOS2, LITAF, and IFIH1 in the Fayoumi versus IL8, CAMP, and CCL4 in the Leghorn. Further analysis identified subline-dependent differences in the pro-inflammatory response within the Fayoumi breed that are likely influenced by the MHC. These results have identified conserved, breed-dependent, and subline-dependent innate immune responses to NDV infection in chickens, and provide a strong framework for the future characterization of the specific roles of genes and pathways that influence the susceptibility of chickens to NDV infection.
Similar content being viewed by others
Introduction
Newcastle Disease Virus (NDV) is an important poultry pathogen, especially in low- and middle- income countries where over 80 percent of the poultry are reared in backyard settings with little to no veterinary care and the increased threat of exposure to pathogens1. Once NDV is introduced into a susceptible flock, mortality rates could be as high as 100 percent, and the disease has the potential to cause large losses in productivity for smallholder farmers who solely rely on their flocks as a source of income1,2. Vaccination against NDV is commonly used in high-income countries and in intensively reared poultry worldwide, but are uncommon in low- to middle-income countries, particularly in the context of backyard poultry, and hence there is a well-documented need to consider development of alternative strategies to control NDV in these settings3.
However, although there are low levels of vaccination in backyard poultry in these settings, many backyard poultry come in regular contact with pathogens, including NDV, in the field, but do not exhibit clinical signs of infection. This natural selection has led to locally adapted ecotypes of backyard poultry commonly reported to be naturally less susceptible to various infectious diseases, including NDV4. Understanding the mechanisms contributing to the relative hardiness of backyard poultry as relates to infectious disease susceptibility is important and has the potential to inform future breeding strategies to improve innate disease resistance, in chickens4.
Recent reports have noted differing levels of susceptibility to NDV amongst inbred Fayoumi and inbred Leghorn lines5. In general, the Fayoumi are considered less susceptible to multiple poultry pathogens, including Salmonella, Eimeria, Marek’s Disease Virus (MDV), and Avian Influenza Virus (AIV), as compared with Leghorn and other breeds6,7,8. While the Fayoumi originates from Egypt and is representative of the hardy backyard type poultry ecotypes common throughout the world, the Leghorn is a European origin breed that provides the foundation stock for many of the modern-day commercial egg-type chickens9,10. These two breeds are genetically distinct, and a recent study showed that the inbred M15.2 Fayoumi subline was less susceptible to NDV than the inbred Ghs6 inbred Leghorn subline, with reduced viral shedding in ocular secretions post-challenge9. However, the mechanisms contributing to these differences remain unclear.
Response to infections in birds, as in other species, involves both the innate and adaptive (humoral and cell-mediated) immune responses11. Given the effective and widespread use NDV vaccines, most studies of immunity to NDV have focused on the antibody and the cell-mediated responses, while much still remains to be learned about the innate immune response12,13. Early studies of the innate immune response of chickens to NDV have suggested high expression of genes involved in the induction of nitric oxide and the interferon response in vitro, in vivo, and in ovo, but much remains to be discovered, including the genetic basis and mechanisms of control of this response14,15.
We have recently described the chicken embryo model to examine the immune response to NDV15. Since chicken embryos become immunocompetent prior to hatch and remain in the protective environment of the shell, they provide a particularly tractable way to study the immune response of chickens while containing costs and reducing confounding variables often associated with animal trials16,17. Our preliminary analyses identified several innate immune genes that were differentially expressed between the Kuroiler and local Tanzanian ecotypes, along with evidence of breed-and subline-dependent expression in the congenic Fayoumi (M5.1 and M15.2) and Leghorn (Ghs6 and Ghs13) sublines that differ (within each line) only at the microchromosome bearing the Major Histocompatibility Complex (MHC)5,9,15,18. Other studies have also revealed that the Fayoumi M15.2 subline is less susceptible to NDV as compared with the Leghorn Ghs6 subline, and identified breed specific transcriptional responses to NDV infection in the trachea, lungs, and Harderian gland using an RNA-seq based approach9,18,19. However, these papers focused on characterizing overall pathways and mechanisms of chicks, not solely focusing on the innate immune response to NDV9,18,19. A closer look at the immune genes critical in the early response to NDV infection of these inbred lines is necessary to further characterize and understand their distinct responses to NDV infection.
We have here used the chick embryo model to elucidate the transcriptional response of the innate immune related genes to NDV infection in highly inbred Fayoumi and Leghorn lines. The results of our investigation provide new insights into the conserved, breed-specific, and subline-specific transcriptional innate immune responses of chickens to NDV infection. In addition, the studies provide a foundation for the future development of rational breeding strategies to improve resistance to NDV based on knowledge of innate immune responsiveness of naturally susceptible and resistant chicken ecotypes.
