Abstract
We here assessed the capability of the MinION sequencing approach to detect and characterize viruses infecting a water yam plant. This sequencing platform consistently revealed the presence of several plant virus species, including Dioscorea bacilliform virus, Yam mild mosaic virus and Yam chlorotic necrosis virus. A potentially novel ampelovirus was also detected by a complimentary Illumina sequencing approach. The full-length genome sequence of yam chlorotic necrosis virus was determined using Sanger sequencing, which enabled determination of the coverage and sequencing accuracy of the MinION technology. Whereas the total mean sequencing error rate of yam chlorotic necrosis virus-related MinION reads was 11.25%, we show that the consensus sequence obtained either by de novo assembly or after mapping the MinION reads on the virus genomic sequence was >99.8% identical with the Sanger-derived reference sequence. From the perspective of potential plant disease diagnostic applications of MinION sequencing, these degrees of sequencing accuracy demonstrate that the MinION approach can be used to both reliably detect and accurately sequence nearly full-length positive-sense single-strand polyadenylated RNA plant virus genomes.
Similar content being viewed by others
Introduction
Metagenomic approaches have enabled the discovery of hundreds previously unknown virus species1,2,3,4. These discoveries have strengthened our understanding of the ecological roles and impacts of viral communities, indicating that viruses are likely essential components of ecosystems as diverse as the human gut5,6 and the oceans7,8. For example, metagenomics approaches that consider the spatial arrangements of plant samples, and the precise environmental contexts of individual sampling sites have revealed the impact of agriculture on the distribution and prevalence of plant viruses at the ecosystem scale9. While the usefulness of viral metagenomics for diagnostics and viral surveillance is still debated10,11,12,13, these approaches have proven effective for the discovery of unknown etiological agents12,14,15. Specifically, in plant virology, viral metagenomics approaches focusing on the analysis of virus-derived small interfering RNAs (siRNA) has gained in popularity for the detection of both known and previously uncharacterized plant viruses within infected plants2,16,17.
Viral metagenomics methodologies, which until now have been primarily based on high-throughput second generation sequencing technologies (Roche 454, Illumina, Ion Torrent, etc.), have generated billions of ~150–300 nucleotides (nt) or of 21–24 nt small RNA (sRNA) reads and, depending on the sequencing coverage, have been successful for detecting and characterizing low-frequency genetic variants within viral populations18. Although viral metagenomics approaches have both expanded our view of global viral biodiversity, and paved the way towards a better understanding of viral ecology and evolution, several methodological biases undermine the effective analysis and taxonomic assignment of the sequence reads and contigs that have been generated using these approaches11,19,20.
One of the most important of these methodological issues relates to the production and analysis of short reads21. Whenever a known related reference sequence is unavailable to guide the mapping of short reads, the reads must be assembled de novo to obtain either full genome sequences, or large genomic sequence “contigs”. These de novo assembled sequences are in most cases likely to be chimaeras of reads from different viral variants22,23: a factor that should exclude these sequences from being used to make phylogenetic and demographic inferences. In addition, differentiation between exogenous and endogenous virus sequences remains difficult to achieve based on de novo assembled short read contigs because the genomic environment of these contigs is usually missing.
A second methodological issue with metagenomic approaches that employ second generation sequencing methods relates to the difficulty of creating scaffolds by chaining reads and contigs together. Failure to build large enough scaffolds or contigs can reduce the proportions of reads that can be reliably identified as being related to known virus species using alignment-based approaches such as BLAST24. This technical problem is compounded by the dearth of viral taxa that are represented in public nucleotide sequence databases such as GenBank. A consequence of this is that, up to 70% or more of the reads that are generated by some environmental viral metagenomic studies, end up being labeled as “dark matter” because they have no detectable homology to sequences within the public nucleotide sequence databases19,25.
Third generation sequencing techniques that are capable of generating much longer reads from individual RNA or DNA molecules promise to eliminate the need to assemble contigs de novo from short sequence reads. These techniques could therefore avoid the problems mentioned above26. Among the third-generation sequencing techniques is that implemented in the Oxford Nanopore MinION. The MinION has recently proved effective for delivering, in real time, average reads lengths of >15 kilobases27,28,29. While the per nucleotide error rate of the MinION long reads is frequently in excess of 20% – substantially higher than that of the shorter Illumina reads30 – the high degrees of genome coverage that are achievable together with post-sequencing data analysis can still yield consensus sequences that are >99% accurate31 and can be effective for identifying recombinant viral variants32. The MinION has been used to detect and determine the full (or nearly full) genome sequences of a range of animal and human viruses, including ebola33, dengue34, zika35, influenza31, cowpox36, and Ross River virus37. The only published applications of the MinION to sequencing plant virus genomes involved the detection of maize streak virus, maize yellow mosaic virus and maize totivirus in maize plants38 and plum pox virus in plum plants39.
Here, we aimed at using the MinION for detecting and characterizing several plant viruses infecting a single yam plant. This sequencing platform proved efficient for detecting Dioscorea bacilliform virus (DBV, Badnavirus genus), yam mild mosaic virus (YMMV, Potyvirus genus) and yam chlorotic necrosis virus (YCNV, Macluravirus genus). The full-length genome sequence of YCNV was determined using a Sanger sequencing approach to enable the comparative quantification of the degrees of sequencing coverage and accuracy using the Illumina and MinION platforms. We show that consensus sequences of YCNV whether obtained by de novo assembly or by mapping of the MinION reads to a reference genome were >99.8% identical to the Sanger-derived sequence of this virus, which indicates that the MinION should in a near future reliably enhance research on, and the monitoring of, plant viruses.
