Abstract
Synthesis of ATP by the F1F0 ATP synthase in mitochondria and most bacteria is energized by the proton motive force (pmf) established and maintained by respiratory chain enzymes. Conversely, in the presence of ATP and in the absence of a pmf, the enzyme works as an ATP-driven proton pump. Here, we investigate how high concentrations of ATP affect the enzymatic activity of the F1F0 ATP synthase under high pmf conditions, which is the typical situation in mitochondria or growing bacteria. Using the ATP analogue adenosine 5′-O-(1-thiotriphosphate) (ATPαS), we have developed a modified luminescence-based assay to measure ATP synthesis in the presence of millimolar ATP concentrations, replacing an assay using radioactive nucleotides. In inverted membrane vesicles of E. coli, we found that under saturating pmf conditions, ATP synthesis was reduced to ~10% at 5 mM ATPαS. This reduction was reversed by ADP, but not Pi, indicating that the ATP/ADP ratio controls the ATP synthesis rate. Our data suggests that the ATP/ADP ratio ~30 in growing E. coli limits the ATP synthesis rate to ~20% of the maximal rate possible at the applied pmf and that the rate reduction occurs via product inhibition rather than an increased ATP hydrolysis rate.
Similar content being viewed by others
Introduction
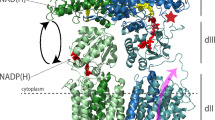
Life is energetically expensive and mostly paid in adenosine triphosphate (ATP). In all aerobic organisms, reducing equivalents from the oxidative breakdown of nutrients are converted to ATP in a process termed oxidative phosphorylation. Located in the cytoplasmic membrane of bacteria or the inner membrane of mitochondria, the members of the respiratory chain create and maintain a proton motive force (pmf) by proton coupled electron transfer reactions, e.g. from NADH to oxygen. The pmf, composed of a proton gradient (ΔpH) and a membrane potential (Δψ), energizes the rotary mechanism of the F1F0 ATP synthase that catalyzes the synthesis of ATP from its substrates ADP and Pi (for a review, see1,2,3,4). The F1F0 ATP synthase, which in most organisms can also work as an ATP driven proton pump, is composed of the membrane embedded F0 part and the soluble F1 part and can be divided into a stator and a rotor part. During ATP synthesis, when the pmf is high, protons are transported through the interface of subunit a and the c-ring converting the pmf into mechanical rotational energy. The c-ring is connected with subunits ε and γ forming the rotor, which transmits the rotary energy to the catalytic sites in the α3β3 headgroup. In the absence of a pmf, the enzyme can hydrolyze cellular ATP to establish an electrochemical gradient (the enzyme rotates in the opposite direction). This hydrolytic function of the F1F0 ATP synthase is vital for some bacteria under anaerobic conditions to maintain the membrane potential, when substrate level phosphorylation is the only source of ATP.
In order to minimize wasteful ATP hydrolysis, organisms have evolved different regulatory mechanisms, such as the inhibitory proteins IF1 in mitochondria5,6. A similar function has been attributed to the ζ subunit of the F1F0 ATP synthase of Paracoccus denitrificans, but is currently debated7. In other bacteria such as Bacillus PS3 and E. coli, the ε subunit has been proposed to ratchet the rotor in the synthesis direction8 by extending its C-terminal domain along the γ subunit9,10,11,12, a conformational change that might be controlled by ATP binding in some bacteria13. An alternative mechanism was suggested recently, in which the inhibitory part of ε works in a similar fashion as IF1 in mitochondria: by blocking both ATP synthesis and hydrolysis in absence of a pmf and being released upon establishment of the pmf 12,14. A further role has been attributed to the ε subunit in the coupling of F1 reactions and F0 proton pumping8,9,15,16,17. In addition to these mechanisms, all F1F0 ATP synthases show a decreased ATP hydrolysis rate in the presence of high ADP concentrations, a phenomenon known as the MgADP-inhibition of ATP hydrolysis18,19. During MgADP inhibition, the enzyme is in a locked state, from which it can be reverted by e.g. the pmf. In some bacteria like E. coli, MgADP inhibition is less pronounced and the threshold pmf for re-activation lowered20,21. Taken together, these mechanisms prohibit complete cellular depletion of ATP in the absence of a sufficient pmf.
In contrast to ATP hydrolysis that can be investigated in detergent solution or even with the F1 part only, ATP synthesis experiments require the presence of a proton tight membrane and a pmf. Quantitative ATP synthesis experiments have been performed with a variety of biological material such as intact mitochondria, submitochondrial particles, chloroplasts, bacteria and inverted bacterial membranes, and purified F1F0 ATP synthase reconstituted into liposomes22,23,24,25,26,27. Depending on the used system, the pmf was established by natural proton pumps23,28 or an acid-base treatment in combination with a potassium/valinomycin diffusion potential to mimic the membrane potential25,29,30. Different methods to quantify ATP have been applied, including the following four. (I) The most popular and sensitive technique is based on the luciferase/luciferin couple, which in the presence of oxygen and ATP emits light that can be detected by a luminometer (for a review, see31). The technique is very sensitive and is applied to determine the total ATP content from all types of samples. However, it cannot be used to measure the synthesis of newly formed ATP in the presence of physiological concentrations of background ATP (mM) which is ~1000-fold too high. Consequently, very little to no ATP is present in kinetic ATP synthesis measurements that are followed by this method. (II) The problem is circumvented by the second method, where radioactively labeled 32PO42− or 33PO42− is added to the solution. Newly synthesized ATP will incorporate radiolabeled phosphate, while the already present ATP is not radioactive. After separation of excess labeled phosphate by extraction, the newly formed ATP can be quantified via autoradiography or scintillation counting. The drawbacks are the use of radioactive substances and the laborious workup that prohibits continuous measurements. (III) The third method is based on the conversion of produced ATP by coupled enzymatic reactions that can be detected by spectroscopy, e.g. addition of D-glucose in the presence of hexokinase and subsequent dephosphorylation by glucose-6-phosphate dehydrogenase in the presence of NAD(P) that is reduced to NAD(P)H and can be followed spectroscopically. Similar to luciferase, this method cannot discriminate between present or freshly synthesized ATP and is less sensitive than the other methods. (IV) A further and more indirect technique originally developed by Chance and colleagues22 to quantify ATP synthesis is to measure the alkalization in the surrounding buffer solution during ATP synthesis. During ATP formation from ADP and Pi, a water molecule is formed that requires uptake of a proton from solution that can be measured either by a pH electrode or a pH sensitive dye such as phenol red. This approach is insensitive towards high concentrations of nucleotides and has successfully been used. However, it comes with limitations that have to be carefully addressed, such as low buffering capacity, limited pH range, and contribution of other proton dependent processes32.
