Abstract
Plants easily experience ammonia (NH4+) toxicity, especially aquatic plants. However, a unique wetland plant species, Myriophyllum aquaticum, can survive in livestock wastewater with more than 26 mM NH4+. In this study, the mechanisms of the M. aquaticum response to NH4+ toxicity were analysed with RNA-seq. Preliminary analysis of enzyme activities indicated that key enzymes involved in nitrogen metabolism were activated to assimilate toxic NH4+ into amino acids and proteins. In response to photosystem damage, M. aquaticum seemed to remobilize starch and cellulose for greater carbon and energy supplies to resist NH4+ toxicity. Antioxidative enzyme activity and the secondary metabolite content were significantly elevated for reactive oxygen species removal. Transcriptomic analyses also revealed that genes involved in diverse functions (e.g., nitrogen, carbon and secondary metabolisms) were highly responsive to NH4+ stress. These results suggested that a complex physiological and genetic regulatory network in M. aquaticum contributes to its NH4+ tolerance.
Similar content being viewed by others
Introduction
As a major nitrogen source, ammonia (NH4+) is important for plant growth and development in soil and freshwater ecosystems1,2. However, high concentrations of NH4+, as a kind of abiotic stress, can damage plant cells and cause a series of syndromes, such as leaf chlorosis, a decrease in net photosynthesis and rhizosphere acidification2,3. As reported in previous studies, abiotic stresses impair productivity, reducing average yields by more than 50% globally4. During the past several decades, a worldwide decline in submerged macrophytes also occurred in many eutrophic lakes, and a high concentration of NH4+ in the aquatic environment was regarded as the main cause1,5. Therefore, NH4+ toxicity is a significant ecological and agricultural issue and an important phenomenon in cell biology3.
Plants have evolved various defense mechanisms at multiple levels in response to unfavourable environments including NH4+ stress1,3. In higher plants, once NH4+ has entered the cells, the glutamine synthetase/glutamate synthase (GS/GOGAT) cycle is the first step in NH4+ assimilation6,7. GS catalyses a reaction that incorporates NH4+ into glutamate and generates glutamine (Gln) as a product8,9. GOGAT transfers the amine group in the amide side chain of Gln to 2-oxoglutarate (2-OG), yielding two molecules of glutamate; one molecule serves as a substrate for GS, whilst the other is used for further metabolism5,8. NH4+ assimilation is closely linked to carbon metabolism since the supply of organic acids, which is maintained by the tricarboxylic acid (TCA) cycle, is indispensable in amino acid synthesis10. Thus, a high availability of carbon skeletons is essential for NH4+ assimilation in plant cells. Moreover, accelerated decomposition of starch is also found in plant cells under abiotic stress (e.g., cold, salt and drought) and provides more carbon skeletons and energy for abiotic stress protection11,12. However, metabolism acceleration produces redundant reactive oxygen species (ROS) and disrupts internal ROS homeostasis, thereafter leading to secondary oxidative stress and intracellular damage in plants1,13. Typically, plant cells upregulate the activity of antioxidant enzymes, such as peroxidase (POD), superoxide dismutase (SOD) and catalase (CAT), to remove excessive ROS14. Nevertheless, as an assistant antioxidant pathway, phenylpropanoid biosynthesis in plants provides substrates for the synthesis of many secondary metabolites, such as phenolics, flavonoids and lignin. These compounds are critical for defence responses to biotic and abiotic stresses15,16,17. However, these results were mainly obtained from terrestrial plants, and less is known about the mechanisms of NH4+ tolerance in wetland (aquatic) plant species.
Normally, the concentrations of NH4+ in natural waters such as rivers and lakes are no more than 14 uM. However, with the input of urban waste water and runoff from farmlands, the concentrations of NH4+ could significantly increase and even up to 26 mM in some constructed wetlands18,19,20. As aquatic plants, most wetland species are sensitive to high concentrations of NH4+ and are likely to experience NH4+ toxicity, which may seriously hinder their growth and ability to remove nutrients16,20. The reported symptoms of NH4+ toxicity range widely, and generally appear with external NH4+ concentrations above 0.1 to 0.5 mM1,5. For example, the growth rate of Azolla filiculoides decreased when the NH4+ concentration was above 0.1 mM, and root damage occurred when the NH4+ concentration was higher than 1 mM21. Ceratophyllum demersum cannot grow well when NH4+ concentrations were higher than 0.4 mM22. Wolffia arrhiza exhibited higher tolerance to NH4+ up to 4 mM23. However, Myriophyllum aquaticum, an important wetland plant species for biomass accumulation and nutrient removal18,24, is entirely different from Myriophyllum spicatum, which is used in ecological engineering for aquatic ecosystem restoration and is sensitive to NH4+ 25. M. aquaticum shows strong potential to resist NH4+ toxicity since it can grow in constructed wetlands with high concentrations of NH4+ (up to 26 mM)24,25,26. For coping with NH4+ toxicity, M. aquaticum has a specialized pathway that can achieve detoxification, and this pathway is highly distinct from that in sensitive species5. For example, M. aquaticum was found to convert accumulated NH4+ nitrogen into nitrate internally to avoid toxicity caused by the accumulation of NH4+ in tissues. Furthermore, the detoxification reaction was found to be closely related to high-quality asparagine synthetase (AS) and GS activities5. In other ways, M. aquaticum can elevate SOD and POD activities to remove redundant ROS and avoid oxidative damage27. However, these studies mainly focused on the leaves of M. aquaticum, and the functions of other key enzymes involved in nitrogen, carbon and antioxidative pathways have not been explored. Furthermore, gene regulation associated with NH4+ tolerance in M. aquaticum is still unknown and thus needs to be further explored.
Within this context, the present work hypothesized that NH4+ detoxification depends on network regulation in M. aquaticum including nitrogen, carbon and secondary metabolisms in roots and leaves. To test this hypothesis, we grew M. aquaticum under two concentrations of NH4+ (12 and 36 mM) that suppress M. aquaticum growth27 for 14 days. These two groups were defined as A12 and A36 group with a negative control group (NC group) under 1 mM NH4+. An integrative approach was applied, first measuring plant biomass (defined as the growth rate) and NH4+ content and then determining nitrogen, carbon and antioxidative metabolites and enzyme activities. Finally, a transcriptomic analysis of roots and leaves was carried out to investigate the complexity of the molecular events underlying the response to NH4+ in this species. Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment were used to identify the important biological processes operating in M. aquaticum roots and leaves.
