Abstract
Virtual Screening (VS) methods can drastically accelerate global drug discovery processes. Among the most widely used VS approaches, Shape Similarity Methods compare in detail the global shape of a query molecule against a large database of potential drug compounds. Even so, the databases are so enormously large that, in order to save time, the current VS methods are not exhaustive, but they are mainly local optimizers that can easily be entrapped in local optima. It means that they discard promising compounds or yield erroneous signals. In this work, we propose the use of efficient global optimization techniques, as a way to increase the quality of the provided solutions. In particular, we introduce OptiPharm, which is a parameterizable metaheuristic that improves prediction accuracy and offers greater computational performance than WEGA, a Gaussian-based shape similarity method. OptiPharm includes mechanisms to balance between exploration and exploitation to quickly identify regions in the search space with high-quality solutions and avoid wasting time in non-promising areas. OptiPharm is available upon request via email.
Similar content being viewed by others
Introduction
The discovery of new drugs is a very expensive process, frequently taking around 15 years with success rates that are usually very low1,2. Many experimental approaches have been used for discovering new compounds with the desired pharmacological properties, ranging from traditional medicine3,4 to High Throughput Screening (HTS) infrastructures5,6. The latter is mostly used by the Pharma Industry, but little by academic research groups; in other words, its application is not widespread outside the industrial domain. In order to avoid these limitations, new techniques based on principles of Physics and Chemistry were developed about three or four decades ago for the computer simulation (mainly using high-performance computing architectures) of systems of biological relevance7,8. Computational chemistry was later applied for processing large compound databases, and also for predicting their bioactivity or other relevant pharmacologic properties. Using this approach, it was shown that it was possible to use such computational methodology to pre-filter compound databases into much smaller subsets of compounds that could be characterized experimentally. This idea was named Virtual Screening (VS), and it reduces the time needed and expenses involved when working on drug discovery campaigns9,10. Nonetheless, the accuracy of the predictions made with VS methods still needs to be improved to avoid discarding promising compounds or providing erroneous signals and the time needed for their calculations still needs to be reduced. The inaccuracies in the predictions of VS methods are mostly due to the simplifications used in their scoring functions11.
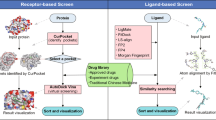
VS methods can be divided into Structure-Based (SBVS) and Ligand-Based (LBVS) methods. When the structure of the protein target is known, SBVS can be applied, and methods such as molecular docking12 and Molecular Dynamics13 are employed. But the number of already resolved crystallographic structures is still insufficient14, so SBVS methods cannot always be applied. Another option is to use LBVS methods, where only data about known compounds with desired properties are used to derive new improved ones. In practice, whether SBVS or LBVS methods should be used, or even both at the same time, will depend on the specific drug discovery project.
This study focuses on LBVS methods, which can be divided into several categories15 such as pharmacophore methods16,17, shape similarity methods (SSM)18, QSAR19, Machine Learning20, atom-based clique-matching such as SQ/SQW21 and Lisica22, property-based (USR23) or atom distribution triplet based (Phase-Shape24).
In SSM, a large database of compounds is processed against a molecular query, to provide information concerning which of the molecules from the database is geometrically similar, in terms of global molecular shape, to the input molecule used. Indeed, different strategies exist for shape calculation. One of the most widely used is the Gaussian25 model. Tools such as ROCS26, WEGA27, SHAFTS28 and Shape-IT29 use it.
The main differences between SSM reside in the accuracy of the predictions. It has been demonstrated that, depending on the compound dataset, some methods perform better than others30, but there is currently no one-size-fits-all approach that can be considered first choice for any molecular dataset. Besides, the computational time needed for the calculations is also of the utmost importance.
Among the previously commented SSM methods, we consider WEGA to be the state of the art in terms of accuracy of the predictions, while ROCS is considered to be the state of the art in terms of computational speed. For achieving such performance, ROCS introduced a number of drastic short-cuts for efficiency for computing overlap volumes between molecules31. For instance, all hydrogen atoms are ignored as they make very small contribution for the overall molecular shape, and all heavy atoms are set with equal radii. Besides, the most critical simplification in ROCS is that the shape density function of each molecule contains only the first-order terms, and all higher order terms in the original Gaussian approach25 are omitted. This significantly simplified ROCS computations but also received criticism for the inaccuracy of this approximation23; mainly that the molecular volumes are significantly overestimated. And since the Gaussian shape algorithms are widely used in various VS methods, it is important to avoid errors introduced to the shape similarity calculation due to this overestimation of the volumes.
WEGA was the first method that partially solved some of these ROCS issues by avoiding the use of only first-order terms and incorporating more terms, at the expenses of increasing computation costs, but increasing accuracy of the calculations, which is desirable in the drug discovery context.
In this work, we introduce a novel SSM method named OptiPharm, which introduces a new optimization scheme that can be adapted through extensive parameterization to relevant features of molecular datasets, such as average size, shape, etc. In other words, OptiPharm is an evolutionary method for global optimization, which can be parametrized to different aims. SSM methods with extensive parameterization at the search level have not been practically explored in the VS context. The most of techniques are local optimizers which do not sufficiently explore the search space. As the results we later show, making an effort to deeply explore the whole search space can be of a great interest to increase hit rates in drug discovery projects.
Method
This section describes the main idea behind shape similarity calculations and its application in the drug discovery process using the new OptiPharm method. Next, the optimization algorithm used in similarity screening calculations is presented and, finally, the benchmarks used in this study are explained in detail.
Shape Similarity
The similarity score between molecules A and B is computed as the overlapping volume of their atoms. In particular, to compare the results obtained by OptiPharm with those achieved by WEGA, the similarity function is implemented as in WEGA27. For the sake of completeness, this function is written in the following form:
where wi and wj are weights associated with the atoms i and j, respectively. This weight is calculated by solving the following mathematical expression:
where k is a universal constant, which is set to 0.8665, vi is the volume of atom i, whose value is computed as \({v}_{i}=\frac{4\pi {\sigma }^{3}}{3}\), similarly to how it was done in the original work of WEGA27.
Finally, the overlapping \({v}_{ij}^{g}\) is represented as a product of Gaussian representations:
where p is a parameter that controls the softness of the Gaussian spheres, i.e., the height of the original Gauss function, and σ is the radius of the atom. More precisely, the radius represents the well-known van der Waals radius. The values associated to those two parameters are obtained by empirical knowledge. For the problem under consideration, the same figures proposed in WEGA27 are considered.
Notice that the score obtained from Equation 1 depends on the number of atoms of the two compared molecules, i.e., the higher this number, the longer the value of \({V}_{AB}^{g}\). In reality, it lies in the interval [0, +inf). To be able to measure the grade of similarity between compounds, independently of the number of atoms that compose them, the Tanimoto Similarity32 value is computed:
where VAA and VBB is the self-overlap volume of molecules A and B, respectively. It has a value in the range \([0,1]\), where 0 means there is no overlapping, and 1 means the shape densities of both molecules are the same.