Results
The gene expression profiles of chicken embryos group into five clusters suggesting a conserved, breed-dependent, and subline-dependent response to NDV
The comparative transcriptional profiles of chicken embryos from the two inbred Leghorn sublines, Ghs6 and Ghs13, and two inbred Fayoumi sublines, M5.1 and M15.2, were examined using a custom OpenArray Technology for gene expression. The viral load of each subline and breed was assessed (Supplementary Fig. 1). The Fayoumi breed had a significantly lower viral load than the Leghorn (p-value < 0.001). When comparing the viral load between the sublines, the M15.2 Fayoumi subline had the lowest viral load and the Ghs6 Leghorn subline had the highest viral load. The table in Supplementary Fig. 1 shows the p-values for the comparisons between the different sublines.
Hierarchical clustering grouped the innate immune related gene expression of the Fayoumi and Leghorn sublines into five clusters (Fig. 1). The approximately unbiased (AU) and bootstrap probability (BP) for all clusters are represented at the node of each major cluster. The AU value indicates there is strong clustering (AU > 95) of the genes with all AU values being statistically significant. Cluster 1 (C1), the blue cluster, contains genes that are highly upregulated in all four sublines in response to NDV, and cluster 2 (C2), the red cluster, are all upregulated but not to the extent of the blue cluster. The genes in these clusters include MX dynamin Like GTPase 1 (Mx1), Interferon Regulatory Factor 1 (IRF1) and Interferon Regulatory Factor 7 (IRF7), and Signal Transducer and Activator of Transcription 1 (STAT1) which have all been previously reported as upregulated in the response to NDV14,15. Clusters 3, 4, and 5, the green, purple, and orange clusters, respectively, show the genes that may be involved in breed- or subline-dependent responses. In C3 and C4, the two Leghorn sublines, Ghs6 and Ghs13, have lower expression of the genes compared to the Fayoumi sublines, with most genes in the green cluster being downregulated in the Leghorns. These genes include Suppressor of Cytokine Signaling 2 (SOCS2), Nuclear Factor Kappa-light-chain-enhancer of activated B cells (NFKB), Toll-like Receptor (TLR5), Janus Kinase (JAK), Jun Proto-Oncogene (JUN), and Mitogen-Activated Protein Kinase 14 (MAPK14). Interestingly, C5 demonstrates the genes from the M5.1 Fayoumi subline are upregulated when compared to all other sublines (M15.2, Ghs6, and Ghs13).
Innate Immune Gene Expression Profiles of the Fayoumi and Leghorn Sublines. The overall innate immune gene expression profiles of the two Fayoumi sublines (M5.1 and M15.2) and two Leghorn sublines (Ghs6 and Ghs13) were clustered using hierarchical clustering in R (pvclust, nboot = 1000). Each cluster is colored (C1 – blue, C2 – red, C3 – green, C4 – purple, and C5 – orange). The AU/BP values are represented at the base node of each cluster. The expression of innate immune genes, log2(Fold Change), is visualized in the heatmap (red - upregulation; green - downregulation). The functional pathways for each gene are highlighted with black dots in the table.
Hierarchical clustering of the sublines based on the individual gene expression profiles demonstrates the clustering of the Leghorn Ghs6 subline and Fayoumi M15.2 subline, while the Leghorn Ghs13 subline appears to cluster more closely with the Fayoumi M5.1 subline (Fig. 1). Although AU values are not statistically significant, there is still a trend toward considerable clustering of these sublines (AU = 94). Examining the overall gene expression profiles shows that the response of these lines may differ in a subline-dependent manner and be influenced by the MHC or other genes located on the chicken microchromosome 16, a locus that has been described to have multiple genes with a role in immunity20.
The analysis reveals a core set of differentially expressed genes in the response of chicken embryos to NDV infection, as well as breed- and subline-dependent responses
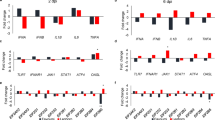
A volcano plot was used to visualize the gene expression profiles of each subline by comparing the normalized Ct values between the experimental versus control groups within each subline and plotting the log 2-fold change versus the negative log p-value. The significantly expressed targets, genes that are significantly expressed in the experimental group over the control group (negative log p-value > 2 equaling a p-value < 0.01), are highlighted in colors corresponding to the legend (Fig. 2). The Ghs6 subline differentially expressed nine genes, the Ghs13 had 11, the M15.2 had 13, and the M5.1 had 22 (Fig. 2). The p-values corresponding to each target within each subline is shown in Supplementary Table 1.
Volcano Plot of the Differentially Expressed Innate Immune Genes in each Subline. The volcano plot visualizes the log2(Fold Change) along the x-axis versus the –log(p-value) along the y-axis for the experimental versus control groups in each subline. The targets significantly expressed in the experimental group versus the controls (p-value < 0.01 or -log(p-value) > 2) are highlighted by the colors that correspond to the legend.