In the present work, we describe a modification of the luciferase/luciferin technique that allows rapid continous measurement of the ATP synthesis rate with a high time resolution at physiological (millimolar range) concentrations of ATP. The principle is based on the properties of a modified ATP, adenosine-5′-O-(1-thiotriphosphate), or ATPαS, which is a hydrolytic substrate for the F1F0 ATP synthase, but is not detected by the luciferin/luciferase system. With the developed method, we were able to measure and compare the ATP synthesis in inverted E. coli membrane vesicles as a function of ADP, ATPαS, and inorganic phosphate (Pi) concentrations. We observe that physiological concentrations of ATP (5 mM), under the form of ATPαS, reduce the ATP synthesis rate to ~10%, and that this rate can be restored using elevated ADP concentrations, but not Pi, suggesting that the ATP synthesis is controlled by the ATP/ADP ratio and not by the ATP/(ADP∙Pi) ratio. The applicability of the technique to more isolated systems was confirmed in ATP synthesis experiments in liposomes containing purified bo3 oxidase and ATP synthase from E. coli.
Results
A modified luciferin/luciferase assay
The widely used reaction of D-luciferin with ATP, catalyzed by the luciferase from firefly, suffers from the drawback that it cannot discriminate between already present and freshly synthesized ATP. In addition, the high sensitivity of the assay impairs the use of ATP concentrations higher than a few micromolar, which is ~1000-fold less than what is found in vivo. The luciferase first catalyzes the binding of D-luciferin to the α-phosphate of ATP-Mg2+, forming PPi-Mg2+ and luciferyl-AMP, which is subsequently oxidized by oxygen to form AMP, CO2 and oxyluciferin in an excited state that emits a photon while returning to the ground state (Supplementary Fig. S1, for a review, see33,34). ATPases like the F1F0 ATP synthase, however, cleave ATP between the β- and the γ-phosphate, leaving ADP and Pi behind. We therefore reasoned that the use of an α-phosphate modified ATP prohibits the reaction with the luciferase but allows the reaction with ATPases. Indeed, commercially available ATPαS, in which the double-bonded oxygen on the α-phosphate is substituted for a sulfur, has been shown not to be a substrate for the luciferase in the deoxy form35, but is hydrolyzed by ATPases including the chloroplast F1F0 ATP synthase36 and the F1 ATPase from Bacillus PS337.
We tested this hypothesis experimentally by comparing the reaction of ATP and ATPαS with the firefly luciferase/luciferin supplied as a mixture in a commercial kit (CLS II, Roche) that is used by many to detect ATP synthesis. A typical amount of the assay was mixed with buffer and varying amounts of normal ATP or ATPαS were added to reach a final nucleotide concentration ranging from 10−12 M up to 5 × 10−3 M. The luminescence produced by the reaction of the luciferase reagent with ATP spanned from ~10° to ~108 arbitrary units (a.u.) between 1 pM and 5 mM whereas ATPαS yielded <104 a.u. at 5 mM (Fig. 1a), representing ~0.01% of the ATP signal response. The observed signal in the reaction with ATPαS is due either to reaction with ATPαS (with a high Km or low Vmax) or to contaminating ATP in the ATPαS preparation. We also found that ATPαS concentrations above 100 µM decreased the overall luciferase signal (<50% at 1 mM), probably due to competition of ATPαS with ATP for the luciferase binding site. Experimentally, this effect is addressed by adding a known concentration of ATP (2.5 µM in our experiments) to relate the arbitrary units of the luminescent signal to the concentration of ATP for the quantification of ATP production. Figure 1b shows that the luminescence signal was linear in the relevant ATP concentration range at all tested ATPαS concentrations.
The reaction of ATPαS vs. ATP with luciferin/luciferase. (a) Varying amounts of ATP (•) and (ATPαS,  ) were mixed with 2 µl luciferin/luciferase (prepared according to the manufacturer), filled up to 100 µl with buffer and measured in triplicate in a tube-type luminometer. (b) Dependency of the luminescence on ATP in the presence of increasing concentrations of ATPαS. The high concentrations of ATPαS used in our experiments quench luminescence to some extent and thus standardization by addition of a small known concentration of ATP is needed (as shown in Fig. 3b). Under all indicated ATPαS concentrations, addition of 0.5, 1, 2.5 and 5 µM ATP resulted in a linearly dependent luminescent signal. The measurements were done in triplicate and linear regressions are shown.
) were mixed with 2 µl luciferin/luciferase (prepared according to the manufacturer), filled up to 100 µl with buffer and measured in triplicate in a tube-type luminometer. (b) Dependency of the luminescence on ATP in the presence of increasing concentrations of ATPαS. The high concentrations of ATPαS used in our experiments quench luminescence to some extent and thus standardization by addition of a small known concentration of ATP is needed (as shown in Fig. 3b). Under all indicated ATPαS concentrations, addition of 0.5, 1, 2.5 and 5 µM ATP resulted in a linearly dependent luminescent signal. The measurements were done in triplicate and linear regressions are shown.
ATPαS is a substrate for the E. coli F1F0 ATP synthase
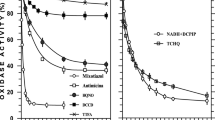
ATP and ATPαS were compared as substrates for hydrolysis by the E. coli F1F0 ATP synthase in detergent solution (Fig. 2a). Using different concentrations of either nucleotide, the ATP hydrolysis rates were determined using phenol red to detect proton consumption during ATP hydrolysis38. The measured ATPαS hydrolysis rate (apparent Km 226 µM, ~21 ATP s−1 enzyme−1 at 1 mM) was 84% of that of ATP at the maximum tested substrate concentration (apparent Km 280 µM, ~25 ATP s−1 enzyme−1 at 1 mM). We also compared the ability of ATP and ATPαS to elicit transmembrane proton pumping, i.e. establishment of a pmf, with ACMA quenching in inverted membrane vesicles (IMV) of E. coli cells (Fig. 2b,c). Both substrates energized proton pumping effectively, indicating that ATPαS hydrolysis by the E. coli F1F0 ATP synthase does not interfere with the rotational mechanism of the enzyme. The lower apparent Km values for proton pumping could be attributed to the different type of preparation (purified enzyme vs. inverted membrane vesicles). Proton pumping was accelerated ~2.5-fold in the absence of a membrane potential (1 µM valinomycin added, Supplementary Fig. S2), indicating that the IMVs were reasonably tight. A lower maximum (~75%) initial quenching rate and total quenching was observed with ATPαS compared to ATP. Both findings can be explained by the decreased ATP hydrolysis rate described above, as the ACMA quench signal is dependent on both the rate of proton pumping and the rate of spontaneous proton leakage through the membrane.