Previous approaches
WEGA is a local optimizer conceived to maximize the overlapping between two molecules A and B, given as input parameters. To direct the search, it computes the derivate of the objective function Tc, which specifically considers Equation 1. It means that WEGA can be only applied when the similarity of two molecules is measured by means of such an equation.
WEGA starts the search with an initial solution and moves it from neighbor to neighbor as long as possible while increasing the objective function value. The main advantage of WEGA is its ability to find a solution in a sufficiently short period of time. On the contrary, its main drawback is its difficulty to escape from local optima where the search cannot find any further neighbor solution that improves the objective function value, i.e., the quality of the final solution closely depends on the considered starting ligand pose, obtained from the conformation of the molecular query. To deal with this drawback and to increase its probability of success, WEGA considers more than a single starting point. More precisely, it applies the local optimizer from four different poses. The first one is obtained by aligning and centering the two input molecules at the origin of the coordinates. The remaining ones are obtained by rotating the first one 180 grades at each axis27.
The interested reader can revise literature33,34,35 for the research progress of WEGA algorithm and some of its applications. In this work, we consider that it is possible to find a better trade-off between quality of the solution and computing time.
Optimization algorithm
OptiPharm is an evolutionary global optimizer, available upon request. It can be considered a general-purpose algorithm, in the sense that it can be used to solve any optimization problem that involves the computation of the similarity of two compounds given as input parameters. In other words, it is independent of the objective function used to measure the similarity between two given molecules. Nevertheless, in this work, its performance is illustrated by solving a maximization problem which consists on finding the s solution which maximizes the Tc function previously defined.
OptiPharm is a global optimization method in the sense that it makes an effort to analyze the whole search space looking for promising areas where the local and global optima can be. In other words, instead of focusing on a set of pre-specified starting points, as WEGA does, it applies procedures to find promising subareas of the search space, which will be deeper analyzed during the optimization procedure. OptiPharm applies procedures based on species evolution to gradually adjust one of the molecules (the query) to the other one (the target), which remain fixed during the optimization procedure.
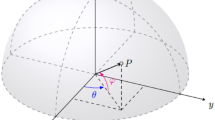
A solution s represents the rotation and translation to be accomplished by the query. More precisely, s is a quaternion of the form s = (θ, c1, c2, Δ), where θ is the rotation angle to be carried out over a rotation edge defined by the points c1 = (x1, y1, z1) and c2 = (x2, y2, z2), and Δ = (Δx, Δy, Δz) represents a displacement vector. It should be borne in mind throughout that a quaternion indicates the rotation and the translation applied to the variable molecule from its initial state.
The parameters associated to a quaternion s are bounded. Since each pair of input compounds can have different sizes, the corresponding limits are dynamically computed by OptiPharm, for each particular instance. To do so, the 3D boxes containing the input compounds are calculated. Then, the bound values for both c1 and c2 are set to the borders of the box containing the variable molecule. Notice that the same axis can be given by an infinite number of two coordinates. In this way, redundancy is prevented, which is very important from an optimization point of view, since exploring the same solutions several times makes the algorithm inefficient. The interval of Δ is set to [−maxD, maxD], being maxD the maximum difference between the boxes. This avoids the evaluation of situations where no overlapping exists between molecules, and the similarity between them is clearly zero (see Fig. 1). Finally, the angle θ is always set in the interval [0, 2π], independently of the compounds considered as input parameters.
The bound values of the quaternion components define a multidimensional search space (or feasible region) with multiple local and global optima.
OptiPharm is a new metaheuristic for global optimization. OptiPharm includes mechanisms to detect promising subareas of the search space and to discard those in which no global optima are expected to be found. In other words, instead of focusing on some fixed starting solutions, OptiPharm attempts to detect new ones which have the potential to become local or global optima. To do so, OptiPharm initially works on a set of M solutions (quaternions), called population. The quaternions can be considered as independent starting points on which OptiPharm applies reproduction procedures based on natural evolution. The term independent signifies that a point has the ability to discover new promising poses (in this work we use the concept of pose as rigid body rotations and translations obtained from starting conformation of query compound) without the participation of the rest of the population. As a consequence, offsprings of new promising solutions can appear. Then, from among all the existing poses, the best M solutions will be promoted to the next stage, where they are improved by means of a local optimizer. This reproduction-replacement-improvement sequence is repeated until a number of iterations tmax is achieved (see Fig. 2).
But the real strength of OptiPharm lies on the concept of radius: each solution in the population has an associated radius value, which determines a multidimensional subarea of the search space. It can be understood as a window, where the reproduction and improvement methods are applied. The radius associated to a pose depends on the iteration i where it has been discovered. More precisely, the radius Ri of a new point, found during the reproduction procedure at iteration i, comes from an exponential function that decreases as the index level (cycles or generations) increases, and which depends on the initial domain landscape (the radius at the first level, R1) and the radius of the smallest candidate solution \({R}_{{t}_{max}}\), which is given as input parameter. This radius mechanism, designed as a balance between exploration and exploitation, is inherited from UEGO, a general optimization method widely used in the literature with promising results36.
During the execution of OptiPharm, several candidate solutions with different radii can coexist simultaneously which means that the method is able to analyze both big and small subregions at the same stage of the optimization procedure as it looks for valuable new solutions (see Fig. 3).
Apart from the maximum number of starting solutions M, the number of iterations tmax and the smallest radius value \({R}_{{t}_{max}}\), OptiPharm has another input given parameter: the maximum number of function evaluations for the whole optimization procedure, N. These function evaluations are distributed among the candidate solutions at each iteration, in such a way that each one has a budget to generate new solutions and to improve them. These budgets are mathematically computed by means of equations that depend on the previously mentioned input parameters. Again, this idea has been borrowed from UEGO36.
In a previous work37 the effects of the different parameters of UEGO and, hence, of OptiPharm were analyzed. Moreover, some guidelines to fine-tune the parameters depending on the problem to be solved were also proposed.
Finally, it should be noted that, unlike most heuristics in the literature, the termination criteria of OptiPharm is not based on the number of function evaluations N, but on the number of iterations tmax. This point is important since the number of function evaluations consumed by OptiPharm depends on the particular case being solved. In other words, OptiPharm adapts itself to the complexity of the problem considered.
In the following subsubsections, the key stages of OptiPharm are explained.