A Venn diagram representing the significantly expressed targets (Fig. 3) and the overlaps between the four sublines identifies 5 genes (IRF1, IRF7, Mx1, Suppressor of Cytokine Signaling 1 (SOCS1), and STAT1) that in common across the 4 sublines (Fig. 3). Of note, the M5.1 Fayoumi subline showed the largest number of genes (10) that were unique to the subline, whereas the Gsh6 Leghorn subline had none (Fig. 3).
Venn Diagram Demonstrating the Common and Uncommon Differentially Expressed Innate Immune Genes in each Subline. The Venn diagram compares the targets significantly expressed in the experimental groups versus the control groups of each of the four sublines. The Ghs6 subline is represented in the green color, Ghs13 in yellow, M5.1 in orange, and M15.2 in blush.
A heatmap depicting the differential expression patterns of the differentially expressed genes for each subline is shown in Fig. 4, including the 5 core genes. The expression of the core genes ranged from 9-fold increases (Ghs13 – SOCS1) to 194-fold increases (Ghs13 – Mx1). Hierarchical clustering of expression profiles of the differentially expressed genes shows the two Leghorn sublines, Ghs6 and Ghs13 closely clustered together (AU = 99), and the M15.2 Fayoumi subline is also clustering more closely with the Leghorns (AU = 93). The M5.1, which seems to be the “high responder” and has the largest amount of differentially expressed genes, is an outlier. However, the M15.2 subline clusters closer to M5.1 with more genes that are significantly upregulated in both of these Fayoumi sublines. This contrasts the clustering using the entire dataset for the gene expression profiles (Fig. 4 versus Fig. 1), identifying both breed- (Fig. 4) and subline-dependent (Fig. 1) expression of genes in the response to NDV, and suggesting a role for the MHC in regulation of the innate immune response.
Heatmap of the Differentially Expressed Innate Immune Genes in each Subline. The fold expression changes, log2(FoldChange), range from low upregulation (light red) to high upregulation (dark red) dark red. There are no genes signifincatly downregulated in the experimental versus control groups in any of the sublines. The gray represents targets that are not differentially expressed over the controls in the respective subline. The expression profiles of the sublines were clustered using hierarchical clustering in R (pvclust, nboot = 1000).
A set of five innate immune genes drives the conserved response to NDV in different chicken lines
The five conserved genes identified among the sublines (Fig. 3) were examined using the Ingenuity Pathway Analysis (IPA) software to define any canonical pathways or diseases/functions involved in the response to NDV. The analysis suggests that each of these five genes is involved in the interferon response (top canonical pathway, p-value 4.21 × 10−8) and the antiviral response (top diseases and function, p-value 1.86 × 10−10) (Fig. 5). The results show that IRF1, IRF7, and STAT1 are each known to activate the antiviral response (Fig. 5, orange arrows), while Mx1 and SOCS1 are also both involved in the antiviral response. However, the Mx1 gene effect is not predicted (Fig. 3, dark grey arrow) and the SCOS1 has inconsistent findings with downstream molecules (Fig. 3, yellow arrow). The interaction network analysis shows that the expression of each of the five genes is dependent upon and interconnected with the others (Fig. 5, light grey arrows). For instance, Mx1 directly interacts with IRF1, IRF7, STAT1, and itself (Fig. 5, solid light grey arrows) and the SOCS1 gene indirectly influences the Mx1 gene (Fig. 5, dashed light grey arrows).
The Conserved Response of chickens to NDV infection. The gene network, including the five core genes upregulated in response to NDV, and the top Disease & Function, the antiviral response, is shown. The data were analyzed through the use of IPA (QIAGEN Inc., https://www.qiagenbioinformatics.com/products/ingenuity-pathway-analysis). The red color demonstrates the average upregulation of the genes, the light grey are the gene-gene interactions (solid = direct, dashed = indirect) the orange color shows activation of that pathway, the dark grey shows the effect is not predicted, and the yellow shows findings not consistent with downstream molecules.
The breed-dependent response to NDV is enriched in the Fayoumi compared with the Leghorn breeds
The differentially expressed genes from the Leghorn and Fayoumi breeds are listed in Table 1 and the gene-gene network is depicted in Supplementary Fig. 2. These genes determined by examining the targets that are differentially expressed in both sublines of the respective breed. Both the Fayoumi and Leghorn responses include the five conserved genes. The responses differ by the expression of Interferon Induced with Helicase C Domain 1 (IFIH1), Lipopolysaccharide Induced TNF Factor (LITAF), Nitric Oxide Synthase 2 (NOS2), and Toll-like Receptor 3 (TLR3) in the Fayoumi and Cathelicidin Antimicrobial Peptide (CAMP), C-C Motif Chemokine Ligand 4 (CCL4), and Interleukin-8 (IL8) in the Leghorn (Table 1, bold text).