Results
Responses to NH4 + stress
NH4 + content
Roots and leaves are important plant organs for nutrient absorption and energy assimilation, respectively. As the NH4+ concentration in the nutrient solution increased, the NH4+ content in the leaves of M. aquaticum increased significantly (p < 0.05) from 69 µg g−1 fresh weight (FW) in the NC group to 162 µg g−1 FW in the A12 group and 357 µg g−1 FW in the A36 group (Fig. 1a). Furthermore, the NH4+ content in roots also significantly increased in the NH4+-stressed groups. Interestingly, the NH4+ content in the leaves of each group was always higher than that in the roots, which indicated that most of the NH4+ was transported to the leaves.
Plant growth
Biomass reduction provoked by NH4+ toxicity is generally associated with the accumulation of NH4+ in tissues of different plant species6,10. With increased NH4+ content, the growth rate significantly decreased by more than 50%, from 0.116 g d−1 in the NC group to 0.053 g d−1 in the A36 group (Fig. 1b). Further, the number of fibrous roots also significantly decreased in the A12 and A36 groups (Fig. 1c). Spearman’s correlation analysis indicated that the changes in growth rate and fibrous root number were significantly correlated (p < 0.01) with the increases in NH4+ content in roots and leaves (Supplementary Table S1).
Chlorophyll (Chl) and maximum quantum yield of photosystem II (Fv/Fm)
Energy metabolism also seems to be damaged by high concentrations of NH4+ 28. Photosynthetic pigment content (such as the Chl a and Chl b contents) decreased significantly (p < 0.05) in the A36 group compared to the NC group (Supplementary Fig. S1a). Moreover, Fv/Fm also significantly decreased from 0.830 in the NC group to 0.796 in the A36 group (Supplementary Fig. S1b). However, there were no significant differences in carotenoid (Car) among the different groups.
Hydrogen peroxide (H2O2), superoxide anion radical (O2 −) and malondialdehyde (MDA)
With the increase in NH4+ content, large amounts of H2O2 and O2− were produced in M. aquaticum roots and leaves (Supplementary Fig. S2a,b) and easily accumulated in the leaves and roots, respectively. As a result of the increased NH4+ content, the MDA content also significantly (p < 0.05) increased in the A12 and A36 groups (Supplementary Fig. S2c). Spearman’s correlation analysis also showed that the MDA content was significantly correlated (p < 0.05) with the increased NH4+ content in roots and leaves.
To better understand the mechanism used by roots and leaves facing NH4+ stress, we further explored their transcriptomic and metabolic responses.
De novo assembly and annotation of the root and leaf transcriptomes
Sequencing and assembling
As shown in Supplementary Table S2, a total of 131.48 G raw nucleotides (124.50 G clean nucleotides), equivalent to 941.20 million raw reads (914.70 million clean reads), were generated from 18 root and leaf samples belonging to the three groups (NC, A12 and A36). All clean reads of M. aquaticum were pooled together and then de novo assembled by Trinity. The transcriptome was de novo assembled to 97338 unigenes, with an N50 = 1187 bp.
Differential expression analyses under different concentrations of NH4 + stress
The transcriptomic changes of M. aquaticum roots and leaves in response to 12 and 36 mM NH4+ were analysed. The number of genes for which the expression was significantly (p < 0.05) altered based on different growth conditions was represented with Venn diagrams (Supplementary Fig. S3a,b). NH4+ treatment led to strong transcriptomic responses in which 4583 genes were differentially expressed in NH4+-stressed roots, while only 965 were differentially expressed in NH4+-stressed leaves. Under 12 mM NH4+, there were only 44 upregulated genes in the leaves (Supplementary Fig. S3b). Nevertheless, the number of upregulated genes in the A36 group was nearly 7-fold greater than that in the A12 group, reaching 351 in the former group, with little difference in the number of upregulated genes in the roots. Thus, different regulatory mechanisms in roots and leaves might have responded to the NH4+ stress.
GO enrichment analysis
GO analysis provided an overview of the processes most affected by NH4+ stress. The most enriched GO terms were similar in roots and leaves responding to 12 and 36 mM NH4+ (Fig. 2). However, there were large differences between the differentially expressed genes (DEGs) in terms of GO terms and the proportion of up- and downregulated genes. Genes related to stress response (e.g., salt, wounding, and cold stresses) were enriched in both roots and leaves under NH4+ stress, which indicated an increased response to NH4+ toxicity and a network response to different stress factors. Furthermore, genes for signalling regulation (e.g., auxin and abscisic acid in both roots and leaves and ethylene in roots) were also enriched, indicating that a complex signal regulation pathway responded to NH4+ stress29. Finally, the genes involved in light harvesting and photosynthesis were significantly (p < 0.05) downregulated in the leaves of the A36 group, indicating downsizing of the light-harvesting apparatus.
GO and KEGG analysis also indicated that strong transcriptional responses were observed for genes involved in processes such as NH4+ assimilation, amino acid synthesis, photosynthesis, starch and sucrose metabolism, ROS removal and phenylpropanoid biosynthesis, as discussed below. The responses of the most strongly regulated genes within these categories under NH4+ stress is shown in Supplementary Table S3. To verify the results of the RNA-seq analysis, qRT-PCR with specific primers was performed on eighteen selected genes involved in different metabolisms (Supplementary Table S4), and the results corresponded to those from RNA-seq (Supplementary Figs S4 and S5).
Nitrogen metabolism
NH4 + assimilation
With increased NH4+ content, significantly increased GS activity was observed in NH4+-stressed leaf cells (Fig. 3a). Interestingly, the GS in roots maintained a higher activity above 690 nmol γ-glutamyl hydroxamate (γ-GHA) min−1 g−1 FW than that in leaves in all groups. Correspondingly, the Gln content also significantly increased in the leaves under 12 mM NH4+, and the content doubled in the leaves of the A36 group compared with that of the NC group, although no significant increase was observed in the roots (Fig. 4a). As shown in Fig. 3b, increased NH4+ in the nutrient medium also caused significant differences in the GOGAT activity in the roots and leaves of M. aquaticum (p < 0.05). GOGAT activity reached a maximum value of 39 nmol NADH min−1 g−1 FW in the roots when plants were treated with 12 mM NH4+, while it peaked at 22 nmol NADH min−1 g−1 FW in the leaves when plants were treated with 36 mM NH4+. In addition, the gene encoding GOGAT was dramatically upregulated in NH4+-stressed root cells (Supplementary Table S3). Furthermore, the gene encoding GDH, another enzyme that catalyses the conversion of 2-OG to glutamate and the reverse, was also significantly upregulated in the roots. Simultaneously, the enzyme activity catalysing the conversion of 2-OG to glutamate was also significantly upregulated in both the roots and leaves of the A12 and A36 groups compared with the NC group (Fig. 3c). Finally, the activity of AS doubled in the roots of the A12 and A36 groups and in the leaves of the A36 group (Fig. 3d), although AS activity decreased slightly in the roots of the A36 group compared with the A12 group. Further, AS activity was significantly higher in the roots than in the leaves of each group, which indicated a faster rate of Asn synthesis in the roots than in the leaves. Correspondingly, the Asn content also increased significantly, reaching 877 µg g−1 FW in the roots of both the A12 and A36 groups, and increased to 443 µg g−1 FW in the leaves of the A36 group (Fig. 4b).
