Abstract
Application of composted tannery sludge (CTS) could promote a shift in the structure of soil microbial communities. Although the effect of CTS on bacterial community has been studied, it is unclear how the composition and diversity of archaeal community respond to CTS amendment and which environmental factors drive the community over time. Here, we hypothesize that the Archaea structure and composition respond to CTS amendment over the time. CTS had been previously applied annually along 6 years and this assessment occurred for 180 days following the application in the 7th year by using different rates (0, 2.5, 5, 10 and 20 ton ha−1). We used amplicon 16S rRNA sequencing to assess the changes in the structure of the archaeal community. Thaumarchaeota and Euryarchaeota were the most abundant phyla found in soils with application of CTS, with Thaumarchaeota dominating the sequences in all samples with relative abundances of >98%. We observed a decreasing trend on the archaeal diversity over the time with increasing CTS application rate, together with an increase in the community similarity. The redundancy analyses (RDA) explained 43% of the total variation in operational taxonomic units and identified Na, pH, Cr and P as the main drivers of the archaeal community over time after application of highest CTS rates. CTS application changes the structure of Archaea community, with significant increase of Thaumarchaeota and Aenigmarchaeota groups, which can be further explored for its biotechnological use in contaminated soils.
Similar content being viewed by others
Introduction
Tannery sludge (TS) is an organic waste, produced in high amount, by tannery industries and contains high concentrations of salts, carbonates, hydroxides and chromium (Cr)1. Usually, this waste is disposed in the soil without previous treatment and, due to its chemical properties, could contribute to environmental pollution by affecting the soil properties, mainly biological ones2. Therefore, it is necessary to find suitable and efficient methods for TS treatment before its disposal in soil. Previous studies have proposed composting as a potential method for treating wastes before their disposal in soils3,4,5,6. However, the application of composted wastes could change the soil chemical and microbiological properties as well5.
Indeed, the application of composted wastes has effects on soil chemical properties. The application of composted tannery sludge (CTS) increases the Cr concentration, salinity and alkalinity1,6. The shifts in chemical properties could influence the soil microbial communities and consequently affect the biological processes in soil.
The knowledge about the factors driving the composition of microbial community in soil after application of waste is important to predict the potential effects on soil functioning since soil microorganisms play important roles in soil ecosystem7. Archaea community plays some fundamental ecological functions in the soil, including oxidation or reduction of metals and alteration of chemical conditions in the environment8. Since the application of CTS promotes changes in the soil environment, such as increased soil pH, salinity, Cr and organic matter content1, it is important to assess the effect of this waste on archaeal community. Previous studies have already evaluated the effect of tannery sludge on fungal and bacterial communities and revealed that these microorganisms are influenced by both characteristics of the waste and soil chemical properties9,10,11, but there are no studies focusing on effect of CTS on archaeal community. Considering the importance of Archaea in nitrogen and carbon turnover, and also their relationship with bacterial and fungal growth rates12, we addressed the following scientific questions in this paper: (1) how do the composition and diversity of archaeal community respond to CTS amendment over time? (2) which environmental factors drive the composition the Archaea community over time?
Results
CTS promoted significant increases in pH, total organic C (TOC), Cr and the content of total organic C, P, Ca, and Mg (Table S1). Cr, TOC, P and Ca increased significantly at all CTS rates. Through Next-Generation Sequencing (NGS) we assessed the structure and diversity of the archaeal community and, after the sequencing pre-processing, more than 360,000 quality sequences were used and grouped in 194 Operational Taxonomic Units (OTU), affiliated to five phyla and 10 classes. The most abundant phyla were Thaumarchaeota (98.5% of the total sequences), followed by Euryarchaeota (1.3%), Woesearchaeota (0.2%) and Aenigmarchaeota (0.05%). The response of the main archaeal phyla over time showed an increase in the relative abundance of Thaumarchaeota and a decrease in the relative abundance of Euryarchaeota; also Aenigmarchaeota increased with the CTS amendment (Fig. 1).
