Abstract
The association between Clostridium species identification from stool samples in preterm neonates and the occurrence of necrotizing enterocolitis has been increasingly reported. To confirm the specific impact of Clostridium butyricum in this pathology, selective culture procedure was used for Clostridia isolation. Whole-genome analysis was employed to investigate genomic relationships between isolates. Stool samples from present study, as well as from previously investigated cases, were implicated including 88 from preterm neonates with necrotizing enterocolitis and 71 from matched controls. Quantitative real-time polymerase chain reaction was performed to evaluate the presence of C. butyricum from stools of new cases. Clostridium species prevalence isolated by culture was compared between patients with necrotizing enterocolitis and controls. By combining results of both culture and quantitative polymerase chain reaction methods, C. butyricum was significantly more frequent in stool samples from preterm neonates with necrotizing enterocolitis than in controls. Whole-genome analysis of 81 genomes including 58 neonates’ isolates revealed that cases were clustered depending on geographical origin of isolation. Controls isolates presented genomic relations with that of patients suggesting a mechanism of asymptomatic carriage. Overall, this suggests an epidemiology comparable to that observed in Clostridium difficile colitis in adults.
Similar content being viewed by others
Introduction
Necrotizing enterocolitis (NEC) is a multifactorial intestinal disease that occurs in 4–12% of preterm neonates, with a high attributable mortality especially in patients with very low birth weight (i.e. 500–1000 g)1,2. Recent studies propounded the idea of bacteriological etiology, mostly microbiota dysbiosis during NEC episodes, leading to the generation of inflammatory process3,4. In recent years, several bacterial species have been suspected of being associated with NEC outbreaks, especially Clostridia like Clostridium butyricum, Clostridium perfringens and Clostridium neonatale4,5. Zhao-Fleming et al. also reported a significantly higher frequency of strictly anaerobic species by using 16S rRNA sequencing to identify bacteria from patients with necrotizing infections6. Cassir et al. described a relationship between the presence of C. butyricum in stool samples from preterm neonates and the occurrence of NEC, using a combination of three techniques (16S rRNA pyrosequencing, culturomics and qPCR). However, neither the pathogenicity nor the epidemiology of this relationship have yet been totally resolved7. In addition to Clostridia, Gram-negative bacterial species were also identified especially, K. pneumoniae, E. cloacae and Uropathogenic Escherichia coli8,9.
In present work, we conducted a multidisciplinary study on a cohort of NEC and control preterm neonates stool samples that we investigated for C. butyricum in stool by qPCR, a small fraction of samples only being previously tested by culture7. The main objective of the current investigation on this cohort was to isolate the qPCR-detected C. butyricum in order to perform strain typing by whole genome sequencing on isolates and to correlate it with the date of sampling as the geographical origin of the isolates. Indeed, in a preliminary work done on 16 strains, we suspected isolates to have a clonal origin isolates10. For this, we used a heat-shock selective culture protocol that allowed us also to isolate C. butyricum but also some other Clostridia previously also suspected to be associated to NEC. We took the opportunity in the present work to add additional cases received between 2015 and 2017. Then, the association between C. butyricum and the occurrence of NEC was reassessed and typing data were analysed together.
Results
Evaluation of Clostridium species community in samples from preterm neonates
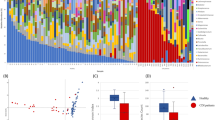
Combination of culture and qPCR results allowed the identification of a strong association between the presence of C. butyricum in stool samples from preterm neonates and the occurrence of NEC, when compared to controls (66/88; 75% vs. 8/71; 11%; p < 0.001 respectively) (Table 1). Overall, 58 strains of C. butyricum were isolated including 52 (59.1%; n = 88) from NEC patients and 6 (8.45%; n = 71) from controls (p < 0.001) (Table 2). By comparing means of cycle thresholds (Ct), the density of C. butyricum was lower in controls stool samples than in NEC (32.938 and 29.182 respectively, p value = 0.0344). The frequency of C. butyricum in this later seems to be associated with the administration of antibiotics (p value = 0.027), which is not the case in controls (p value = 0.838) (Supplementary Table S1). As for the other previously proposed NEC-associated Clostridium species, we isolated four strains (4.5%) of C. neonatale from NEC patients and one (1.14%) from controls (p = 0.26). Clostridium perfringens was detected equally in patients with NEC and controls (respectively 8; 9% and 6; 8.4%). This was also the case for Clostridium paraputrificum (1; 1.1% and 1; 1.4% respectively). Finally, we identified other Clostridia from the stools of patients with NEC: Clostridium sardiniense (1; 1.1%), Clostridium tertium (1; 1.1%) and Clostridium difficile (2; 2.3%). In particular, 7/8 and 1/6 of C. perfringens from the NEC patients and controls, respectively, were co-isolated with C. butyricum in the same sample.
