Abstract
The rockpool shrimp Palaemon elegans is considered an important crustacean species within the European coastline fauna. This species is experiencing an ongoing geographical expansion beyond its native distribution range due to unintentional human introductions. A better knowledge of the genetic diversity, geographic structure and connectivity of its populations is necessary. In the present study, microsatellite loci were isolated using the Illumina MiSeq platform. The microsatellite-enriched library sequencing produced 3.9 million raw reads. Reads were processed and primer pairs were designed for microsatellite sequences amplification. Ninety-six microsatellite loci were preliminary screened in individuals from Atlantic and Mediterranean localities. From them, 21 loci exhibited reliable polymorphism and were thoroughly characterized in 30 individuals from a Cantabrian locality (Spain). No linkage disequilibrium between pairs of loci was detected. Number of alleles per locus ranged from 2 to 12. Observed and expected heterozygosities ranged from 0.033 to 0.833 and from 0.033 to 0.869 respectively. No significant departure from the Hardy-Weinberg equilibrium was detected in most of loci. This is the first time that microsatellite markers have been developed for P. elegans. This characterized microsatellite suite provides new suitable tools for further analyses, facilitating the understanding of population genetics both in natural and introduced populations.
Similar content being viewed by others
Introduction
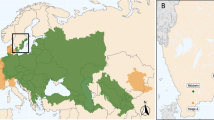
The rockpool shrimp Palaemon elegans Rathke, 1837 is a crustacean decapod with a native geographical distribution ranging from the Atlantic Ocean (from Scotland and Norway to Mauritania, including the Azores, Madeira and Canary Islands) to the entire Mediterranean Sea and the Black Sea1. Nowadays, this shrimp also inhabits the Caspian Sea and the Aral Sea because of unintentional introductions2. Similarly, this species was introduced in the Baltic Sea where it is replacing the native congeneric P. adspersus2,3. Occurrence of P. elegans in the northeast coast of the United States was recently reported4, so its distribution range is currently extended beyond European waters.
This shrimp is common in tidal rockpools and in Zostera, Posidonia and Cymodocea sea grasses meadows. It also can be found in hypersaline lagoons and in slightly brackish water close to river mouths5. Palaemon elegans is characterized by its capability to adapt to highly variable environmental conditions. Therefore, due to its broad ecological niche and its recent and ongoing geographic expansion, it is considered an important species within the European coastline fauna6.
Population genetics analyses in P. elegans are scarce and exclusively based on mitochondrial DNA (mtDNA) markers. Reuschel et al.6 carried out a phylogeographic analysis with two mtDNA markers, revealing the existence of three haplogroups, one of them in the Atlantic localities (type I) and two from the Mediterranean localities (types II and III). Genetic differentiation between the Atlantic populations (type I) and the Mediterranean populations (type II) was observed as well as the putative occurrence of a cryptic species within P. elegans (type III). This population genetic pattern was supported again with mtDNA markers in later phylogeographic studies in the Mediterranean Sea7,8. These findings highlighted the need to carry out further genetic studies with polymorphic nuclear markers, such as microsatellites, in order to clarify the population biology of P. elegans.
Given its codominant nature, biparental mode of inheritance and high levels of polymorphism9, microsatellite markers have been used in a wide range of applications in population genetics, ecological, conservation and evolutionary studies10. Indeed, microsatellites loci are extremely valuable tools in population genetics because they might reveal the existence of genetically isolated populations even in fine-scale studies11. Polymorphic microsatellite loci have been developed for three Palaemon species12,13,14. In the case of P. serratus14, thirteen loci showed positive cross-species amplification in P. elegans. Nevertheless cross-amplification from congeneric species is not generally feasible because inherent problems like allele size homoplasy, polymorphisms biases, null alleles presence, broken repeats motifs or amplification of non-orthologous loci could arise15,16,17,18. Thus, de novo development of species-specific microsatellite markers is strongly recommended.
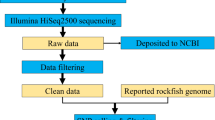
Classically, microsatellite development required the construction of an enriched library by cloning and Sanger sequencing, a laborious, time-consuming and expensive strategy19,20. This drawback could be overcome with the advent of next-generation sequencing (NGS) technologies, which produce a large amount of sequences, providing a faster and cost-effective approach for microsatellite loci discovery21. The first microsatellite markers developed using NGS were based on Roche 45422, however Illumina has demonstrated its capability for microsatellite isolation23,24,25 and it is currently the platform used for this purpose.
