Abstract
In the recent years, new molecular methods have been proposed to discriminate multicentric hepatocellular carcinomas (HCC) from intrahepatic metastases. Some of these methods utilize sequencing data to assess similarities between cancer genomes, whilst other achieved the same results with transcriptome and methylome data. Here, we attempt to classify two HCC patients with multi-centric disease using the recall-rates of somatic mutations but find that difficult because their tumors share some chromosome-scale copy-number alterations (CNAs) but little-to-no single-nucleotide variants. To resolve the apparent conundrum, we apply a phasing strategy to test if those shared CNAs are identical by descent. Our findings suggest that the conflicting alterations occur on different homologous chromosomes, which argues for multi-centric origin of respective HCCs.
Similar content being viewed by others
Introduction
Hepatocellular carcinomas (HCC) with intra-hepatic metastases (IM) are profoundly different in their development and clinical outcome from multi-centric tumors1. The clinical discrimination between these subtypes has historically been challenging and is usually based on tumor location, blood vessel involvement of the primary tumor, and/or hemodynamics in CT/MRI imaging before resection2. Once tumors are resected, pathological evaluation is performed to discriminate between the IMs and MCs. However, there is no consensus at present on how to discriminate the two and it is advised to use complementary molecular tests to reduce the uncertainty. For example, in a recent study Furuta et al. showed similarities between genomes of multi-centric tumors using recall-rates of somatic alterations3, while others achieved the same with transcriptome and methylome data4,5. In this study, we extend Furuta’s strategy by accounting for convergent evolution of copy-number alterations (CNA) in multi-centric tumors that can, in certain situations, complicate data interpretation and clinical reasoning.
Results
We generated testing data from multiple-regions of chemotherapy-naïve multi-centric HCCs of two HBV/HCV-negative patients (Table 1). In each patient, somatic mutations were identified from multi-region exome sequencing data from three synchronous tumors (Fig. 1A). Prior to the analysis, sequencing reads were mapped onto the human genome hs37d5 and GATK haplotype caller was used to identify single-nucleotide (SNV) mutations and indels simultaneously. On average, 42.8/21.0 somatic SNVs and 106/15.3 indels were detected per tumor region per patient (Fig. 1B, Supplementary Table 2), of which six and four variants per patient were cancer driver mutations (Fig. 1C). Pairwise comparison of somatic SNVs across all regions of a given tumor yielded average recall rates of 65% and 85% per patient, while across different tumors 10.9% and 7.3% (Fig. 2; Supplementary Tables 3–6).
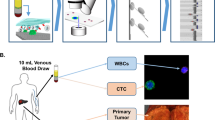
Multi-region exome sequencing of multi-centric hepatocellular carcinoma. (A,B) Sampling scheme and mutation burden of tumor regions (grey: single-nucleotide variants, blue: indels). (C) Cancer driver mutations. Abbreviation: T: tumor, L: left, R: right, (C) control, matched normal tissue/blood, CNA: copy-number alteration, SNV: single-nucleotide variant.
We therefore ensured that there was not a significant enrichment of false positive variants in our analysis by replicating 21/23 (91%) selected somatic mutations and re-assessing hematoxylin-eosin stains of tumor tissues to ensure that they contained at least 80% of tumor cells (Supplementary Table 7). We then undertook a down-sampling approach to equalize read depth across all tumor regions and repeated the analysis with additional cancer-specialized calling algorithms, including MuTect26 (i.e. GATK4 implementation of MuTect) and Strelka7. Nevertheless, read-depth equalizing or the use of different variant calling algorithms did not improve the recall rates and a gap between the degree of inter- and intra-tumor heterogeneity remained.
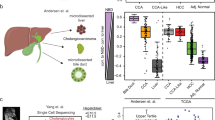
Most somatic SNVs occurred at low allelic frequencies irrespectively of whether they were recalled in different regions of the same tumor or not (Fig. 3; Supplementary Tables 8–9). Since cancer driver mutations were no exception and thus no clearly clonal candidate was found, we wondered if some other type of mutations could help us to classify multi-centric tumors of our patients. For example, there were four possibly identical copy-number alterations (CNA; Fig. 4) occurring in two tumors of each patient. To determine if these CNAs were not identical by descent, we used a combination of breakpoint analysis and chromosome phasing. The specific phasing algorithm is detailed in material and methods and on a Fig. 4 and is, at heart, a Welch’s two sample t-test that compares allele-frequency changes at polymorphic sites of candidate CNAs of tumor samples and a tumor-free tissue, on the assumption that allele frequencies for each chromosomal haplotype are random variables with mean μ and variance σ2. Our null hypothesis was that CNAs occurred at the same homologous chromosome in different tumors. After reviewing each case, the null hypothesis was rejected on a significance level of 0.05 for cases of CNAs of chromosome 21q, 8q and 17p, whilst accepted for the case of 6q. In this case, however, we used the breakpoint information to argue that 6qter alterations were independent mutation events.