A subline-dependent response suggests a role for the MHC in innate immunity to NDV infection
The results show considerable differences in innate immune gene expression profiles in response to NDV infection between the Fayoumi sublines. For instance, the M5.1 subline significantly upregulated 13 unique genes as compared to only 4 in the M15.2 subline. Differentially expressed genes in the M5.1 subline include multiple interleukin genes (IL1B, IL4, IL6, IL8, and IL18) along with other cytokines, whereas the genes upregulated in the M15.2 subline includes myosin heavy chain 15 (MYH15), cluster of differentiation 40 (CD40), CAMP, and NFKB (Table 1, Supplementary Fig. 3).
A few genes within the Leghorn sublines were also identified as subline-specific. For instance, the Ghs13 subline significantly upregulates 3 genes, CD80, IFIH1, and myeloid differentiation primary response 88 (MYD88). This is in contrast with the Ghs6 subline in which only NOS2 was significantly upregulated (Table 1, Supplementary Fig. 4).
Discussion
NDV ranks amongst the most significant diseases that limit poultry production worldwide2. One approach to reduce the impact of NDV infection for smallholder farmers may be through the identification and introgression through selective breeding of innate immune related genes that reduce susceptibility of chickens to NDV, with the potential of, in the long-term, improving the livability and “hardiness” of backyard poultry and, indirectly, the productivity of smallholder farmers.
To begin to understand mechanisms underlying the innate immune response of chickens to NDV, our studies have begun to identify breed- and subline-dependent differences in gene expression profiles of both inbred and outbred chicken populations with known differential susceptibilities to NDV15. Our results demonstrate differences in viral load amongst the lines examined here. The M15.2 Fayoumi subline had the lowest viral load, followed by the M15.2 Fayoumi subline, then the two Leghorn sublines Ghs6 and Ghs13, respectively. Overall, the Fayoumi breed also had significantly lower expression than the Leghorn. Further, the use of congenic lines suggests a role for the MHC in modulating the innate immune response to NDV. Importantly, as evident from the clustering patterns of the sublines based on the expression profiles of significantly expressed genes, the two Leghorn sublines clustered with the Fayoumi M15.2 subline. The M5.1 Fayoumi subline that was somewhat of an outlier suggests the existence of both breed- and subline-depended transcriptional responses of chickens to NDV infection (Figs 1 and 4).
The analysis identified a set of genes that were uniformly upregulated – or a “conserved response” - across all sublines. This finding is consistent with earlier reports on the immune response of chickens to NDV and other avian pathogens7,14,15,21,22,23,24,25. For instance, the response of SPF White Leghorn chickens post-virulent NDV challenge identified Mx1, IRF1, IRF7, and SOCS1 were upregulated14. Thus, our studies with highly characterized and inbred lines confirms a role for these genes as part of the conserved response to NDV infection, as well as provides compelling evidence of conservation of the innate immune response between hatched chickens and the developing chicken embryo, further validating the use of the chick embryo model to examine the innate immune response. The five genes with the conserved responses across the sublines appear to be part of an interconnected network (Fig. 5), involving the activation of the Antiviral Response (p-value 8.16 × 10−13), and are likely to influence the expression of each other through multiple innate immune related pathways, including the interferon regulatory pathway, a significantly regulated canonical pathway (p-value < 6 × 10−10) from the Core Analysis performed in IPA. For instance, Mx1 is known to be induced by type I and III interferons and has antiviral activity against RNA viruses26. IRF1 and IRF7 are both regulators of the interferon response and act as an activator/repressor of multiple genes and the induction of signaling cascades of pattern recognition receptors, respectively27,28. In contrast, STAT1 is involved in the upregulation of genes once induced by type I, II, and III interferons and has direct interactions with the IRF genes29. SOCS1 is a negative regulator of cytokines and directly regulates the JAK-STAT pathway30. Thus, these studies suggest that these five conserved innate immune system related genes are involved in the interferon response of chickens, and likely have significant roles in the responsiveness of chickens to NDV regardless of breed. This is consistent as well with multiple other studies that have noted the importance of interferon signaling in the response to various poultry pathogens, including Marek’s Disease (MD), Infectious Bronchitis Virus (IBV), and AIV9,14,18,22,24,25,31,32.
Since the conserved response seems to be important regardless of breed or infectious agent, it is tempting to speculate that the breed- and subline-dependent differences in expression patterns may provide candidate markers for further investigation that may be plausible for selective breeding for genetic improvement of response (lower susceptibility) to NDV infection.