Distribution of the most abundant archaeal phyla between treatments and time after application of different rates of composted tannery sludge (CTS). Center lines in the boxplots show the medians; box limits indicate the 25th and 75th percentiles. Asterisks indicate significant differences within a treatment based on Tukey’s test (P < 0.05).
We observed a decreasing diversity over the time (Fig. 2A) and with the increased CTS application (Fig. 2B). An increase in the archaeal community similarity was observed with increasing CTS rate (Fig. S1), showing a homogenization of the community diversity.
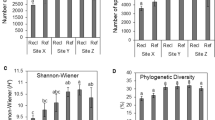
Shannon’s diversity index based on archaeal 16S rRNA genes of soil treated with composted tannery sludge (CTS) at different rates. (A) Variation of archaeal diversity according to time (0 to 180 days) after CTS application. (B) Average diversity for each treatment. Center lines in the boxplots show the medians; box limits indicate the 25th and 75th percentiles.
Physico-chemical properties explained 43% of the total variation in community structure based on OTU level, where 65% and 15% were displayed on the first (horizontal) and second (vertical) axes, respectively (Fig. 3). Treatments and sampling period were used as supplementary environmental variables. Soil samples were separated according to the treatments, with a clear pattern from the unamended soil to amended soils with increased CTS rate application, as confirmed by ANOSIM analysis (Table S2). This result revealed Na, pH, Cr and P as the most important factors driving the archaeal community composition over time.
Redundancy analysis diagram (RDA) of correlations between significant physico-chemical proprieties and OTU’s. (A) Rates of composted tannery sludge (CTS - 2.5, 5, 10 and 20 ton/ha); Time of sampling in days (0, 45, 75, 150 and 180); Total organic carbon – TOC; P – phosphorus; Ca – calcium; Cr – Chromium; pH – soil pH; EC – electric conductivity. All OTU’s are denoted by a small x-mark.
To further explore the correlation between archaeal phyla and soil chemical parameters, we calculated all possible Spearman’s rank correlation (Fig. 4). Thaumarchaeota, Woesearchaeota and Aenigmarchaeota showed a positive correlation with most of the variables while Euryarchaeota and MEG (Miscellaneous Euryarchaeotic Group) correlated negatively.
The Unweighted UniFrac distance metric with Principal Coordinates Analysis (PcoA) clustered the samples according to the CTS rate (Fig. S2). A pairwise comparison at OTU level between each treatment with the control revealed that the increased rate of CTS application increased the number of OTUs, where T2 presented three OTUs with increased abundance, followed by T3 with 15 OTUs, T4 with 22 and T5 with 23 OTUs with increased abundance in comparison with the control (Fig. S3).
The principal response curve (PRC) analysis at OTU level showed 10% of the total variance being attributed to the sampling time, while 45% was attributed to CTS rates (Fig. 5). Two OTUs presented a high positive response to the treatment (OTU802497748 and OTU477570063) with bk greater than 4; while three OTUs showed a high negative response (OTU64862401, OTU90781807, and OTU644603813) with bk lower than −4. Blast was used to better identify each responsive OTU showing extreme bk values in the PRC analysis and all of them belong to uncultured archaeon affiliated to the Thaumarchaeota phylum (Table S3).
Principal response curve (PRC) diagram of OTU data set indicating the effects of the composted tannery sludge (CTS) into the soil. The lines represent the course of the treatment levels in time. The OTU weight (bk) can be interpreted as the affinity of the OTU’s with the PRC. The Monte Carlo permutation test indicated that a significant part of the variance explained by treatment is displayed in the diagram (p = 0.014). Only OTUs with a bk value >1 or <−1 are shown for clarity. Asterisks indicate which treatments are significantly different from the controls (Williams test on PCA coordinates, see materials and methods section).
Co-occurrence network analysis showed an increase in complexity from T1 to T4, with a relatively low complexity in T5 (Fig. S4). The T4 treatment presents the highest complexity, with 165 correlations, and average degree of 9.16 (Table S4).