The reaction of ATPαS with the E. coli F1F0 ATP synthase. (a) Varying amounts of ATP (•) and (ATPαS,  ) were incubated with purified E. coli F1F0 ATP synthase and ATP hydrolysis was measured using a phenol red assay. Shown are average values and standard deviation from three measurements. (b) ATP and ATPαS hydrolysis-driven proton pumping by E. coli F1F0 ATP synthase in inverted membrane vesicles was followed using ACMA fluorescence quenching. Shown are typical traces, here for 1 µM ATP (black), 250 µM ATPαS (red) and 250 µM ATP (green). (c) Experiment like b, but varying amounts of ATP (•) and ATPαS(
) were incubated with purified E. coli F1F0 ATP synthase and ATP hydrolysis was measured using a phenol red assay. Shown are average values and standard deviation from three measurements. (b) ATP and ATPαS hydrolysis-driven proton pumping by E. coli F1F0 ATP synthase in inverted membrane vesicles was followed using ACMA fluorescence quenching. Shown are typical traces, here for 1 µM ATP (black), 250 µM ATPαS (red) and 250 µM ATP (green). (c) Experiment like b, but varying amounts of ATP (•) and ATPαS( ) were used, taking into account the initial ACMA quenching (within 5 sec) of the different measurements. Shown are average values and standard deviation from three measurements.
) were used, taking into account the initial ACMA quenching (within 5 sec) of the different measurements. Shown are average values and standard deviation from three measurements.
ATP synthesis kinetics of inverted E. coli vesicles at physiological ATP concentrations
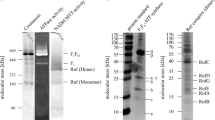
In this work, we applied the novel method described above to follow ATP synthesis in the presence of high ATP concentration and high pmf, reflecting the physiological conditions of growing bacteria. As a model system, we used inverted membrane vesicles (IMVs) of E. coli, produced by a French Press39. In contrast to mitochondria, E. coli membranes do not contain nucleotide and phosphate carriers that control the availability and distribution of their substrates in the mitochondrial matrix. In IMVs, the orientation of all proteins is homogenously inverted40, exposing the F1 part of the F1F0 ATP synthase to the buffer solution, allowing to readily manipulate the relevant substrates ATP, ADP and Pi. For prospective studies investigating mutant variants, we used an E. coli F1F0 ATP synthase knockout strain (DK8), transformed with the plasmid pBWU13, constitutively expressing F1F0 ATP synthase. This combination has been shown to yield F1F0 ATP synthase overexpression when grown in minimal media using glycerol as carbon source41. If grown in rich media, however, F1F0 ATP synthase production levels were comparable to a strain BL21 with chromosomally encoded F1F0 ATP synthase, as judged by Western blot (Supplementary Fig. S3). As complex I is expressed predominantly under anaerobic conditions42, we have used dithiothreitol (DTT) and ubiquinone Q1 as synthetic electron donor and acceptor, respectively, to energize the quinol oxidase bo3 as the primary proton pump to establish a maximal pmf (Fig. 3a). The required amount of ubiquinone Q1 (≥20 µM) to obtain maximal ATP synthesis was titrated (Supplementary. Fig. S4) and in all measurements, a ubiquinone Q1 concentration of 80 µM was used to ensure sufficient pmf for maximal ATP synthesis. Figure 3b shows a typical continuous ATP synthesis measurement in IMV of E. coli as described26,43, with almost no free ATP present (a defined concentration of ATP was added prior to measurement to quantify the signal), that was started by the addition of 80 µM ubiquinone Q1. ATP synthesis was immediately observed as a linear increase of the luminescence signal and was followed for at least 30 s. Addition of 400 nM potassium cyanide, a potent inhibitor of the bo3 oxidase, rapidly blocks ATP generation. Similarly, <10% ATP synthesis was observed in the presence of the proton ionophore CCCP (10 µM). Using 2 mM DTT and 80 µM ubiquinone Q1 as electron donor, the ATP synthesis rate was about 1.5 to 2-fold higher compared to energization with 500 µM NADH, both in the DK8/pBWU13 and BL21 strains (Supplementary Table 1).
Respiratory chain driven ATP synthesis in inverted membranes of E. coli. (a) Cartoon illustrating the experimental setup. Inverted membrane vesicles of E. coli were energized using DTT and ubiquinone Q1 to initiate inward proton pumping of the quinol bo3 oxidase, generating a pmf (see Supplementary Fig. S4). ATP synthesis was catalyzed by the F1F0 ATP synthase with its F1 head directed toward to the outside and detected using a luminescent based assay. (b) Raw data from continuous measurement of ATP synthesis in IMVs using luminescence in the presence of 50 μM ADP and 10 mM Pi. Proton pumping by the bo3 oxidase was supported by DTT as electron donor and ubiquinone Q1 (UQ1) as electron mediator. Addition of the cytochrome oxidase inhibitor potassium cyanide (KCN) stops ATP synthesis. The arrows indicate the different point of additions. From such data, the slope of the linear range and the ATP standard addition were used to calculate ATP synthesis rates in the following figures. Luminescence values were for periods of 30 s, allowing addition of reagents when needed (indicated by short gaps in the trace). (c) Dependency of ATP synthesis rate from the Pi concentration in the presence of 50 μM ADP and in the absence of ATP. (d) Dependency of ATP synthesis rate from the ADP concentration in the presence of 10 mM Pi and in the absence of ATP. (e) Dependency of the ATP synthesis rate from the ATPαS concentration in the presence of 50 μM ADP and 10 mM Pi. The ATP/ADP ratio is indicated in red.
In a first set of experiments, we characterized our experimental system by varying concentrations of inorganic phosphate (Fig. 3c) and ADP (Fig. 3d). Apparent Km values for ADP and phosphate were 11 µM and 104 µM, respectively. Under fully saturating conditions using 50 μM ADP and 10 mM phosphate, a maximal ATP synthesis rate of ~250 ± 50 nmol ATP min−1 mg−1 was obtained at 25 °C, which is in good accordance to a recent report for E. coli IMVs7. At maximal tested ADP (5 mM) and Pi (100 mM) concentrations, the ATP synthesis rate declined to ~70%.
Next, we tested the influence of increasing concentrations of ATPαS on the rate of ATP synthesis in the presence of a constantly high pmf. Keeping ADP (50 µM) and phosphate (10 mM) constant, we varied the ATPαS concentration between 1 µM and 5 mM, resulting in a distinct decrease in the synthesis rate. As depicted in Fig. 3e, ATP synthesis was maximal below <25 µM ATPαS (100%) but started declining beyond 50 μM ATPαS, reaching a residual rate of 10% to 20% in the physiologically relevant concentration range of 2 to 5 mM.