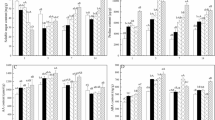
NH4+ assimilation-related enzyme activity in roots and leaves of Myriophyllum aquaticum. Enzyme activities were determined for glutamine synthetase (a), glutamate synthase (b), glutamate dehydrogenase (c), and asparagine synthetase (d). Values (means ± SDs) were determined from five biological replicates (n = 5). Different letters above the bars indicate a significant difference at p < 0.05.
Biosynthesis of amino acids
Changes in the fifteen most abundant free amino acids in response to NH4+ stress were quantified, and the results are shown in Fig. 5. The most abundant amino acid under normal conditions was glutamic acid (Glu), which accounted for 30% of the total free amino acid content in the leaves of M. aquaticum. However, serine (Ser) became the most abundant, accounting for 25% and 16% of the total free amino acid content in the leaves of the A12 and A36 groups, respectively. Phenylalanine (Phe) and tyrosine (Tyr), which are aromatic amino acids and substrates for phenol and flavonoid biosynthesis15,16, were also significantly increased in the roots and leaves under NH4+ stress. Interestingly, proline (Pro), an osmoticum and ROS scavenger often found to increase in response to NH4+ stress30,31, also showed a significant increase in the leaves but no significant change in the roots under NH4+ stress. Furthermore, the histidine, threonine, arginine, valine, isoleucine, leucine and lysine contents also increased significantly in response to NH4+ stress.
Protein biosynthesis
Protein, the main nitrogen-containing compound in plant cells, was measured in all groups. Protein synthesis seemed to be significantly accelerated in the tissues of the A12 and A36 groups, as the protein content was significantly higher in the A12 and A36 groups than in the NC group (Fig. 4c), which indicated that more free amino acids were used for protein synthesis under NH4+ stress.
Carbon metabolism
Photosynthesis
As photosynthesis mainly occurs in leaves, we considered the gene regulation related to photosynthesis in leaves but not roots. In response to NH4+ stress, the expression levels of many genes encoding enzymes involved in light-harvesting and photosynthesis pathways were significantly repressed in the A36 group compared to the NC group (Supplementary Table S3), including four genes encoding light-harvesting complex (LHC) proteins in leaves. The downregulated expression of genes encoding Chl-binding proteins, Chl a and Chl b had fewer binding proteins to fix, which might have led to the significant decrease in the Chl pigment content and Fv/Fm (Supplementary Fig. S1a,b). Moreover, the expression of a gene encoding phosphoenolpyruvate carboxylase (PPC), which catalyses CO2 fixation, was 2.8-fold downregulated in the A36 group compared to the NC group, which indicated a reduced CO2 fixation efficiency in the leaves of the former group.
Starch cellulose and sucrose metabolism
In response to the reduced light energy and carbon acquisition efficiency in leaves, some genes associated with starch and cellulose decomposition were upregulated in NH4+-stressed cells. In particular, a gene encoding β-glucosidase was significantly upregulated 110.8-fold and 215.2-fold in the NH4+-stressed root cells of the A12 and A36 groups (Supplementary Table S3), respectively, indicating accelerated cellulose decomposition and glucose production for further utilization. Furthermore, a gene encoding β-amylase3 (BAM3) was upregulated more than 3.7-fold in the NH4+-stressed root cells in the A36 group. M. aquaticum seemed to upregulate a number of genes to decompose macromolecular organics under NH4+ stress. To verify this hypothesis, we subsequently measured the enzyme activity of amylase and β-glucosidase. The results revealed increased amylase activity in both the roots and leaves of the NH4+-stressed groups (Fig. 6a). Additionally, the β-glucosidase activity also significantly increased in the roots of the NH4+-stressed groups but decreased in the leaves of these groups (Fig. 6b). We further explored the amount of soluble carbohydrates that supply carbon skeletons for subsequent NH4+ assimilation into Gln and Asn and behave as signalling molecules. The amount of soluble carbohydrates significantly increased in the leaves of both the A12 and A36 groups but increased in the roots of only the A36 group (Supplementary Fig. S6a). Another significantly upregulated gene, namely, one encoding sucrose synthase, which catalyses sucrose synthesis from fructose and glucose, was observed in NH4+-stressed root cells. Correspondingly, a significant increase in sucrose content was observed in both the roots and leaves of the A36 group (Supplementary Fig. S6b). Additionally, the fructose content also significantly decreased in the NH4+-stressed groups (Supplementary Fig. S6c) as a result of fructose consumption for sucrose synthesis. Interestingly, the glucose content seemed to remain relatively stable in the NH4+-stressed groups (Supplementary Fig. S6d), even though the degradation of starch and cellulose was accelerated.
Decomposition of carbohydrate
KEGG analysis also indicated the promotion of several genes encoding enzymes potentially involved in glycolysis metabolism, pyruvate metabolism and the TCA cycle (Supplementary Table S3). Interestingly, significant upregulation of the gene encoding phosphoenolpyruvate carboxykinase (PckA) was observed in the roots, which indicated an accelerated transformation from oxaloacetate to phosphoenolpyruvate. Furthermore, the upregulation of pyruvate kinase (PK), which catalyses the transformation from phosphoenolpyruvate to pyruvate, seemed to increase the production of pyruvate, which was then transformed into acetyl-CoA for the TCA cycle. Several key genes connected to the TCA cycle were also upregulated in NH4+-stressed cells, such as the genes encoding citrate synthase (CS), aconitate hydratase (ACO) and adenosine triphosphate (ATP) citrate (pro-S)-lyase (ACLY). Interestingly, the gene encoding 2-OG dehydrogenase (OGDH), an important enzyme catalysing the oxidation of 2-OG and the synthesis of succinyl coenzyme A, was also upregulated in the roots and leaves under NH4+ stress. An accelerated TCA cycle could generate more ATP for plants to resist NH4+ toxicity.