The aim of this study was the isolation and characterization of novel polymorphic microsatellite loci for P. elegans using Illumina high-throughput sequencing. Given that there are no microsatellite loci previously developed for this shrimp species, these markers will provide new suitable tools in order to assess the genetic diversity, geographic structure and connectivity of populations at smaller geographic scales, among other possible future applications.
Results and Discussion
The development of microsatellite markers in non-model species has become a rapid and cost-effective process thanks to the advances in high-throughput sequencing technologies26. Thousands of microsatellite loci can be identified in the large amount of sequence data. Illumina is currently the platform used to accomplish microsatellite isolation and it produces more reads at lower prices than Roche 45427. Paired-end sequencing is frequently the preferred strategy for microsatellite isolation as longer reads are generated. In this study the P. elegans microsatellite-enriched library was sequenced using Illumina MiSeq PE, an approach already used for microsatellite discovery in other organisms, e.g. Ewers-Saucedo et al.28, Landínez-García & Márquez29 and Gaeta et al.30. Here, the microsatellite-enriched library sequencing produced 3,902,540 paired-end reads. These paired-end reads were processed and overlapped into 1,766,031 sequences, with a mean length of 171 bp (range: 50–538 bp). Tandem repeats were identified and 500 sequences containing microsatellite motifs were used for primer design. For the preliminary screening, ninety-six out of these 500 primer pairs were tested in five individuals from the different Atlantic and Mediterranean localities that already showed genetic divergence with mitochondrial markers in Reuschel et al.6. From them, primer pairs that produced no amplification or unexpected size PCR products were discarded. Likewise, monomorphic loci were excluded, as well as loci that showed stuttering patterns of ambiguous interpretation. A final suite of 21 microsatellite loci (perfect tri- and tetranucleotide repeats) yielded consistent amplification and reliable polymorphism. Sequences containing these 21 microsatellite markers were deposited in GenBank under accession numbers MH078079-MH078099 (Supplementary Material S1).
The microsatellite loci were characterized through the genotyping of 30 individuals collected in the Santoña locality (Table 1). Successful amplification and scoring was achieved for the 21 microsatellite loci in all individuals. No evidence of linkage disequilibrium between pairs of loci was detected after Bonferroni correction. Hence, all microsatellites were considered as independent markers. Regarding to the genetic variability, the number of alleles per locus ranged from 2 to 12, with a grand mean of 4.6 alleles per locus. The observed heterozygosity (Ho) for each locus ranged from 0.033 to 0.833, and the expected heterozygosity (He) for each locus ranged from 0.033 to 0.869 (Table 1). Ho and He averaged 0.390 and 0.464, respectively. These levels of polymorphisms are in line with those found in the counterpart common littoral shrimp P. serratus in other Atlantic locality14. Significant departures from Hardy-Weinberg equilibrium (HWE) were found in four loci (Pe11, Pe14, Pe18 and Pe19) after sequential Bonferroni correction (Table 1). A heterozygote deficit was detected for these four loci. Similarly, heterozygosity deficiency has been reported in other shrimp species12,14,31,32,33. Among the frequent causes for HWE deviations, inbreeding is quite common. The four loci out of HWE were accompanied with positive FIS values (Table 1). A global FIS value of 0.174 also suggested a heterozygosity deficit in the Santoña locality. However, null alleles could also be invoked to explain heterozygote deficits. Although microsatellite null alleles are widespread, marine invertebrates have demonstrated particularly higher frequencies than other groups34,35. Due to the instability of the flanking sequences of microsatellites, some alleles could not be amplified and consequently dropped out, resulting in a homozygote excess. The four loci that deviated from HWE showed evidence of occurrence of null alleles even though mainly in a low proportion (Table 1). Thus, presence of null alleles is the most likely explanation for those departures from HWE. Overall, most of the 21 loci showed low null allele frequencies (Table 1). Only one locus, the locus Pe11, exhibited a high null allele frequency, >0.2 according to Chapuis & Estoup36. This locus Pe11 is precisely one of the four loci that showed significant deviation from HWE. Therefore, given that the locus Pe11 could be potentially problematic, its inclusion in future analyses might be carefully considered.