Initialization method
In the initialization phase, the two input molecules are aligned and centered at the origin of the coordinates (see Fig. 4). Then, from this initial situation, a population of M poses is composed. The first pose represents this initial stage, i.e. the former candidate solution will be equal to s1 = (θ, c1, c2, Δ) = (0, (0, 0, 0), (0, 0, 0), (0, 0, 0)), indicating than the molecule to be optimized is not moved with respect to the target, which remains fixed. Three more initial poses are obtained by rotating the variable molecule π radians at each axis (always from the initial state), resulting in the following candidate solutions s2 = (π, (1, 0, 0), (0, 0, 0), (0, 0, 0)), s3 = (π, (0, 1, 0), (0, 0, 0), (0, 0, 0)) and s4 = (π, (0, 0, 1), (0, 0, 0), (0, 0, 0)). Finally, in order to introduce some randomness and prevent a possible drift to local optima, M − 4 molecular poses, with all their randomly obtained parameters, are also included.
Initially both molecules are aligned and centered at the origin of the coordinates (see figure above). The variable molecule is depicted in green, while the target is represented in red. Then, OptiPharm applies procedures based on species evolution to gradually adjust the variable molecule to the target. The two figures below show intermediate solutions obtained by OptiPharm when, from the initial state (top), a rotation is carried out (left) and a consecutive translation is accomplished (right).
Figure 5 shows the five initial solutions achieved for a particular instance with M = 5. As can be seen, there is always some overlap between both molecules. Consequently, the objective function is always greater than zero, while the radius value associated to all the initial poses is equal to R1. Notice that such a value is equal to the diameter of the search space.
Initial solutions for a case with M = 5: (a) s1, initial situation; (b) s2, obtained when rotating s1 π rad at x-axis; (c) s3, obtained when rotating s1 π rad at y-axis; (d) s4, obtained when rotating s1 π rad at z-axis; (e) s5, all the parameter (θ, c1, c2, Δ) are randomly computed in the limits dynamically calculated by OptiPharm, for this particular instance.
Reproduction method
The reproduction method is in charge of exploring the different subareas defined by the radius of each pose s in the population (see Fig. 3). The idea is to find new promising solutions which can evolve toward local or global optima at later phases of the algorithm. Each subarea is analyzed independently of the remaining ones. The process is as follows:
From each pose si in the population, new candidate solutions sij are randomly computed in the area defined by its radius (see Fig. 6(a)). Additionally, for each pair of trial solutions (sij and sik), the middle point (Mid(sij, sik)) of the segment connecting the pair is computed (see Fig. 6(b)). Then, the objective function value of the extreme points (f(sij) and f(sik)), as well as the middle point (f(Mid(sij, sik))), is computed. If any objective function value of these new generated points is better of the original solution si, it will be updated, i.e., the centre of that subarea si will be the one with the best objective function value. Additionally, if the objective function value in the middle solution is better than that of the extreme points, it may mean that it is in a hill (see Fig. 6(b)), so that it is considered a candidate to be included in the population list. On the contrary, the endpoints will be inserted as new poses. The radius of the new pose in the population will be that one associated with the current iteration. Figure 6(c) shows a summary of the whole process by keeping the references to the names in Fig. 6(a,b).
Replacement method
After the reproduction method has been applied, it is highly probable that the size of the population will be greater than the population size given by the input parameter M. Therefore, a mechanism for selecting the surviving solutions must be applied. Different types of replacements exist but, in this work, a deterministic and highly elitist one has been implemented: the original population and their corresponding offspring are grouped in an intermediate population, and then the M best solutions, i.e., the best poses, are selected as members of the population. The remaining ones are eliminated.
The implementation of this direct replacement involves the use of a sorting procedure whereby the poses are sorted according to their shape similarity value.
Improvement method
In order to introduce some noise into the search process, and hence avoid the convergence to local optima, a mutation operator is usually applied to the new offspring. Then, in most evolutionary or genetic algorithms, mutation mechanisms are included in the optimization procedure, which runs small random changes to the new individuals. However, for the present problem, the use of improvement methods has better shown to better approximate the poses towards the optima.
The improvement method implemented in OptiPharm is the local search method SASS, initially proposed by Solis and Wets38. It has been chosen mainly because it is a derivative-free optimization algorithm that can be applied to maximize any arbitrary function over a bounded subset of \({{\mathbb{R}}}^{N}\).
Several modifications have been included to adapt SASS to the problem at hand. In the following they are briefly described.
Algorithm SASS internally assumes that the range in which each variable is allowed to vary is the interval \([0,1]\). Since this is not our case, when necessary we use a function to rescale (normalize) the variable values to the interval \([0,1]\), and the function denorm to invert this process. In SASS, the new points are generated using a Gaussian perturbation \(\xi \in {{\mathbb{R}}}^{3}\) over the search point (x,α) and a normalized bias term \(b\in {{\mathbb{R}}}^{3}\) to direct the search. The standard deviation σ specifies the size of the sphere that most likely contains the perturbation vector. In this work, its upper bound σub should have the same value as the normalized radius of the caller solution. Then, the parameter σub is also considered an argument of SASS. Hence, any single step taken by the optimizer is no longer than the radius of the calling candidate solution. Finally, the stopping rules are determined by a maximum number of function evaluations (femax) and by the maximum number of consecutive failures (Maxfcnt).
OptiPharm applies SASS to every pose in the population. See Fig. 7 for an illustrative example of its performance.
Example. The local optimizer SASS has been used as Improvement method. This figure shows the performance of SASS for a 2D case. SASS is a derivative-free optimization algorithm that can be applied to maximize an arbitrary function over a bounded subset of \({{\mathbb{R}}}^{N}\). It looks for an improving direction and moves the starting point along it by making changes of different sizes (if the number of consecutive successes is larger than a pre-specified value, then the advance along the suggested searching direction will be longer; otherwise, the size of the step will be reduced. The area of action of the optimizer is limited by the corresponding radius. In OptiPharm, the stopping rule of SASS is determined by a maximum number of function evaluations and by the maximum number of consecutive failures.
Computational Experiments Framework
Hardware setup
All the experiments carried out in this work have been executed in a Bullx R424-E3, which consists of 2 Intel Xeon E5 2650v2 (16 cores), 128 GB of RAM memory and 1 TB HDD.
Methodology to test the performance of the algorithms
OptiPharm is a computer program which implements an evolutionary optimization algorithm which includes randomness in the search procedure. Then, in order to test its performance, we run each particular instance several times and we provide some statistical metrics, as usually is done when testing any heuristic algorithm in works in literature39,40,41,42,43. From a statistic point of view, a minimum number of 30 samples need to be considered for this44. Nevertheless, in this work, each particular instance has been run 100 times to increase confidence in the results. Then, figures as the average value and the standard deviation are computed to analyze its effectiveness and efficiency. It is important to highlight that executing several times a particular instance is only a methodology to analyze the robustness of the algorithm, but in the real world scenario, OptiPharm only needs a single run to provide reliable results.
Regarding WEGA, it is only run once for each particular instance, since it is deterministic (it uses a descent gradient method) and different executions always produce the same result.