ROS removal and secondary metabolism
ROS removal
In response to increased ROS (Supplementary Fig. S2a,b), the activity of antioxidative enzymes was elevated in both the A12 and A36 groups. SOD activity in the roots and leaves under NH4+ stress was significantly higher than that in the NC group (Fig. 7a). However, POD activity in the leaves was significantly higher in only the A36 group (Fig. 7b). Further, POD activity in the roots initially significantly increased in the A12 group but slightly decreased in the A36 group. Furthermore, CAT activity was also observed to increase in both the roots and leaves of the NH4+-stressed groups, especially in the roots of the A12 group and the leaves of the A36 group (Fig. 7c). In fact, CAT activity also slightly decreased in the roots of the A36 group compared to the A12 group, although the activity in the A36 group was still significantly higher than that in the NC group. Interestingly, NH4+ stress provoked the increased expression of many genes in the stress response category, which are related to abiotic and biotic stress (Fig. 2, Supplementary Table S3).
Secondary metabolites
With the upregulation of multi-genes involved in phenylpropanoid biosynthesis (Supplementary Table S3), the total phenolic content in leaves was significantly increased when plants were treated with 12 mM NH4+, although the increase in the content in roots was not significant (Fig. 8a). However, the total phenolic content in both the roots and leaves of the A36 group was significantly higher than that of the NC group. Total phenolics seemed to accumulate primarily in the leaves of all groups, as the content in leaves was always higher than that in roots in the same group. Similarly, the expression of genes in the flavonoid pathway encoding chalcone synthase (CHS), chalcone isomerase (CHI), naringenin 3-dioxygenase (N3D) and flavonol synthase (FLS) was also increased. Due to the increased expression of genes involved in the flavonoid biosynthesis pathway, the flavonoid content also significantly increased in the roots of the A12 and A36 groups by more than 2-fold (Fig. 8b). Interestingly, flavonoids mainly accumulated in the roots of all groups, which was different from the pattern observed for total phenolics. In fact, the flavonoid content in roots was more than ten times higher than that in leaves in the same group, especially when plants were treated with 12 or 36 mM NH4+.
Discussion
Previous studies have proven that M. aquaticum is highly tolerant of aquatic environments with high concentrations of NH4+ 5,26,27. However, the physiological and genetic regulations associated with this tolerance are poorly understood. This is the first study on the gene regulation mechanism of high tolerance of NH4+ toxicity in M. aquaticum using RNA-seq with the goal of constructing a physiological and genetic regulatory network for this species (Fig. 9). To the best of our knowledge, these regulations have not previously been systematically investigated in aquatic and wetland plant species.
Nitrogen metabolism responses to NH4 + toxicity
In our study, NH4+ content in leaves is always higher than that in roots under the same NH4+ concentration (Fig. 1a). Because of the toxic nature of NH4+, NH4+ should be assimilated into amino acids in roots once it was absorbed from sediment and then amino acids (Asp, Glu, Asn and Gln) were transported to shoots and leaves via xylem and phloem. However, when plants were treated with ammonium as the sole nitrogen, NH4+ content in shoots and leaves seemed to rise and could reach concentration in the millimolar range32,33. Excess NH4+ might be loaded into the xylem and transported to leaves via ammonium transporters or passively via aquaporins or nonselective cation and K+-specific channels34,35,36. Due to an increase in the NH4+ content in roots and leaves, many processes connected to nitrogen metabolism were affected. Stimulated by high concentrations of NH4+, the activity of GS and GOGAT was significantly increased in leaves under NH4+ stress3,9. Surprisingly, GS activity in the roots of M. aquaticum maintained a high level even without NH4+ stress, which indicated that an immediate mechanism of NH4+ assimilation occurs in M. aquaticum roots31,37. However, Gln content in leaves was always lower than that in roots under the same NH4+ concentration, although GS activity in roots was higher than that in leaves. The reason could be that synthesised Gln in roots normally transported to shoots and leaves for storage and signalling, which led to the higher Gln content in leaves than that in roots36,38,39. Interestingly, expression of the gene encoding GDH and GDH activity were both significantly increased in both the roots and leaves of both NH4+-stressed groups, which indicated that GDH might also participate in an important anabolic reaction to scavenge NH4+ in NH4+-stressed cells of M. aquaticum. This scavenging could be an alternative strategy for transforming NH4+ into glutamate to avoid NH4+ damage to plant cells40,41. Previous research has shown that AS plays an important role in nitrogen reactivation and can be used to improve nitrogen use efficiency5,10. In the present study, NH4+ stress was found to significantly stimulate AS activity in M. aquaticum roots, which indicated that AS is closely associated with the detoxification of NH4+ in M. aquaticum, as reported in a previous study5. Thus, GS was the main enzyme responsible for NH4+ assimilation in M. aquaticum under normal concentrations of NH4+, while many enzymes (such as GDH, GOGAT, AS and GS) were activated and involved in NH4+ detoxification and scavenging under NH4+ stress.
Free amino acids play crucial roles in an array of cellular functions, including nitrogen storage, osmotic regulation, and free-radical scavenging under various stresses12,42. Several amino acids are also precursors for certain phytohormones that regulate plant growth and development42. As a result of NH4+ stress, the expression of key enzymes involved in Glu, Tyr, histidine, arginine, Ser, threonine and valine synthesis was increased (Supplementary Table S3)9,12, and the synthesis of most free amino acids was correspondingly accelerated (Fig. 5). As a large amount of Glu was consumed for Gln synthesis, upregulation of the activity of GDH and GOGAT seemed to lead to the synthesis of more glutamate to guarantee the availability of glutamate for biosynthesis of amino acids. These results suggest that Glu may play a key role in NH4+ tolerance in M. aquaticum, which is consistent with the many known roles of these amino acids1,9. Glu is also the precursor of Pro, which serves as an osmoticum in response to different types of stress42. Interestingly, we did not find an increase in free Pro content in roots under NH4+ stress. However, significantly increased Pro content were observed in NH4+-stressed leaves, which indicated that Pro mainly plays a role in the leaves of M. aquaticum. In addition to these functions, different free amino acids are also basic substrates for protein biosynthesis. Thus, M. aquaticum might try to avoid NH4+ toxicity by accelerating the conversion of NH4+ to free amino acids and subsequently to proteins2,9.
Carbon metabolism responses to NH4 + toxicity
Chlorosis is one of the main responses of M. aquaticum to NH4+ toxicity27,43. The reduced Chl a and Chl b in NH4+-stressed cells are probably caused by the repression of their biosynthesis. In addition, the coordinated decrease in Chl and Chl-binding proteins under NH4+ toxicity suggests that the biosynthesis of Chl is coordinated with Chl-binding proteins, which was also observed in a previous study44,45. These results clearly indicate that NH4+ toxicity reduces photosynthetic efficiency, which is in agreement with the observed reduction in the Fv/Fm. Organic acids play a special role in C4 photosynthetic metabolism as intermediate pools of fixed carbon46,47. Since PPC is essential for the fixation of CO2 into oxaloacetate, its downregulation (Supplementary Table S3) might significantly repress the efficiency of CO2 fixation in the C4-dicarboxylic acid pathway48 and might lead to the production of more ROS because of excessive energy around the photosystem43,45. As a result of the decreased efficiency of carbon and light energy acquisition, M. aquaticum seemed to have less carbon and energy for NH4+ resistance.