Materials and Methods
Plant material
A water yam plant (Dioscorea alata) infected by YMMV and YCNV was collected in Kerala (India), in 2010 and maintained in vivo at the DSMZ Plant Virus Department (PV-1066).
Small RNA extraction and Illumina sequencing
Small RNA (sRNA) molecules, including small 21–24 nucleotides (nt) interfering RNAs, were extracted from symptomatic leaves of the water yam sample using a RNAzolRT kit (WAK Chemie, Steinbach, Germany) following the manufacturers protocol (RNAzol®RT Brochure, 2010, Molecular Research Center, Inc. Cincinnati, OH). The small RNA fraction was quantified in a Qubit fluorimeter using a Qubit RNA HS Assay Kit (Thermo Fischer Scientific, Waltham, USA) and checked for quality in a bioanalyser 2100 (Agilent). RNA passing the quality check was used for library preparation and subjected to high-throughput sequencing on an Illumina “Hi-Seq 2000” instrument using the services of a commercial company (Fasteris SA, Plan-les-Ouates, Switzerland).
In silico screening of publically available expressed sequence tag (EST) data from yam
A systematic search of viral sequences in de novo assembled sequences (using the CAP3 sequence assembly program40) from publicly available D. alata expressed sequence tag (EST) resources (GenBank accession numbers: HO809681-HO825421, HO825422-HO840419 and HO850622-HO864016) was performed using BLASTn and BLASTx searches against the GenBank nucleotide collection (nt/nr) sequence databases implemented in the software KoriBlast 3.1 (KoriLog, Muzillac, France) with a maximum e-value threshold of 10−3.
Characterization of the full-length genome sequence of yam chlorotic necrosis virus using Sanger sequencing
Total RNA from the water yam sample was extracted using the Qiagen® RNeasy Plant Mini Kit (Qiagen, Valencia, CA) as described by the manufacturer. A set of 30 overlapping primers (Supplementary Table 1) were designed based on an alignment (using MUSCLE41 with default parameters) of the consensus sequence of YCNV obtained using the sRNA Illumina approach, one 2306 nt long sequence with a high degree of similarity (80.7%) to YCNV recovered from D. alata EST sequences obtained from a plant grown in Nigeria42, and sequences of known macluravirus species (Chinese yam necrotic mosaic virus (CYNMV) and yam chlorotic necrotic mosaic virus (YCNMV)). RT-PCR reactions were performed using the Qiagen OneStep RT-PCR Kit (Qiagen, Valencia, CA). The 25 μL RT-PCR reaction mix consisted of 1 μL of eluted RNA, 14 μL of RNAse-free water, 5 μL of RT-PCR buffer (5X), 1 μL of dNTP mix (10 mM), 1.5 μL of each primer (10 μM) and 1 μL of RT-PCR enzyme mix. The RT-PCR program was as follows: 50 °C for 30 min, 95 °C for 15 min, 35 cycles at 94 °C for 1 min, 55 °C for 1 min and 72 °C for 1 or 2 min with a final 72 °C extension for 10 min. PCR products were analyzed by electrophoresis on a 1.2% agarose gel in TAE buffer stained with ethidium bromide and visualized under UV light. Since the extreme terminal ends were not covered, the exact termini were analyzed by 5′- and 3′-RACE. To amplify and sequence the 5′ region of YCNV genome, cDNAs were produced with the virus specific primer YamMac31R (Supplementary Table 1) using SuperScript III Reverse Transcriptase (Thermo Fischer Scientific, Waltham, USA). These cDNAs were subsequently tailed in parallel with polyA, G or C using a recombinant terminal transferase TdT enzyme mini kit (New England Biolabs, Ipswich, USA). The 5′RACE-PCR was performed as described in Knierim et al.43. The sequence of the 3′ end was determined with a 3′RACE-PCR procedure (Supplementary Table 1). Amplicons generated with poly-T primer and virus specific primer YAMMAC4F (Supplementary Table 1) were directly sequenced using automated Sanger sequencing (Genewiz, South Plainfield, USA). Assembly and alignment of the nucleotide sequences and the identification of ORFs was performed as previously described44. The polyA tail was further removed from the YCNV reference sequence to improve the reliability of alignment.
Nanopore MinION library preparation and sequencing
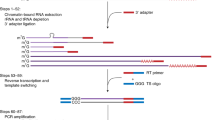
One-hundred mg of yam leaves were ground with a mortar and pestle in liquid nitrogen and 450 µL of RLC buffer (Qiagen) with 0.1% of ß-mercaptoethanol (Sigma Aldrich) were added in a 1.5 ml tube, which was then vortexed twice and kept for five minutes at room temperature. Total RNA was isolated using the RNeasy Plant Mini Kit (Qiagen). A concentration of 25 ng/μl of total RNA was quantified by the Qubit fluorometer (Life technologies). The MinION sequencing library was prepared using the SQK-PCS108 cDNA-PCR kit (Oxford Nanopore Technologies Ltd). Briefly, cDNA was first generated using RT and the strand-switching method (Supplementary Fig. 1) and was then amplified by PCR using primers supplied in the SQK-PCS108 cDNA-PCR kit. Adapters also supplied in the SQK-PCS108 cDNA-PCR kit were then ligated to the PCR product (Supplementary Fig. 1). The eluted library was loaded onto a R9.4 Flow Cell (FLO-MIN106 R9.4). Sequencing was performed using the MinION Mk1B device (MIN-101B). One strand of the duplex was sequenced at a time, producing 1D reads. The flow cell was run for 48 h using the standard MinKNOW software (version 1.11.5) and reads generated were base-called in real-time using the cloud-based Metrichor service provided by Oxford Nanopore Technologies.