Benchmarks
Unlike OptiPharm, WEGA does not consider the hydrogen atoms in the shape similarity calculations. To be able to compare the results provided by both algorithms, OptiPharm has been configured to omit the hydrogens when computing the shape similarity score. Additionally, as WEGA does27, all the heavy atom radii have been set to 1.7 Å. Furthermore, all compound pairs are centred and aligned in the same way. Consequently, the molecule centroids have been located at the coordinates centre of the search space. Finally, each molecule has been aligned in such a way that its longest axis has been oriented at X-axis and the shortest along the Z-axis.
The underlying OptiPharm algorithm is parameterizable, which means that it can be fine-tuned depending on the user’s preferences. So users may prefer to obtain high-quality solutions at the expense of slightly increasing the computational effort, while others may want an acceptable solution with reasonable computing time. In this work, the parameters that control OptiPharm were tuned by trying several combinations of parameter values with a reduced set of problems, and following the guidelines described in a previous work37. As a consequence, two different sets of input parameters are proposed, given rise to two versions of OptiPharm with different aims:
-
(i)
OptiPharm Robust (OpR). In this case, the set of input parameters is chosen to make OptiPharm reliable and robust; in other words, to allow OptiPharm to deeply explore and exploit the search space in the search for the best possible pose. In particular, the following values were considered: N = 200000 function evaluations, M = 5 starting poses, tmax = 5 iterations and \({R}_{{t}_{max}}=1\) as the smallest possible radius.
-
(ii)
OptiPharm Fast (OpF). On this occasion, the parameters are tuned so that the running times are lower or similar to those of WEGA, enabling a fair comparison between both algorithms. The following values were considered: N = 1000 function evaluations, M = 5 starting poses, tmax = 5 iterations and a minimum radius of \({R}_{{t}_{max}}=5\).
From the previous paragraphs, one could infer that the number of starting poses, M = 5, and the number of iterations, tmax = 5, can be fixed independently of the goal pursued, while the smallest radius \({R}_{{t}_{max}}\), and most importantly, the number of function evaluations N have a bigger influence in both the effectiveness and the efficiency of the algorithm.
Four computational studies were designed by considering the well-known Food and Drug Administration (FDA)45, Directory of Useful Decoys (DUD)46, Directory of Useful Decoys - Enhanced (DUD-E)47 and Maybridge datasets. In the following sections, they are briefly described.
FDA
The FDA, a federal agency of the United States Department of Health and Human Services, is responsible for protecting and promoting public health by controlling, among other things, prescription and over-the-counter pharmaceutical drugs (medications). This agency provides a data set containing 1751 molecules, which represents approved medicines that can be used with safety in humans in the USA. It is a common practice48, in the current scenario, to identify which compound pairs in the FDA database share a high degree of shape similarity. To compare the performance of both OptiPharm and WEGA, a set of 40 query compounds were randomly selected from this database. In order to obtain a representative set of samples, the FDA dataset was initially sorted according to the number of atoms of the compounds, and divided into 24 intervals (see Fig. 8). Then, a subset of compounds was randomly chosen for each interval. The number of selected samples in each interval was proportional to the number of compounds it included.
DUD
Tests were also carried out applying shape similarity calculations and using different sets of molecules that are known to be active or inactive, and standard VS benchmark tests, such as the DUD46, whereby VS methods check how efficient they are at differentiating ligands that are known to bind to a given protein target, from non-binders or decoys. Input data for each molecule of each set contain its molecular structure and information about whether it is active or not. Information about active molecules for each protein of the DUD set was taken from experimental data. Decoys were prepared in order to resemble active ligands physically, but at the same time, to be chemically different from active molecules, making it very unlikely that they would act as binders. On average, for each ligand it is possible to find 36 decoy molecules that are very similar in physical terms, but with a very different topology. Details about how decoys were prepared (selected from already existing molecules in the ZINC database) can be found in the literature46, so that we shall only mention here the principal details required to understand the present study.
-
1.
The initial database was built using 3.5 million Lipinski-compliant molecules from the ZINC database of commercially available compounds (version 6, December 2005).
-
2.
Feature key fingerprints were calculated using the default type 2 substructure keys of CACTVS49 and the fingerprint-based similarity analysis was performed with the program SUBSET. Compounds with Tc values lower than 0.9 to any annotated ligand (named as actives) were selected. This reduced the number of ZINC compounds to 1.5 million molecules topologically dissimilar to the ligands.
-
3.
The program QikProp (Schrodinger, LLC, New York, NY) was used to calculate 32 physical properties of all the annotated ligands and selected ZINC compounds from the previous step, and QikSim (Schrodinger, LLC, New York, NY) was applied to prioritize ZINC compounds possessing similar physical properties to any of the ligands.
-
4.
A weight of 4 was used to emphasize the druglike descriptors (molecular weight, number of hydrogen bond acceptors, number of hydrogen bond donors, number of rotatable bonds, and log P), while the rest of the descriptors were ignored (weight 0) during the similarity analysis procedure.
-
5.
Finally, thirty-six decoy compounds were selected for each ligand, leading to a total of 95316 decoys that were similar in terms of physical properties but topologically dissimilar to the 2950 annotated ligands. The total number of decoys is less than 36 times the number of annotated ligands because some ligands had the same decoys.
The original DUD database downloaded from http://zinc.docking.org has been used.
DUD-E
The DUD-E47 is a well-known benchmark for structure-based virtual screening methods from the Shoichet Lab at UCSF47. The methodology of the DUDE benchmark is fully described in its original work47. Briefly, the benchmark is constructed by first gathering diverse sets of active molecules for a set of target proteins. Analogue bias is mitigated by removing similar actives; similar actives are eliminated by first clustering the actives based on scaffold similarity, then selecting exemplar actives from each cluster. Then, each active molecule is paired with a set of property-matched decoys (PMD)50. PMD are selected to be similar to each other and to known actives with respect to some 1-dimensional physicochemical descriptors (e.g., molecular weight) while being topologically dissimilar based on some 2D fingerprints (e.g., ECFP51). The enforcement of the topological dissimilarity supports the assumption that the decoys are likely to be inactive because they are chemically different from any know active. The benchmark consists of 102 targets, 22,886 actives (an average of 224 actives per target) and 50 PMD per active52. The original DUD-E database downloaded from http://dude.docking.org/ has been used in this work.
Maybridge
Maybridge53 Screening Hit Discovery collection (over 53,000 compounds) is a commercial library of small hit-like and lead-like organic compounds of high diversity (Tanimoto Clustering at 0.9)54, that covers ca. 87% of the 400,000 theoretical drug pharmacophores with general compliance with the Lipinsky rule of five and of good ADMET properties. The HitCreatorTM Collection (selection of 14,400 of Maybridge screening compounds) aims to represent the diversity of the main collection covering the drug-like chemical space. Maybridge also offers a fragment library (30,000 fragments), a hit-to-lead building block collection, and a Ro3 2500 diversity fragment library (2500 fragments) with a Tanimoto similarity index of 0.66 (based on standard Daylight fingerprinting), assured solubility, optimized for SPR and Ro3 compliant. It provides special collections of Fluoro55, Fluoro and Bromo-fragment libraries56. The original Maybridge database downloaded from https://www.maybridge.com has been used in this study.