Starch and cellulose are the major storage carbohydrates in plants and are most commonly associated with storage organs such as roots, rhizomes, stems and seeds18,49,50. A number of studies have shown that many plant species, including several important crop species, can remobilize starch and cellulose reserves to release carbon and energy to resist abiotic or biotic stress11,12,50. Hydrolysis of starch to maltose by BAM represents the predominant pathway of starch degradation. In the A36 group, the activity of amylase significantly increased, and the gene encoding BAM3 was upregulated at the same time in roots and leaves. In addition to starch metabolism, cellulose degradation also seemed to be accelerated by the significant upregulation of β-glucosidase activity to generate more glucose. Moreover, the biosynthesis of sucrose was promoted by the upregulation of sucrose synthase. Glucose and sucrose can help stabilize proteins and cell structures, particularly when the stress becomes severe or persists for longer periods49,50. These compounds can also act as free-radical scavengers, protecting against oxidation by removing excessive ROS, re-establishing the cellular redox balance51 and acting as signalling molecules connected to the ABA-dependent signalling pathway to activate downstream stress-response regulation11,29,49.
Accelerated degradation of starch and cellulose releases more glucose, which then enters the glycolysis and pyruvate pathways11,49. With many genes involved in glycolysis and pyruvate pathways being upregulated under NH4+ stress, more acetyl-CoA, the final product of those pathways and the original substrate for the TCA cycle, enters the TCA cycle in the mitochondria45,52. Regarding the three key catalytic enzymes in the TCA cycle37,52, genes encoding CS and OGDH were significantly upregulated in the NH4+-stressed root cells of both the A12 and A36 groups, whilst the expression of the isocitrate dehydrogenase (ICDH)-encoding gene was significantly upregulated in the leaves of both the A12 and A36 groups. The TCA cycle can regulate the energy and NADH levels in plant tissues and supply substrates for amino acid synthesis47. In response to high concentrations of NH4+, 2-OG in the TCA cycle furnishes the GS/GOGAT cycle with the carbon skeleton required for amino acid biosynthesis37,52. The NH4+-dependent enhancement of carbon input to the TCA cycle is associated with increased NH4+ assimilation, which is supported by the higher concentrations of amino acids and proteins observed under excess NH4+ supply52,53. Thus, increased degradation of starch and cellulose, along with induction of glycolysis and the TCA cycle, could provide NH4+-stressed cells with enough energy and NADH as well as many other substrates for NH4+ assimilation and amino acid and protein biosynthesis47,52.
ROS removal and secondary metabolisms responses to NH4 + toxicity
Generally, environmental stresses disrupt internal ROS homeostasis, thereafter leading to oxidative stress and cellular damage in plants1,13. In this study, M. aquaticum differentially regulated the expression level of genes encoding several representative antioxidant enzymes, such as SOD, POD and CAT, under NH4+ stress for ROS removal1. However, NH4+-induced inhibition of antioxidative enzyme activities was observed in the A36 group, which indicated a threshold for NH4+ tolerance in M. aquaticum20,27. Although redundant ROS damage the nucleic acids, proteins and lipids in NH4+-stressed cells54, some researchers have suggested that ROS accumulation induced by NH4+ stress is important for defence against a multitude of environmental stimuli, such as pathogens, salinity stress and cadmium-induced oxidative damage13,54,55. Our transcriptomic results also showed that NH4+ stress modulated the expression of many genes involved in abiotic and biotic stress responses (e.g., oxidation, cold, heat, and bacterium defences) (Fig. 2), which also confirms the evidence that NH4+-triggered antioxidant defence might be a universal strategy for adaptive plant responses to multiple environmental stimuli14,30,55,56.
Phenols perform a wide range of functions to mitigate the effect of oxidative stress by acting as ROS scavengers and lipid peroxidation inhibitors, thereby protecting important cellular components such as photosynthetic apparatuses15,31. In this study, the phenolic and flavonoid contents were both observed to increase under NH4+ stress and mainly accumulated in different plant tissues. KEGG analysis also revealed that most of the genes involved in the phenylpropanoid biosynthesis pathway were upregulated under NH4+ stress (Supplementary Table S3), especially the genes encoding the key enzymes that catalyse multiple reactions in this pathway, such as 4-coumarate-CoA ligase (4CL), cinnamoyl-CoA reductase (CCR) and cinnamyl-alcohol dehydrogenase (CAD), which was also reported in previous studies15,16. The end products of the phenylpropanoid biosynthesis pathway are different kinds of lignin originating from Phe10. Flavonoids may constitute a ‘secondary’ antioxidant system in response to severe stress to complement the role of antioxidant enzymes in combating that stress10,16,17. Thus, flavonoids accumulated in M. aquaticum tissues may also act as osmolytes to protect plants from NH4+ stress damage17. Interestingly, genes for anthocyanin biosynthesis, such as dihydroflavonol 4-reductase (DFR), anthocyanidin synthase (ANS), anthocyanidin reductase (ANR) and leucoanthocyanidin reductase (LAR), were also upregulated under NH4+ stress (Supplementary Table S3). The demonstration of the accelerated synthesis of phenylpropanoids and their derivatives such as phenolics, flavonoids, anthocyanins and lignin in NH4+-stressed M. aquaticum further emphasized the importance of these secondary compounds for the adaptation of M. aquaticum to high-NH4+ environments.
Conclusions
The present work indicated that a complex physiological and genetic regulatory network including nitrogen metabolism, carbon metabolism, abiotic stress response and secondary metabolism at the root and leaf levels was involved in NH4+ resistance in M. aquaticum. The synthesis of amino acids and proteins was accelerated to assimilate NH4+ in plant tissues. Furthermore, starch and cellulose were decomposed for greater supplies of carbon and energy. Antioxidative enzyme activity and the secondary metabolite content were also significantly elevated to remove redundant ROS.
