Abstract
Alcoholic liver disease (ALD) is a significant health hazard and economic burden affecting approximately 10 million people in the United States. ALD stems from the production of toxic-reactive metabolites, oxidative stress and fat accumulation in hepatocytes which ultimately results in hepatocyte death promoting hepatitis and fibrosis deposition. Monocyte-derived infiltrating Ly6Chi and Ly6Clow macrophages are instrumental in perpetuating and resolving the hepatitis and fibrosis associated with ALD pathogenesis. In the present study we isolated liver infiltrating macrophages from mice on an ethanol diet and subjected them to metabolomic and proteomic analysis to provide a broad assessment of the cellular metabolite and protein differences between infiltrating macrophage phenotypes. We identified numerous differentially regulated metabolites and proteins between Ly6Chi and Ly6Clow macrophages. Bioinformatic analysis for pathway enrichment of the differentially regulated metabolites showed a significant number of metabolites involved in the processes of glycerophospholipid metabolism, arachidonic acid metabolism and phospholipid biosynthesis. From analysis of the infiltrating macrophage proteome, we observed a significant enrichment in the biological processes of antigen presentation, actin polymerization and organization, phagocytosis and apoptotic regulation. The data presented herein could yield exciting new research avenues for the analysis of signaling pathways regulating macrophage polarization in ALD.
Similar content being viewed by others
Introduction
Alcoholic liver disease (ALD) affects approximately 10 million people in the United States and is a significant economic burden and public health hazard1. The pathogenesis of ALD stems from the production of toxic-reactive metabolites, reactive oxygen and nitrogen species (ROS and RNS), and oxidative stress associated with the metabolism of ethanol in hepatocytes2. Fat accumulation in hepatocytes (steatosis) is the earliest histopathological change in the liver associated with alcohol intake3. Continued steatosis results in hepatocyte death via apoptosis and necrosis which promotes inflammation and fibrosis formation4,5. A large number of individuals who develop fatty liver suffer no further complications while others progress from steatosis to hepatitis (liver inflammation). Persistent hepatitis and hepatocyte death can result in scar formation in the liver (cirrhosis) resulting in impaired liver function and altered architecture6. Persistent cirrhosis can ultimately lead to hepatocellular carcinoma and liver failure7.
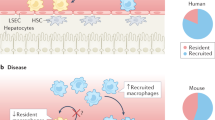
Macrophages are instrumental in promoting and resolving the hepatitis and fibrosis associated with ALD as evidenced by clinical observations that macrophage inflammatory genes are upregulated in ALD and cirrhosis patients8. Furthermore, hepatic macrophage activation and enhanced production of tumor necrosis factor α (TNFα), interleukin (IL)-6, chemokine (C-C motif) ligand 2 (CCL2) and ROS is elicited with ethanol administration in ALD animals9,10. Kupffer cells (KC), the liver resident macrophages, account for approximately 90% of the macrophage population in the healthy liver11. KC are primarily involved in the maintenance of tissue homeostasis by serving as immune sentinels sensing pathogens, antigens or damaged cells through interactions with numerous cell surface receptors to initiate and potentiate the inflammatory response12. The immune response to liver injury is initiated through the production of pro-inflammatory cytokines, IL-1β and TNFα by KC. Additionally, KC produce chemokines, such as CCL2, which induces the recruitment of additional inflammatory cells, such as monocytes, to the site of injury13. Inflammation progresses with the chemotactic recruitment of Ly6C+ monocytes to inflamed tissue that differentiate into Ly6Chi infiltrating macrophages (IMs)13. During acute or chronic liver injury, the macrophage subtype promoting inflammation in the liver are Ly6Chi monocyte-derived macrophages14,15. Ly6Chi macrophages exert pro-inflammatory, tissue-destructive responses as well as releasing pro-fibrotic mediators, such as IL-1β, platelet-derived growth factor (PDGF), connective tissue growth factor (CTGF) and transforming growth factor (TGF) β which activate hepatic stellate cells to deposit extracellular matrix and stimulate fibrosis formation16,17,18,19. While Ly6Chi macrophages initially exert pro-fibrotic and pro-inflammatory function they can differentiate into Ly6Clow macrophages to facilitate tissue repair and inflammation resolution20,21.
Macrophages represent an incredibly diverse cell type which, depending on tissue micro-environmental cues, switch from a pro- to anti-inflammatory phenotype in the progression of various diseases. The remarkable heterogeneity of macrophages is exemplified by their often opposing roles in a variety of diseases. For instance, pro-inflammatory macrophages are important in the elimination of extracellular pathogens, but are instrumental in the pathogenesis of atherosclerosis, autoimmune and metabolic diseases22. Anti-inflammatory macrophages are instrumental in wound healing and inflammation resolution but when not properly regulated, factor into the pathogenesis of asthma, fibrosis and cancer development23,24. During the progression of ALD, macrophages actively promote and resolve the inflammatory response, rendering therapeutic targeting of macrophages a significant challenge. Therefore, a thorough analysis of the metabolic and protein differences between Ly6Chi and Ly6Clow infiltrating macrophages following ethanol consumption is imperative in understanding the signaling pathways governing macrophage phenotypic switching. This mechanism could be harnessed for targeted therapeutic manipulation of macrophage populations in the liver.