Discussion
We assessed the structure and diversity of archaeal community in response to application of CTS for seven years in soil. For this, we sequenced the V4 region of the 16S rRNA gene to describe patterns in taxonomical composition of soil archaeal communities and correlated them with CTS induced changes in soil parameters. As expected, Thaumarchaeota and Euryarchaeota were the most abundant phyla found in soils with application of CTS (Fig. 1), which corroborates previous studies that reported these phyla as dominant in several soils13,14,15. Interestingly, the phylum Thaumarchaeota was dominant in all samples and matches the fact that this phylum is found in high abundance in both marine and soils environments16,17. Members of this phylum play important roles in biogeochemical cycles, such as nitrogen and carbon18. This phylum represents a group of organisms that are able to oxidize ammonia aerobically and is important for the nitrification process in soils19. Considering that nitrogen is a key element controlling terrestrial productivity, the assessment of the effects of CTS on this community is of paramount importance.
In line with our first hypothesis, the relative abundance of archaeal phyla showed different responses to CTS amendment. The phylum Thaumarchaeota increased in relative abundance over time, while Euryarchaeota decreased (Fig. 1) as a response to short-term CTS application (180 days) in the seventh consecutive year. Probably, this increasing abundance is associated with the increase in soil pH over time (Table S2). Soil pH is the main driver of the abundance of Thaumarchaeota and this phylum contains groups adapted to different pH range, with Nitrososphaera as the group found under alkaline conditions20. In contrast, Euryarchaeota decreased probably as response to increasing Cr accumulation21, as confirmed by the strong negative correlation with CTS rate (Fig. 4). On the other hand, Aenigmarchaeota increased with increasing CTS rates, indicating that the waste stimulated the abundance of this phylum. This phylum was recently described22 and their members were found in wastewater from mines and sediments of hot springs23. Interestingly, they are adapted to extreme environments such as high pH variation, temperature and heavy metals23, which may explain the increase of this phylum in the samples with high CTS rates. Our results reinforce the observation that changes in soil chemical properties imposed by the CTS application have affected the different archaeal groups distinctly. The structure of archaeal communities is influenced by changing environmental factors, such as soil pH, moisture content, temperature and organic carbon, among others24.
The diversity of archaeal community suggests a detrimental effect of CTS, with a trend to decrease the diversity with the increase of CTS rate. The application of CTS changed some soil chemical properties, herewith selecting specific groups of Archaea, resulting in more homogenous communities with increased CTS rate (Fig. S1B)25.
The redundancy analysis (RDA) indicates that changes in soil chemical parameters over time due to CTS application resulted in changes in the composition of the archaeal community (Fig. 5). This finding is in agreement with previous studies that have shown that the archaeal communities are significantly affected by environmental variables, such as Na26, P27, pH28 and Cr21. Thus, our results confirm that different environmental drivers affect different groups of Archaea. Considering that most of the sequences were affiliated to Thaumarchaeota phylum, we suggest that the effects of CTS application may be better characterized at the phylogenetic level. Recently, Gubryn-Rangin et al.29 demonstrated links between the phylogeny of soil Thaumarchaeota and pH. The analysis based on the Unifrac distance corroborates our previous analysis revealing a differentiation of the archaeal community at the phylogenetic level, i.e. a difference of the species within the phylum Thaumarchaeota. In a study with 46 soils presenting 29 soils parameters, Oton et al.30 showed that adaptation to pH and organic matter content induced strong ecological coherence at various taxonomic level resolutions for Thaumarchaeota. Indeed, a pairwise comparison in our data revealed that the increased rate of CTS increased the number of OTUs that were selected and increased in relative abundance (Fig. S3).
As an important method for evaluating the response of community composition over time, the PRC method is a multivariate technique based on the redundancy analysis ordination technique. The PRC analysis pointed to specific OTUs that responded positively and negatively to the treatments compared with the control as they develop over time. Thus, it shows a pattern of archaeal OTUs presenting greater or lower sensitivity to CTS. This result resembles the effect of CTS on bacterial community as previously reported by Miranda et al.9. It means that the CTS amendment increased the abundance of specific OTUs that could show adaptation to pollution, i.e. high concentration of Cr, as reported before.