Methods
Experimental design
This study was conducted in a greenhouse at a stable temperature (25–30 °C) and under a 12/12 h dark/light photoperiod at the Research Center for Eco-Environmental Sciences, Chinese Academy of Sciences, Beijing. Polyethylene plastic containers (0.5 m × 0.4 m × 0.4 m) were filled with approximately 10 cm of sterile quartz sand (Φ = 1–3 mm). M. aquaticum seedlings with a uniform length of 50 ± 3 cm were planted in plastic containers. Before the experiments, plants were cultured for two weeks in 50% modified Hoagland solution, as described previously27. After a two-week preincubation, plants were transplanted into plastic containers. The culture solution in each container consisted of 25 L 50% modified Hoagland solution amended with various concentrations of NH4Cl. Plants treated with 12 mM NH4Cl were defined as the A12 group, whilst plants treated with 36 mM NH4Cl were defined as the A36 group. A negative control (NC) group was treated with 1 mM NH4Cl. Each group contained five containers as replicates, and each container housed 30 seedlings. The pH of the culture solution in each container was adjusted to 6.0 ± 0.1 with 1 M KOH. The culture solution was exchanged every 2 days to maintain the pH and nutrient concentrations. After 14 days of cultivation, plants in each container were harvested. Root and leaf samples rinsed with distilled water were ground with liquid nitrogen and fully mixed12,15. Part of the mixture was used for tissue and enzyme analysis, whilst the remainder was used for RNA extraction.
Phenotype analysis
To determine plant growth, plants were harvested on day 14 and then measured for growth rate as previously described25. Fv/Fm was measured using an LI-6400 portable photosynthesis system (LI-COR Inc., Lincoln, Nebraska, USA), whilst the Chl a (Chl a), Chl b (Chl b) and total Car contents in plant leaves were measured with a spectrophotometer (DR6000, HACH, USA) after extraction with 80% acetone57. The NH4+ content in plant tissues was measured by using the method described5 and expressed as µg g−1 FW. The level of lipid peroxidation in plant leaves was evaluated by determining the MDA content using the thiobarbituric acid test58. The H2O2 content was determined according to the method59 by analysing the titanium-hydroperoxide complex production at 410 nm. The O2- content was measured by the oxidation of hydroxylamine at 530 nm as described previously60. The amounts of Gln, Asn and free amino acids were determined by an L-8900 high-speed amino acid analyser (Hitachi, Japan)15. Protein was extracted and determined by using the method described61. The soluble carbohydrate, glucose, fructose and sucrose contents in plant tissues were determined as described previously10. Total phenolics and flavonoids were expressed as µg g−1 dry weight (DW) using gallic acid and rutin as standards, respectively62,63.
Enzyme activity analysis
GS activity was expressed as nmol γ-GHA min−1 g−1 FW at 25 °C and tested at 540 nm, whilst GDH activity was expressed as nmol NADH consumption min−1 g−1 FW at 25 °C and monitored spectrophotometrically at 340 nm61. GOGAT activity was determined by using the method48. AS activity was expressed as nmol NH4+ consumption min−1 g−1 FW at 25 °C and determined with a spectrophotometer64. Total amylase activity was determined as the speed of maltose production measured at 540 nm65,66. β-glucosidase activity was expressed as nmol p-nitrophenol min−1 g−1 FW at 25 °C and tested at 400 nm67. The SOD and POD activities were measured by the method described previously27. CAT activity was computed by calculating the speed of H2O2 decomposition68. One unit of CAT activity was defined as the amount of 1 nmol H2O2 decomposition in one minute.
RNA extraction and library construction
Root and leaf samples were fully ground with liquid nitrogen, and 200 mg sample was used for total RNA extraction and purified with an EASYspin Plus Complex Plant RNA Kit (Aidlab Biotech, Beijing, China). Three biological replicates of each group were used for RNA-seq analysis. The NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (#E7530L, NEB, USA) was used for sequencing library construction46.
Sequencing, assembly and annotation of the transcriptome
After the index-coded samples were clustered, the libraries were sequenced on a HiSeq X Ten platform, and 150 bp paired-end reads were generated. The raw RNA-seq reads were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (BioProject accession no. PRJNA497710). Raw reads were processed with Perl scripts to ensure the quality of data used for further analysis. Trinity software was used for de novo assembly46. Transcripts and predicted peptides were annotated by the NR, Swiss-Prot, Pfam, UniProt and eggNOG databases.
Differentially expressed genes and enrichment analysis
HTSeq v0.6.0 was used to count reads for each gene in each sample, and the reads per kilobase per million mapped reads (RPKM) were calculated to estimate the expression level of genes in each sample69. An estimated false discovery rate (FDR) was assigned to each gene (RPKM ≥ 1) and adjusted with the Benjamini and Hochberg approach to control the FDR70. Genes with an FDR ≤ 0.05 and an |log2 ratio| ≥ 1 were identified as DEGs. GO and KEGG enrichment of DEGs was implemented to draw pathway maps of the molecular interaction and reaction networks7.
Gene expression validation
To further evaluate the reliability of the RNA-seq results, eighteen genes of M. aquaticum were selected for qRT-PCR to assess their expression in response to NH4+ stress. The β-actin gene was used as an internal control. The relative expression levels were calculated as previously described71.
Data analysis
IBM SPSS 20.0 software (IBM Corp.) was used for data analysis. One-way analysis of variance with Tukey’s test was applied to identify significant differences between groups and between tissues (p < 0.05). A Spearman two-tailed test was used to identify significant correlations (p < 0.05) between growth characteristics and NH4+ content in the plant samples.
References
Britto, D. T. & Kronzucker, H. J. NH4 + toxicity in higher plants: a critical review. Journal of Plant Physiology 159, 567–584 (2002).
Liu, Y. & von Wiren, N. Ammonium as a signal for physiological and morphological responses in plants. Journal of Experimental Botany 68, 2581–2592 (2017).
Li, B. H., Li, G. J., Kronzucker, H. J., Baluska, F. & Shi, W. M. Ammonium stress in Arabidopsis: signaling, genetic loci, and physiological targets. Trends in Plant Science 19, 107–114 (2014).
de Zelicourt, A., Colcombet, J. & Hirt, H. The role of MAPK modules and ABA during abiotic stress signaling. Trends in Plant Science 21, 677–685 (2016).
Zhou, Q. Y., Gao, J. Q., Zhang, R. M. & Zhang, R. Q. Ammonia stress on nitrogen metabolism in tolerant aquatic plant-Myriophyllum aquaticum. Ecotoxicology and Environmental Safety 143, 102–110 (2017).
Konishi, N. et al. Contributions of two cytosolic glutamine synthetase isozymes to ammonium assimilation in Arabidopsis roots. Journal of Experimental Botany 68, 610–625 (2017).
Goel, P. et al. Transcriptome and co-expression network analyses identify key genes regulating nitrogen use efficiency in Brassica juncea L. Scientific Reports 8, 18 (2018).
Tobin, A. K. & Yamaya, T. Cellular compartmentation of ammonium assimilation in rice and barley. Journal of Experimental Botany 52, 591–604 (2001).
Guan, M., de Bang, T. C., Pedersen, C. & Schjoerring, J. K. Cytosolic glutamine synthetase Gln1;2 is the main isozyme contributing to GS1 activity in Arabidopsis shoots and can be up-regulated to relieve ammonium toxicity. Plant Physiology 171, 1921–1933 (2016).