In the current study, we isolated Ly6Chi and Ly6Clow macrophages from the livers of ethanol-fed mice and subjected the isolated cells to metabolomic and proteomic analysis to achieve an integrated bioinformatics approach. Here, we present an in-depth analysis of the altered metabolome and proteome between Ly6Chi and Ly6Clow liver infiltrating macrophages following ethanol consumption. The data herein elucidates novel signaling mechanisms governing macrophage phenotypic switching, with the potential for opening new avenues for therapeutic targeting macrophage polarization in ameliorating ALD progression.
Results
Comparative Metabolomic Analysis of Ly6Chi and Ly6Clow Infiltrating Macrophages Following Ethanol Administration
Infiltrating Ly6Chi and Ly6Clow liver macrophage populations from ethanol fed mice were isolated by flow sorting (Fig. 1). Following macrophage isolation, metabolites were separated from proteins using cold methanol extraction. Following methyl-tert-butyl ether (MTBE) liquid-liquid extraction, metabolites were analyzed by mass spectrometry (Fig. 2). After performing statistical analysis of the peak height intensities in Mass Profiler Professional, the ANOVA significant metabolites were uploaded to Metaboanalyst. We identified a number of metabolites with significant fold change differences between the Ly6Chi and Ly6Clow macrophages (Fig. 3). From the metabolite analysis, we observed 102 significantly altered metabolites between the macrophage subtypes (Table 1). In the lipid positive fraction, we detected 58 differentially regulated metabolites with 39 upregulated and 19 downregulated in the Ly6Clow compared to the Ly6Chi macrophages. From the lipid negative fraction, we measured 30 differentially regulated metabolites with 15 upregulated and 15 downregulated in the Ly6Clow compared to the Ly6Chi macrophages. In the aqueous fraction, we detected 14 differentially regulated metabolites with 8 being upregulated and 6 downregulated in Ly6Clow compared to the Ly6Chi macrophages. Following analysis with Metaboanalyst, we performed Metabolites Biological Role (MBROLE) analysis for pathway enrichment. From the 102 significantly altered metabolites we observed 6 pathways significantly enriched of which glycerophospholipid metabolism, arachidonic acid metabolism and phospholipid biosynthesis were further analyzed for their potential role in regulating macrophage polarization. (Table 2). Ly6Chi and Ly6Clow macrophages are significantly enriched for glycerophospholipid metabolism, metabolic pathways, arachidonic acid metabolism, linoleic metabolism and phospholipid biosynthesis with differential regulation of the metabolites involved in each functional pathway (Supplementary Table S1).
Liver macrophage flow sorting schematic. CD45 was used to select for myeloid cells. CD11b and SiglecF were used to gate out eosinophils (Eos, CD11b+ SiglecF+) from macrophages (Mϕ, CD11b+ SiglecF−). Macrophages F4/80 and CD11b were used to identify infiltrating macrophages (IM, CD11bhi F4/80Int) from Kupffer cells (KC, CD11bInt F4/80hi). Mixture of V450 conjugated anti-Ly6G, CD3, CD19, NK1.1 were used to gated out the neutrophils, lymophocytes and Nature Killer cells. IM were finally separated into the two infiltrating macrophage phenotypes based on expression level of Ly6C: Ly6Chi and Ly6Clow.
Significantly alter metabolites between Ly6Chi and Ly6Clow macrophages. (A) Log2 fold change of significantly altered metabolites (n = 3 in each experiment). The pink dots represent the significant metabolites. (B) Heat map of significantly altered metabolites (n = 3 in each experiment). Metabolites are significant with a fold change +/− 1.5 and t-test p < 0.05 when comparing Ly6Chi and Ly6Clow.
Comparative Proteomic Analysis of Ly6Chi and Ly6Clow Infiltrating Macrophages Following Ethanol Administration
Following methanol extraction of metabolites, the remaining protein pellet was subjected to protein extraction and tryptic digested for mass spectrometry proteomics analysis. Peptides detected by mass spectrometry were searched in Spectrum Mill to determine the protein identification. We detected 1,304 proteins in Ly6Chi and Ly6Clow macrophages with 340 and 214 proteins, respectively, uniquely expressed between macrophage subtypes (Fig. 4A). The 1,304 protein found in the Ly6Chi and Ly6Clow macrophages were subjected to DAVID analysis. From the 1,304 proteins analyzed, we observed 429 biological processes of which 105 were unique for Ly6Clow and 75 for Ly6Chigh macrophages (Fig. 4B). Furthermore, we detected 200 molecular functions from the 1,304 proteins of which 23 are unique for Ly6Clow and 28 for Ly6Chigh macrophages (Fig. 4C). The UniProt accession numbers for the common and unique proteins, biological processes and molecular functions are listed in the Supplementary Information Section (Supplementary Tables S3–S5). Protein quantitative analysis of significantly altered proteins was obtained from Mass Profiler Professional and we detected 47 differentially regulated proteins between the Ly6Chi and Ly6Clow macrophages (Table 3). The significantly altered proteins between the Ly6Chi and Ly6Clow macrophages were analyzed using the DAVID bioinformatics resource and we observed a total of 21 biological processes and 9 molecular functions from DAVID analysis of the protein quantification obtained (Supplementary Table S2). Of the significantly enriched biological processes and molecular functions, immune processes, actin polymerization and organization, phagocytosis, apoptotic processes and antigen presentation were selected for additional literature based analysis in their potential role for regulating macrophage polarization (Table 4).