Copy-number alterations (CNAs) and chromosome phasing. (A) An algorithmic overview. (B) Copy-number profile of multi-centric HCCs. Candidate CNAs are indicated by arrow. Chromosomal gains are blue, losses are red. (C) Haplotype inference using polymorphic sites and their respective allelic frequency changes. Abbreviation: (C) control; blood sample or non-tumor tissue, SNV: single-nucleotide variant, T: tumor, H: haplotype.
Discussion
In this study, we attempted to classify multi-centric hepatocellular carcinomas using the information that may be not be available for strategies that either compare tumor morphology or mutation burden. Especially, the use of the latter as proposed by Furuta et al. can be sometimes misleading because the assignment of single or multiple origin category is possible only for tumors which share most or no somatic mutations, but otherwise there is no clear threshold to distinguish them. Given that cancer driver mutations can occur in cells that do not undergo malignant transformation8,9, it could well be that multi-centric tumors originating independently can be difficult to distinguish from IMs that have arisen very early10. It is also possible that these tumors behave differently in development and clinical outcome from IMs that originated late in progression and are classified as true IMs. Thus, if disease prognosis and/or clinical classification of IMs is the end-point of an analysis, molecular strategies utilizing somatic mutations may not be precise enough to distinguish the differing clinical categories. Whilst our study illustrates how refinements of the existing methods can be used for better data interpretation, it also outlines some of the greater limitations that must be overcome before molecular strategies are implemented clinically.
Methods
Patients and samples
The study was approved by the ethical committee of the Medical University of Graz (ethical approval 26-170-ex-13/14) on 20th January 2014. All methods were performed according to the relevant guidelines and regulations. Informed consent for study participation was obtained from both patients. DNA was extracted from six fresh-frozen tumors and each tumor was sampled at least twice. Pathological examinations of tumor samples were performed by experienced pathologists to ensure that each biopsy contained at least 80% of tumor cells. DNA derived of 20 samples was subjected to Illumina HiSeq. 2000 pair-end sequencing, including six tumor-free (cirrhotic) liver parenchyma and two blood samples.
Illumina sequencing
Whole exome sequencing (WES) was performed using the Illumina HiSeq. 2000 platform after short insert ~400 bp libraries were constructed, flow cells prepared, and clusters generated. For WES target DNA was enriched by bait capture (Agilent SureSelect ver. 4) and short-insert libraries were sequenced to average depth of 118X (IQR 56, Supplementary Table 1). Sequencing data are freely available from the authors after a reasonable request.
Semiconductor sequencing
Ion torrent sequencing was selected for technical replication of somatic variants. Used in conjunction with the AmpliSeq Library Kit 2.0, the Ion AmpliSeq™ Comprehensive Cancer Panel was selected to capture exons of 409 genes from the Cancer Gene Census database and the resulting libraries were ran on the Ion PGM™ Sequencer. The raw reads were processed using the Ion Reporter software with recommended settings.
Sanger sequencing
Following PCR amplification, Sanger sequencing was used to detect TERT promoter mutations g.5:1295228 G > A/T and g.5:1295250 G > A. The primer sequences were 5′-CCAGGGCTTCCCACGTGC-3′ for the forward primer and 5′-ACTGGGGACCCGGGCACC-3′ for the reverse primer.
Variant Detection and Filtering
Raw sequencing reads were quality-checked (fastqc ver. 0.11.7), adapter-trimmed, duplicate-removed (Picard tools ver. 2.9) and mapped onto the hs37d5 version of the human genome (BWA ver. 0.7). The GATK pipeline (ver. 3.7) was used to perform base-quality score recalibration and variant calling. Specifically, we used GATK haplotype caller algorithm with standard settings, followed by Variant Quality Score Recalibration (VQSR) for single-base substitution identification and the Scalpel algorithm (ver. 0.5.3) for indels. Variant files containing too few variants were specifically filtered (QD > 10.0, MQ > 40.0, FS < 30.0, SOR < 3.0, MQRankSum > −12.5, ReadPosRankSum > −8.0) to extract high-quality variants. We used the human genome assembly hs37d5 and 2017 versions of ANNOVAR databases to annotate variants. Germline or somatic origin of the variants and indels were determined based on their presence or absence in the matched tumour-free tissue.