Vega-Mas, I. et al. Elevated CO2 induces root defensive mechanisms in tomato plants when dealing with ammonium toxicity. Plant and Cell Physiology 58, 2112–2125 (2017).
Thalmann, M. et al. Regulation of leaf starch degradation by abscisic acid is important for osmotic stress tolerance in plants. Plant Cell 28, 1860–1878 (2016).
Cao, T., Xie, P., Ni, L. Y., Zhang, M. & Xu, J. Carbon and nitrogen metabolism of an eutrophication tolerative macrophyte, Potamogeton crispus, under NH4 + stress and low light availability. Environmental and Experimental Botany 66, 74–78 (2009).
Xie, Y. et al. Heme-hemeoxygenase 1 system is involved in ammonium tolerance by regulating antioxidant defence in Oryza sativa. Plant Cell and Environment 38, 129–143 (2015).
Fujita, M. et al. Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Current Opinion in Plant Biology 9, 436–442 (2006).
Wang, L. X. et al. A multilevel investigation to discover why Kandelia candel thrives in high salinity. Plant Cell and Environment 39, 2486–2497 (2016).
Wang, W., Li, R., Zhu, Q., Tang, X. & Zhao, Q. Transcriptomic and physiological analysis of common duckweed Lemna minor responses to NH4 + toxicity. Bmc Plant Biology 16, 92 (2016).
Pandey, A. et al. Genome-wide expression analysis and metabolite profiling elucidate transcriptional regulation of flavonoid biosynthesis and modulation under abiotic stresses in banana. Scientific Reports 6, 13 (2016).
Liu, F. et al. Purification and reuse of non-point source wastewater via Myriophyllum-based integrative biotechnology: a review. Bioresource Technology 248, 3–11 (2018).
Zhang, S. N., Liu, F., Xiao, R. L., He, Y. & Wu, J. S. Nitrogen removal in Myriophyllum aquaticum wetland microcosms for swine wastewater treatment: N-15-labelled nitrogen mass balance analysis. Journal of the Science of Food and Agriculture 97, 505–511 (2017).
Wang, Y., Wang, J., Zhao, X., Song, X. & Gong, J. The inhibition and adaptability of four wetland plant species to high concentration of ammonia wastewater and nitrogen removal efficiency in constructed wetlands. Bioresource Technology 202, 198–205 (2016).
Kitoh, S., Shiomi, N. & Uheda, E. The growth and nitrogen fixation of Azolla filiculoides Lam. in polluted water. Aquatic Botany 46, 129–139 (1993).
Gao, J. et al. Ammonia stress on the carbon metabolism of Ceratophyllum demersum. Environmental Toxicology and Chemistry 34, 843–849 (2015).
Monselise, E. B. I. & Kost, D. Different ammonium-ion uptake, metabolism and detoxification efficiencies in two Lemnaceae. Planta 189, 167–173 (1993).
Liu, F. et al. Nitrogen removal and mass balance in newly-formed Myriophyllum aquaticum mesocosm during a single 28-day incubation with swine wastewater treatment. Journal of Environmental Management 166, 596–604 (2016).
Zhang, L. et al. Physiological response of a submerged plant (Myriophyllum spicatum) to different NH4Cl concentrations in sediments. Ecological Engineering 58, 91–98 (2013).
Li, X. et al. Seasonality distribution of the abundance and activity of nitrification and denitrification microorganisms in sediments of surface flow constructed wetlands planted with Myriophyllum elatinoides during swine wastewater treatment. Bioresource Technology 248, 89–97 (2018).
Wang, R. et al. The adaptability of a wetland plant species Myriophyllum aquaticum to different nitrogen forms and nitrogen removal efficiency in constructed wetlands. Environmental Science and Pollution Research 25, 7785–7795 (2018).
Apudo, A. A., Cao, Y., Wakibia, J., Li, W. & Liu, F. Physiological plastic responses to acute NH4 +–N toxicity in Myriophyllum spicatum L. cultured in high and low nutrient conditions. Environmental and Experimental Botany 130, 79–85 (2016).
Shen, J. B. et al. The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Scientific Reports 7, 14 (2017).
Zhu, J. K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Bittsanszky, A., Pilinszky, K., Gyulai, G. & Komives, T. Overcoming ammonium toxicity. Plant Science 231, 184–190 (2015).
Finnemann, J. & Schjoerring, J. K. Translocation of NH4 + in oilseed rape plants in relation to glutamine synthetase isogene expression and activity. Physiologia Plantarum 105, 469–477 (1999).
Vanbeusichem, M. L., Kirkby, E. A. & Baas, R. Influence of nitrate and ammonium nutrition on the optake, assimilation and distribution of nutrients in Ricinus communis. Plant Physiology 86, 914–921 (1988).
Balkos, K. D., Britto, D. T. & Kronzucker, H. J. Optimization of ammonium acquisition and metabolism by potassium in rice (Oryza sativa L. cv. IR-72). Plant Cell and Environment 33, 23–34 (2010).
Scherzer, S. et al. The Dionaea muscipula ammonium channel dmAMT1 provides NH4 + uptake associated with venus flytrap’s prey digestion. Current Biology 23, 1649–1657 (2013).
Tegeder, M. & Masclaux-Daubresse, C. Source and sink mechanisms of nitrogen transport and use. New Phytologist 217, 35–53 (2018).
Miller, A. J. & Cramer, M. D. Root nitrogen acquisition and assimilation. Plant and Soil 274, 1–36 (2005).
Mokhele, B., Zhan, X. J., Yang, G. Z. & Zhang, X. L. Review: Nitrogen assimilation in crop plants and its affecting factors. Canadian Journal of Plant Science 92, 399–405 (2012).
Xuan, W., Beeckman, T. & Xu, G. H. Plant nitrogen nutrition: sensing and signaling. Current Opinion in Plant Biology 39, 57–65 (2017).
Setien, I. et al. High irradiance improves ammonium tolerance in wheat plants by increasing N assimilation. Journal of Plant Physiology 170, 758–771 (2013).
Skopelitis, D. S. et al. Abiotic stress generates ROS that signal expression of anionic glutamate dehydrogenases to form glutamate for proline synthesis in tobacco and grapevine. The Plant Cell 18, 2767–2781 (2006).
Szabados, L. & Savoure, A. Proline: a multifunctional amino acid. Trends in Plant Science 15, 89–97 (2010).
Zhao, P. et al. ATHB17 enhances stress tolerance by coordinating photosynthesis associated nuclear gene and ATSIG5 expression in response to abiotic stress. Scientific Reports 7, 15 (2017).