Venn Diagrams of unique protein, biological processes and molecular functions between Ly6Chi and Ly6Clow macrophages. (A) Number of common and unique proteins between Ly6Chi and Ly6Clow macrophages. (B) Number of common and unique biological processes between Ly6Chi and Ly6Clow macrophages. (C) Number of common and unique molecular functions between Ly6Chi and Ly6Clow macrophages. Lists of common and unique protein, biological processes and molecular functions can be found in Supplementary Tables S3–S5.
Discussion
Alcoholic liver disease is a major public health issue and accounts for approximately 48% of liver cirrhosis related deaths1. As infiltrating macrophages are known to mediate the pathogenesis of ALD from steatosis to cirrhosis8,9,10, analysis of the altered signaling pathways between the different subsets of these cells in response to ethanol is of the utmost importance in developing treatment options to prevent the progression of ALD or promote the reversal of scar tissue formation in the liver. Macrophages display a remarkable capacity to adapt their phenotype based on tissue micro-environmental cues such as lipid exposure, hypoxia, cytokines, and efferocytosis of apoptotic cells21,25. To date, no studies have been conducted providing analysis of the cellular metabolome and proteome of infiltrating liver macrophages isolated from an in vivo model of ALD. While several studies have utilized immortalized mouse macrophages (RAW264.7) for transcriptomic26 and lipidomic26,27,28 analysis following inflammatory stimuli, this study is the first to look at in vivo polarized macrophages in an ALD model, therefore allowing for the natural effects of the tissue microenvironment, such as the gut-liver signaling axis, and ethanol metabolism on regulating liver infiltrating macrophage phenotypes.
It has previously been shown that following phagocytosis of apoptotic hepatocytes, Ly6Chi macrophages differentiate into Ly6Clow macrophages which express higher levels of phagocytosis related genes after alcohol intake21. In healthy or control diet fed mouse livers, infiltrating macrophages are limited until liver insult elicits the recruitment of Ly6C+ monocytes into the liver tissue.11,21,25 Therefore, the analysis done in this study was focused on the difference between Ly6Chi and Ly6Clow macrophages from ethanol fed mice without comparison to control diet fed animals. In our present study, we observed a significant increase in phagocytosis and engulfment related proteins (Table 4). We detected an upregulation of phagocytosis related proteins in Ly6Clow macrophages; this is expected as phagocytosis of apoptotic cells induces an anti-inflammatory phenotype29,30. Additionally, we saw a significant enrichment in proteins involved in regulating the apoptotic process. Furthermore, we observed a significant enrichment in actin polymerization and cytoskeletal reorganization in Ly6Clow macrophages. Alterations in actin contractility, cytoskeletal organization and cellular elongation have been shown to induce macrophages to an anti-inflammatory phenotype as evidenced by increased arginase-1 and YM-1 expression, hallmarks of anti-inflammatory macrophage polarization31. Additionally, defects in actin polymerization have been shown to attenuate macrophage phagocytic ability32. This suggests further in vivo analysis of actin polymerization and cytoskeletal organization in murine macrophages may elucidate a novel therapeutic strategy in modulating macrophage phenotypes in ALD by affecting macrophage phagocytosis and response to apoptotic stimuli.