To check whether the decrease in rate was due to the absolute ATPαS concentration or not, we investigated how the initial apparent mass-action ratio for the ATP synthesis reaction Γ’ = [ATP]/([ADP]∙[Pi]) influences the ATP synthesis rate. At chemical equilibrium, where no net ATP synthesis takes place the mass-action ratio is equal to the equilibrium constant K44. In equilibrium thermodynamics, the Gibbs free energy change is defined by the equation
where R is the gas constant and T the temperature. While thermodynamic considerations cannot be used to predict kinetic rates, the data of Fig. 3e show that varying the initial Γ’ (100 M−1 at 50 µM ATPαS and 104 M−1 at 5 mM ATPαS) directly affect ATP synthesis rates. It was thus interesting to observe whether the effect caused by increased ATPαS concentrations could be reverted by increasing ADP or phosphate simultaneously, thus leaving Γ’ constant (Γ’ = 100 M−1). First, we tested this by increasing both ADP and ATPαS concentrations in parallel at a fixed Pi concentration (10 mM) to keep Γ’ = 100 M−1 and ATPαS/ADP = 1 constant (Fig. 4a). If ATPαS and ADP were below 50 µM, non-maximal activity was observed due to non-saturating ADP levels (compare to Fig. 3d). Maximal ATP synthesis rate was observed between 50 and 200 μM nucleotides (100%, 196 ± 33 nmol ATP min−1 mg−1), and slowly decreased to 60% (116 ± 18 nmol ATP min−1 mg−1) at 5 mM ATPαS and ADP. Considering the decrease of activity by 5 mM ADP itself (75%, see Fig. 3d), ATP synthesis was recovered to ~85%, demonstrating that increasing ADP concentrations can revert the effect observed above. In a second scenario, we adjusted ATP and Pi in parallel in the presence of a fixed ADP concentration (50 µM), to keep Γ’ = 100 M−1 and ATP/Pi = 0.005 constant. The data from Fig. 4b show that in contrast to ADP, inorganic phosphate cannot reverse the inhibitory effect of ATPαS. The curve decreases more than the one seen in Fig. 3e, declining from a maximum of 185 ± 8 nmol ATP min−1 mg−1 at 50 μM ATPαS (ATP/ADP = 1) to 37 ± 3 nmol ATP min−1 mg−1 (20 ± 3% of maximum observed rate) at 1 mM ATPαS (ATP/ADP = 20). This pronounced decrease is probably due to the inhibitory effect observed at high phosphate concentration (Fig. 3c).
Dependency of ATP synthesis and hydrolysis on nucleotide concentration. The values of ATP synthesis rate at increasing adenosine 5′-O-(1-thiotriphosphate) (ATPαS) and ΔG (constant 50 μM ADP and 10 mM Pi, Fig. 3e) are indicated as  for comparison. (a) An equimolar concentration of ADP was used to keep a constant ATP/ADP ratio of 1 and a ΔG of 40 kJ/mol, in the presence of 10 mM Pi. (b) Like a, but a constant ATPαS/Pi ratio of 0.005 was used to keep constant a ΔG of 40 kJ/mol (Pi concentrations are indicated in millimolar below each circle), in the presence of 50 μM ADP. (c) ATP hydrolysis as a function of the ADP concentration, with an ATP concentration of 50 μM
for comparison. (a) An equimolar concentration of ADP was used to keep a constant ATP/ADP ratio of 1 and a ΔG of 40 kJ/mol, in the presence of 10 mM Pi. (b) Like a, but a constant ATPαS/Pi ratio of 0.005 was used to keep constant a ΔG of 40 kJ/mol (Pi concentrations are indicated in millimolar below each circle), in the presence of 50 μM ADP. (c) ATP hydrolysis as a function of the ADP concentration, with an ATP concentration of 50 μM  or 2.5 mM (•). The activity was followed by measuring the initial ACMA fluorescence quenching (5 sec) during proton pumping.
or 2.5 mM (•). The activity was followed by measuring the initial ACMA fluorescence quenching (5 sec) during proton pumping.
We have repeated the crucial experiments also with proteoliposomes containing F1F0 ATP synthase and a proton pump prepared as described28. To this end, the purified E. coli F1F0 ATP synthase and bo3 oxidase were co-reconstituted into liposomes made of 4:1 DOPC:DOPG (w/w). Proton pumping was initiated with DTT/Q1 and ATP synthesis was followed as described for the inverted membrane vesicle. First, ATP synthesis rates in the presence of 50 µM ADP, and either 10 µM or 2 mM ATPαS were measured. As depicted in Supplementary Fig. S5, ATP synthase rate dropped to ~6% in the presence of 2 mM ATPαS, making the inhibitory effect even stronger than with IMVs (~18%) at the same ATPαS concentration. In the presence of 2 mM ADP, the activity was recovered to ~50% (75% in the IMVs). Taken together, the measurements support our findings with the membrane vesicles and reinforce the notion that the effect is directly connected to the F1F0 ATP synthase.
Discussion
ATPαS can be used as a substrate analogue for ATP hydrolysis in the E. coli F1F0 ATP synthase
In order to allow continuous detection of ATP synthesis by a luciferase/luciferin assay in the presence of high ATP concentrations, we have used a modified ATP analogue that is not detected by the luciferase system but is a suitable substrate for ATP hydrolysis with similar kinetic properties as ATP. We show that the sulfur-substituted alpha phosphate indeed barely reacts with luciferin. The observed signal increase upon rising ATPαS concentration (~104 times lower than ATP) is due either to a very slow reaction of ATPαS with luciferin or to slight ATP contamination in the ATPαS preparation. ATPαS binding to luciferase/luciferin as a competitive inhibitor is likely, as the overall signal for a constant amount of ATP is decreasing with increasing concentrations of ATPαS. Technically, this effect was compensated by the addition of a precisely known, suitable amount of ATP for every measurement to quantify the luminescence signal. In contrast to its reaction with luciferin/luciferase, ATPαS behaved similarly to ATP when used to drive ATP hydrolysis with the E. coli F1F0 ATP synthase. As with the F1F0 enzyme in chloroplasts and the isolated F1 part of the ATP synthase of Bacillus PS3, ATP hydrolysis and proton pumping kinetics with the E. coli enzyme were comparable. In the reaction with ATPαS, Vmax was ~75% of the one found for ATP and both had a similar Km’ of ~200–300 µM in the absence of ADP. Earlier, Löbau et al. reported an apparent Km(MgATP) = 140 µM for E. coli F1F0 ATP synthase45. Data for ATPαS with intact F1F0 ATP synthase is missing, but in elegant single-molecule experiments following the rotation of the gamma subunit, Noji and coworkers have found an apparent Km (91 µM) with the purified F1 ATPase from Bacillus PS3 lacking the regulatory epsilon subunit37. In comparison to published values, the Km values reported here were ~2-fold higher, which is likely due to different assay conditions (e.g. buffer, Mg2+ concentration) and different enzyme preparations from different organisms. However, given the similar behavior within our preparation, it was concluded that ATPαS can be used to replace background ATP in ATP synthesis experiments. In the past, ATP synthesis experiments in the presence of high ATP were performed following the incorporation of radioactive 32PO42- or 33PO42- into newly formed ATP.