Interestingly, when examining the genes that are differentially regulated in both Fayoumi sublines versus both Leghorn sublines, there are differences with four genes (IFIH1, LITAF, NOS2, and TLR3) in the Fayoumi and three genes (CAMP, CCL4, and IL8) in the Leghorn that are unique. The response of the Fayoumi and Leghorns both include the conserved genes described above. Further interaction network analysis with the breed-dependent responses revealed a significantly more interconnected pathway for the Fayoumi versus the Leghorn (Supplementary Fig. 2). This core analysis also revealed the activation of the interferon response (p-values < 6 × 10−10) in both breeds, consistent with the genes involved in the conserved response. IFIH1 has been shown to be a positive antiviral gene and is also correlated with an enhanced response alongside purposive selection in some breeds33. The other genes upregulated in the Fayoumi all play key roles in the antiviral response as well, for example LITAF is a key regulator of the inflammatory response, NOS2 has been shown to be upregulated in response to virulent NDV infection, and TLR3 induces the interferon response14,34,35. The upregulation of these genes, combined with the generally higher response of the five conserved genes in the Fayoumis, may contribute to the overall enhanced response as compared to the Leghorn breed.
Although there were differences between the two breeds when examining the breed-dependent response, there are also differences seen in the subline-dependent response. Overall, the Fayoumi M5.1 subline has 12 genes unique to the subline as compared to four genes in the M15.2 subline. Interaction network analysis was also performed to examine the gene-gene networks of the sublines (Supplementary Figs 3 and 4). These responses are of interest since they may be influenced by the MHC or other genes on microchromosome 16.
Intriguingly, when investigating the gene-specific differences between the Fayoumi sublines, many interleukin genes are significantly expressed in the M5.1 subline, but not M15.2. The interleukins are secreted cytokines, particularly important in signaling the immune responses and inflammatory response36. One study on MD found that the expression of IL6 and IL18 was associated with the level of susceptibility of four Leghorn lines differing in susceptibility to MD23. The upregulation of the interleukins and other pro-inflammatory cytokines significantly influences many pathways as well, including the M5.1 subline represses the LXR/RXR Pathway and activates many others, including the IL6 and IL17 signaling pathways, the TLR pathway, and the role of pattern recognition receptors (PRRs) in recognition of virus and bacteria (Supplementary Fig. 3C). These cytokines, particularly the pro-inflammatory cytokines, play key roles in the avian immune response to pathogens and the control of infectious diseases in poultry37.
The differences seen between the Leghorn are one (Ghs6) and three (Ghs13) genes being subline-dependent, however, these genes may still be influencing the response to NDV (Supplementary Fig. 4). The NOS2 gene, significantly upregulated in the Ghs6 subline, is also significantly upregulated in both Fayoumi sublines, but is just below the cutoff of significance in the Ghs13 subline (p = 0.018, cutoff = 0.01). Demonstrating that this gene, although below our significance cut off, is still important in the response, and may be involved in the conserved response, which is consistent with previous studies showing the importance of the iNOS pathway in response to NDV infection14. The CD80 gene, significantly upregulated in the Ghs13 subline, has been found to be differentially regulated as part of the MHCII system in two meat-type broiler lines differing in susceptibility to Salmonella infection21. Although previous studies have not examined the relation of MYD88 and IFIH1 to the MHC, this study suggests a possible role for these genes in the immune response to NDV that need to be studied further.
Overall, our study is consistent with other studies regarding the core conserved gene expression response to NDV, as well as other poultry pathogens. However, the data suggest breed- and subline-dependent expression of innate immune genes that may serve as genetic markers associated with reduced susceptibility to NDV. Future studies are required to assess whether there are similar response patterns in outbred populations of chickens, as well as with more pathogenic field strains in the chick embryo. Studies associating these responses with the level of susceptibility of hatched chicks to NDV and other important avian pathogens are needed to better understand the chicken innate immune response. This study provides a framework for future efforts to improve the health and productivity of chickens through genetic selection for reduced susceptibility to NDV and other major diseases, particularly in low- and middle income countries where NDV remains endemic and continues to represent a major threat.
Methods
Ethics statement and animal use
The animal use protocol was approved by the Pennsylvania State University IACUC committee (protocol number 47175). All methods were performed in accordance with the relevant guidelines and regulations outlined in this protocol. Embryonated eggs from two inbred Leghorn lines, Ghs6 and Ghs13, and two inbred Fayoumi lines, M5.1 and M15.2, were provided from the Iowa State University (Ames, IA, USA). The eggs were received and incubated at 37.5 °C, 55% humidity, rotating hourly. The eggs were temporarily removed from the incubators to candle for viability and perform inoculations with virus.