Finally, the co-occurrence network analysis showed that the network complexity increased with increasing CTS application rate (Fig. S4). The network analysis revealed that the CTS application directly affected the dynamics of the archaeal community, mainly the Thaumarchaeota group, by increasing the number of interaction among them. These effects on the community dynamics may be explained by the increased abundance of specific organisms (defined here at OTU level). We also identified the archaeal OTUs with more betweeness centrality, which may represent key taxa for a connected community31. Interestingly, for all the networks, the key OTU was affiliated to the genus Nitrososphaera (Table S5). This genus belongs to the group of ammonia oxidizing Archaea, an important player in the transformation of nitrogen32. Also, Nitrososphaera presents high metabolic versatility and can adapt to different environmental conditions33. A previous study has found biochemical characteristics of Nitrososphaera, such as chromate reductase activity, that contribute to its resistance to Cr34. This could explain the key role of this genus in the community dynamics in CTS-treated soils with high accumulation of Cr (Fig. S4 and Table S5). A genomic analysis of the species Nitrosphaera gargensis revealed the presence of two NADPH-dependent FMN reductases and four nitroreductases that could have a chromate reductase activity, which are suggested to be determinants of chromate resistance in bacteria33,34. In this sense, further studies could explore the role of specific members of Thaumarchaeota in bioremediation of soils contaminated with high rates of chromium.
Our results indicated a CTS-induced alteration of the archaeal communities with a decreasing diversity and increasing community similarity with CTS application rates. More specifically, we demonstrated that the phyla Thaumarchaeota and Aenigmarchaeota were the most responsive groups, which are able to resist to and potentially oxide contaminants, and can be further used as biotechnological products for contaminated sites.
Methods
This study was performed at the Federal University of Piauí, Brazil. CTS was obtained by mixing TS with sugarcane straw and cattle manure (ratio 1:3:1; v:v:v) after a process of composting over three months. The properties of CTS were assessed according APHA35 and USEPA36 (Table S6). This field experiment presents a history of CTS application, one time per year, for 6 years (2009–2014) in five doses: 0 (without CTS application), 2.5, 5, 10, and 20 ton/ha of CTS (dry basis). Detailed information of this experiment with CTS can be found in Sousa et al.37.
In 2015 (7th year of application), CTS was applied on the soil surface and incorporated into the 0–20 cm layer with a harrow. In order to measure the time-dependent effect of CTS on the diversity and structure of archaeal community, soils were sampled at 0, 45, 75, 150 and 180 days after CTS application from January to June, 2015. Thus, four replicates were sampled in each plot (0–20 cm), sieved (2 mm), and stored at −20 °C. The chemical properties of the soil (Table S1) were determined according to methods described in Embrapa38.
To assess the archaeal community profile we used the same procedures as described in Araujo et al.15, as follow: soil DNA was extracted from 0.5 g (field moist soil) of soil using the PowerLyzer PowerSoil DNA Isolation Kit (MoBIO Laboratories, Carlsbad, CA, USA), according to the manufacturer’s instructions. The DNA extraction was performed in triplicate for each soil sample. The quality and concentration of the extracted DNA was estimated using a Nanodrop 1000 (Thermo Scientific, Waltham, MA, USA). The amplicon library of the archaeal 16S rRNA genes was amplified using the V4 region-specific primers 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′)39. The first step amplification comprised 25 μL reaction containing the following solutions: 14.8 μL of nuclease-free water (Certified Nuclease-free, Promega, Madison, WI, USA), 2.5 μL of 10X High Fidelity PCR Buffer (Invitrogen, Carlsbad, CA, USA), 1.0 μL of 50 mM MgSO4, 0.5 μL of each primer (10 μM concentration, 200 pM final concentration), 1.0 unit of Platinum Taq polymerase High Fidelity (Invitrogen, Carlsbad, CA, USA), and 4.0 μL of template DNA (10 ng). The conditions for PCR were as follows: 94 °C for 4 min to denature the DNA, with 25 cycles at 94 °C for 45 s, 60 °C for 60 s, and 72 °C for 2 min, with a final extension of 10 min at 72 °C. In the second step, a unique pair of Illumina Nextera XT indexes (Illumina, San Diego, CA) was added to both ends of the amplified products. Each 50 μL reaction contained the following solutions: 23.5 μL of nuclease-free water (Certified Nuclease-free, Promega, Madison, WI, USA), 5.0 μL of 10X High Fidelity PCR Buffer (Invitrogen, Carlsbad, CA, USA), 4.8 μL of 25 mM MgSO4, 1.5 μL of dNTP (10 mM each), 5.0 μL of each Nextera XT index (Illumina, San Diego, CA, USA), 1.0 unit of Platinum Taq polymerase High Fidelity (Invitrogen, Carlsbad, CA, USA), and 5.0 μL of each product from previous PCR. The conditions for this second round PCR were as follows: 95 °C for 3 min to denature the DNA, with 8 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 min, with a final extension of 5 min at 72 °C.