Illumina and MinION reads analyses
Illumina and MinION reads and contigs were compared to sequences in the GenBank database using DIAMOND45 searches for MinION reads or BLASTn and BLASTx24 searches for Illumina reads with a maximum e-value threshold of 10−3. Individual macluravirus-related MinION reads were then pairwise aligned with the YCNV Sanger reference genome (hereafter called the reference sequence) using a kmer-based approach (k = 8). Specifically, for each read (hereafter called a query sequence) identified as having detectable homology to the reference sequence, the coordinates of all perfectly matching 8 nt long query sequence fragments were identified on the reference sequence. All perfectly matching 8 nt fragments that were within 20 nucleotides of at least one other perfectly matching 8 nt long fragment on either the query or reference sequences were flagged as being accurately mapped and each of the nucleotides within these fragments was identified as being accurately sequenced (i.e. all eight nucleotides were denoted as “matches”). When 8mer fragments of the query sequence that contained nucleotides that were not denoted as matches in this initial scan (i.e. those nucleotides that were not within any 8 mer fragment that was accurately mapped), were bounded by two accurately mapped 8 mer fragments, the portion of the reference sequence between the mapping coordinates of these accurately mapped fragments were pairwise aligned (using ClustalW46; with default settings) with the unmapped query sequence nucleotides. Matching nucleotide positions, mismatched nucleotide positions and the numbers and lengths of insertions and deletions in the query sequence fragment were then enumerated from these pairwise alignments. The 3′ and 5′ unmapped fragments of the query sequences were not used to enumerate numbers of matching nucleotides, mismatching nucleotides, insertions and deletions.
De novo assembly of Illumina sRNA and MinION sequencing data
Illumina reads cleanups and corrections were performed using the CutAdapt version 1.8.1 program47 and de novo assemblies were generated using SPAdes 3.6.2 software48 and CAP340. MinION reads de novo assembly was done using Canu v1.649 with the suggested parameters for Nanopore R9 1D sequencing (-nanopore-raw overlapper = mhap utgReAlign = true) using reads >1 kb (minReadLength = 1000) and a genome size of 20 kb (genomeSize = 20 k).
Illumina sRNA and MinION sequencing coverage and accuracy
Both sets of reads were independently and jointly mapped to the complete Sanger YCNV genome with Bowtie2 v2.2.950 using the end-to-end and very sensitive option. All reads mapped by Bowtie2 were kept in the analysis. Resulting bam alignment files were then processed with samtools 1.351 to produce mpileup files. Bases with a phred score lower than 13 were filtered out. The resulting mpileup files were then parsed with custom python and R scripts to extract positions and coverages (second and fourth column, respectively), while the similarity was computed using the fifth column of the file, which displayed the numbers of matches, mismatches and indels at each position in the original bam files. Similarity was reported as percentage of match over the coverage. Means of similarity across 100 nt long sliding windows (step size: one nucleotide), were used for clarity of display. Dataset and scripts are available in a github repository at the address: https://github.com/loire/roumagnac2018_figs. Consensus sequences of the YCNV genome were obtained from mapped sRNA Illumina reads, mapped MinION reads, and jointly mapped sRNA Illumina plus MinION reads. Pairwise identity analyses of the reference YCNV Sanger sequence and de novo Minion contigs, mapped sRNA Illumina reads-, mapped MinION reads-, and mapped sRNA Illumina/MinION reads- consensus sequences were carried out using the MUSCLE-based pairwise alignment option implemented in SDT v1.252.
Pairwise identity and phylogenetic analyses
All available macluravirus full genomes were downloaded from GenBank and aligned using MUSCLE41. Maximum-likelihood phylogenetic trees were inferred using PhyML 3.153 implemented in MEGA version 6.0654 with the nucleotide substitution model GTR + G + I selected as best-fit models by jModelTest55. Branch supports were tested using 1000 bootstrap replicates.
Results
Taxonomic assignment of virus-related reads produced by the MinION sequencing technology
With the intention of comparing the potential utility of Illumina and MinION sequencing platforms within a virus discovery context, we attempted to determine the complement of viruses present within a single yam plant using a Nanopore MinION device. In total, 2,036,598 reads were generated of which 41,487 (2%) were identified as likely being virus-derived using DIAMOND searches (Table 1). While most of these viral reads were assigned to YMMV (22,055 reads, 53.2%) and YCNV (19,252 reads, 46,4%), 156 reads (0.4%, 1.2 × 10−4 < DIAMOND e-values < 1.7 × 10−34) were related to DBV, 17 to pestiviruses (0.04%, 7.5 × 10−6 < DIAMOND e-values < 7.4 × 10−15), 5 to bymoviruses (0.01%, 3.8 × 10−19 < DIAMOND e-values < 3.4 × 10−37) and 2 to begomoviruses (0.005%, 8 × 10−4 < DIAMOND e-values < 1.2 × 10−10). Interestingly, DBV is reportedly the most prevalent virus in yam56 and transcriptionally active endogenous geminiviral (EGV) sequences have also been identified integrated within the genomes of many yam species57. The two begomovirus MinION reads recovered in this study share 67% and 89% identities with previously identified EGVs in D. alata. Five reads were assigned to the Bymovirus genus: a sister genus of the Macluravirus genus in the Potyviridae family. These reads may have been misassigned due to the potentially high rate of MinION sequencing errors and may actually be macluravirus-related reads.