Virus
The lentogenic LaSota strain of NDV was kindly provided by Dr. Siba Samal at the University of Maryland, College of Veterinary Medicine (College Park, MD, USA). Viral titrations were performed with the final titer of the undiluted viral suspension of 107 50% egg infectious dose (EID50)/mL. The viral suspension was stored at −80 °C until further use.
Embryonated egg inoculations and tissue harvest
Embryonated eggs from each subline, at 18 days of embryonic development, were candled and the airsacs were marked. Small holes were generated just above the airsac in the eggs using an egg punch, and 0.1 mL of the viral suspension was deposited directly to the allantoic fluid. The eggs were sealed with adhesive glue and placed back in the incubators until death or removal for tissue harvest. Controls (uninfected eggs) were treated similarly and inoculated with 0.1 mL of sterile 1X phosphate buffered saline (PBS).
The eggs were removed from the incubator 72 hours post inoculation, as close to hatch as possible, and placed at 4 °C for 3–4 hours to avoid opening eggs with viable chick embryos. The chick embryos were removed from the eggs and washed with 1X PBS. The lung tissues were harvested and stored at −80 °C prior to further use.
RNA extraction
RNA from the lung tissues of 30 experimental and 10 control embryos was extracted using the RNeasy Plus Kit (QIAGEN Inc., Germantown, MD, USA) following the recommended protocols after homogenization. The lung tissue was placed in a tube with 1 mL of RLT Lysis buffer (provided in the RNeasy kit) and 10–15 1.5 mm silica beads (Biospec Products, Bartlesville, OK, USA). The tissues were homogenized using a Mini-Beadbeater-96 (Biospec Products, Bartlesville, OK, USA) for 1 min. 600 uL of the homogenate was added to a clean microcentrifuge tube and centrifuged at 10,000 rpm for 2 min to remove any debris. The RNeasy Kit protocol was then followed with the supernatant after centrifugation.
cDNA synthesis
cDNA synthesis was performed immediately following RNA extraction using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Carlsbad, CA, USA). The manufacturer protocols were followed using 2 µg of each respective RNA sample. cDNA was stored at −20 °C.
OpenArray format and analysis
The OpenArray (Applied Biosystems, Carlsbad, CA, USA) format with 56 assays per sample and 48 samples per plate was selected (Supplementary Tables 2 and 3). The array contained 50 chicken innate immune genes, 2 viral genes, and 4 housekeeping genes. These genes were selected from our previous study in the chick embryo of different lines, other studies that found significance of these genes in the response to NDV and other pathogens, and availability of the TaqMan Gene Expression Assay from the Thermo Fisher Scientific inventory. Samples were sent to the Genomics Core Facility at the Pennsylvania State University (University Park, PA, USA) and the OpenArray was run on the QuantStudio 12 K Flex Real-Time PCR System (Applied Biosystems, Calrlsbad, CA, USA). Three technical replicates were run for each sample. The.eds files were opened using the ThermoFisher Cloud software (https://www.thermofisher.com/us/en/home/cloud.html) and exported as excel files that were used for downstream analysis.
Gene expression was analyzed using the △△Ct method comparing infected △Ct values (normalized with the Ct of the 4 housekeeping genes) with the average of the control △Ct values (normalized with the average Ct of the 4 housekeeping genes).
Figures were generated in R38. Hierarchical clustering was performed using pvclust package (nboot = 1000, hclust.method = “complete”, dist.method = “maximum”) and visualized with ggtree39,40. The volcano plot was generated using ggplot2 by plotting the log2(fold change) versus the -log(p-value)41, and the Venn diagram was generated with the VennDiagram package42.
Data were analyzed through the use of IPA (QIAGEN Inc., https://www.qiagenbioinformatics.com/products/ingenuitypathway-analysis). IPA was performed through inputting the datasets for each subline individually as well for each breed using the fold expression changes of the targets and the p-values from comparing the experimental versus control chick embryos. Core Analysis was performed for each dataset in IPA to describe the top canonical pathways and diseases/functions regulated by these genes. Comparison analyses of the core analysis for each were also generated in IPA. The gene-gene networks were generated using the Interaction Network Analysis through the build function when examining the pathways43. Although IPA is generally used for a larger set of genes, we were still able to examine the pathways, with a bias toward immune pathways (due to the input of only the expression of innate immune genes), that are significantly regulated post-NDV infection.
Accession codes
The dataset is publicly available at: https://www.ncbi.nlm.nih.gov/bioproject/479417.
References
Ashraf, A. & Shah, M. Newcastle disease: present status and future challenges for developing countries. African Journal of Microbiology Research 5, https://doi.org/10.5897/AJMR2013.6540 (2014).