After indexing, the PCR products were cleaned up using Agencourt AMPure XP–PCR purification beads (Beckman Coulter, Brea, CA, USA), according to the manufacturer’s manual, and quantified using the dsDNA BR assay Kit (Invitrogen, Carlsbad, CA, USA) on a Qubit 2.0 fluorometer (Invitrogen, Carlsbad, CA, USA). Once quantified, different volumes of each library were pooled into a single tube in equimolar concentration. After quantification, the molarity of the pool was determined and diluted to 2 nM, denatured, and then diluted to a final concentration of 8.0 pM with a 20% PhiX (Illumina, San Diego, CA, USA) spike for loading into the Illumina MiSeq sequencing device (Illumina, San Diego, CA, USA).
Sequence data were processed using QIIME 1.0 following the UPARSE standard pipeline according to Brazilian Microbiome Project40 to produce an OTU table and a set of representative sequences. Briefly, the reads were truncated at 130 bp and quality-filtered using a maximum expected error value of 1. Pre-filtered reads were dereplicated and singletons were removed using USEARCH 7.0. These sequences were clustered into OTUs at a 97% similarity cutoff following the UPARSE pipeline41. After clustering, the sequences were aligned and taxonomically assigned against the Silva database42.
We used analysis of similarity ANOSIM to test whether the treatments harbored significantly different microbial community structure. Alpha diversity was calculated from a matrix of richness, using the Shannon index. ANOSIM and alpha diversity index were calculated with the software PAST 343. Further, in order to test the significance of the treatment in clustering of the samples, we analyzed the community structure based on the OTU phylogenetic distance using unweight UniFrac dissimilarity44.
In order to visualize the differential archaeal community composition among treatments, the Statistical Analysis of Metagenomic Profiles (STAMP) software was used45. For this, the OTU table generated by QIIME was used as input and the P-values were calculated using the ANOVA with post-hoc test Tukey-Kramer, followed by Benjamini-Hochberg false discovery rate correction46. The analysis of similarity (SIMPER) was used to evaluate the community similarity within/ between treatments and time using the software Primer6 (PrimerE, Ivybridge, United Kingdom). Spearman’s rank correlation coefficient were calculated to explore the relationship between archaeal community with environmental factors according to the different treatment using the ‘multtest’ package in R47 and the correction was made using Benjamini-Hochberg false discovery rate. For visualization, heatmaps were constructed based on z-score transformed phylum abundance to improve normality and homogeneity of the variances. The z-score was calculated as the deviation from row mean in units of standard deviations above or below the mean.
In order to assess the complexity of the interactions among archaeal taxa, we performed network analysis according to Mendes et al.48. Briefly, non-random co-occurrence analyses were performed using SparCC49 based on a OTU table clustered at 97% identity. For each network, P-values were obtained by 99 permutations of random selections of the data table, subjected to the same analytical pipeline. In the networks, we included SparCC correlations statistically significant (P < 0.01) and with a magnitude of >0.7 or <−0.7. The nodes in the reconstructed network represent taxa at OTU level, whereas the edges represent significantly positive or negative correlations between nodes. The network graphs were made based on a set of measures, as number of nodes, number of edges, modularity, number of communities, average node connectivity, average path length, diameter and cumulative degree distribution. Co-occurrence analyses were carried out using the Python module ‘SparCC’ and networks visualization and properties measurements were calculated with the interactive platform Gephi50.