The ATP synthesis rate is strongly reduced by an increasing ATP/ADP ratio
The physiological nucleotide concentration ratio (ATP/ADP ~20–1000) strongly favors ATP hydrolysis (ATP/ADP at equilibrium is ~10−5). However, the presence of respiratory proton pumps (e.g. bo3 oxidase in E. coli) generating and maintaining a pmf are able to shift the net reaction towards ATP synthesis. Typical in vitro ATP synthesis studies nevertheless employ conditions where no ATP is present due to the incompatibility with the luciferase assay, while ADP is used in excess to maximize the observed rates. In this report, we describe the use of ATPαS to overcome this limitation and have investigated the effect of cellular concentrations of ATPαS (as a substrate analogue of ATP) on the enzymatic reaction of the E. coli F1F0 ATP synthase directly. Similar measurements have been performed in intact mitochondria, but the interpretation of results is complicated by the presence of pmf driven phosphate and nucleotide carriers from the SLC25 family that transport these substrates from the bulk to the site of ATP synthesis in the mitochondrial matrix. Consequently, the different substrates added to the outside affect the F1F0 ATP synthase only indirectly. By contrast, no such transporters are present in E. coli and the complete inversion of protein orientation in IMVs allows direct access of the experimentally added substrates ATP, ADP and Pi to the catalytic sites in the F1 part of the ATP synthase.
The influence of the cytoplasmic ATP/ADP ratio on the ATP synthesis rate had been investigated with intact mitochondria by e.g. Küster et al.24 and LaNoue et al.46. Küster et al. report that the extra-mitochondrial ATP/ADP ratio controls the ATP synthesis rate, dropping steeply after a ratio of ~5 with essentially no ATP synthesis at a ratio of 100. LaNoue et al. report that ATP synthesis and ATP hydrolysis are only in equilibrium at low pmf, while at high pmf values, the increased net ATP synthesis is due to nearly blocked ATP hydrolysis rather than an increased ATP synthesis rate. Both studies confirm a kinetic control of the ATP synthesis rate with the ATP/ADP translocase being the controlling module46. Omitting the added complexity of the ATP/ADP translocase, Ferguson and colleagues investigated cells and membranes of the gram negative bacterium Paracoccus denitrificans47. In their studies, they find that membrane vesicles of P. denitrificans are not capable of ATP hydrolysis, neither under high or low pmf conditions, and conclude that the kinetic control of ATP synthesis is an intrinsic property of the F1F0 ATP synthase. There is an ongoing discussion to determine if the backwards reaction in P. denitrificans is blocked by the ζ subunit or not6,7,48, or if the inhibitory effect is caused by Mg-ADP inhibition, as proposed earlier49.
With the present study, we can add information to the situation in E. coli, which represents a mixed situation. Like P. denitrificans, no ATP/ADP translocase is present, allowing a more direct investigation of the properties of the F1F0 ATP synthase. However, in contrast to P. denitrificans, membranes of E. coli show no respiratory coupling and the F1F0 ATP synthase is fully active in ATP hydrolysis direction32,50,51. In our measurements with a constant amount of ADP and phosphate, an increasing ATPαS concentration (and thus ATP/ADP ratio) provokes a sharp decrease of ATP synthesis over an ATP/ADP ratio of 1, with 50% activity at ratio of ~20 (1 mM ATPαS) and <10% activity at a ratio of 100 (5 mM ATPαS), which is in good agreement with the data from mitochondria. As the pmf in our experiments is unchanged (no notable effect of increasing nucleotide concentrations on respiratory activity were found), the data indicates that the observed effect is either due to product inhibition by the presence of ATPαS or by an increased rate of ATP hydrolysis. Since our assay directly detects newly formed ATP with a very high time resolution and an immediate decrease in ATP synthesis rate is observed, we can confirm that a direct competition of ATPαS and ADP indeed takes place. We cannot rule out an increased ATP hydrolysis rate, but the effective ATPαS concentrations (>1 mM) are well above the apparent Km of ATP for ATP hydrolysis, which however could be increased in the presence of a pmf. In similar experiments, where ADP was raised in parallel to keep constant the ATPαS/ADP ratio and mass-action ratio (Γ’ = 100 M−1), the inhibitory effect of ATPαS was only mildly observed, suggesting that the competition of ADP and ATPαS indeed control the rate of ATP synthesis. A more detailed interpretation of these results is complicated by the presence of three catalytic sites that have different affinities for ATP and ADP depending on their conformation state (tight, loose, and empty)52. Finally, by varying the ATP and Pi concentrations in parallel, resulting in the same Γ’ = 100 M−1, but increasing the ATP/ADP ratio, we found that a variation of the phosphate concentration could not prevent the inhibitory effect, indicating that ATP synthesis is not controlled by the phosphorylation potential variable [ATP]/([ADP]∙[Pi]), but rather by the ATP/ADP ratio. The inhibitory effect of ATPαS at higher phosphate concentrations was even more pronounced (<5% residual activity at 5 mM ATPαS), which can be attributed to a slight inhibitory effect of Pi concentrations >100 mM.
In respiring E. coli, the ATP and ADP concentrations have been found to be ~3.5 mM and 0.12 mM, respectively, yielding an ATP/ADP ratio of ~3053. From our measurements, we can thus assume that despite saturating pmf, the E. coli synthesizes ATP with a rate of only ~20% of vmax. Etzold et al. determined a maximal turnover rate of 270 ATP s−1 per ATP synthase in E. coli at 37 °C54. However, in their measurements, 95% of the ATP synthases were blocked to maximize the available driving force per F1F0 ATP synthase. Furthermore, their experiments were performed in the presence of 1 mM ADP and the total absence of ATP (ATP/ADP = 0). It can thus be estimated that the ATP synthesis rates is only 5 to 10 ATP s−1 per F1F0 ATP synthase in growing E. coli. The observed necessary concentrations of ADP >1 mM to restore maximal ATP synthesis are not physiologically relevant. However, the nearly universal ATP hydrolysis inhibition mechanism by Mg-ADP52,55,56,57,58,59 could be responsible for the apparent absence of ATP hydrolysis in our experiments. We have thus measured the inhibitory effect of ADP on proton pumping on the same vesicles as used for ATP synthesis. The data from Fig. 4c show that the apparent Ki of ADP increases from ~100 µM to ~400 µM in the presence of 50 µM ATP or 2.5 mM ATP, respectively. An even higher Ki is expected during ATP synthesis under high pmf, that has been shown to counteract ADP inhibition of ATP hydrolysis19,57, and hence being considerably higher compared to the ADP concentrations found in growing cells (~120 µM)53. ADP concentrations >400 µM, however will be easily reached when the pmf is low and ATP is converted to ADP, and the mechanism thus serves as effective protection against cellular depletion of ATP. Finally, our results seem to be independent of the reported inhibition of ATP hydrolysis by the ε subunit, as for the E. coli enzyme it has been shown to be absent in the presence of a pmf and independent of the ATP concentration14. In the future, it will be interesting to investigate, if ATP synthesis in other organisms than E. coli is similarly dependent on the ATP concentration.