Recently Zhang et al. provided a comprehensive analysis profiling lipid species during in vitro differentiation of mouse and human macrophages cell lines. They reported a significant increase in the composition of glycerophospholipid species during macrophage differentiation. Furthermore, they saw a significant increase in the levels of lysophospholipids in anti-inflammatory macrophages compared to pro-inflammatory macrophages suggesting that modulation of glycerophospholipid metabolism could be a vital signaling component in differentiation of liver macrophage phenotypes33. In our study, we found a significant enrichment in glycerophospholpid metabolism with differential metabolite regulation between Ly6Chi and Ly6Clow macrophages. Additionally, we observed enrichment for arachidonic acid metabolism and phospholipid biosynthesis (Table 2). In each of the enriched pathways, we detected a massive upregulation in multiple phosphatidylcholine (PC) species in Ly6Clow macrophages. PCs has been shown to promote an anti-inflammatory phenotype in macrophages through modulating actin assembly and increasing mycobacterium growth in RAW264.7 and J774 macrophages34. Likewise, we observed a substantial upregulation in phosphatidylethanolamine (PE(18:2/18:2) in Ly6Clow macrophages. Following stimulation with nonsteroidal anti-inflammatory agents, macrophages have been shown to display an increase in multiple PE species and take on an anti-inflammatory phenotype35. Therefore, the observed changes we see in PC and PE species correlate with in vitro studies highlighting the anti-inflammatory properties of PC and PE glycerophospholipid species in modulating macrophage phenotypes. Also of interest in regard to PE(18:2/18:2) is the linonleic acid (18:2) constituents present at the sn-1 and sn-2 positions, as linoleic acid has been shown to promote an anti-inflammatory phenotype in macrophages36. These results suggest the involvement of phospholipase A2 (PLA2) in regulating macrophage polarization in ALD. PLA2 is involved in the hydrolysis of sn-2 fatty acids from membrane glycerophospholipids yielding a free fatty acid, arachidonic acid, and a lysophospholipid37. The functions of PLA2 in modulating the inflammatory response have been well established in a variety of inflammatory contexts38,39,40,41,42. Ishihara et al. have shown that targeting cytosolic PLA2 activity in non-alcoholic fatty liver disease models proved beneficial in preventing hepatic fibrosis formation and reducing hepatocyte death43,44. Rodrigues et al. showed that using diethylcarbmazine, which modulates arachidonic acid metabolism and cyclooxygenase-2 (COX-2) mediated prostaglandin production, elicited an anti-inflammatory and protective response in ALD45. In addition to COX-2 mediated arachidonic acid metabolism and prostaglandin synthesis, arachidonic acid can be metabolized via cytochrome P450 epoxygenase mediated pathway to generate epoxyeicosatrienoic acids (EETs)46. We found a significant increase in EETs in the Ly6Clow phenotype. Endogenous EETs have been shown to regulate the ability of in vitro THP-1 monocytes to differentiate into pro-inflammatory macrophages in response to pro-inflammatory stimuli (lipopolysaccharide (LPS) and interferon γ (IFNγ) as well as preventing differentiation under anti-inflammatory stimuli (IL-4)46. Additionally, it has been shown that the immunomodulatory effect of EETs on inducing pro-inflammatory macrophage differentiation was facilitated through attenuation of NF-κB signaling47. Finally, studies have shown that eicosatrienoic acid inhibits LPS induced inflammatory gene expression in macrophages48. We detected an upregulation of eicosatrienoic acid metabolites in the anti-inflammatory, Ly6Clow macrophages after alcohol consumption. These studies coupled with the observed increase in arachidonic acid, glycerophospolipid metabolism and phospholipid biosynthesis as well as increased calcium ion binding suggest future investigation of the role of calcium dependent and independent PLA2 activity for therapeutic targeting of macrophage polarization in ALD.
The present study provides a framework for future studies utilizing multi-omics approaches for analyzing signaling difference between pro- and anti-inflammatory macrophages isolated from ALD mouse models. We detected a number of metabolic and protein mediated pathways that were significantly altered between the two macrophage subtypes, validating a number of in vitro studies analyzing the lipid, metabolite, and protein profile of polarized macrophages26,27,28,33,48. While the present study utilized an ALD model in which the degree of inflammation is not as evident histopathologically as more aggressive models, such as the NIAAA model, it allowed for the sufficient isolation of infiltrating liver macrophages not normally present in the healthy liver. We identified a number of metabolic pathways significantly altered due to the early onset of alcohol-induced hepatic inflammation (arachidonic acid metabolism, glycerophospholipid metabolism and phospholipid biosynthesis), which suggests that PLA2 enzymes play a critical role in modulating macrophage phenotypes. To explore the impact of PLA2 on ALD, future studies could utilize whole body PLA2 knockout mice or known PLA2 pharmacological inhibitors to elucidate the impact of PLA2 on macrophage polarization in ALD models. Overall, the data presented here justifies a further need to investigate numerous signaling mechanisms implicated in the modulation of macrophage phenotypes during ALD.
Materials and Methods
Animal Model
Female C57BL/6 J mice (The Jackson Laboratory, Bar Harbor, ME, USA) (n = 30) were maintained under pathogen-free conditions in the Center for Laboratory Animal Care at the University of Colorado Anschutz Medical Campus (Aurora, CO, USA). All experiments were performed using an Institutional Animal Care and Use Committee (IACUC) approved protocol and in accordance to the guidelines of the IACUC at the University of Colorado Anschutz Medical Campus. To elicit infiltrating macrophage recruitment to the liver, mice were fed an ethanol-containing Lieber-Decarli liquid diet (Bio-Serv, Flemington, NJ, USA). Ethanol content was introduced gradually by increasing 1.6% (v/v) every 2 days until 5%. All mice were then fed the liquid diet containing 5% ethanol for 4 weeks, as described previously49,50.