The structure of the Archaea community (as OTU’s) in the soil samples and its correlation with biological and environmental parameters was determined using Redundancy Analysis (RDA; length of gradient = 1.6). Since we analyzed relative abundance data, the data were arcsine transformed before analysis. The significance of each physico-chemical parameters for explaining the variation in community composition was tested with Monte Carlo permutation tests using sampling date as blocks. For data set an RDA analysis was performed using the significant physico-chemical parameters as explanatory variables and treatment and sampling date as passive explanatory variables. In addition, the dataset at the OTU level was analyzed by the principal response curve (PRC) method51 to show and test temporal changes in archaeal community of the soil caused by different rates of CTS as compared with those in the control (CTS-free soil) and also to identify specific taxa that responded positively or negatively to CTS application. The RDA and PRC analyses were performed using the CANOCO Software package, version 552.
The overall significance of the CTS treatment regime on the variation in OTU (p ≤ 0.05) was tested by performing 999 Monte Carlo permutations53. The NOECs of the tannery sludge treatment per sampling date was calculated by applying the Williams test54 to the sample scores of the first principal component of each sampling date53, using the Community Analysis computer program, version 4.3.0555 using a significance level of 0.0556.
References
Araujo, A. S. F., Silva, M. D. M., Leite, L. F. C., Araujo, F. F. & Dias, N. S. Soil pH, electric conductivity and organic matter after three years of consecutive applications of composted tannery sludge. African Journal of Agricultural Research 8, 1204–1208 (2013).
Patel, A. & Patra, D. D. Influence of heavy metal rich tannery sludge on soil enzymes vis-à-vis growth of Tagetes minuta, an essential oil bearing crop. Chemosphere 112, 323–332 (2014).
Araujo, A. S. F., Monteiro, R. T. R. & Carvalho, E. M. S. Effect of composted textile sludge on growth, nodulation and nitrogen fixation of soybean and cowpea. Bioresource Technology 98, 1028–1032 (2007).
Madrid, F., Lopez, R. & Cabrera, F. Metal accumulation in soil after application of municipal solid waste compost under intensive farming conditions. Agriculture, Ecosystems and Environment 119, 249–256 (2007).
Yuksel, O. Influence of municipal solid waste compost application on heavy metal content in soil. Environmental Monitoring and Assessment 187, 313–320 (2015).
Araujo, A. S. F. et al. Soil microbial properties after 5 years of consecutive amendment with composted tannery sludge. Environmental Monitoring and Assessment 187, 4153–4160 (2015).
Mendes, L. W. et al. Using metagenomics to connect microbial community biodiversity and functions. Current Issues in Molecular Biology 24, 103–118 (2017).
Bini, E. et al. Archaeal transformation of metals in the environment. FEMS Microbiology Ecology 73, 1–16 (2010).
Miranda, A. R. L. et al. Responses of soil bacterial community after seventh yearly applications of composted tannery sludge. Geoderma 318, 1–8 (2018).
Nakatani, A. S. et al. Changes in the genetic structure of Bacteria and microbial activity in an agricultural soil amended with tannery sludge. Soil Biology & Biochemistry 43, 106–114 (2011).
Nakatani, A. S. et al. Effects of tannery sludge application on physiological and fatty acid profiles of the soil microbial community. Applied Soil Ecology 61, 92–99 (2012).
Bengtson, P., Sterngren, A. & Rousk, J. Archaeal abundance across a pH gradient in an arable soil and its relationship with bacterial and fungal growth rates. Applied and Environmental Microbiology 78, 5906–5911 (2012).
Tripathi, B. M. et al. Soil pH and biome are both key determinants of soil archaeal community structure. Soil Biology & Biochemistry 88, 1–8 (2015).