The AUC metric
In this work, to measure the goodness of the algorithms when distinguishing between ligands and decoys, the Area Under a ROC Curve (AUC) was computed, as previously done in other related papers27. See57 for an in-depth description of calculation. Broadly speaking, the AUC of a set of elements is computed by considering a descriptor value that is associated to each element.
For the problem at hand, such a descriptor is given by Equation 4, which measures the shape similarity between two molecules, A and B. However, before computing the AUC, given a query molecule and a set of molecules the similarity to which is to be computed, a optimization problem must be solved to obtain the shape similarity scores for each molecule in the set. Then, the list is sorted in descending order according to the shape similarity values. Without going into detail, an AUC value equal to 1 means that such a particular algorithm has been able to differentiate perfectly between two datasets - in our case, between ligands and decoys. In other words, it is possible to determine a cut-off point (a real value) which divides the list into two intervals that contain all the decoys and ligands, respectively. When it is not possible to determine only two intervals, more cut-off points should be considered in an incremental way. Of course, the larger the number of intervals, the smaller the AUC value. However, AUC values smaller than or equal to 0.5 mean the algorithm has poor effectiveness, i.e., a random method would have achieved a similar classification.
Results and Discussion
Results obtained for FDA database
It is important to mention that for all the algorithms and all the instances, a score equal to 1 is obtained when a molecule is compared to itself. Thus, from here on, when we mention “the molecule with the highest shape similarity to a query compound”, and noted by BestComp, we exclude the case where target molecule and query are equal.
Table 1 shows, for each query compound, its number of atoms (nA), the other compound from the FDA database with the highest shape similarity (BestComp) and the associated function score (Tc), according to OpR, OpF and WEGA. As can be seen, the OpR algorithm provides the highest shape similarity values Tc, although it is also the most time-consuming method according to Table 2. This means that better predictions can be accomplished by using OpR when there are no time constraints. However, if lower execution times are required, algorithms such as OpF or WEGA should be considered.
To the best of our knowledge, no algorithm, method or program exists that is able to provide with certainty the most similar molecule to a given query compound. Until this work, WEGA was the algorithm providing the most optimal shape similarity values27,34. Now, as can be seen in Table 1, OpR improves on WEGA in terms of the ability to find higher values of shape similarity when processing a query compound against a ligand database. Therefore, to analyze the effectiveness of OpF and WEGA in term of their predictions, the solutions provided by OpR will be considered the optimal ones.
As can be seen in Table 1, the predictions of WEGA coincide with those of OpR in 22 out of 40 cases, while OpF does it in 30 out of 40 occasions. This represents a small advantage to OpF against WEGA in terms of success in the predictions. Additionally, from Table 2, which shows the computing times, one can appreciate that OpF is quicker than WEGA.
Furthermore, it is important to study the instances where the predictions of OpF and WEGA do not coincide with those achieved by OpR. This occurs in 18 out of 40 cases for WEGA, and 10 times for OpF. Then, for each particular query, the 1751 compounds are sorted in descending order according to the shape similarity value obtained by OpR. Next, it is computed the position i in the list where the BestComp achieved by OpF (resp. WEGA) is, and which one shape similarity value, Tc(OpR). This information is shown in Table 1, columns 6 and 9 for OpF and WEGA, respectively. Broadly speaking, in most of cases the predictions carried out by OpF are located in a better position in the OpR list than the predictions proposed by WEGA.
It is important to mention that, in general, OptiPharm is designed to maintain population diversity and to investigate many promising poses in parallel, avoiding the genetic drift towards a single (local or global) optimal pose. However, depending on the selected set of parameters, the accuracy when approximating to the optima may be higher or lower. For this reason, OpF has been fine-tuned to explore the search space looking for the most promising poses, but without wasting time by “polishing” them. In optimization terms, the input parameters are selected to determine the highest peaks in the search space, but not to actually reach the top of the highest peak. Even when OpF proposes as BestComp the same compound as OpR (or even WEGA), its shape similarity value may be smaller. If the algorithm is allowed to run longer, as with OpR, the identified poses can be polished, increasing the score value. In this case we prioritize the computational effort. Figure 9 depicts a graphical example of this fact, specifically the query DB09236 from the FDA database, whose result can be seen in Table 1. Considering this query, OpR reveals that DB00270 is the compound which maximizes the shape similarity function, with a score value equal to Tc = 0.672 (see Fig. 9(a)). OpF reveals that molecule DB01115 maximises the shape similarity function with a score value equal to Tc = 0.615. Finally, WEGA reveals that the molecule DB01433 maximizes the shape similarity function with Tc = 0.662. Apparently, WEGA achieves a more similar compound than OpF, since it provides as solution a compound with a higher score than the one proposed by OpF. However, when OpR optimizes the query with the molecule DB01115 proposed by OpF, it provides a score value of 0.669 (see Fig. 9(b)). By contrast, OpR gives a value of 0.662 when it optimizes the query with the compound DB01433 given by WEGA, (see Fig. 9(c)). This means that the solution provided by OpF is more similar in terms of shape than that of WEGA.
Table 2 shows performance values among the different methods. Clearly, the slowest algorithm is OpR, since it has been fine-tuned to be robust and accurate. Even so, the time values are not extremely high when compared against the other two methods. In fact, taking into account the possibility of using high-performance computing to accelerate it (please, see Future Work Section), it would be perfectly justifiable to use Robust mode to increase the percentage success in the predictions. For its part, OpF is the fastest algorithm, reducing on average the computational effort of WEGA almost 3.5 times. Besides, as can be appreciated in the Speedup column, the lower the number of atoms, the greater the increase in speed obtained by OpF. Additionally, it is important to mention that OpF is able to adapt itself to the complexity of the problem to solve.
Finally, it is interesting to remark that, in spite of the randomness included at some stages of the OptiPharm algorithm, its variability is almost negligible, as can be appreciated from the standard deviation values provided in Table 2.
Results obtained for DUD and DUD-E databases
Tables 3 and 4 show the results of testing the shape-based VS performance of both OptiPharm (in its two versions) and WEGA against the DUD and DUD-E databases, respectively. Metrics of AUC values and execution time have been computed. As previously was mentioned, to test the OptiPharm reliability, each particular instance has been run 100 times and average values have been computed. Furthermore, the corresponding SD has also been provided. Regarding WEGA, since it is deterministic, only one single execution has been carried out for each particular instance and the corresponding values have been shown.