De novo assembly of Illumina and MinION reads
A total of 15,365,074 small RNA - including small interfering RNA - raw Illumina sequence reads were generated from the water yam sample. In agreement with the analysis of the MinION reads, 56 contigs >100 nt long that were obtained by de novo assembly of Illumina reads showed significant degrees of similarity to badnaviruses, macluraviruses and potyviruses based on BLASTx searches (Table 2). Overall, these contigs were shorter (maximum length of 2424 nt) than the MiniION reads. While contigs sharing identities to begomoviruses, bymoviruses and pestiviruses were not recovered using de novo assembly of Illumina reads, 4 contigs related to ampeloviruses were identified (5.0 × 10−5 < BLASTx e-values <3.0 × 10−14).
De novo assembly of MinION reads yielded two large contigs. BLASTn comparisons between these two contigs and all sequences in GenBank indicated that the highest similarity scores were detected with YMMV (accession number KJ125479, highest percent nucleotide identity = 85%, query coverage = 99%, e-value = 0) for contig #1 (8791 bp) and YCNV (accession number MG755240, highest percent nucleotide identity = 82%, query coverage = 99%, e-value = 0) for contig #2 (7277 bp).
Characterization of the full-length genome sequence of yam chlorotic necrosis virus
A 8263-nt full length genome sequence of YCNV isolate Kerala (accession number MH341583) was further assembled and annotated, confirming that it possesses a typical macluravirus genome organization (Fig. 1B), including a large open reading frame (ORF) predicted to translate into a 2628 aa long polyprotein (Fig. 1B). This large unique ORF, which starts at position 147 and ends at position 8033, was predicted by analogy with other macluraviruses to encode nine functional products after cleavage, including a helper component-proteinase (HC-Pro), P3 (protein of unknown function), 7 K (protein of unknown function), a cylindrical inclusion putative helicase (CI), 9 K (protein of unknown function), a genome-linked protein (VPg), a nuclear inclusion protease (NIa-Pro), a nuclear inclusion polymerase (NIb) and a coat protein (CP)58,59. Like the other known annotated macluravirus genomes, this virus lacks a P1 proteinase, and is likely to use its HC-Pro and NIa proteinases for polyprotein cleavage. The HC-Pro self-cleavage site between HC-Pro and P3 is likely the FVG/V site located at aa positions 259 to 262 of the polyprotein.
Comparison of the (A) coverage and (C) similarity of MinION and Illumina reads when aligned to the reference yam chlorotic necrosis virus (YCNV) genome sequence. (B) Genome organization of YCNV. The ORFs that are likely to represent genes expressing characteristic macluravirus proteins were identified based on comparisons with other members of the genus Macluravirus, including the helper component-proteinase (HC-Pro), P3, 7 K, the cylindrical inclusion putative helicase (CI), 9 K, the genome-linked protein (VPg), the nuclear inclusion protease (NIa-Pro), the nuclear inclusion polymerase (NIb) and the coat protein (CP).
Based on the alignment of the 2628 aa long polyprotein with artichoke latent virus (ArLV) Chinese yam necrotic mosaic virus (CYNMV) and YCNV isolate YCNV-YJish from China60, five putative NIa proteinase cleavage sites were identified that have the canonical dipeptide Q(E)/M(A) with a conserved L residue at the −2 position from the cleavage site. These potential cleavage sites were located at positions 569 (Q/A, cleaves between P3 and 7 K), 630 (Q/M, 7K-CI), 1289 (Q/A, CI-9K), 1369 (E/M, 9K-NIa), and 2340 (E/M, NIb-CP) of the polyprotein (Fig. 1B). Two additional putative cleavage sites containing noncanonical dipeptides, including E/I (1551, VPg-NIa) and Q/H (1768, NIa-NIb) were also identified. Cleavage of the polyprotein therefore yields mature proteins of the following sizes: HC-Pro (261 aa), P3 (309 aa), 7 K (62 aa), CI (660 aa), 9 K (81 aa), VPg-NIa (400 aa, and 183 for VPg alone), NIb (573 aa) and CP (289 aa). As identified in other macluraviruses, a P3N-PIPO ORF that is classically found in Potyviridae members61 was identified within the P3 ORF. The consensus polymerase slippage GAAAAAA sequence (nt positions 1358 to 1364) is present, followed by a short PIPO region of 59 aa, ending at a stop codon at positions 1538–1540 of the genome.
A phylogenetic tree based on whole genome sequences of known macluraviruses indicated that YCNV isolate Kerala clusters with the other described macluraviruses (Fig. 2A), its closest relatives being YCNV isolate YJish from China and two other macluraviruses isolated from yam: CYNMV and yam chlorotic necrotic mosaic virus (YCNMV)59,62. The virus sharing the highest genome-wide pairwise identity with YCNV isolate Kerala (81.9%) is YCNV isolate YJish (Fig. 2B).
Total error rate assessment of YCNV-related MinION reads
The total mean sequencing error rate of the 19,252 YCNV-related MinION reads was 11.25%, including substitutions, insertions and deletions. In addition, the average numbers of insertions per nucleotide per read and the average number of deletions per nucleotide per read were 0.003% and 0.015%, respectively. Noteworthy, the alignment-based approach that was used for the calculation of the total mean sequencing error rate revealed that several MinION reads were composed of non-contiguous regions of the YCNV genome, suggesting that either defective viral variants were sequenced or that the MinION generated chimeric sequences.