Concluding Remarks
In this work, we have introduced a novel, alternative, non-radioactive method to measure ATP synthesis in vitro under physiological nucleotide concentrations and high pmf, using ATPαS as a replacement for ATP. The method allows to selectively measure newly produced ATP and is unaffected by the presence of high concentrations of ATPαS. The luciferase setup allows rapid and continuous measurements, avoiding the necessity of taking samples at distinct time points, thus considerably increasing time resolution in measurements.
We have applied this method to inverted membrane vesicles of E. coli and proteoliposomes containing F1F0 ATP synthase and bo3 oxidase and find that the ATP/ADP ratio, but not ATP/(ADP∙Pi), controls the rate of ATP synthesis. The effect is based on a direct competition between of ATPαS and ADP for the catalytic binding sites. From this, we can conclude that the physiological ATP concentration prevents a maximal ATP synthesis rate despite saturating pmf conditions. Finally, our results suggest that Mg-ADP inhibition of ATP hydrolysis is not relevant to maximize ATP synthesis rates in E. coli, but is an effective tool to prevent total loss of cellular ATP during energy starvation.
Material and Methods
Chemicals and kits were purchased from Sigma-Aldrich and Thermo Scientific if not otherwise stated.
Growth conditions of bacterial cells
E. coli strain DK8, lacking the unc operon coding for its ATP synthase, was transformed with a pBWU13 plasmid and grown at 37 °C in LB-Lennox medium containing 50 μg/ml ampicillin.
Preparation of Escherichia coli DK8:pBWU13 inverted membrane vesicles
The following procedure was performed at 4 °C. Bacterial cells were suspended in 10 ml/g HMG buffer (20 mM Hepes pH 7.5, 2 mM MgCl2, and 10% glycerol), supplemented with a tiny spatula tip of Pefabloc serine protease inhibitor, phenylmethanesulfonyl fluoride and DNAse I, and broken with a French pressure cell press or a Microfluidizer. Unbroken cells were collected by centrifugation at 8000 g for 10 min. The supernatant containing the membrane vesicles was centrifuged at high speed (200′000 g) for 1 h, resuspended in 5 ml HMG supplemented with 0.5% cholate per gram of initial cells, to remove proteins loosely associated with the membrane, and centrifuged again at high speed. The membrane vesicles were resuspended in HMG and centrifuged a third time at high speed to remove traces of cholate. They were then resuspended in HMG to a concentration of 10 mg/ml total proteins (quantified with a bicinchoninic acid assay), aliquoted and flash-frozen with liquid nitrogen for storage at −80 °C.
Purification of F1F0 ATP synthase from E. coli
The F1F0 ATP synthase of E. coli strain DK8:pBWU13 was purified as previously described26.
ATP hydrolysis assay by ACMA quenching
With this method we indirectly measure the hydrolytic activity of the F1F0-ATPase by following the establishment of the resulting proton gradient, using the self-quenching property of the fluorescent dye 9-Amino-6-Chloro-2-Methoxyacridine (ACMA). The fluorescence of this membrane permeable dye quenches when a proton gradient forms across a lipid compartment60.
The fluorescence of the dye 9-Amino-6-Chloro-2-Methoxyacridine (ACMA) (excitation wavelength: 410 nm; emission wavelength: 480 nm) was monitored with a Cary Eclipse Fluorescence Spectrophotometer from Agilent Technologies. A cuvette with stirring magnet was filled with HMK buffer (10 mM Hepes-KOH pH 7.5, 1 mM MgCl2, 10 mM KNO3), 150 μg/ml inverted bacterial membrane vesicles and 2 μM ACMA in a final volume of 1 ml. The fluorescence baseline, taken as 100% fluorescence to determine the percentage of quenching, was measured and hydrolysis was started by the addition of 1–500 μM ATP or ATPαS. The pH gradient established by the ATP synthase quenched the ACMA dye proportionally to the hydrolytic activity and was eventually dissipated by the addition of 20 mM NH4Cl.
To assess the effect of valinomycin and the coupling of IMVs, a buffer containing 10 mM MOPS-KOH pH 7.5, 100 mM KCl, and 2.5 mM MgCl2 was used instead of HMK. The measurement was carried as above, with or without 1 μM valinomycin, and started with 500 μM ATP.
ATP hydrolysis assay by phenol red spectrophotometry
Phenol red is a pH-sensitive dye which can be used to detect the dissociation of protons in the ATP hydrolysis reaction38, at pH > 7.2: ATP4− + H2O ⇌ ADP3− + Pi2− + H+ (see for example61).
Phenol red buffer (0.5 mM Hepes-KOH pH 8, 100 mM KCl, 4 mM MgCl2, 0.1 mM EDTA, 10% glycerol, 0.02% triton X-100) was mixed with 25 μM phenol red and desired concentrations of ADP, ATP and ATPαS, to a final volume of 1 ml. The pH was adjusted to 8 with KOH and the baseline absorbance was recorded at 557 nm with a Cary 60 UV-Vis spectrophotometer. 20 nmol HCl were used for standardization and 5 μg E. coli purified F1F0 ATP synthase were added to start the reaction.
Luciferin/luciferase reaction with ATP versus ATPαS
The firefly enzyme luciferase drives an ATP-dependent reaction in two steps to produce light that is proportional to the amount of ATP present and can be measured with a luminometer.
To compare the reactivity of ATP versus ATPαS with the luciferin/luciferase system, 200 μg/ml luciferase reagent (containing luciferin, luciferase and magnesium) of an ATP Bioluminescence Assay Kit CLS II from Roche Applied Science was added to 50 mM Hepes-NaOH pH 7.5, 10 mM total phosphate and 3 mM MgCl2, and the baseline was measured. 1 μM to 5 mM ATP or ATPαS was then added and the luminescence measured.
Luciferin/luciferase ATP synthesis assay
The assay was performed in clear 1.5 ml Eppendorf tubes in a final volume of 100 μl. 50 mM Hepes-NaOH pH 7.5 were used as buffer and the assay mixture contained 10 mM MgCl2, 200 μg/ml luciferase reagent, 40 μg/ml inverted bacterial membrane vesicles, 2 mM (saturating) DTT and variable concentrations of ATPαS, ADP (typically 50–100 μM) and total phosphate (typically 10 mM). The luminescence was monitored with a Promega GloMax luminometer in steps of 30 s, allowing additions of relevant reagents to the tube when required.
The baseline was first measured (30 s), before 250 pmol ATP were added as a calibration standard (30 s). Ubiquinone Q1, an ethanol soluble analogue of ubiquinone that can be reduced by DTT, was used to energize the membrane by bo3 oxidase proton pumping and start the ATP synthesis (3 × 30 s). ATP synthesis could be interrupted by the addition of 400 nM potassium cyanide or 10 μM CCCP (2 × 30 s).