In general terms, the SD values obtained for OpR and OpF are quite small, which indicates that their variability is small, and that (i) they converge toward the same optima in spite of the included randomness and (ii) the computing time is practically the same when different executions of the same instance are carried out.
Focusing now on Table 3, it is possible to infer that the three algorithms are equivalent in terms of accuracy of the predictions, i.e. they obtain about the same AUC values regardless of the considered instance. In fact, the average of the AUC values is practically equal, as can be seen in the last row of the table. Nevertheless, OpF is almost 5 times faster than WEGA and more than 16 times quicker than OpR.
Finally, similar conclusions than previously can be obtained for the DUD-E database (see Table 4). In terms of effectiveness, OpR and WEGA are comparable, since they obtain practically the same mean AUC value. On the contrary, OpF obtains an average AUC value slightly smaller. Nevertheless, OpF is more than 17 times faster than WEGA and more than 38 times quicker than OpR.
Results obtained when hydrogen atoms are considered
By default, WEGA does not consider hydrogen atoms during optimization, which is a common practice for most tools in the current scenario, since evaluation without hydrogens is less time-consuming. However, this simplification may have serious consequences in a VS process. In this work, the effect of excluding the hydrogens of the molecules when optimizing is analyzed. Table 5 shows number of atoms for the 40 query molecules selected from the FDA database when the hydrogens are not taken into account and when they are considered (columns 2 and 6 respectively). Additionally, the molecule BestComp from the FDA dataset, which maximizes the shape similarity and the corresponding score value Tc, both when the input molecules include the hydrogens and when they not, is shown. Notice that these experiments were accomplished using OpR since, according to the previous results, it is the most efficient algorithm. For the sake of completeness, the average execution time (in seconds), in both cases, has also been included. As it can be seen, in 15 out of 40 cases, the BestComp molecule differs, depending on whether the hydrogens are considered or not. Additionally, and as expected, the computing time decreases when hydrogens are not considered (see columns 5 and 9). This means that excluding the hydrogens of the molecules is not an appropriate simplification; although the computing effort is shorter, the molecule that which maximizes the shape similarity can change.
Finally, for a fair comparison in terms of score value, the optimized BestComp obtained by OpR when no hydrogens are considered is re-evaluated, but considering now the hydrogens. As we can see, the obtained score value is always smaller than the one obtained when the hydrogens are included (compare columns 8 and 10). This means that the BestComp molecule found by OpR when the hydrogens are considered is indeed more similar than the one proposed when the hydrogens are excluded. The Fig. 10 illustrates this fact.
Query compound DB06439 is represented by the red structure. Hydrogens are white atoms. Colours remain fixed. (a) Tc = 0.515 where the compound DB00207 is the yellow structure. (b) Tc = 0.591 where the compound DB00207 is the green structure. (c) Tc = 0.533 where the compound DB00207 is the orange structure. (d) The three previous compounds are optimized with respect to the query.
In addition, the impact on the classification when the hydrogen atoms are considered has also been evaluated when DUD and DUD-E databases are considered as input. The algorithms OpR and OpF have been selected to this aim. The corresponding results are shown in Tables 6 and 7. Notice that WEGA has not been included in the study since it never considers the hydrogens.
Broadly speaking, the mean AUC value increases slightly when the hydrogen atoms are considered in DUD database, for both OpR and OpF algorithms. See last row of Tables 3 and 6. In particular, an increment of 0.03 (resp. 0.01) has been obtained for OpR (resp. OpF). In addition, for 23 out of 40 cases, OpR obtains better AUC values when the hydrogens are considered. Regarding OpF, it happens for 20 out of 40 instances.
The same increasing tendency can be appreciated in the mean AUC value when the DUD-E database is considered. Please, see Tables 4 and 7. In this case, an increment of 0.02 has been obtained for both OpR and OpF algorithms. Both OpR and OpF obtain better AUC values in more than half of the cases (58 out of 102 for OpR and 67 out of 102 for OpF).
In general terms, considering the hydrogens increases the average computing time. Compare again Tables 3 and 6 for DUD database, and Tables 4 and 7 for DUD-E benchmark. As can be seen, the time increases 2.9x times for both OpR and OpF when DUD is considered as input. For the DUD-E case, the increase is of 4.8x and 5.7x for OpR and OpF, respectively. Of course, the larger the number of atoms considered for a compound, the higher the computing time associated to its evaluation, but the more realistic the associated scoring function value.
Therefore, based on the results, it can be concluded that a more realistic classification of compounds can be obtained if hydrogen atoms are considered. In such a case, the computing time can be reduced by using high-performance computing approaches.
Results obtained for Maybridge database
Finally, a study has been conducted to show the utility of OpR, i.e. it can find good quality solutions when possible.
The effectiveness of OpR has been analyzed when it is executed with the Maybridge database considering hydrogens. In particular, a set of query compounds were selected from such a database. The choice procedure was carried out as follows: the Maybridge dataset was initially sorted according to the number of atoms of the compounds and split into 38 intervals. Then, a single compound was randomly chosen for each interval. Table 8 summarizes the obtained results. In particular, it is shown: (i) the number nC of compounds with a number of atoms included in the interval \({\rm{nA}}\in [i,j)\); (ii) the randomly selected query from such an interval, and (iii) the other molecule from Maybridge (BestComp) with the highest shape similarity value (Tc) according to OpR. The last row of the table shows the total number of compounds with nA < 95 (resp. nA ≥ 95) and the average Tc value. Notice that there exist intervals with 0 compounds, we note those cases by including ‘−’ in the corresponding columns.
As can be seen in Table 8, OpR obtains an average Tc value of 0.940 for queries with nA < 95. This is not rare since the number of compounds with less than 95 atoms is equal to 53370, so the probability of finding similar molecules is relatively high. On the contrary, the average Tc value obtained by OpR for molecules with more than 95 atoms is equal to 0.637, which is not a bad figure if we consider that only 29 out of 53399 molecules have more than 95 atoms. Even so, OpR obtains good quality solutions for queries with more than 95 atoms. See for example, the instances JFD0120 and JFD0063, with 96 and 104 atoms, respectively. For those two cases, OpR has found compounds with Tc values of 0.930 and 0.875, even when the number of molecules with similar sizes is not high. Let focus now on the worst cases, i.e. those where OpR obtains the lowest Tc values. They are JFD02950 and JFD02946 with 180 and 135 atoms, respectively. Notice that there are not molecules in the database with similar sizes. More precisely, there are just 10 molecules, including JFD02950 and JFD02946, with \({\rm{nA}}\in [135,190)\). Therefore, the probability of discovering similar molecules in terms of shape is very low, since the most likely is that they do not exist. Then, from the results, it is possible to infer that OpR finds a high-quality solution to a given query when it exists in the corresponding database.