Genomic coverage and genome sequencing depth of YCNV sequences obtained using de novo assembly or mapping of Nanopore and Illumina reads
De novo assembly of the MinION reads, yielded a large contig (contig #2) that covered 87.5% of the full-length YCNV genome and shared 99.889% pairwise identity with the reference Sanger sequence of this genome (Table 3). The 5′ part of the YCNV sequence was missing from this MinION contig. De novo assembly of the Illumina reads yielded 18 contigs that were further chained together to form a single scaffold that covered 88.1% of the full-length YCNV genome and shared 99.739% pairwise identity with the reference Sanger sequence of this genome (Table 3).
A total of 5713 and 293,986 of the MinION and Illumina reads were further mapped to the complete YCNV Sanger genome, respectively. The similarity of the Illumina reads mapped to the reference YCNV genome was higher than the similarity of the mapped MinION reads (Fig. 3A). In addition, the degree of genome sequencing depth was higher for the Illumina technology (615-fold for Illumina vs. 366-fold for MinION, Fig. 3B). The distribution of the Illumina and MinION reads mapped to the reference YCNV genome also differed. While the Illumina reads were quite evenly distributed on the reference sequence with some hot spots of deeper coverage, the MinION reads were predominantly distributed towards the 3′ part of the reference genome (Fig. 1A). In both the MinION and Illumina derived macluravirus genome sequences, sequencing errors were quite evenly distributed across the YCNV genome (Fig. 1C). Mapping of the MinION reads against the Sanger YCNV reference sequence yielded a consensus YCNV sequence that covered 96.5% of the reference genome and shared 99.837% pairwise identity with it (Table 3). On the other hand, mapping of the Illumina reads against the YCNV Sanger reference yielded a consensus that covered the entire YCNV sequence and shared 99.891% pairwise identity with it (Table 3). Finally, mapping both MinION and Illumina reads together against the YCNV reference yielded a consensus sequence covering the entire YCNV genome and sharing 99.927% pairwise identity with it (Table 3).
Discussion
While the MinION sequencing approach has recently proved to be a reliable real-time portable technology that enhances research and monitoring of human and animal viruses31,33,34,35,36,37, to date this technology has seldom been used in plant virology38,39. Here we assessed the capability of this sequencing approach to detect and characterize viruses infecting a water yam plant. While our overarching goal was to assess whether the MinION should be used in plant virology to enhance research on, and the monitoring of, plant viruses, we challenged the MinION technology with the well-established Illumina sequencing technology and compared the results obtained using these two sequencing approaches.
Three viruses, including a badnavirus, a macluravirus, and a potyvirus were detected using both approaches. Other viral sequences were revealed by only one or the other of the two sequencing approaches, including pestivirus- and begomovirus-related sequences revealed by the MinION and ampelovirus-related sequences revealed by Illumina. To date, the Pestivirus genus is not known to contain any plant virus species. Hence, it is plausible that the occurrence of pestivirus-related reads is a consequence of contamination by genetic material from an animal virus. The two begomovirus-related MinION reads may indicate the presence of transcribed endogenous geminiviral sequences57. This is because the MinION protocol developed in this study specifically targeted poly(A) sequences: which would have presumably been plant or plant-associated bacterial/fungal derived polyadenylated mRNA transcripts (Supplementary Fig. 1). Interestingly, it is possible that the analysed yam sample was infected by a novel ampelovirus that went undetected by the MinION approach. Specifically, ampeloviruses have positive sense ssRNA genomes lacking a 3′ poly(A) tract and were probably not amplified using the MinION amplification strategy used in this study (Supplementary Fig. 1) and the transcripts of this non-polyadenylated virus were maybe far too few to be detected by the MinION.
Overall, the present study shows that the MinION is an efficient means of detecting and characterizing positive sense ssRNA viral genomes that have 3′ poly(A) tracts. It also indicates that the de novo assembly of MinION reads can yield nearly complete high-quality genome sequences for plant virus species. However, the study also reveals that plant viruses lacking a 3′ poly(A) tract (e.g. ampeloviruses) could potentially go undetected using the MinION protocol that was applied here. This bias would probably not have been an issue if viral genomes were randomly amplified. Interestingly, a recent study has shown that MinION enables rapid detection of enteric viruses following the ligation of sequencing adapters to randomly pre-amplified viral genomes63. Specifically, viral metagenomics approaches based on random amplification of virion-associated nucleic acids (VANA) purified from virus-like particles have proven effective in the discovery of novel RNA and DNA plant viruses2,64,65, which paves the way towards sequencing all types of plant viruses using the MinION technology. In addition, MinION 1D2 libraries should be used because they offer increased accuracy advantages relative to the existing 1D approach. However, PCR-free MinION detection of plant viruses may be problematic because of how technically difficult and time-consuming it is to purify large-enough quantities of plant virus DNA/RNA from plant tissue samples.
Sanger sequencing of the yam-infecting macluravirus provided a standard against which we could compare the degree of genome coverage and sequencing accuracy achieved by the MinION and Illumina approaches. However, before using the genome of this macluravirus as a guiding sequence, we tentatively taxonomically assigned it. The virus sharing the highest genome-wide pairwise identity with YCNV isolate Kerala (81.9%) is YCNV isolate YCNV-YJish (Fig. 2B). This degree of similarity is above the species demarcation thresholds recommended for all of the different Potyviridae genera (nt sequence identity less than 76% either in the coat protein gene or over the whole genome), suggesting that the water yam plant used in this study is infected by YCNV. While the genome coverage of the mapped Illumina reads was better than that of the mapped MinION reads, the consensus YCNV sequences generated by both approaches shared more than 99.83% identity with the YCNV reference Sanger sequence (Table 3). Combining both the MinION and Illumina reads yielded a consensus YCNV sequence sharing 99.93% identity with the Sanger determined YCNV sequence (Table 3). This degree of accuracy translates to a maximum of six errors over a 8263 nt genome. Considering the known intra-isolate variability of RNA viruses, and that sequences were determined from different plant parts collected at different times for the MinION, Illumina and Sanger sequencing approaches, this “maximum error rate” is remarkably low as some of the identified differences likely reflect actual differences between the sequenced viral populations and not sequencing errors.