Rodrigues, T., Belmok, A., Catao, E. & Kyaw, C. M. Archaea in natural and impacted brazilian environments. Archaea 1259608 (2016).
Araujo, A. S. F. et al. Archaea diversity in vegetation gradients from the Brazilian Cerrado. Brazilian Journal of Microbiology 49, 522–528 (2018).
Pester, M., Schleper, C. & Wagner, M. The Thaumarchaeota: an emerging view of their phylogeny and ecophysiology. Current Opinion in Microbiology 14, 300–306 (2011).
Fierer, N. Embracing the unknown: disentangling the complexities of the soil microbiome. Nature Reviews Microbiology 15, 579–590 (2017).
Walker, C. B. et al. Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine crenarchaea. Proceedings of the National Academy of Sciences USA 107, 8818–8823 (2010).
Könneke, M. et al. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437, 543–546 (2005).
Navarro-Noya, Y. E. et al. Archaeal Communities in a Heterogeneous Hypersaline-Alkaline Soil. Archaea 2015, 646829 (2015).
Li, B., Bao, Y., Xu, Y., Xie, S. & Huang, J. Vertical distribution of microbial communities in soils contaminated by chromium and perfluoroalkyl substances. Science of the Total Environment 599–600, 156–164 (2017).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Takai, K., Moser, D. M., DeFlaun, M., Onstott, T. C. & Fredrickson, J. K. Archaeal diversity in waters from deep South African gold mines. Applied and Environmental Microbiology 67, 5750–5760 (2001).
Lu, X., Seuradge, B. J. & Neufeld, J. D. Biogeography of soil Thaumarchaeota in relation to soil depth and land usage. FEMS Microbiology Ecology 93, fiw246 (2017).
Goss-Souza, D. et al. Soil microbial community dynamics and assembly under long-term land use change. FEMS Microbiology Ecology 93, 1–13 (2017).
Yan, L. et al. Distribution of archaeal communities along the coast of the gulf of finland and their response to oil contamination. Frontiers in Microbiology 9, 15–25 (2018).
Lin, X. et al. Microbial community stratification linked to utilization of carbohydrates and phosphorus limitation in a boreal peatland at Marcell Experimental Forest, Minnesota, USA. Applied and Environmental Microbiology 80, 3518–3530 (2014).
Siles, J. A. & Margesin, R. Abundance and diversity of bacterial, archaeal, and fungal communities along an altitudinal gradient in alpine forest soils: what are the driving factors? Microbial Ecology 72, 207–220 (2016).
Gubry-Rangin et al. Niche specialization of terrestrial archaeal ammonia oxidizers. Proceedings of National Academy of Science USA 108, 21206–21211 (2011).
Oton, E. V., Quince, C., Nicol, G. W., Prosser, J. I. & Gubry-Rangin, C. Phylogenetic congruence and ecological coherence in terrestrial Thaumarchaeota. The ISME Journal 10, 85–96 (2016).
Borgatti, S. P. Centrality and network flow. Social Networks 27, 55–71 (2005).
Tourna, M. et al. Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proceedings of the National Academy of Sciences 108, 8420–8425 (2011).
Spang, A. et al. The genome of the ammonia-oxidizing Candidatus Nitrososphaera gargensis: insights into metabolic versatility and environmental adaptations. Environmental Microbiology 14, 3122–3145 (2012).
Ramírez-Díaz et al. Mechanisms of bacterial resistance to chromium com- pounds. Biometals 21, 321–332 (2008).
APHA. Standard methods for the examination for water and wastewater. Washington: American Public Health Association. 1600 pp (2005).
USEPA. Method 3050B-Acid digestion of sediments, sludges and soils. (United States Environmental Protection Agency (USEPA), United States (1996).
Sousa, R. S. et al. Time-dependent effect of composted tannery sludge on the chemical and microbial properties of soil. Ecotoxicology 26, 1366–1377 (2017).
EMBRAPA - Manual de métodos de análise de solo. Rio de Janeiro, 212 p (1997).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proceedings of the National Academy of Sciences 108, 4516–4522 (2011).