Isolation of Liver Non-Parenchymal Cells (NPCs)
Liver NPCs were isolated following a previously described method51. Briefly, a 20-G catheter was put through the mouse superior vena cava, the inferior vena cava was clamped, and the portal vein cut. The liver was perfused with Hank’s balanced salt solution (HBSS), followed by a digestion buffer [1 × HBSS, supplemented with 0.04% collagenase (type IV; Sigma, St. Louis, MO, USA), 1.25 mM CaCl2, 4 mM MgSO4, and 10 mM HEPES]. After digestion, the liver was disrupted in ACD solution (1 × HBSS, supplemented with 0.5% FBS, 0.6% citrate-dextrose solution, and 10 mM HEPES). Single cells were passed through a 100-μm cell strainer, and the cells were fractionated using 30% (w/v) Nycodenz (Axis-Shield PoC AS, Oslo, Norway) at 1.155 g/mL to yield liver NPCs and further purified using 30% Percoll (Sigma) at 1.04 g/mL.
Flow Cytometry Assisted Cell Sorting (FACS)
To purify KCs, Ly6Chi and Ly6Clow IMs, liver NPCs were incubated with normal rat serum (Sigma) and anti-mouse FcγRII/III (Becton Dickinson, Franklin Lakes, NJ, USA) to minimize nonspecific antibody binding. Subsequently, the cells were stained with anti-CD45, anti-Ly6C, anti-Ly6G, anti-CD19, anti-SiglecF (Becton Dickinson) and anti-F4/80, anti-CD11b, anti-NK1.1 and anti-CD3 (eBioscience, San Diego, CA, USA), and sorted using a BD FACSAria II Cell Sorter (BD Bioscience, San Jose, CA, USA).
Metabolomics Sample Preparation and Analysis
Cell pellets from different sort dates were combined in order to get 3 technical replicates of approximately 400–500 K cells per sample type (Ly6Chi and Ly6Clow). Extractions were performed using volumes of 70% MeOH/water and 100% MeOH based on cell numbers. Cold methanol was used to precipitate proteins prior to liquid-liquid extraction of metabolites. Proteins pellets were saved for future proteomics analysis. Liquid-liquid extraction was performed on the supernatant using water and methyl tert-butyl ether (MTBE).The aqueous and lipid fractions were retained for analysis. Lipid fractions were analyzed using SB-C18 HPLC analytical column in positive and negative ionization mode on the Agilent 6560 IM-QTOF (in QTOF mode only). Aqueous fractions were analyzed using a HILIC column in positive ionization mode on the Agilent 6560 IM-QTOF (in QTOF mode only). A pooled sample was used as instrument QCs to monitor the entire instrument analysis. Initial data QC, peak threshold evaluation, retention time variation, and charge carrier evaluation was performed in Agilent MassHunter Qualitative Analysis, version B.07.00. Data extraction was performed in MassHunter Profinder, version B.08.00. Differential Analysis was performed in Agilent Mass Profiler Professional (MPP), version 14.5. Compound annotation (database searches and molecular formula generation) was performed in MassHunter ID Browser software, version 14.5. Raw MS data were checked for quality and reproducibility. Appropriate spectral and chromatogram peak height thresholds were determined by careful examination of the raw data. Appropriate charge carriers to be allowed during data extraction were determined after preliminary extraction on selected samples. The Agilent “recursive workflow” was used to prepare data. This workflow includes the following steps: 1) untargeted extraction using the Find-by-Molecular Feature algorithm, 2) mass and time alignment of extracted compounds, 3) targeted extraction using the Find-by-Ion algorithm (using the list of ions prepared in step 1), 4) final mass and time alignment of extracted compounds.
Metaboanalyst and Metabolites Biological Role (MBROLE) Analysis
For Metaboanalyst comparison the following analysis parameters were used: Mass Tolerance: 0.05, No Missing Value Imputation, Data Filtering: Mean Intensity Value, Sample Normalization: Normalization by Sum, Data Transformation: None, Data Scaling: Mean Centering, Fold Change Threshold: 2, T-test: Group Variance Equal. For MBROLE metabolite functional enrichment analysis, pathways were considered significant with a p < 0.05.
Proteomics Sample Preparation, nHPLC-MS and nHPLC-MS/MS Analysis
Following methanol extraction for metabolomics, the remaining cell pellets from each technical replicate were processed using the PreOmics iST 8x Kit (Cat # 00001) following the included protocol. Digested macrophage samples were loaded onto a 2 cm PepMAP 100, nanoviper trapping column and chromatographically resolved on-line using a 0.075 × 250 mm, 2.0 µm Acclaim PepMap RSLC reverse phase nano column (Thermo Scientific) using a 1290 Infinity II LC system equipped with a nanoadapter (Agilent). Mobile phases consisted of water + 0.1% formic acid (A) and 90% aq. acetonitrile + 0.1% formic acid (B). Samples were loaded onto the trapping column at 3.2 μL/min for 2.5 minutes at initial condition before being chromatographically separated at an effective flow rate of 345 nl/min using a gradient of 3–8.5% B over 4.0 minutes, 8.5–26% B over 48.5 minutes, and 26–35% over 7.5 minutes for a total 60 minute gradient. The gradient method was followed by a column wash at 70% B for 5 minutes. For nHPLC-MS, data was collected with a 6550 QTOF equipped with a nano source (Agilent) operated in MS mode. For nHPLC-MS/MS, data was collected with a 6550 QTOF equipped with a nano source (Agilent) operated using Data Dependent Acquisition CID Auto MS/MS. The capillary voltage, drying gas flow, and drying gas temperature were set to 1300 V, 11.0 l/min, and 200 C, respectively. Data was collected in positive ion polarity over mass ranges 290–1700 m/z at a scan rate of 1.5 spectra/s. MS/MS scans were collected over mass ranges 50–1700 m/z at a scan rate of 3 spectra/s. Singly charged species were excluded from being selected during MS/MS acquisition. Following data acquisition in MS/MS mode, sample data was searched in SpectrumMill to identify proteins.