Besides this promise, the potential utility of the MinION platform for generating sequencing reads in excess of 10 Kb could have a major impact on viral metagenomics studies. Reads of this length would encompass the genomes of most viruses with RNA or ssDNA genomes. This would correct two of the most important biases that arise during metagenomics studies11,20. First, long reads would solve the assembly chimaera problems that have plagued attempts to use short-read sequencing platforms such as Illumina for sequencing genomes directly from non-clonal viral populations. Secondly, long-read sequencing platforms like MinION will yield smaller numbers of unassigned reads (i.e. will reduce the amount of dark matter) by increasing the efficiency with which reads can be assembled into contigs. While the sizes of these contigs will be larger, the probability that highly variable genes (such as silencing suppressor or movement protein encoding genes) will be accurately linked within contigs to more conserved genes (such as coat proteins, polymerases or replication-associated genes) will be increased.
However, the per-read error rate of the MinION remains very high (most often exceeding 10%) and the total numbers of sequenced nucleotides are lower than Illumina and other shorter reads platforms. Also, it remains to be determined whether the MinION platform is capable of differentiating between exogenous and endogenous viral sequences, between the different components of segmented virus genomes or between viral variants. These limitations could hinder reliable de novo assembly of MinION reads, which might still result in chimeric consensus sequences that mix reads originating from different viral variants. Consequently, there are several areas in which improvement is still needed before sequences generated by this technology can be used reliably for downstream analyses such as to test for evidence of genetic recombination, adaptive evolution, or resistance breaking mutations.
Nevertheless, successfully demonstrating that the MinION approach can be used to both reliably detect and accurately sequence nearly full-length plant virus genomes is an important first step towards the application of highly portable sequencing platforms like the MinION to field-based plant virus diagnostics such as has already been attempted for the identification of closely-related plant species66 and the mobile real-time surveillance of Zika viruses in Brazil67. While field-monitoring of plant viral diseases is a still-missing first step towards identifying and controlling the emergence of new plant diseases, widespread use of highly portable sequencing platforms like the MinION is likely to revolutionize plant disease diagnostics and is already poised to enable the implementation of reliable epidemiosurveillance networks.
References
Dutilh, B. E., Reyes, A., Hall, R. J. & Whiteson, K. L. Editorial: Virus Discovery by Metagenomics: The (Im)possibilities. Front Microbiol 8, 1710, https://doi.org/10.3389/fmicb.2017.01710 (2017).
Roossinck, M. J., Martin, D. P. & Roumagnac, P. Plant Virus Metagenomics: Advances in Virus Discovery. Phytopathology 105, 716–727, https://doi.org/10.1094/PHYTO-12-14-0356-RVW (2015).
Delwart, E. L. Viral metagenomics. Rev.Med.Virol. 17, 115–131 (2007).
Lipkin, W. I. Microbe hunting. Microbiol Mol Biol Rev 74, 363–377 (2010).
Scarpellini, E. et al. The human gut microbiota and virome: Potential therapeutic implications. Digestive and liver disease: official journal of the Italian Society of Gastroenterology and the Italian Association for the Study of the Liver 47, 1007–1012, https://doi.org/10.1016/j.dld.2015.07.008 (2015).
Stecher, B. et al. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc Natl Acad Sci USA 109, 1269–1274, https://doi.org/10.1073/pnas.1113246109 (2012).
Alberti, A. et al. Viral to metazoan marine plankton nucleotide sequences from the Tara Oceans expedition. Sci Data 4, https://doi.org/10.1038/Sdata.2017.93 (2017).
Sunagawa, S. et al. Ocean plankton. Structure and function of the global ocean microbiome. Science 348, 1261359, https://doi.org/10.1126/science.1261359 (2015).
Bernardo, P. et al. Geometagenomics illuminates the impact of agriculture on the distribution and prevalence of plant viruses at the ecosystem scale. The ISME journal 12, 173–184, https://doi.org/10.1038/ismej.2017.155 (2018).
Lewandowska, D. W. et al. Metagenomic sequencing complements routine diagnostics in identifying viral pathogens in lung transplant recipients with unknown etiology of respiratory infection. Plos One 12, e0177340, https://doi.org/10.1371/journal.pone.0177340 (2017).
Massart, S., Olmos, A., Jijakli, H. & Candresse, T. Current impact and future directions of high throughput sequencing in plant virus diagnostics. Virus Res 188, 90–96, https://doi.org/10.1016/j.virusres.2014.03.029 (2014).
Candresse, T. et al. Appearances can be deceptive: revealing a hidden viral infection with deep sequencing in a plant quarantine context. Plos One 9, e102945 (2014).
Nieuwenhuijse, D. F. & Koopmans, M. P. Metagenomic Sequencing for Surveillance of Food- and Waterborne Viral Diseases. Front Microbiol 8, 230, https://doi.org/10.3389/fmicb.2017.00230 (2017).
Feng, H. C., Shuda, M., Chang, Y. & Moore, P. S. Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 319, 1096–1100 (2008).
Schlaberg, R. et al. Viral Pathogen Detection by Metagenomics and Pan-Viral Group Polymerase Chain Reaction in Children With Pneumonia Lacking Identifiable Etiology. J Infect Dis 215, 1407–1415, https://doi.org/10.1093/infdis/jix148 (2017).