Core-genome and core-genome single-nucleotide polymorphism phylogenetic analysis
The average length of genomes included in this study was 4,676,116 bp. The greatest genome size was 5,214,902 bp (strain NEC23), the smallest genome size was 4,014,159 bp (strain DORA). An average of 4,363 open reading frames was predicted (Supplementary Table S2). Although strains isolated from Marseille (NICU-1 and NICU-2) were over-represented in this study, the eBURST analysis of core-SNP enabled the identification of three main C. butyricum clonal groups (a, b, c): group (b) included one cluster; cluster (b), thus both group (a) and (c) included two clusters; respectively cluster (a1, a2) and cluster (c1, c2) (Figs 1, 2 and Supplementary Figure S1–S3). The majority of NEC-associated C. butyricum (28/52, 54%) were clustered within the group (b). Genomic relationships were identified between NEC and control isolates, also grouped depending on geographic source of isolation. Strains isolated from the same NICUs were close to each other i.e. cluster (a1), (a2), (b), (c1) isolates detected from NICU-1 and 2 in Marseille, also, C. butyricum strains from NICU-3 and NICU-4 on the other hand in cluster (a1) and (b) (Fig. 1). Core-SNP temporal clustering revealed that highly-related strains were identified between 2009 and 2010 in group (b) (73.5%; 25/34), also, during 2010 in cluster (c1) (Fig. 2). Nearly all strains were grouped in clusters revealing their clonality whereas strains from other origin were dispersed and far from each another. The comparative analysis allowed the identification of hemolysin-encoding genes (hemolysin A, B, C and beta) in NEC and control isolates from all clusters.
Clostridium butyricum geographic relationship based on core-genome single-nucleotide polymorphism phylogenetic analysis. The color of strain names represents the geographic zone of isolation. (Marseille: NICU-1, NICU-2; Nice: NICU-3; Montpellier: NICU-4). Red strain names represents the 16 isolates sequenced Cassir et al.7. NEC: necrotizing enterocolitis, NICU: neonatal intensive care units.
Clostridium butyricum temporal relationship based on core-genome single-nucleotide polymorphism phylogenetic analysis. The color of strain names represents the timing of isolation. Red strain names represents the 16 isolates sequenced Cassir et al.7. NEC: necrotizing enterocolitis, C: control.
Genotyping using multispacer sequence typing phylogeny
MST phylogeny identified four sequence types (ST) where the strains had the same MST: NEC1, NEC3, NEC10 and NEC17 (Supplementary Figure S4 and S5). We compared the eBURST analysis of MST with that of core-SNP. Isolates were clustered into three groups centered on the Kwashiorkor strain (KW10): Group a’ included the same cluster as Group a, except for NEC1, NEC8, NEC35, NEC72 and NEC74; Group b’ included the same strains as Group b except for NEC29, NEC52, NEC64, NEC65 from Group (a) and NEC88 from Group (c); finally, Group c’ included NEC50.