We applied the following exclusion filters to somatic variants: (i) presence in a segmental duplication region; (ii) variant present in any read from paired normal sample; (iii) fewer than ten reads in total at the variant site in the normal sample; (iv) fewer than eight reads in total in the tumor; (v) fewer three variant reads in the tumor; variant allele frequency <3% in the tumor; and (vi) presence of variant in the Exome Aggregation Consortium dataset (release 22.6.2017) at a frequency >2%. Variants identified in constitutional DNA from any of the other local, non-cancer sequencing project at a frequency of 5% (for example, 29 million variants across 284 samples from the Oxford-Illumina WGS500 consortium) were discarded as being more likely due to systematic error in our pipeline than genuine somatic mutations.
Copy Number Calling
Nexus Copy Number Discovery software (ver. 9.0) was used to identify CNAs from sequencing data and for independent validation of these alterations with Affymetrix Oncoscan arrays. The raw array data were processed with Affymetrix ChaS software, (ver. 3.2) and imported as segmental reports into the software. Only CNAs larger than 50Kb with a minimum support of 21 probes were considered for analysis.
Chromosome Phasing
Allele frequencies (AF) of polymorphic sites of candidate CNAs were extracted and AF differences between tumors and a normal tissue were calculated. For each case, corresponding chromosomal haplotypes were inferred from SNVs with AFs shifted in the same direction (i.e. either positively or negatively). Welch two sample t-test was then used to compare AF values of respective haplotypes across different tumors on the assumption that AF observations are random variables with mean \({\mu }\) and variance σ2.
References
Bosman, F., Carneiro, F., Hruban, R. & Theise, N. WHO Classification of Tumours of the Digestive System, (WHO Press, Geneva, 2010).
Toyoda, H. et al. Non-hypervascular hypointense nodules detected by Gd-EOB-DTPA-enhanced MRI are a risk factor for recurrence of HCC after hepatectomy. J Hepatol 58, 1174–1180 (2013).
Furuta, M. et al. Whole genome sequencing discriminates hepatocellular carcinoma with intrahepatic metastasis from multi-centric tumors. J Hepatol 66, 363–373 (2017).
Nomoto, S. et al. Hypermethylation of multiple genes as clonal markers in multicentric hepatocellular carcinoma. Br J Cancer 97, 1260–1265 (2007).
Nakata, T. et al. Identification of genes associated with multiple nodules in hepatocellular carcinoma using cDNA microarray: multicentric occurrence or intrahepatic metastasis? Hepatogastroenterology 55, 865–872 (2008).
do Valle, I. F. et al. Optimized pipeline of MuTect and GATK tools to improve the detection of somatic single nucleotide polymorphisms in whole-exome sequencing data. BMC Bioinformatics 17, 341 (2016).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, (1811–1817 (2012).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Ammerpohl, O. et al. Distinct DNA methylation patterns in cirrhotic liver and hepatocellular carcinoma. Int J Cancer 130, 1319–1328 (2012).
Zhao, Z. M. et al. Early and multiple origins of metastatic lineages within primary tumors. Proc Natl Acad Sci USA 113, 2140–2145 (2016).
Acknowledgements
The authors would like to express their sincere gratitude to Dr. Diego Mallo of Arizona State University for useful comments and other contributions. We also thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics for generation of sequencing data. MK was generously supported by the Krebsliga beider Basel (grant no. KLBB-12-2013) and the University of Basel (“Förderung exzellenter Nachwuchsforschender”). S.P. is supported by Swiss National Science Foundation (PZ00P3_168165). V.P. is supported by the Swiss Centre for Applied Human Toxicology (SCAHT).
Author information
Authors and Affiliations
Contributions
M.K., C.L. and L.T. conceived the study. M.K. designed experiments and analyzed data with help of SR, B.A., K.H., W.C. and M.Ko. L.Q. with S.P. supervised and V.P. performed laboratory experiments. C.L. provided and L.Q. with C.Q. processed tissue samples. C.L., L.T. and D.B. performed histological assessment of tumor specimens. M.K. wrote the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lackner, C., Quagliata, L., Cross, W. et al. Convergent Evolution of Copy Number Alterations in Multi-Centric Hepatocellular Carcinoma. Sci Rep 9, 4611 (2019). https://doi.org/10.1038/s41598-019-40843-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-40843-9
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.