Donaire, L. et al. Deep-sequencing of plant viral small RNAs reveals effective and widespread targeting of viral genomes. Virology 392, 203–214 (2009).
Kreuze, J. F. et al. Complete viral genome sequence and discovery of novel viruses by deep sequencing of small RNAs: a generic method for diagnosis, discovery and sequencing of viruses. Virology 388, 1–7 (2009).
Li, J. Z. et al. Comparison of illumina and 454 deep sequencing in participants failing raltegravir-based antiretroviral therapy. Plos One 9, e90485, https://doi.org/10.1371/journal.pone.0090485 (2014).
Rosario, K. & Breitbart, M. Exploring the viral world through metagenomics. Curr Opin Virol 1, 1–9 (2011).
Rose, R., Constantinides, B., Tapinos, A., Robertson, D. L. & Prosperi, M. Challenges in the analysis of viral metagenomes. Virus Evolution 2, vew022–vew022, https://doi.org/10.1093/ve/vew022 (2016).
Wommack, K. E., Bhavsar, J. & Ravel, J. Metagenomics: read length matters. Appl Environ Microbiol 74, 1453–1463, https://doi.org/10.1128/AEM.02181-07 (2008).
Pignatelli, M. & Moya, A. Evaluating the Fidelity of De Novo Short Read Metagenomic Assembly Using Simulated Data. Plos One 6, https://doi.org/10.1371/journal.pone0019984 (2011).
Roux, S., Emerson, J. B., Eloe-Fadrosh, E. A. & Sullivan, M. B. Benchmarking viromics: an in silico evaluation of metagenome-enabled estimates of viral community composition and diversity. Peerj 5, https://doi.org/10.7717/peerj.3817 (2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic Local Alignment Search Tool. Journal of Molecular Biology 215, 403–410 (1990).
Brum, J. R. et al. Illuminating structural proteins in viral “dark matter” with metaproteomics. Proc Natl Acad Sci USA 113, 2436–2441, https://doi.org/10.1073/pnas.1525139113 (2016).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet 17, 333–351, https://doi.org/10.1038/nrg.2016.49 (2016).
Jain, M., Olsen, H. E., Paten, B. & Akeson, M. The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome Biol 17, 239, https://doi.org/10.1186/s13059-016-1103-0 (2016).
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nat Nanotechnol 6, 615–624, https://doi.org/10.1038/nnano.2011.129 (2011).
Pomerantz, A. et al. Real-time DNA barcoding in a rainforest using nanopore sequencing: opportunities for rapid biodiversity assessments and local capacity building. Gigascience 7, https://doi.org/10.1093/gigascience/giy033 (2018).
Greninger, A. L. et al. Rapid metagenomic identification of viral pathogens in clinical samples by real-time nanopore sequencing analysis. Genome Med 7, 99, https://doi.org/10.1186/s13073-015-0220-9 (2015).
Wang, J., Moore, N. E., Deng, Y. M., Eccles, D. A. & Hall, R. J. MinION nanopore sequencing of an influenza genome. Front Microbiol 6, 766, https://doi.org/10.3389/fmicb.2015.00766 (2015).
Sauvage, V. et al. Early MinION (TM) nanopore single-molecule sequencing technology enables the characterization of hepatitis B virus genetic complexity in clinical samples. Plos One 13, https://doi.org/10.1371/journal.pone.0194366 (2018).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228–232, https://doi.org/10.1038/nature16996 (2016).
Yamagishi, J. et al. Serotyping dengue virus with isothermal amplification and a portable sequencer. Scientific reports 7, https://doi.org/10.1038s41598-017-03734-5 (2017).
Quick, J. et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc 12, 1261–1276, https://doi.org/10.1038/nprot.2017.066 (2017).
Kilianski, A. et al. Bacterial and viral identification and differentiation by amplicon sequencing on the MinION nanopore sequencer. Gigascience 4, https://doi.org/10.1186/s13742-015-0051-z (2015).
Batovska, J., Lynch, S. E., Rodoni, B. C., Sawbridge, T. I. & Cogan, N. O. I. Metagenomic arbovirus detection using MinION nanopore sequencing. J. Virol. Methods 249, 79–84, https://doi.org/10.1016/j.jviromet.2017.08.019 (2017).
Adams, I. P. et al. Characterising maize viruses associated with maize lethal necrosis symptoms in sub Saharan Africa. bioRxiv, 161489, https://doi.org/10.1101/161489 (2017).
Bronzato Badial, A. et al. Nanopore Sequencing as a Surveillance Tool for Plant Pathogens in Plant and Insect Tissues. Plant Dis. 102, 1648–1652, https://doi.org/10.1094/PDIS-04-17-0488-RE (2018).
Huang, X. Q. & Madan, A. CAP3: A DNA sequence assembly program. Genome Research 9, 868–877 (1999).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797 (2004).
Narina, S. S. et al. Generation and analysis of expressed sequence tags (ESTs) for marker development in yam (Dioscorea alata L.). BMC Genomics 12, 100, https://doi.org/10.1186/1471-2164-12-100 (2011).
Knierim, D., Maiss, E., Menzel, W., Winter, S. & Kenyon, L. Characterization of the complete genome of a novel Polerovirus infecting Sauropus androgynus in Thailand. Journal of Phytopathology 163, 695–702, https://doi.org/10.1111/jph.12365 (2015).