Discussion
Inclusion of additional cases and the use of a selective culture procedure for Clostridium isolation confirmed our previous results showing a significant association between C. butyricum and NEC in preterm neonates7,10. Indeed, 75% of NEC were positive using qPCR and culture, while this rate was only 11% in controls (p < 0.001). Culture technique emphasizes the specificity of qPCR, as 3.4% of NEC-associated C. butyricum were negative by qPCR and positive by culture. Reciprocally, qPCR validated the culture by detection of C. butyricum in 75.75% (50/66) from all positive samples. The advantage of this selective culture protocol allowed easy isolation of C. butyricum during NEC outbreaks as compared to culturomics technique which is highly fastidious7. The drawback of included culture method was the possibility to skip isolation of in vivo non-spore-form of Clostridia species. As well, the density of C. butyricum was higher in NEC than in controls (p value = 0.0344). The association between C. butyricum-linked NEC with antibiotics administration (p value = 0.027), suggested a mechanism of pathogenicity comparable to that of C. difficile, where cases of pseudomembranous colitis were significantly associated with previous antibiotic administration, especially lincosamides, third generation cephalosporins and fluoroquinolones. It would be interesting, in future studies, to evaluate the influence of antibiotics families in the occurrence of NEC. Additionally, culture enabled the isolation of other Clostridia (n = 25) previously suggested in linked with NEC that were not evaluated by Cassir et al.7. Among these, C. neonatale and C. perfringens were specifically suspected to be associated with NEC4,5,11,12,13,14,15. These species were infrequently isolated, the most predominant being C. perfringens, with a total of 14 isolates observed equally in both NEC and control groups. Moreover, it was co-isolated with C. butyricum in 62.5% of patients. This result differs from that of Sim et al., where the authors used a metagenomic approach to detect C. perfringens type A (3/12; 25%) from NEC samples16. It has been demonstrated that early colonization by C. perfringens affected the onset of NEC14. Furthermore, unbalanced gut microbiota in Lebanese neonates suffering from NEC have been reported, including C. perfringens widely identified in patients, despite the limited number of samples tested in this study17. C. neonatale was less prevalent than C. butyricum and C. perfringens and not significantly associated with NEC (NEC and controls, n = 4, 4.5% versus n = 1, 1.4% respectively, p = 0.5). However, it is difficult to draw conclusion from a limited number of isolates. C. neonatale was first identified by Alfa et al., during a NEC outbreak in a Canadian NICU. However, no control neonates were tested, and its pathogenicity was not fully investigated15, only its microbiologic features were studied18, particularly through comparative phenotypic features with C. butyricum19.
Genomic relations were previously reported by Benamar et al. on a limited number of strains (n = 32) including 16 isolated from NEC and controls10. Three clonal clades were identified. In the present work, 42 additional preterm neonates’ strains isolated from different geographic origin and time of isolation were analysed. Core-genome, core-SNP and MST phylogenetic analysis on an exceptional collection of C. butyricum strains (n = 81) confirmed genomic relationship between NEC-associated and controls isolates as the discrimination of the prior identified clades. Therefore, three main epidemic clonal groups were identified, including a large clonal cluster occurring between 2009 and 2010 (cluster (b)). This last clustered the majority of NEC-associated C. butyricum (28/52, 54%) identified as clade C in Benamar et al.10. In some cases, especially during September/October 2010, NEC occurred as an outbreak (n = 10) with highly related isolates. Other small epidemics clusters were detected with a limited number of strains (Groups a1, a2, c1 and c2). Additionally, strains isolated from the same NICU were highly related suggesting the existence of geographic-associated C. butyricum clonal lineages and epidemic transmission. This was confirmed by similarities observed between controls and NEC-associated isolates and highlighted a C. butyricum asymptomatic carriage of these genetically-related strains. Clusters (a2) and (c2) were not observed in Benamar et al.10 and corresponded to strains isolated after 2015. This finding confirms the temporal clustering.