DAVID Bioinformatics Analysis
Functional pathway enrichment of significantly altered proteins between Ly6Chi and Ly6Clow macrophage population was analyzed using the Database for Annotation, Visualization and Integrated Discovery (DAVID) Bioinformatics Resource 6.8. For pathway enrichment, significantly altered proteins were compared to the whole mouse background. Pathways were considered significant with at least 3 proteins involved, a fold enrichment >2, and a p < 0.05.
Statistical Analysis
Statistical analysis of significantly altered metabolites and proteins was determined using Mass Profiler Professional Software. For Metaboanalyst, significantly altered metabolites were determined based of the difference in peak height intensity between the analyze metabolites with a p < 0.05. For MBROLE analysis for metabolite functional pathway enrichment, pathways were considered significant with a p < 0.05. For DAVID pathway enrichment, significantly altered proteins were compared to the whole mouse background. Pathways were considered significant with at least 3 proteins involved, a fold enrichment >2, and a p < 0.05.
References
Gao, B. & Bataller, R. Alcoholic liver disease: pathogenesis and new therapeutic targets. Gastroenterology 141, 1572–1585, https://doi.org/10.1053/j.gastro.2011.09.002 (2011).
Rubio, A. et al. Identification of a gene-pathway associated with non-alcoholic steatohepatitis. J Hepatol 46, 708–718, https://doi.org/10.1016/j.jhep.2006.10.021 (2007).
Baraona, E. & Lieber, C. S. Effects of ethanol on lipid metabolism. J Lipid Res 20, 289–315 (1979).
Feldstein, A. E. et al. Hepatocyte apoptosis and fas expression are prominent features of human nonalcoholic steatohepatitis. Gastroenterology 125, 437–443 (2003).
Wieckowska, A. et al. In vivo assessment of liver cell apoptosis as a novel biomarker of disease severity in nonalcoholic fatty liver disease. Hepatology 44, 27–33, https://doi.org/10.1002/hep.21223 (2006).
Marra, F., Gastaldelli, A., Svegliati Baroni, G., Tell, G. & Tiribelli, C. Molecular basis and mechanisms of progression of non-alcoholic steatohepatitis. Trends Mol Med 14, 72–81, https://doi.org/10.1016/j.molmed.2007.12.003 (2008).
Capece, D. et al. The inflammatory microenvironment in hepatocellular carcinoma: a pivotal role for tumor-associated macrophages. Biomed Res Int 2013, 187204, https://doi.org/10.1155/2013/187204 (2013).
Tapia-Abellan, A. et al. The peritoneal macrophage inflammatory profile in cirrhosis depends on the alcoholic or hepatitis C viral etiology and is related to ERK phosphorylation. BMC Immunol 13, 42, https://doi.org/10.1186/1471-2172-13-42 (2012).
Petrasek, J. et al. IL-1 receptor antagonist ameliorates inflammasome-dependent alcoholic steatohepatitis in mice. J Clin Invest 122, 3476–3489, https://doi.org/10.1172/JCI60777 (2012).
Enomoto, N. et al. Role of Kupffer cells and gut-derived endotoxins in alcoholic liver injury. J Gastroenterol Hepatol 15(Suppl), D20–25 (2000).
Ju, C. & Tacke, F. Hepatic macrophages in homeostasis and liver diseases: from pathogenesis to novel therapeutic strategies. Cell Mol Immunol 13, 316–327, https://doi.org/10.1038/cmi.2015.104 (2016).
Sica, A., Erreni, M., Allavena, P. & Porta, C. Macrophage polarization in pathology. Cell Mol Life Sci 72, 4111–4126, https://doi.org/10.1007/s00018-015-1995-y (2015).
Serbina, N. V., Shi, C. & Pamer, E. G. Monocyte-mediated immune defense against murine Listeria monocytogenes infection. Adv Immunol 113, 119–134, https://doi.org/10.1016/B978-0-12-394590-7.00003-8 (2012).
Karlmark, K. R. et al. Hepatic recruitment of the inflammatory Gr1+ monocyte subset upon liver injury promotes hepatic fibrosis. Hepatology 50, 261–274, https://doi.org/10.1002/hep.22950 (2009).
Tacke, F. & Randolph, G. J. Migratory fate and differentiation of blood monocyte subsets. Immunobiology 211, 609–618, https://doi.org/10.1016/j.imbio.2006.05.025 (2006).