Palanga, E. et al. Metagenomic-Based Screening and Molecular Characterization of Cowpea- Infecting Viruses in Burkina Faso. Plos One 11(10), e0165188, https://doi.org/10.1371/journal.pone.0165188 (2016).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nature methods 12, 59–60 (2015).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948, https://doi.org/10.1093/bioinformatics/btm404 (2007).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet 17(1), 10 (2011).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19, 455–477, https://doi.org/10.1089/cmb.2012.0021 (2012).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27, 722–736, https://doi.org/10.1101/gr.215087.116 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Muhire, B. M., Varsani, A. & Martin, D. P. SDT: a virus classification tool based on pairwise sequence alignment and identity calculation. Plos One 9, e108277, https://doi.org/10.1371/journal.pone.0108277 (2014).
Guindon, S., Delsuc, F., Dufayard, J. F. & Gascuel, O. Estimating maximum likelihood phylogenies with PhyML. Methods in molecular biology 537, 113–137, https://doi.org/10.1007/978-1-59745-251-9_6 (2009).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Molecular Biology and Evolution 30, 2725–2729 (2013).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nature methods 9, 772, https://doi.org/10.1038/nmeth.2109 (2012).
Seal, S. et al. The prevalence of badnaviruses in West African yams (Dioscorea cayenensis-rotundata) and evidence of endogenous pararetrovirus sequences in their genomes. Virus Res 186, 144–154, https://doi.org/10.1016/j.virusres.2014.01.007 (2014).
Filloux, D. et al. The genomes of many yam species contain transcriptionally active endogenous geminiviral sequences that may be functionally expressed. Virus Evolution 1, 1–17 (2015).
Minutillo, S. A. et al. Complete Nucleotide Sequence of Artichoke latent virus Shows it to be a Member of the Genus Macluravirus in the Family Potyviridae. Phytopathology 105, 1155–1160, https://doi.org/10.1094/Phyto-01-15-0010-R (2015).
Zhang, P. et al. Complete genome sequence of yam chlorotic necrotic mosaic virus from Dioscorea parviflora. Arch Virol 161, 1715–1717, https://doi.org/10.1007/s00705-016-2818-7 (2016).
Lan, P. et al. Complete genome sequence of yam chlorotic necrosis virus, a novel macluravirus infecting yam. Arch Virol, https://doi.org/10.1007/s00705-018-3851-5 (2018).
Chung, B. Y., Miller, W. A., Atkins, J. F. & Firth, A. E. An overlapping essential gene in the Potyviridae. Proc Natl Acad Sci USA 105, 5897–5902, https://doi.org/10.1073/pnas.0800468105 (2008).
Kondo, T. & Fujita, T. Complete nucleotide sequence and construction of an infectious clone of Chinese yam necrotic mosaic virus suggest that macluraviruses have the smallest genome among members of the family Potyviridae. Arch Virol 157, 2299–2307, https://doi.org/10.1007/s00705-012-1429-1 (2012).
Theuns, S. et al. Nanopore sequencing as a revolutionary diagnostic tool for porcine viral enteric disease complexes identifies porcine kobuvirus as an important enteric virus. Scientific reports 8, 9830, https://doi.org/10.1038/s41598-018-28180-9 (2018).
Palanga, E. et al. Metagenomic-based screening and molecular characterization of cowpea-infecting viruses in Burkina Faso. Plos One 11, e0165188 (2016).
Claverie, S. et al. From Spatial Metagenomics to Molecular Characterization of Plant Viruses: A Geminivirus Case Study. Advances in virus research 101, 55–83, https://doi.org/10.1016/bs.aivir.2018.02.003 (2018).
Parker, J., Helmstetter, A. J., Devey, D., Wilkinson, T. & Papadopulos, A. S. T. Field-based species identification of closely-related plants using real-time nanopore sequencing. Scientific reports 7, 8345, https://doi.org/10.1038/s41598-017-08461-5 (2017).
Faria, N. R. et al. Mobile real-time surveillance of Zika virus in Brazil. Genome Med 8, 97, https://doi.org/10.1186/s13073-016-0356-2 (2016).
Acknowledgements
D.P.M. has received a research grant from the National Research Foundation of South Africa. This study was funded by the Agropolis Fondation (E-Space flagship program). Data available at NCBI. GenBank accessions #8263-nt full length genome sequence of yam chlorotic necrosis virus isolate Kerala, Accession Number MH341583. Cleaned Illumina and MinION reads have been deposited in the sequence read archive of GenBank (Accession Number: SRP158898).
Author information
Authors and Affiliations
Contributions
Plant sampling M.L.J., T.M. and S.W. Molecular biology D.F., E.F., L.C., S.G. and S.W. Analysis and interpretation of data D.F., E.L., T.C., S.W., D.P.M. and P.R. Manuscript preparation D.F., E.L., T.C., S.W., D.P.M. and P.R. Study supervision D.F., M.L.J., T.M., S.W. and P.R.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Filloux, D., Fernandez, E., Loire, E. et al. Nanopore-based detection and characterization of yam viruses. Sci Rep 8, 17879 (2018). https://doi.org/10.1038/s41598-018-36042-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-36042-7
This article is cited by
-
Identification of an emerging cucumber virus in Taiwan using Oxford nanopore sequencing technology
Plant Methods (2022)
-
Metagenomic sequencing for detection and identification of the boxwood blight pathogen Calonectria pseudonaviculata
Scientific Reports (2022)
-
How do emerging long-read sequencing technologies function in transforming the plant pathology research landscape?
Plant Molecular Biology (2022)
-
Molecular characterization of Cordyline virus 1 isolates infecting yam (Dioscorea spp)
Archives of Virology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.