Geider, R. J., Roche, J., Greene, R. M. & Olaizola, M. Response of the photosynthetic apparatus of Phaeodactylum tricornutum (Bacillariophyceae) to nitrate, phosphate, or iron starvation1. Journal of Phycology 29, 755–766 (1993).
Alipanah, L., Rohloff, J., Winge, P., Bones, A. M. & Brembu, T. Whole-cell response to nitrogen deprivation in the diatom Phaeodactylum tricornutum. Journal of Experimental Botany 66, 6281–6296 (2015).
Yang, Q. Y., Shohag, M. J. I., Feng, Y., He, Z. L. & Yang, X. E. Transcriptome comparison reveals the adaptive evolution of two contrasting ecotypes of Zn/Cd hyperaccumulator Sedum alfredii hance. Frontiers in Plant Science 8, 12 (2017).
Igamberdiev, A. U. & Eprintsev, A. T. Organic acids: the pools of fixed carbon involved in redox regulation and energy balance in higher plants. Frontiers in Plant Science 7, 15 (2016).
Sarasketa, A., Gonzalez-Moro, M. B., Gonzalez-Murua, C. & Marino, D. Nitrogen source and external medium pH interaction differentially affects root and shoot metabolism in Arabidopsis. Frontiers in Plant Science 7, 12 (2016).
Thalmann, M. & Santelia, D. Starch as a determinant of plant fitness under abiotic stress. New Phytologist 214, 943–951 (2017).
Dong, S. Y., Zhang, J. & Beckles, D. M. A pivotal role for starch in the reconfiguration of C-14-partitioning and allocation in Arabidopsis thaliana under short-term abiotic stress. Scientific Reports 8, 12 (2018).
Miller, G., Suzuki, N., Ciftci-Yilmaz, S. & Mittler, R. Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell and Environment 33, 453–467 (2010).
Hachiya, T. & Sakakibara, H. Interactions between nitrate and ammonium in their uptake, allocation, assimilation, and signaling in plants. Journal of Experimental Botany 68, 2501–2512 (2017).
Sato, S. & Yanagisawa, S. Characterization of metabolic states of Arabidopsis thaliana under diverse carbon and nitrogen nutrient conditions via targeted metabolomic analysis. Plant and Cell Physiology 55, 306–319 (2014).
Zinta, G., Khan, A., AbdElgawad, H., Verma, V. & Srivastava, A. K. Unveiling the redox control of plant reproductive development during abiotic stress. Frontiers in Plant Science 7, 700 (2016).
Patterson, K. et al. Distinct signalling pathways and transcriptome response signatures differentiate ammonium- and nitrate-supplied plants. Plant Cell and Environment 33, 1486–1501 (2010).
Laloum, T., Martin, G. & Duque, P. Alternative splicing control of abiotic stress responses. Trends in Plant Science 23, 140–150 (2018).
Zarcotejada, P. et al. Assessing vineyard condition with hyperspectral indices: leaf and canopy reflectance simulation in a row-structured discontinuous canopy. Remote Sensing of Environment 99, 271–287 (2005).
Dhindsa, R. S., Plumb-Dhindsa, P. & Thorpe, T. A. Leaf senescence: correlated with increased levels of membrane permeability and lipid peroxidation, and decreased levels of superoxide dismutase and catalase. Journal of Experimental Botany 32, 93–101 (1981).
Patterson, B. D., Macrae, E. A. & Ferguson, I. B. Estimation of hydrogen peroxide in plant extracts using titanium(IV). Analytical Biochemistry 139, 487–492 (1984).
Wang, A. G. Quantitative relation between the reaction of hydroxylamine and superoxide anion radicals in plants. Plant Physiology Commu. nications 26, 55–57 (1990).
Sarasketa, A., Gonzalez-Moro, M. B., Gonzalez-Murua, C. & Marino, D. Exploring ammonium tolerance in a large panel of Arabidopsis thaliana natural accessions. Journal of Experimental Botany 65, 6023–6033 (2014).
Djeridane, A. et al. Antioxidant activity of some algerian medicinal plants extracts containing phenolic compounds. Food Chemistry 97, 654–660 (2006).
Jia, Z., Tang, M. C. & Wu, J. M. The determination of flavonoid contents in mulberry and their scavenging effects on superoxide radicals. Food Chemistry 64, 555–559 (1999).
Seiffert, B., Zhou, Z. W., Wallbraun, M., Lohaus, G. & Mollers, C. Expression of a bacterial asparagine synthetase gene in oilseed rape (Brassica napus) and its effect on traits related to nitrogen efficiency. Physiologia Plantarum 121, 656–665 (2004).
Vijayan, J. et al. Transcriptomic and physiological studies identify cues for germination stage oxygen deficiency tolerance in rice. Environmental and Experimental Botany 147, 234–248 (2018).
Bernfeld, P. Amylases, α and β. Methods in Enzymology 1, 149–158 (1955).
Kilker, R. D., Saunier, B., Tkacz, J. S. & Herscovics, A. Partial purification from Saccharomyces cerevisiae of a soluble glucosidase which removes the terminal glucose from the oligosaccharide Glc3Man9GlcNAc2. Journal of Biological Chemistry 256, 5299–5603 (1981).
Aebi, H. Catalase in vitro. Methods in Enzymology 105, 121–126 (1984).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B 57, 289–300 (1995).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We wish to thank Annoroad Gene Technology Co., Ltd, Beijing, China for their help with the data analysis. This research was supported by the National Natural Science Foundation of China (Nos 31670507, 31700429), the Interdisciplinary Innovation Team Program of the Chinese Academy of Sciences (No. 2015, Team of Directional Biotransformation of Environmental Pollutants) and the Major Science and Technology Program for Water Pollution Control and Treatment of China (Nos 2015ZX07206-006 and 2018ZX07105001).
Author information
Authors and Affiliations
Contributions
X.Z., R.W. and S.X. conceived the project and designed the study; R.W., H.S., C.J., S.Z. and S.W. performed the experiments; R.W., G.Z. and B.C. analyzed the data; R.W., S.F. Z.B. and B.C. wrote the manuscript, with input from all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, R., Xu, S., Sun, H. et al. Complex regulatory network allows Myriophyllum aquaticum to thrive under high-concentration ammonia toxicity. Sci Rep 9, 4801 (2019). https://doi.org/10.1038/s41598-019-41236-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41236-8
This article is cited by
-
Ascorbic acid alleviates the autolysis of the sea cucumber Apostichopus japonicus via the activation of antioxidant enzymes to remove reactive oxygen species during live storage
Aquaculture International (2023)
-
Genome-wide identification of resistance genes and transcriptome regulation in yeast to accommodate ammonium toxicity
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.