Kolb, M., Margetts, P. J., Anthony, D. C., Pitossi, F. & Gauldie, J. Transient expression of IL-1beta induces acute lung injury and chronic repair leading to pulmonary fibrosis. J Clin Invest 107, 1529–1536, https://doi.org/10.1172/JCI12568 (2001).
Kinnman, N. et al. PDGF-mediated chemoattraction of hepatic stellate cells by bile duct segments in cholestatic liver injury. Lab Invest 80, 697–707 (2000).
Seki, E. et al. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat Med 13, 1324–1332, https://doi.org/10.1038/nm1663 (2007).
Paradis, V. et al. Effects and regulation of connective tissue growth factor on hepatic stellate cells. Laboratory Investigation 82, 767–773, https://doi.org/10.1097/01.Lab.0000017365.18894.D3 (2002).
Dal-Secco, D. et al. A dynamic spectrum of monocytes arising from the in situ reprogramming of CCR2+ monocytes at a site of sterile injury. J Exp Med 212, 447–456, https://doi.org/10.1084/jem.20141539 (2015).
Wang, M. et al. Chronic alcohol ingestion modulates hepatic macrophage populations and functions in mice. J Leukoc Biol 96, 657–665, https://doi.org/10.1189/jlb.6A0114-004RR (2014).
Lumeng, C. N., Bodzin, J. L. & Saltiel, A. R. Obesity induces a phenotypic switch in adipose tissue macrophage polarization. J Clin Invest 117, 175–184, https://doi.org/10.1172/JCI29881 (2007).
Wynn, T. A. IL-13 effector functions. Annu Rev Immunol 21, 425–456, https://doi.org/10.1146/annurev.immunol.21.120601.141142 (2003).
Movahedi, K. et al. Different tumor microenvironments contain functionally distinct subsets of macrophages derived from Ly6C(high) monocytes. Cancer Res 70, 5728–5739, https://doi.org/10.1158/0008-5472.CAN-09-4672 (2010).
Marentette, J. & Ju, C. In Molecules, Systems and Signaling in Liver Injury Cell Death in Biology and Diseases (eds W. X. Ding & X. M. Yin) 53–74 (Springer, Cham, 2017).
Maurya, M. R. et al. Analysis of inflammatory and lipid metabolic networks across RAW264.7 and thioglycolate-elicited macrophages. J Lipid Res 54, 2525–2542, https://doi.org/10.1194/jlr.M040212 (2013).
Andreyev, A. Y. et al. Subcellular organelle lipidomics in TLR-4-activated macrophages. J Lipid Res 51, 2785–2797, https://doi.org/10.1194/jlr.M008748 (2010).
Dennis, E. A. et al. A mouse macrophage lipidome. J Biol Chem 285, 39976–39985, https://doi.org/10.1074/jbc.M110.182915 (2010).
Huynh, M. L., Fadok, V. A. & Henson, P. M. Phosphatidylserine-dependent ingestion of apoptotic cells promotes TGF-beta1 secretion and the resolution of inflammation. J Clin Invest 109, 41–50, https://doi.org/10.1172/JCI11638 (2002).
Pupjalis, D., Goetsch, J., Kottas, D. J., Gerke, V. & Rescher, U. Annexin A1 released from apoptotic cells acts through formyl peptide receptors to dampen inflammatory monocyte activation via JAK/STAT/SOCS signalling. EMBO Mol Med 3, 102–114, https://doi.org/10.1002/emmm.201000113 (2011).
McWhorter, F. Y., Wang, T., Nguyen, P., Chung, T. & Liu, W. F. Modulation of macrophage phenotype by cell shape. Proc Natl Acad Sci USA 110, 17253–17258, https://doi.org/10.1073/pnas.1308887110 (2013).
Ninkovic, J. & Roy, S. Morphine decreases bacterial phagocytosis by inhibiting actin polymerization through cAMP-, Rac-1-, and p38 MAPK-dependent mechanisms. Am J Pathol 180, 1068–1079, https://doi.org/10.1016/j.ajpath.2011.11.034 (2012).
Zhang, C. et al. Quantitative profiling of glycerophospholipids during mouse and human macrophage differentiation using targeted mass spectrometry. Sci Rep 7, 412, https://doi.org/10.1038/s41598-017-00341-2 (2017).
Treede, I. et al. Anti-inflammatory effects of phosphatidylcholine. J Biol Chem 282, 27155–27164, https://doi.org/10.1074/jbc.M704408200 (2007).
Zhang, G., Zhao, L., Zhu, J., Feng, Y. & Wu, X. Anti-inflammatory activities and glycerophospholipids metabolism in KLA-stimulated RAW 264.7 macrophage cells by diarylheptanoids from the rhizomes of Alpinia officinarum. Biomed Chromatogr 32, https://doi.org/10.1002/bmc.4094 (2018).
de Gaetano, M., Alghamdi, K., Marcone, S. & Belton, O. Conjugated linoleic acid induces an atheroprotective macrophage MPhi2 phenotype and limits foam cell formation. J Inflamm (Lond) 12, 15, https://doi.org/10.1186/s12950-015-0060-9 (2015).