Alexander, D. J., Aldous, E. W. & Fuller, C. M. The long view: a selective review of 40 years of Newcastle disease research. Avian Pathology 41, 329–335, https://doi.org/10.1080/03079457.2012.697991 (2012).
Brown, V. R. & Bevins, S. N. A review of virulent Newcastle disease viruses in the United States and the role of wild birds in viral persistence and spread. Veterinary Research 48, https://doi.org/10.1186/s13567-017-0457-9 (2017).
Muchadeyi, F. C. & Dzomba, E. F. In Poultry Science (ed. Milad Manafi) (2017).
Zhou, H. & Lamont, S. J. Genetic characterization of biodiversity in highly inbred chicken lines by microsatellite markers. Animal Genetics 30, 256–264 (1999).
Kim, D. K. et al. Immune-related gene expression in two B-complex disparate genetically inbred Fayoumi chicken lines following Eimeria maxima infection. Poultry Science 87, 433–443 (2008).
Cheeseman, J. H., Kaiser, M. G., Ciraci, C., Kaiser, P. & Lamont, S. J. Breed effect on early cytokine mRNA expression in spleen and cecum of chickens with and without Salmonella enteritidis infection. Developmental and Comparative Immunology 31, 52–60 (2007).
Wang, Y., Lupiani, B., Reddy, S. M., Lamont, S. J. & Zhou, H. RNA-seq analysis revealed novel genes and signaling pathways associated with disease resistance to avian influenza infection in chickens. Poultry Science 93, 485–493 (2014).
Deist, M. S. et al. Novel mechanisms revealed in the trachea transcriptome of resistant and susceptible chicken lines following infection with Newcastle disease virus. Clinical and Vaccine Immunology 24, https://doi.org/10.1128/CVI.00027-17 (2017).
Kim, D. K. et al. Immune-related gene expression in two B-complex disparate genetically inbred Fayoumi chicken lines following Eimeria maxima infection. Poultry Science 87, https://doi.org/10.3382/ps.2007-00383 (2008).
Sharma, J. M. Overview of the avian immune system. Veterinary Immunology and Immunopathology 30, 13–17 (1991).
Kapczynski, D. R., Afonso, C. L. & Miller, P. J. Immune responses of poultry to Newcastle disease virus. Developmental and Comparative Immunology 41, 447–453 (2013).
Ahmed, K. A. et al. Immune response to Newcastle disease virus in chicken lines divergently selected for cutaneous hypersensitivity. International Journal of Immunogenetics 34, 445–455 (2007).
Rue, C. A. et al. Virulent Newcastle disease virus elicits a strong innate immune response in chickens. Journal of General Virology 92, 931–939 (2011).
Schilling, M. A. et al. Transcriptional innate immune response of the developing chicken embryo to Newcastle disease virus infection. Frontiers in Genetics 9, https://doi.org/10.3389/fgene.2018.00061 (2018).
Davison, T. The immunologists’ debt to the chicken. British Poultry Science 44, https://doi.org/10.1080/0007166031000085364 (2010).
Ribatti, D. The chick embryo chorioallantoic membrane in the study of angiogenesis and metastasis. 1 edn, (Springer Netherlands, 2010).
Deist, M. S. et al. Resistant and susceptible chicken lines show distinctive responses to Newcastle disease virus infection in the lung transcriptome. BMC Genomics 18, https://doi.org/10.1186/s12864-017-4380-4 (2017).
Deist, M. S. et al. Novel analysis of the Harderian gland transcrptome response to Newcastle disease virus in two inbred chicken lines. Scientific reports 8, https://doi.org/10.1038/s41598-018-24830-0 (2018).
Miller, M. M. & Taylor, R. L. Brief review of the chicken Major Histocompatibility Complex: the genes, their distribution on chromosome 16, and their contributions to disease resistance. Poultry Science 95, https://doi.org/10.3382/ps/pev379 (2016).
Chiang, H.-I. et al. Gene expression profiling in chicken heterophils with Salmonella enteritidis stimulation using a chicken 44K Agilent microaray. BMC Genomics 9, https://doi.org/10.1186/1471-2164-9-526 (2008).
Hu, Z. et al. Strong innate immune response and cell death in chicken splenocytes infected with genotype VIId Newcastle disease virus. Virology Journal 9 (2012).
Kaiser, P., Underwood, G. & Davison, F. Differential cytokine responses following Marek’s Disease Virus infection of chicks differing in resistance to Marek’s Disease. Journal of Virology 77, https://doi.org/10.1128/JVI.77.1.762.2003 (2003).
Matulova, M. et al. Chicken innate immune response to oral infection with Salmonella enterica serovar Enteritidis. Veterinary Research 44 (2013).
Ranaware, P. B. et al. Genome wide host gene expression analysis in chickens lungs infected with Avian Influenza viruses. PLOSone 11, https://doi.org/10.1371/journal.pone.0153671 (2016).