Pylro, V. S. et al. Data analysis for 16S microbial profiling from different benchtop sequencing platforms. Journal of Microbiological Methods 107, 30–37 (2014).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research 41, D590–D596 (2013).
Hammer, O., Harper, D. A. T. & Ryan, P. D. PAST: Paleontological Statistic software package for education and data analysis. Paleontologia Eletronica 4, 1–9 (2001).
Lozupone, C. & Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Applied and Environmental Microbiology 71, 8228–8235 (2005).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Benjamini, Y. & Hochberg, Y. Controling the false Discovery rate – a practical and powerful approach to multiple testing. Journal of the Royal Statistical 57, 289–300 (1995).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2014).
Mendes, L. W., Raaijmakers, J. M., Hollander, M., Mendes, R. & Tsai, S. M. Influence of resistance breeding in common bean on rhizosphere microbiome composition and function. The ISME Journal 12, 212–224 (2018).
Friedman, J. & Alm, E. J. Inferring Correlation Networks from Genomic Survey Data. Plos Computational Biology 8, e1002687 (2012).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. ICWSM 8, 361–362 (2009).
Van den Brink, P. J. & Ter Braak, C. J. F. Principal response curves: analysis of time-dependent multivariate responses of a biological community to stress. Environmental Toxicology and Chemistry 18, 138–148 (1999).
Ter Braak, C. J. F. & Šmilauer, P. Canoco reference manual and user’s guide: software for ordination, version 5.0. Microcomputer Power (Ithaca, New York, USA, 2012).
Van den Brink, P. J., Van Wijngaarden, R. P. A., Lucassen, W. G. H., Brock, T. C. M. & Leeuwangh, P. Effects of the insecticide Dursban®4E (a.i. chlorpyrifos) in outdoor experimental ditches. II. Invertebrate community responses. Environmental Toxicology and Chemistry 15, 1143–1153 (1996).
Williams, D. A. The comparison of several dose levels with zero dose control. Biometrics 28, 519–531 (1972).
Hommen, U., Düllmer, U. & Vith, D. A computer program to evaluate plank-ton data from freshwater field tests. In: Hill, I. R., Heimbach, F., Leeuwangh, P. & Matthiesen, P. (Eds), Fresh water Field Tests for Hazard Assessment of Chemicals (Lewis Publishers, Baca Raton, USA, 1994).
Araujo, A. S. F., Lima, L. M., Santos, V. M. & Schmidt, R. Repeated application of composted tannery sludge affects differently soil microbial biomass, enzymes activity, and ammonia-oxidizing organisms. Environmental Science and Pollution Research International 23, 19193–19200 (2016).
Acknowledgements
Ana Roberta Lima Miranda thanks CAPES for her scholarship for going to Wageningen University. Ademir Sergio Ferreira de Araujo, Vania Maria Maciel Melo and Wanderley Jose de Melo thank CNPq for their fellowship of research. This research is supported by CNPq (Grant 305069/2018-7).
Author information
Authors and Affiliations
Contributions
A.R.L.M., J.E.L.A., L.W.M. and L.N.L. conducted the experiment(s) and analyzed the data. A.S.F.A. designed the experiment. V.M.M.M., L.W.M., L.N.L., W.J.M. and P.J.V.B. analyzed the results, discussed, wrote and reviewed the manuscript. All the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Miranda, A.R.L., Mendes, L.W., Lemos, L.N. et al. Dynamics of archaeal community in soil with application of composted tannery sludge. Sci Rep 9, 7347 (2019). https://doi.org/10.1038/s41598-019-43478-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-43478-y
This article is cited by
-
Applications of Cr-rich composted tannery sludge in the soil decrease microbial biomass and select specific bacterial groups
Environmental Science and Pollution Research (2022)
-
Dynamics of bacterial and archaeal communities along the composting of tannery sludge
Environmental Science and Pollution Research (2021)
-
Long-term effect of composted tannery sludge on soil chemical and biological parameters
Environmental Science and Pollution Research (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.