References
von Ballmoos, C., Wiedenmann, A. & Dimroth, P. Essentials for ATP synthesis by F1F0 ATP synthases. Annu Rev Biochem 78, 649–72 (2009).
Walker, J. E. The ATP synthase: the understood, the uncertain and the unknown. Biochem Soc Trans 41(1), 1–16 (2013).
Igamberdiev, A. U. & Kleczkowski, L. A. Optimization of ATP synthase function in mitochondria and chloroplasts via the adenylate kinase equilibrium. Frontiers in Plant Science 6, 10 (2015).
Junge, W. & Nelson, N. ATP Synthase. Annual Review of Biochemistry 84(1), 631–657 (2015).
Pullman, M. E. & Monroy, G. C. A Naturally Occurring Inhibitor of Mitochondrial Adenosine Triphosphatase. J Biol Chem 238, 3762–9 (1963).
Garcia-Bermudez, J. & Cuezva, J. M. The ATPase Inhibitory Factor 1 (IF1): A master regulator of energy metabolism and of cell survival. Biochim Biophys Acta 1857(8), 1167–1182 (2016).
Varghese, F. et al. Deleting the IF(1)-like ζ subunit from Paracoccus denitrificans ATP synthase is not sufficient to activate ATP hydrolysis. Open Biology 8(1), 170206 (2018).
Klionsky, D. J., Brusilow, W. S. & Simoni, R. D. In vivo evidence for the role of the epsilon subunit as an inhibitor of the proton-translocating ATPase of Escherichia coli. Journal of Bacteriology 160(3), 1055–1060 (1984).
Feniouk, B. A., Suzuki, T. & Yoshida, M. The role of subunit epsilon in the catalysis and regulation of FOF1-ATP synthase. Biochim Biophys Acta 1757(5–6), 326–38 (2006).
Cingolani, G. & Duncan, T. M. Structure of the ATP synthase catalytic complex (F(1)) from Escherichia coli in an autoinhibited conformation. Nat Struct Mol Biol 18(6), 701–7 (2011).
Shirakihara, Y. et al. Structure of a thermophilic F1-ATPase inhibited by an epsilon-subunit: deeper insight into the epsilon-inhibition mechanism. Febs j 282(15), 2895–913 (2015).
Shah, N. B. & Duncan, T. M. Aerobic Growth of Escherichia coli Is Reduced, and ATP Synthesis Is Selectively Inhibited when Five C-terminal Residues Are Deleted from the Subunit of ATP Synthase. J Biol Chem 290(34), 21032–41 (2015).
Kato-Yamada, Y. & Yoshida, M. Isolated epsilon subunit of thermophilic F1-ATPase binds ATP. J Biol Chem 278(38), 36013–6 (2003).
Sielaff, H., Duncan, T. M. & Borsch, M. The regulatory subunit epsilon in Escherichia coli FOF1-ATP synthase. Biochim Biophys Acta Bioenerg 1859(9), 775–788 (2018).
Turina, P. et al. Physiological ligands ADP and Pi modulate the degree of intrinsic coupling in the ATP synthase of the photosynthetic bacterium Rhodobacter capsulatus. Biochemistry 43(34), 11126–34 (2004).
Cipriano, D. J. & Dunn, S. D. The Role of the ϵ Subunit in the Escherichia coli ATP Synthase: The C-Terminal Domain is Required for Efficient Energy Coupling. Journal of Biological Chemistry 281(1), 501–507 (2006).
D’Alessandro, M. et al. Modulation of coupling in the Escherichia coli ATP synthase by ADP and Pi: Role of the epsilon subunit C-terminal domain. Biochim Biophys Acta 1858(1), 34–44 (2017).
Hirono-Hara, Y. et al. Pause and rotation of F1-ATPase during catalysis. Proceedings of the National Academy of Sciences 98(24), 13649–13654 (2001).
Feniouk, B. A., Suzuki, T. & Yoshida, M. Regulatory interplay between proton motive force, ADP, phosphate, and subunit epsilon in bacterial ATP synthase. J Biol Chem 282(1), 764–72 (2007).
Senior, A. E. et al. Catalytic properties of Escherichia coli F1-ATPase depleted of endogenous nucleotides. Archives of Biochemistry and Biophysics 297(2), 340–344 (1992).
Lapashina, A. S. & Feniouk, B. A. ADP-Inhibition of H+-FOF1-ATP Synthase. Biochemistry (Mosc) 83(10), 1141–1160 (2018).
Nishimura, M., Ito, T. & Chance, B. Studies on bacterial photophosphorylation. III. A sensitive and rapid method of determination of photophosphorylation. Biochim Biophys Acta 59, 177–82 (1962).
Racker, E. & Stoeckenius, W. Reconstitution of purple membrane vesicles catalyzing light-driven proton uptake and adenosine triphosphate formation. J Biol Chem 249(2), 662–3 (1974).
Kuster, U., Bohnensack, R. & Kunz, W. Control of oxidative phosphorylation by the extra-mitochondrial ATP/ADP ratio. Biochim Biophys Acta 440(2), 391–402 (1976).
Fischer, S. & Gräber, P. Comparison of Delta pH- and Delta phi-driven ATP synthesis catalyzed by the H(+)-ATPases from Escherichia coli or chloroplasts reconstituted into liposomes. FEBS Lett 457(3), 327–32 (1999).
Wiedenmann, A., Dimroth, P. & von Ballmoos, C. Functional asymmetry of the F(0) motor in bacterial ATP synthases. Mol Microbiol 72(2), 479–90 (2009).
Nilsson, T. et al. Lipid-mediated Protein-protein Interactions Modulate Respiration-driven ATP Synthesis. Sci Rep 6, 24113 (2016).
von Ballmoos, C. et al. Mimicking respiratory phosphorylation using purified enzymes. Biochim Biophys Acta 1857(4), 321–31 (2016).
Jagendorf, A. T. & Uribe, E. ATP formation caused by acid-base transition of spinach chloroplasts. Proc Natl Acad Sci USA 55(1), 170–7 (1966).
Petersen, J. et al. Comparison of the H+/ATP ratios of the H+-ATP synthases from yeast and from chloroplast. Proc Natl Acad Sci USA 109(28), 11150–5 (2012).
Lundin, A. Optimization of the firefly luciferase reaction for analytical purposes. Adv Biochem Eng Biotechnol 145, 31–62 (2014).
Zharova, T. V. & Vinogradov, A. D. ATPase/synthase activity of Paracoccus denitrificans Fo.F1 as related to the respiratory control phenomenon. Biochim Biophys Acta 1837(8), 1322–9 (2014).
Marques, S. M. & Esteves da Silva, J. C. Firefly bioluminescence: a mechanistic approach of luciferase catalyzed reactions. IUBMB Life 61(1), 6–17 (2009).