Conclusions and Future Work
This work has introduced the SSM OptiPharm, based on novel metaheuristic approaches and illustrated its performance in terms of prediction accuracy and running time when processing well-known benchmarks such as DUD, and in addition FDA dataset. Comparison made with WEGA show that OptiPharm offers the same predictive accuracy but at a much lower computational cost (average speedup is 5x). Another of the advantages of the method compared with WEGA is that its optimization algorithm is easily parameterizable so that very different heuristic schemes can be tested, and so it adapts itself to a given database depending on the average molecular size and topology, to name a few. Also, bearing in mind that OptiPharm, unlike WEGA, allows optimizing including the hydrogen atoms of the compounds. Results have shown that its consideration improves the predictions, although it is more costly from a computational point of view. High-performance computing approaches may be a good alternative to deal with this drawback.
OptiPharm has been designed with parallelism in mind. Notice that each pose in the population can generate a new offspring without the participation of the remaining quaternions in the population, meaning that the Reproduction method can be easily parallelized by dividing the poses in the population among the available processing units. Similarly, the poses can also be enhanced by distributing them in the Improvement procedure. This means that OptiPharm can be drastically accelerated by using high-performance computing with practically no effort. In the future, several programming paradigms based on both shared and distributed memory architectures will be implemented and analyzed. In particular, a parallel version of OptiPharm will be implemented to be executed on GPUs, and compared with the GPU-accelerated WEGA58.
Availability of Data and Materials
The databases belong to their authors and access to them depends on any applicable restrictions. OptiPharm software is available upon request via email.
References
Drews, J. Drug discovery: a historical perspective. Science 287, 1960–1964 (2000).
Ban, F. et al. Best practices of computer-aided drug discovery: lessons learned from the development of a preclinical candidate for prostate cancer with a new mechanism of action. Journal of Chemical Information and Modeling 57, 1018–1028 (2017).
Qiu, J. Traditional medicine: a culture in the balance. Nature 448, 126–128 (2007).
Fu, X. et al. Toward understanding the cold, hot, and neutral nature of chinese medicines using in silico mode-of-action analysis. Journal of Chemical Information and Modeling 57, 468–483 (2017).
White, R. E. High-throughput screening in drug metabolism and pharmacokinetic support of drug discovery. Annual review of pharmacology and toxicology 40, 133–157 (2000).
Glick, M., Jenkins, J. L., Nettles, J. H., Hitchings, H. & Davies, J. W. Enrichment of high-throughput screening data with increasing levels of noise using support vector machines, recursive partitioning, and laplacian-modified naive bayesian classifiers. Journal of chemical information and modeling 46, 193–200 (2006).
Terstappen, G. C. & Reggiani, A. In silico research in drug discovery. Trends in pharmacological sciences 22, 23–26 (2001).
Karplus, M. & McCammon, J. A. Molecular dynamics simulations of biomolecules. Nature Structural & Molecular Biology 9, 646–652 (2002).
McInnes, C. Virtual screening strategies in drug discovery. Current opinion in chemical biology 11, 494–502 (2007).
Geppert, H., Vogt, M. & Bajorath, J. Current trends in ligand-based virtual screening: molecular representations, data mining methods, new application areas, and performance evaluation. Journal of chemical information and modeling 50, 205–216 (2010).
Bohm, H.-J. & Stahl, M. The use of scoring functions in drug discovery applications, vol. 18 (John Wiley & Sons, 2003).
Yuriev, E. & Ramsland, P. A. Latest developments in molecular docking: 2010–2011 in review. Journal of Molecular Recognition 26, 215–239 (2013).
Ganesan, A., Coote, M. L. & Barakat, K. Molecular dynamics-driven drug discovery: leaping forward with confidence. Drug discovery today 22, 249–269 (2017).
Lipinski, C. A. Rule of five in 2015 and beyond: target and ligand structural limitations, ligand chemistry structure and drug discovery project decisions. Advanced drug delivery reviews 101, 34–41 (2016).
Leelananda, S. P. & Lindert, S. Computational methods in drug discovery. Beilstein Journal of Organic Chemistry 12, 2694–2718 (2016).
Seidel, T., Bryant, S. D., Ibis, G., Poli, G. & Langer, T. 3D pharmacophore modeling techniques in computer-aided molecular design using LigandScout (John Wiley & Sons, 2017).
Sperandio, O. et al. MED-SumoLig: a new ligand-based screening tool for efficient scaffold hopping. Journal of Chemical Information and Modeling 47, 1097–1110 (2007).
Yan, X. et al. Chemical structure similarity search for ligand-based virtual screening: methods and computational resources. Current drug targets 17, 1580–1585 (2016).
Debnath, S., Debnath, T., Majumdar, S., Arunasree, M. & Aparna, V. A combined pharmacophore modeling, 3D QSAR, virtual screening, molecular docking, and ADME studies to identify potential HDAC8 inhibitors. Medicinal Chemistry Research 25, 2434–2450 (2016).
Ain, Q. U., Aleksandrova, A., Roessler, F. D. & Ballester, P. J. Machine-learning scoring functions to improve structure-based binding affinity prediction and virtual screening. Wiley Interdisciplinary Reviews: Computational Molecular Science 5, 405–424 (2015).
Miller, M. D., Sheridan, R. P. & Kearsley, S. K. SQ: A program for rapidly producing pharmacophorically relevent molecular superpositions. Journal of Medicinal Chemistry 42, 1505–1514 (1999).
Lešnik, S. et al. LiSiCa: a software for ligand-based virtual screening and its application for the discovery of butyrylcholinesterase inhibitors. Journal of Chemical Information and Modeling 55, 1521–1528 (2015).
Ballester, P. J. & Richards, W. G. Ultrafast shape recognition to search compound databases for similar molecular shapes. Journal of Computational Chemistry 28, 1711–1723 (2007).
Sastry, G. M., Dixon, S. L. & Sherman, W. Rapid shape-based ligand alignment and virtual screening method based on atom/feature-pair similarities and volume overlap scoring. Journal of Chemical Information and Modeling 51, 2455–2466 (2011).
Grant, J. A., Gallardo, M. A. & Pickup, B. T. A fast method of molecular shape comparison: a simple application of a gaussian description of molecular shape. Journal of Computational Chemistry 17, 1653–1666 (1996).
ROCS, OpenEye Scientific Software, Santa Fe, NM. http://www.eyesopen.com.
Yan, X. et al. Enhancing molecular shape comparison by weighted gaussian functions. Journal of Chemical Information and Modeling 53, 1967–1978 (2013).
Li, S., Song, Y., Liu, X. & Li, H. A rapid python-based methodology for target-focused combinatorial library design. Combinatorial chemistry & high throughput screening 19, 25–35 (2016).
Shape-it, Silicos-it: chemoinformatics services and software. http://silicos-it.be.s3-website-eu-west-1.amazonaws.com/
Lagarde, N., Zagury, J.-F. & Montes, M. Benchmarking data sets for the evaluation of virtual ligand screening methods: review and perspectives. Journal of chemical information and modeling 55, 1297–1307 (2015).