The first NEC-associated C. butyricum outbreak was reported by Howard et al. in 1977, based on clinical examination and microbiologic analysis, since no genotyping methods were employed20. Gorham et al. reported 152 cases of C. butyricum hand-carriage isolated from 152 healthcare workers during a NEC outbreak21. Several studies showed that C. butyricum is a beneficial microorganism, especially MIYAIRI 588 strain, that prevents entero-hemorrhagic E. coli, C. difficile infections and gastric ulcers22. Conversely, other strains might contribute to the pathogenesis of several infectious diseases, such as botulism and NEC22,23. Recently, Wydau-Dematteis et al. reported the importance of the dlt operon of C. butyricum associated NEC by establishing the resistance of C. butyricum to antimicrobial peptides, lysozymes and vancomycin24. A high dose of butyric acid has also been demonstrated to have a cytotoxic effect on diverse cell lines, notably Caco-2 cells monolayers25. Genomic identification in C. butyricum of a toxin comparable to β-hemolysin secreted by Brachyspira hyodysenteriae7 suggests a similar possible mechanism for cytotoxicity26. Moreover, we previously reported a cytotoxic activity of C. butyricum supernatant tested by flow cytometry on Jurkat cells7. Such a toxigenic mechanism of Clostridia in NEC disease is supported by the findings of Heida et al., who observed a low density of Clostridia by fluorescent in situ hybridization27. Other toxins-related NEC cases have been suggested in association with different bacterial species such as toxigenic E. coli, Pseudomonas aeruginosa, Serratia spp., delta toxin-producing strains of coagulase-negative Staphylococci, S. aureus and pore-forming type A toxin Clostridium perfringens28,29.
The picture observed in C. butyricum-associated NEC resembles that of C. difficile- associated pseudomembranous colitis30. Similarly to C. butyricum-associated NEC, C. difficile hospital-associated isolates were found to be highly related with asymptomatic carriage, supporting the idea of a plausible transmission amongst hospitals as from environmental reservoirs31,32. This work is the first to note that the WGS was used to investigate probable C. butyricum-associated NEC outbreak, during a period of 8 years using SNP analysis. WGS is a revolutionizing large-scale genomic-based technique, highlighting the clonal transmission between reservoirs and patients33, as the identification of virulence factors34. In addition, we observed that MST techniques represent an alternative method as clusters were approximately validated by core-SNP analysis, but due to continuous reduction of costs, WGS will be probably preferred for routine use in future diagnosis.
Conclusion
The key findings of this study were first, the confirmation of significant association between C. butyricum strains and the occurrence of NEC, then the genetic correlation between NEC and non-NEC-associated strains of C. butyricum. This suggested that cases emerge from a small number of genetically heterogeneous sources, validated the possibility of asymptomatic carriage and confirmed that most strains are hospital-acquired. These findings and significant association with the administration of antibiotics in NEC patients resembles epidemiology of C. difficile in adults.
Methods
Study design and patients
One hundred and fifty-nine preterm neonates with NEC and matched controls were enrolled, including neonates from the study of Cassir et al. (NEC n = 82, control n = 67)7, and 10 (NEC n = 6, control n = 4) received between 2015 and 2017 (Supplementary Table S3). The stool samples collected from 88 NEC and 71 controls were from four south-eastern regions of France (Marseille, Nice, Nîmes and Montpellier), including five neonatal intensive care units (NICUs) (Table 2, Supplementary Table S2 and Figure S6). Cases were defined as patients with suspected, definite or advanced NEC corresponding to Bell stages I, II or III. When feasible, stool samples were collected on the day of symptoms onset and stored at −80 °C. Controls were matched with patients by sex, gestational age, birth weight, days of life, feeding strategies, mode of delivery, and previous antibiotic therapy and NICU. All patients with NEC and controls were preterm neonates (under 37 weeks completed weeks of gestation). Since the dates of antibiotic administration were not recorded, patients were not distinguished for the duration of antibiotic therapy. None of the selected patients had received probiotics or been included in therapeutic protocols prior to stool collection and all were negative for routine microbial investigation. Agreements from the ethics committee of the “Institut Fédératif de Recherche, IFR48” and the “Institut Hospitalo-Universitaire, IHU-2017-007” were obtained to confirm the study procedure, as well as a written informed consent from parents of all patients7. All methods were performed in accordance with the relevant guidelines and regulations.