Nelson, D. L. & Cox, M. M. Lehninger Principles of Biochemistry: W. H. Freeman (2008).
Ronaghi, M. et al. Real-time DNA sequencing using detection of pyrophosphate release. Anal Biochem 242(1), 84–9 (1996).
Strotmann, H. et al. Thiophosphate analogs of ADP and ATP as substrates in partial reactions of energy conversion in chloroplasts. Biochim Biophys Acta 545(1), 122–30 (1979).
Yukawa, A., Watanabe, R. & Noji, H. Effects of an ATP analogue, adenosine 5′-[alpha-thio]-triphosphate, on F1-ATPase rotary catalysis, torque generation, and inhibited intermediated formation. Biochem Biophys Res Commun 458(3), 515–9 (2015).
Ishmukhametov, R. R., Galkin, M. A. & Vik, S. B. Ultrafast purification and reconstitution of His-tagged cysteine-less Escherichia coli F1Fo ATP synthase. Biochim Biophys Acta 1706(1–2), 110–6 (2005).
Rosen, B. P. & McClees, J. S. Active transport of calcium in inverted membrane vesicles of Escherichia coli. Proc Natl Acad Sci USA 71(12), 5042–6 (1974).
Futai, M. Orientation of membrane vesicles from Escherichia coli prepared by different procedures. The Journal of Membrane Biology 15(1), 15–28 (1974).
Moriyama, Y. et al. One-step purification of Escherichia coli H(+)-ATPase (F0F1) and its reconstitution into liposomes with neurotransmitter transporters. J Biol Chem 266(33), 22141–6 (1991).
Unden, G. & Bongaerts, J. Alternative respiratory pathways of Escherichia coli: energetics and transcriptional regulation in response to electron acceptors. Biochim Biophys Acta 1320(3), 217–34 (1997).
Solomon, K. A. & Brusilow, W. S. Effect of an uncE ribosome-binding site mutation on the synthesis and assembly of the Escherichia coli proton-translocating ATPase. J Biol Chem 263(11), 5402–7 (1988).
Nicholls, D. G. & Ferguson, S. J. Quantitative Bioenergetics: The Measurement of Driving Forces. Bioenergetics (Fourth edition), p. 27–51 (2013).
Lobau, S., Weber, J. & Senior, A. E. Catalytic site nucleotide binding and hydrolysis in F1F0-ATP synthase. Biochemistry 37(30), 10846–53 (1998).
LaNoue, K. F., Jeffries, F. M. & Radda, G. K. Kinetic control of mitochondrial ATP synthesis. Biochemistry 25(23), 7667–75 (1986).
Ferguson, S. J. et al. The ATPase as an irreversible component in electron transport linked ATP synthesis. FEBS Lett 62(3), 272–5 (1976).
Mendoza-Hoffmann, F. et al. The Biological Role of the zeta Subunit as Unidirectional Inhibitor of the F1FO-ATPase of Paracoccus denitrificans. Cell Rep 22(4), 1067–1078 (2018).
Zharova, T. V. & Vinogradov, A. D. Energy-dependent transformation of F0.F1-ATPase in Paracoccus denitrificans plasma membranes. J Biol Chem 279(13), 12319–24 (2004).
Burstein, C., Tiankova, L. & Kepes, A. Respiratory control in Escherichia coli K 12. Eur J Biochem 94(2), 387–92 (1979).
Zharova, T. V. & Vinogradov, A. D. Oxidative phosphorylation and respiratory control phenomenon in Paracoccus denitrificans plasma membrane. Biochemistry (Mosc) 77(9), 1000–7 (2012).
Boyer, P. D. Catalytic site occupancy during ATP synthase catalysis. FEBS Lett 512(1–3), 29–32 (2002).
Buckstein, M. H., He, J. & Rubin, H. Characterization of nucleotide pools as a function of physiological state in Escherichia coli. J Bacteriol 190(2), 718–26 (2008).
Etzold, C., Deckers-Hebestreit, G. & Altendorf, K. Turnover number of Escherichia coli F0F1 ATP synthase for ATP synthesis in membrane vesicles. Eur J Biochem 243(1–2), 336–43 (1997).
Harris, D. A. et al. Specificity of nucleotide binding and coupled reactions utilising the mitochondrial ATPase. Biochim Biophys Acta 504(3), 364–83 (1978).
Drobinskaya, I. Y. et al. Tightly bound adenosine diphosphate, which inhibits the activity of mitochondrial F1-ATPase, is located at the catalytic site of the enzyme. FEBS Letters 182(2), 419–424 (1985).
Hirono-Hara, Y. et al. Activation of pausing F1 motor by external force. Proc Natl Acad Sci USA 102(12), 4288–93 (2005).
Singh, D. & Gruber, G. Crystallographic and enzymatic insights into the mechanisms of Mg-ADP inhibition in the A1 complex of the A1AO ATP synthase. J Struct Biol 201(1), 26–35 (2018).
Boyer, P. D. Catalytic site forms and controls in ATP synthase catalysis. Biochim Biophys Acta 1458(2-3), 252–62 (2000).
Casadio, R. Measurements of transmembrane pH differences of low extents in bacterial chromatophores. European Biophysics Journal 19(4), 189–201 (1991).
Manchester, K. L. Free energy ATP hydrolysis and phosphorylation potential. Biochemical Education 8(3), 70–72 (1980).
Acknowledgements
We thank Dr. Andrea Eberle (University of Bern) for inspiring discussions on next generation sequencing techniques that promoted the use of ATPαS. This work was supported by the Swiss National Science Foundation (SNSF).
Author information
Authors and Affiliations
Contributions
C.v.B. and A.M. designed functional experiments. A.M. performed all experiments, and together with C.v.B. analyzed the data. C.v.B. and A.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meyrat, A., von Ballmoos, C. ATP synthesis at physiological nucleotide concentrations. Sci Rep 9, 3070 (2019). https://doi.org/10.1038/s41598-019-38564-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-38564-0
This article is cited by
-
A novel pyrazolone complex P-FAH-Cu-bpy induces death of Escherichia coli and Staphylococcus aureus by disrupting cell structure and blocking energy
Archives of Microbiology (2023)
-
The assembly, regulation and function of the mitochondrial respiratory chain
Nature Reviews Molecular Cell Biology (2022)
-
Mechanism of AAA+ ATPase-mediated RuvAB–Holliday junction branch migration
Nature (2022)
-
Dietary supplementation with inosine-5′-monophosphate improves the functional, energetic, and antioxidant status of liver and muscle growth in pigs
Scientific Reports (2022)
-
Rationally designed mariner vectors for functional genomic analysis of Actinobacillus pleuropneumoniae and other Pasteurellaceae species by transposon-directed insertion-site sequencing (TraDIS)
Animal Diseases (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.