Rastogi, P. & McHowat, J. Inhibition of calcium-independent phospholipase A2 prevents inflammatory mediator production in pulmonary microvascular endothelium. Respir Physiol Neurobiol 165, 167–174, https://doi.org/10.1016/j.resp.2008.11.006 (2009).
Marentette, J., Kolar, G. & McHowat, J. Increased susceptibility to bladder inflammation in smokers: targeting the PAF-PAF receptor interaction to manage inflammatory cell recruitment. Physiol Rep 3, https://doi.org/10.14814/phy2.12641 (2015).
Marentette, J. & McHowat, J. Recruitment of inflammatory cells to the bladder endothelium exposed to cigarette smoke extract. Faseb J 28 (2014).
Sharma, J. et al. Lung endothelial cell platelet-activating factor production and inflammatory cell adherence are increased in response to cigarette smoke component exposure. Am J Physiol-Lung C 302, L47–L55, https://doi.org/10.1152/ajplung.00179.2011 (2012).
Kispert, S. E. et al. Cigarette smoke-induced urothelial cell damage: potential role of platelet-activating factor. Physiol Rep 5, https://doi.org/10.14814/phy2.13177 (2017).
Sharma, J. et al. Mice with Genetic Deletion of Group VIA Phospholipase A2beta Exhibit Impaired Macrophage Function and Increased Parasite Load in Trypanosoma cruzi-Induced Myocarditis. Infect Immun 84, 1137–1142, https://doi.org/10.1128/IAI.01564-15 (2016).
Ishihara, K., Kanai, S., Tanaka, K., Kawashita, E. & Akiba, S. Group IVA phospholipase A(2) deficiency prevents CCl4-induced hepatic cell death through the enhancement of autophagy. Biochem Biophys Res Commun 471, 15–20, https://doi.org/10.1016/j.bbrc.2016.01.186 (2016).
Ishihara, K. et al. Group IVA phospholipase A2 participates in the progression of hepatic fibrosis. Faseb J 26, 4111–4121, https://doi.org/10.1096/fj.12-205625 (2012).
Rodrigues, G. B. et al. Diethylcarbamazine: possible therapeutic alternative in the treatment of alcoholic liver disease in C57BL/6 mice. Clin Exp Pharmacol Physiol 42, 369–379, https://doi.org/10.1111/1440-1681.12369 (2015).
Bystrom, J. et al. Endogenous epoxygenases are modulators of monocyte/macrophage activity. PLoS One 6, e26591, https://doi.org/10.1371/journal.pone.0026591 (2011).
Dai, M. et al. Epoxyeicosatrienoic acids regulate macrophage polarization and prevent LPS-induced cardiac dysfunction. J Cell Physiol 230, 2108–2119, https://doi.org/10.1002/jcp.24939 (2015).
Huang, W. C., Tsai, P. J., Huang, Y. L., Chen, S. N. & Chuang, L. T. PGE2 production is suppressed by chemically-synthesized Delta7-eicosatrienoic acid in macrophages through the competitive inhibition of COX-2. Food Chem Toxicol 66, 122–133, https://doi.org/10.1016/j.fct.2014.01.031 (2014).
Horiguchi, N. et al. Cell type-dependent pro- and anti-inflammatory role of signal transducer and activator of transcription 3 in alcoholic liver injury. Gastroenterology 134, 1148–1158, https://doi.org/10.1053/j.gastro.2008.01.016 (2008).
Zhang, X. et al. Interleukin-6 is an important mediator for mitochondrial DNA repair after alcoholic liver injury in mice. Hepatology 52, 2137–2147, https://doi.org/10.1002/hep.23909 (2010).
Holt, M. P., Cheng, L. L. & Ju, C. Identification and characterization of infiltrating macrophages in acetaminophen-induced liver injury. J Leukocyte Biol 84, 1410–1421, https://doi.org/10.1189/jlb.0308173 (2008).
Acknowledgements
We thank the Mass Spectrometry Core in the Skaggs School of Pharmacy for their assistance with proteomics and metabolomics. Supported by the National Institute of Alcohol Abuse and Alcoholism (NIAAA) (Grants U01AA021723, R21AA024636 and R01DK10957 to C.J.), NIAAA (Grant R01AA022146 to K.S.F.).
Author information
Authors and Affiliations
Contributions
J.O.M. performed data analysis, data interpretation and drafted the manuscript. M.W. performed animal experiments and isolated macrophages. C.R.M. and R.P. performed proteomic sample preparation and analysis. X.Z. performed metabolomics sample preparation and data analysis. N.R. performed data analysis and interpretation. K.S.F. assisted in data interpretation and funded the proteomic analysis. C.J. conceived the study and funded animal experiments and metabolomics analysis. The authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marentette, J.O., Wang, M., Michel, C.R. et al. Multi-omics Analysis of Liver Infiltrating Macrophages Following Ethanol Consumption. Sci Rep 9, 7776 (2019). https://doi.org/10.1038/s41598-019-43240-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-43240-4
This article is cited by
-
Alterations in cellular and organellar phospholipid compositions of HepG2 cells during cell growth
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.