Verhelst, J., Parthoens, E., Schepens, B., Fiers, W. & Saelens, X. Interferon-inducible protein Mx1 inhibits influenza virus by interferring with functional viral ribonucleoprotein complex assembly. Journal of Virology 86, https://doi.org/10.1128/JVI.01682-12 (2012).
Escalante, C. R., Yie, J., Thanos, D. & Aggarwal, A. K. Structure of IRF-1 with bound DNA revelas determinatnts of interferon regulation. Nature 391, https://doi.org/10.1038/34224 (1998).
Ning, S., Pagano, J. S. & Barber, G. N. IRF7: activation, regulation, modificaiton, and function. Genes and Immunity 12, https://doi.org/10.1038/gene.2011.21 (2011).
Lu, H. F. et al. Diallyl disulfide induced signal transducer and activator of transcription 1 expression in human colon cancer colo 205 cells using differential display RT-PCR. Cancer Genomics and Proteomics 4 (2007).
Du, C. et al. SOCS-1 is involved in TNF-a induced mitochondrial dysfunction and apoptosis in renal tubular epithelial cells. Tissue and Cell 49, https://doi.org/10.1016/j.tice.2017.06.005 (2017).
Heidari, M. et al. Marek’s disease virus-induced immunosuppression: Array analysis of chicken immune response gene expression profiling. Viral Immunology 23, https://doi.org/10.1089/vim.2009.0079 (2010).
Smith, J., Sadeyen, J.-R., Cavanagh, D., Kaiser, P. & Burt, D. W. The early immune response to infection of chickens with Infectious Bronchitis Virus (IBV) in susceptible and resistant birds. BMC Veterinary Research 11, https://doi.org/10.1186/s12917-015-0575-6 (2015).
Li, J. et al. Genotypes of IFIH1 and IFIT5 in seven chicken breeds indicated artifical selection for commercial traits influenced antiviral genes. Infection, Genetics, and Evolution 56, https://doi.org/10.1016/j.meegid.2017.10.019 (2017).
Hong, Y. H., SLillehoj, H. S., Lee, S. H., Park, D. W. & Lillehoj, E. P. Molecular cloning and characterization of chicken lipopolysaccharide-induced TNF-a factor (LITAF). Developmental and Comparative Immunology 30, https://doi.org/10.1016/j.dci.2005.12.007 (2006).
Karpala, A. J., Lowenthal, J. W. & Bean, A. G. Activation of the TLR3 pathway regulates IFNbeta produciton in chickens. Developmental and Comparative Immunology 32, https://doi.org/10.1016/j.dci.2007.08.004 (2008).
Britannica, T. E. O. T. E. In Encyclopedia Britannica (Encyclopedia Britannica, Inc., 1998).
Wigley, P. & Kaiser, P. Avian cytokines in health and disease. Rev Bras Cienc Avic 5, https://doi.org/10.1590/S1516-635X2003000100001 (2003).
Team, R. C. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2017).
Suzuki, R. & Shimodaira, H. pvclust: Hierarchical clustering with p-values via multiscale bootstrap resampling. R package version 2.0-0 (2015).
Yu, G., Smith, D., Zhu, H., Guan, Y. & Lam, T. T.-Y. ggtree: an R package for visualization and annotation of phylogenetic trees with thier covariates and other associated data. Methods in Ecology and Evolution 8, 28–36 (2017).
Wickham, H. ggplot2: Elegant graphics for data analysis. Springer-Verlag New York (2009).
Chen, H. VennDiagram: Generate high-resolution Venn and Euler plots. R package version 1.6.18 (2017).
IPA: Casual analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30 (2014).
Acknowledgements
The authors would like to thank the Penn State Genomics Core Facility for their support in performing the QuantStudio 12K Flex runs for the OpenArray plates. This research was supported by grant (OPP1083453) from the Bill and Melinda Gates Foundation (to J.J.B., V.K. and M.S.) and USDA-NIFA-Animal Health Funds (to S.J.L.) and training grant USDA-NIFA 2013-38420-20496 (M.S.D.).
Author information
Authors and Affiliations
Contributions
M.S. and V.K. developed the study concept and design. M.S., S.M. and M.C. performed experiments. M.S. analyzed data and drafted the manuscript. R.K., M.S.D., S.J.L., J.J.B. and V.K. critically revised the content. All authors have read and approved manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schilling, M.A., Memari, S., Cavanaugh, M. et al. Conserved, breed-dependent, and subline-dependent innate immune responses of Fayoumi and Leghorn chicken embryos to Newcastle disease virus infection. Sci Rep 9, 7209 (2019). https://doi.org/10.1038/s41598-019-43483-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-43483-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.