Clostridium species isolation, C. butyricum qPCR detection
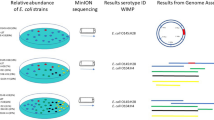
Stools were cultured for Clostridium species isolation using HS treatment based on the thermal resistance of spores35. Briefly, samples were suspended (v/v) in sterile phosphate-buffered saline by vortexing and heated for 20 minutes at 80 °C. Stools were then cultured on 5% Columbia sheep blood agar (Becton Dickinson®, USA) at 37 °C for 48 hours in an anaerobic atmosphere inside an anaerobic chamber. Provisional identification of all strains was performed by MALDI-TOF, then the amplification and sequencing of 16S rRNA gene was used to confirm this identification, as previously described36. Simultaneously, specific detection of C. butyricum qPCR was performed on DNA extracts obtained from stool samples of cases received after 2015, as described previously7. Data of qPCR and culture-based technique were analysed, the same 30 stool samples tested by culturomics method in Cassir et al. study being used as positive controls to validate the ability of spore-based method in the isolation of C. butyricum7.
Core-genome and core-genome single-nucleotide polymorphism analysis
The genomic DNA of isolated C. butyricum was sequenced on MiSeq sequencer (Illumina, San Diego, CA, USA) using the paired-end strategy. Reads were assembled and scaffolded using SPAdes version 3.4.037. Overall, 81 genomes were analysed, including 42 from strains isolated and sequenced in this study, 16 from previous isolates7, 5 strains isolated in our laboratory from humans as part of gut microbiota analysis38 and 18 from the Genbank database. Analysed genomes and their references are summarized in Supplementary Table S2. Protein sequences and nucleotide coordinates were predicted using the GeneMarkS platform (Genemark™, USA)39. Orthologous proteins analysis were performed using the ProteinOrtho software40. Core-genome coding DNA sequences were inferred from the pan-genome, concatenated and aligned using a Python script. SNPs were obtained from the core-genome using SNP-sites software41. Phylogenetic trees were generated using the maximum-likelihood method within PhyML42 and edited by Treegraph 2 software43. PHYLOViZ version 2.0 was used to generate a core-SNP eBURST phylogenetic tree based on expected numbers of nucleotide substitutions per variable site44. Scaled phylogenetic trees were procured using Adobe Photoshop CC 2018. BLASTP was retrieved to allow the identification of potential hemolysin sequences within the draft genomes of C. butyricum, their protein sequences are in Supplementary Material 1.
Genotyping using multispacer sequence typing
MST was performed as described by Benamar et al.10. Briefly, DNA from patients’ isolates was extracted and amplified by standard PCR and then sequenced using 16 capillary sequencer 3130 XL (Applied Biosystems®, USA) and the BigDye Terminator v1.1 Cycle Sequencing kit (Applied Biosystems®, USA). For C. butyricum genomes available online, intergenic spacers were extracted by in Silico BLASTn.
Statistical analysis
Statistical analysis was performed using the SPSS® statistics software 2016 (IBM, NY, USA). Mean and standard deviation were used to describe continuous variables. Percentage and number of events were used for quantitative variables. Student t-test or Mann-Whitney U test was used to perform two-group comparisons for quantitative variables. The chi-square (Mantel-Haenszel) test was used to perform two-group comparisons for qualitative variables.
References
Cassir, N., Simeoni, U. & La Scola, B. Gut microbiota and the pathogenesis of necrotizing enterocolitis in preterm neonates. Future Microbiol. 11, 273–92 (2016).
Seeman, S. M., Mehal, J. M., Haberling, D. L., Holman, R. C. & Stoll, B. J. Infant and maternal risk factors related to necrotising enterocolitis-associated infant death in the United States. Acta Paediatr. Int. J. Paediatr. 105, e240–e246 (2016).
Neu, J. & Pammi, M. Pathogenesis of NEC: Impact of an altered intestinal microbiome. Seminars in Perinatology 41, 29–35 (2017).
Hosny, M., Cassir, N. & La Scola, B. Updating on gut microbiota and its relationship with the occurrence of necrotizing enterocolitis. Hum. Microbiome J. 4, 14–19 (2017).
Roze, J.-C. et al. Nutritional strategies and gut microbiota composition as risk factors for necrotizing enterocolitis in very-preterm infants. Am. J. Clin. Nutr. 36, 821–830 (2017).