Nicholls, A., MacCuish, N. E. & MacCuish, J. D. Variable selection and model validation of 2D and 3D molecular descriptors. Journal of computer-aided molecular design 18, 451–474 (2004).
Jaccard, P. Distribution de la flore alpine dans le bassin des Dranses et dans quelques régions voisines. Bulletin de la Société Vaudoise des Sciences Naturelles 37, 241–272 (1901).
Ding, P. et al. PTS: a pharmaceutical target seeker. Database 2017, bax095 (2017).
Ge, H. et al. Scaffold hopping of potential anti-tumor agents by WEGA: a shape-based approach. Med. Chem. Commun. 5, 737–741 (2014).
Shang, J., Dai, X., Li, Y., Pistolozzi, M. & Wang, L. HybridSim-VS: a web server for large-scale ligand-based virtual screening using hybrid similarity recognition techniques. Bioinformatics 33, 3480–3481 (2017).
Jelásity, M., Ortigosa, P. M. & Garca, I. UEGO, an abstract clustering technique for multimodal global optimization. Journal of Heuristics 7, 215–233 (2001).
Ortigosa, P. M., Garca, I. & Jelásity, M. Reliability and performance of UEGO, a clustering-based global optimizer. Journal of Global Optimization 19, 265–289 (2001).
Solis, F. J. & Wets, R. J.-B. Minimization by random search techniques. Mathematics of Operations Research 6, 19–30 (1981).
Redondo, J. L., Fernández, J., Garca, I. & Ortigosa, P. M. Solving the multiple competitive location and design problem on the plane. Evolutionary Computation 17, 21–53 (2009).
Redondo, J. L., Ortigosa, P. M. & Zilinskas, J. Multimodal evolutionary algorithm for multidimensional scaling with city-block distances. Informatica 23, 601–620 (2012).
Petering, M. E. & Hussein, M. I. A new mixed integer program and extended look-ahead heuristic algorithm for the block relocation problem. European Journal of Operational Research 231, 120–130 (2013).
Ivorra, B. et al. Modelling and optimization applied to the design of fast hydrodynamic focusing microfluidic mixer for protein folding. Journal of Mathematics in Industry 8, 4 (2018).
Fernández, J., G.-Tóth, B., Redondo, J. L. & Ortigosa, P. M. The probabilistic customer’s choice rule with a threshold attraction value: effect on the location of competitive facilities in the plane. Computers and Operations Research 101, 234–249 (2019).
Johnson, R. A. & Bhattacharyya, G. K. Statistics: principles and methods (John Wiley & Sons, 2014).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic acids research 34, D668–D672 (2006).
Huang, N., Shoichet, B. K. & Irwin, J. J. Benchmarking sets for molecular docking. Journal of medicinal chemistry 49, 6789–6801 (2006).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. Journal of Medicinal Chemistry 55, 6582–6594 (2012).
den Haan, H., Morante, J. J. H. & Perez-Sanchez, H. Computational evidence of a compound with nicotinic α4β2-Ach receptor partial agonist properties as possible coadjuvant for the treatment of obesity. bioRxiv (2016).
Ihlenfeldt, W. D., Takahashi, Y., Abe, H. & Sasaki, S.-I. Computation and management of chemical properties in CACTVS: an extensible networked approach toward modularity and compatibility. Journal of chemical information and computer sciences 34, 109–116 (1994).
Wallach, I. & Lilien, R. Virtual decoy sets for molecular docking benchmarks. Journal of Chemical Information and Modeling 51, 196–202 (2011).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. Journal of Chemical Information and Modeling 50, 742–754 (2010).
Wallach, I., Dzamba, M. & Heifets, A. AtomNet: a deep convolutional neural network for bioactivity prediction in structure-based drug discovery. arXiv preprint arXiv:1510.02855 (2015).
Maybridge. Available online: http://www.maybridge.com, (Accessed on 10 october 2018).
Butina, D. Unsupervised data base clustering based on daylight’s fingerprint and tanimoto similarity: A fast and automated way to cluster small and large data sets. Journal of Chemical Information and Computer Sciences 39, 747–750 (1999).
Monge, A., Arrault, A., Marot, C. & Morin-Allory, L. Managing, profiling and analyzing a library of 2.6 million compounds gathered from 32 chemical providers. Molecular Diversity 10, 389–403 (2006).
Pérez-Regidor, L., Zarioh, M., Ortega, L. & Martn-Santamara, S. Virtual screening approaches towards the discovery of toll-like receptor modulators. International Journal of Molecular Sciences 17 (2016).
Fawcett, T. An introduction to ROC analysis. Pattern Recognition Letters 27, 861–874 (2006).
Yan, X., Li, J., Gu, Q. & Xu, J. gWEGA: GPU-accelerated WEGA for molecular superposition and shape comparison. Journal of Computational Chemistry 35, 1122–1130 (2014).
Acknowledgements
This work was funded by grants from the Spanish Ministry of Economy and Competitiveness (TIN2015-66680-C2-1-R, CTQ2017-87974-R), Junta de Andalucía (P12-TIC301), Fundación Séneca–Agencia de Ciencia y Tecnología de la Región de Murcia under Projects 19419/PI/14 and 18946/JLI/13. Powered@NLHPC: This research was partially supported by the supercomputing infrastructure of the NLHPC (ECM-02). The authors also thankfully acknowledge the computer resources and the technical support provided by the Plataforma Andaluza de Bioinformática of the University of Málaga. This work was partially supported by the computing facilities of Extremadura Research Centre for Advanced Technologies (CETA–CIEMAT), funded by the European Regional Development Fund (ERDF). CETA–CIEMAT belongs to CIEMAT and the Government of Spain. Savns Puertas Martn is a fellow of the Spanish ‘Formación de profesorado universitario’ program, financed by the Spanish Ministry of Education, Culture and Sport.
Author information
Authors and Affiliations
Contributions
S. Puertas-Martín, J. L. Redondo, P. M. Ortigosa and H. Pérez-Sánchez contributed equally to this work and discussed the results and implications and commented on the manuscript at all stages.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Puertas-Martín, S., Redondo, J.L., Ortigosa, P.M. et al. OptiPharm: An evolutionary algorithm to compare shape similarity. Sci Rep 9, 1398 (2019). https://doi.org/10.1038/s41598-018-37908-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37908-6
This article is cited by
-
Improving drug discovery through parallelism
The Journal of Supercomputing (2023)
-
A two-layer mono-objective algorithm based on guided optimization to reduce the computational cost in virtual screening
Scientific Reports (2022)
-
Electrostatic-field and surface-shape similarity for virtual screening and pose prediction
Journal of Computer-Aided Molecular Design (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.