Zhao-Fleming, H., Dissanaike, S. & Rumbaugh, K. Are anaerobes a major, underappreciated cause of necrotizing infections? Anaerobe 45, 65–70 (2017).
Cassir, N. et al. Clostridium butyricum Strains and Dysbiosis Linked to Necrotizing Enterocolitis in Preterm Neonates. Clin. Infect. Dis. 61, 1107–1115 (2015).
Boccia, D., Stolfi, I., Lana, S. & Moro, M. L. Nosocomial necrotising enterocolitis outbreaks: Epidemiology and control measures. Eur. J. Pediatr. 160, 385–391 (2001).
Ward, D. V. et al. Metagenomic sequencing with strain-level resolution implicates uropathogenic E. coli in necrotizing enterocolitis and mortality in preterm infants. Cell Rep 14, 2912–2924 (2016).
Benamar, S. et al. Multi-spacer typing as an effective method to distinguish the clonal lineage of Clostridium butyricum strains isolated from stool samples during a series of necrotizing enterocolitis cases. J. Hosp. Infect. 95, 300–305 (2016).
Dittmar, E. et al. Necrotizing enterocolitis of the neonate with Clostridium perfringens: Diagnosis, clinical course, and role of alpha toxin. Eur. J. Pediatr. 167, 891–895 (2008).
Heida, F. H. et al. A necrotizing enterocolitis-associated gut microbiota is present in the meconium: results of a prospective study. Clin Infect Dis 62, 863–870 (2016).
Haque, K. Necrotizing enterocolitis - Some things old and some things new: A comprehensive review. J. Clin. Neonatol. 5, 79 (2016).
De la Cochetière, M.-F. et al. Early intestinal bacterial colonization and necrotizing enterocolitis in premature infants: The putative role of Clostridium. Pediatr. Res. 56, 366–370 (2004).
Alfa, M. J. et al. An Outbreak of necrotizing enterocolitis associated with a novel Clostridium species in a neonatal intensive care unit. Clin Infect Dis 35, 101–105 (2002).
Sim, K. et al. Dysbiosis anticipating necrotizing enterocolitis in very premature infants. Clin. Infect. Dis. 60, 389–397 (2015).
Itani, T. et al. Preterm infants with necrotising enterocolitis demonstrate an unbalanced gut microbiota. Acta Paediatr. 40–47, https://doi.org/10.1111/apa.14078 (2017).
Ferraris, L. et al. One-step multiplex PCR assay for differentiating proposed new species ‘clostridium neonatale’ from closely related species. J. Clin. Microbiol. 53, 3621–3623 (2015).
Schönherr-Hellec, S. et al. Comparative phenotypic analysis of “Clostridium neonatale” and Clostridium butyricum isolates from neonates. Anaerobe 48, 76–82 (2017).
Howard, F., Bradley, J., Flynn, D., Noone, P. & Szawatkowski, M. Outbreak of necrotising enterocolitis caused by Clostridium butyricum. Lancet 310, 1099–1102 (1977).
Gorham, P., Millar, M. & Godwin, P. G. R. Clostridial hand-carriage and neonatal necrotising enterocolitis. Journal of Hospital Infection 12, 139–141 (1988).
Cassir, N., Benamar, S. & La Scola, B. Clostridium butyricum: from beneficial to a new emerging pathogen. Clin. Microbiol. Infect. 22, 37–45 (2015).
Aureli, P. et al. Two cases of type e infant botulism caused by neurotoxigenic Clostridium butyricum in Italy. 54 (1986).
Wydau-dematteis, S., Louis, M. & Saubam, B. Anaerobe The functional dlt operon of Clostridium butyricum controls the D-alanylation of cell wall components and influences cell septation and vancomycin-induced lysis. Anaerobe 35, 105–114 (2015).
Peng, L., He, Z., Chen, W., Holzman, I. R. & Lin, J. Effects of butyrate on intestinal barrier function in a caco-2 cell monolayer model of intestinal barrier. Pediatr. Res. 61, 37–41 (2007).
Hsu, T., Hutto, D. L., Chris Minion, F., Zuerner, R. L. & Wannemuehler, M. J. Cloning of a beta-hemolysin gene of Brachyspira (Serpulina) hyodysenteriae and its expression in Escherichia coli. Infect. Immun. 69, 706–711 (2001).
Heida, F. H. et al. Identification of bacterial invasion in necrotizing enterocolitis specimens using fluorescent in situ hybridization. J. Perinatol. 1–6, https://doi.org/10.1038/jp.2016.165 (2016).
Scheifele, D. W. Role of bacterial toxins in neonatal necrotizing enterocolitis. J.Pediatr. 117, S44–S46 (1990).
Gohari, I. M. et al. A novel pore-forming toxin in type A Clostridium perfringens is associated with both fatal canine hemorrhagic gastroenteritis and fatal foal necrotizing enterocolitis. PLoS One 10 (2015).
Janoir, C. Virulence factors of Clostridium difficile and their role during infection. https://doi.org/10.1016/j.anaerobe.2015.10.009 (2016).
Curry, S. R. et al. Use of multilocus variable number of tandem repeats analysis genotyping to determine the role of asymptomatic carriers in Clostridium difficile transmission. Clin. Infect. Dis. 57, 1094–1102 (2013).
Eyre, D. W. et al. Diverse sources of C. difficile infection identified on whole-genome sequencing. N. Engl. J. Med. 369, 1195–205 (2013).
Price, J. R. et al. Transmission of Staphylococcus aureus between health-care workers, the environment, and patients in an intensive care unit: a longitudinal cohort study based on whole-genome sequencing. Lancet Infect. Dis. 17, 207–214 (2017).
Samore, M. H. Clinical and molecular epidemiology of sporadic and clustered cases of nosocomial Clostridium difficile diarrhea. Am. J. Med. 100, 32–40 (1996).
Marler, L. M. et al. Comparison of five cultural procedures for isolation of Clostridium difficile from stools. Journal of Clinical Microbiology 30, 514–516 (1992).
Hosny, M. et al. Description of Clostridium phoceensis sp. nov., a new species within the genus Clostridium. New Microbes New Infect. 14, 85–92 (2016).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Lagier, J. C. et al. Current and past strategies for bacterial culture in clinical microbiology. Clin. Microbiol. Rev. 28, 208–236 (2015).
Besemer, J. GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 29, 2607–2618 (2001).
Lechner, M. et al. Proteinortho: Detection of (Co-) orthologs in large-scale analysis. BMC Bioinformatics 12, 124 (2011).
Keane, J. A. et al. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genomics 2 (2016).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Stöver, B. C. & Müller, K. F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinformatics 11, 7 (2010).
Nascimento, M. et al. PHYLOViZ 2.0: Providing scalable data integration and visualization for multiple phylogenetic inference methods. Bioinformatics 33, 128–129 (2017).
Acknowledgements
We are grateful to the medical teams of the Assistance publique - Hôpitaux de Marseille (AP-HM), CHU Nîmes, CHU Montpellier and CHU Nice for providing us the stool samples. We thank the genomics platform at the Institut Hospitalo-Universitaire (IHU) - Méditerranée Infection for technical assistance. This work was supported by the French Government under the “Investissements d’avenir” program managed by the Agence Nationale de la Recherche (ANR), [reference: Méditerranée-Infection 10-IAHU-03], by Région Provence-Alpes-Côte d’Azur and European funding FEDER PRIMI. M. Hosny was supported by the Fondation de Coopération Scientifique, Méditerranée-Infection [Infectiopole Sud 2016].
Author information
Authors and Affiliations
Contributions
M.H. performed the microbiological and molecular biology analysis, M.H., A.C., A.L. and P.C. conducted the genomic analysis, M.H., R.A.A., N.C. and J.B.K. performed the statistical analysis, B.L. and M.H. wrote the paper and designed the study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hosny, M., Bou Khalil, J.Y., Caputo, A. et al. Multidisciplinary evaluation of Clostridium butyricum clonality isolated from preterm neonates with necrotizing enterocolitis in South France between 2009 and 2017. Sci Rep 9, 2077 (2019). https://doi.org/10.1038/s41598-019-38773-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-38773-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.