Microsatellite markers have proven to be crucial in understanding population genetics and ecology of several shrimp species as Penaeus setiferus37, P. vannamei38, P. monodon39, or Aristeus antennatus40. Assessment of genetic diversity and population structure provides interesting data for conservation and fishery management measures. The microsatellite loci characterized here provide new suitable tools available for the scientific community to study Palaemon elegans population dynamics. Specifically, future studies using these microsatellite loci will shed light about P. elegans phylogeography at smaller geographic scales. In fact, there are ongoing projects that aim to analyze Atlantic and Mediterranean localities along the European coastlines using these markers to address the genetic differentiation between these basins. This microsatellite set will be also greatly useful for inferring the introduction routes and the genetic profiles of the introduced populations of this species in European and North America waters41,42,43.
Material and Methods
Specimens of P. elegans used in this study were collected from Ártabro Gulf (Galicia, Spain, 43°18′48.1″N, 8°33′51.3″W) in 2012 and ethanol-stored at laboratory. Morphological identification was carried out according to González-Ortegón & Cuesta5. Genomic DNA was extracted from 25 mg of muscle abdominal tissue using NZY Tissue gDNA Isolation kit (NZYTech) following the manufacturer’s instructions. Extracted DNA quality and concentration was determined with the NanoDrop ND-1000 spectrophotometrer (Thermo Fisher Scientific).
A microsatellite-enriched genomic library from this sample was constructed at AllGenetics & Biology SL (A Coruña, Spain). The library was prepared using Nextera XT DNA Library Preparation kit (Illumina) following the manufacturer’s instructions. Microsatellite motifs AC, AG, ACG and ATCT were used for the library enrichment. The enriched library was sequenced in the Illumina MiSeq PE300 platform (Macrogen Inc., Seoul, Korea). Reads were processed using Geneious 10.0.544 and in-house developed scripts. Sequences containing microsatellite loci were selected for primer design. Primer pairs were designed using Primer345 for PCR amplification of 500 microsatellite loci, with primers hybridizing at the flanking regions of tandem repeats.
A preliminary panel of 94 primer pairs was screened in individuals from five different Atlantic and Mediterranean localities (one individual per locality): Ré Island (France, 46°11′36.2″N, 1°30′10.2″W), Santoña (Spain, 43°28′04.2″N, 3°28′30.6′′W), Benijo (Spain, 28°34′21.7″N, 16°11′47.0″W), Llança (Spain, 42°21′06.3″N, 3°11′15.1″E) and Collioure (France, 42°31′43.8′′N, 3°05′07.6′′E). All genomic DNA extractions were accomplished following the aforementioned protocol. PCR reactions were performed following Schuelke46 so for each microsatellite locus three primers were used: a specific forward primer, a specific 5′-tailed reverse primer and a fluorescently-labeled oligonucleotide identical to the 5′-tail of the reverse primer. As 5′-tails we used the universal sequences M13 (5′-GGAAACAGCTATGACCAT-3′) and CAG (5′-CAGTCGGGCGTCATCA-3′). M13 oligonucleotides were labeled with the HEX dye, meanwhile CAG oligonucleotides were labeled with the 6-FAM dye. Nested PCR reactions were conducted in a final reaction volume of 12.5 µL, containing 1 µL of DNA (10 ng/µL), 6.25 µL of the Type-it Microsatellite PCR Kit (Qiagen), 4 µL of PCR-grade water, and 1.25 µL of the primer mix (2 µM for both forward primer and HEX-M13 or 6-FAM-CAG oligonucleotides and 0.2 µM in the case of M13- or CAG-tailed reverse primers). The optimal PCR conditions consisted in an initial denaturation step at 95 °C for 5 min, followed by 30 cycles of 95 °C for 30 s, 56 °C for 90 s, 72 °C for 30 s; 8 cycles of 95 °C for 30 s, 52 °C for 90 s, 72 °C for 30 s; and a final extension step at 68 °C for 30 min. Amplification reactions were performed on a T100 Thermal Cycler (Bio-Rad). Fluorescently labeled PCR products were run on a 3130xl Genetic Analysis System (Applied Biosystems) for fragment analysis in the Scientific Research Support Services (University of A Coruña), using GeneScan 500 (−250) ROX internal size standard (Applied Biosystems). Geneious 8.0.544 was used for fluorescent profiles analyzing and allele peaks calling.
All microsatellite loci that yielded consistent amplification and reliable polymorphism were further assessed by genotyping 30 individuals from the Santoña locality (Cantabria, Spain). Singleplex PCRs were carried out as mentioned above. For each locus characterization, number of alleles, observed heterozigosity (Ho), expected heterozigosity (He) and deviations from Hardy-Weinberg equilibrium (HWE) were estimated using GenAlEx 6.547. GENEPOP48 was used to estimate inbreeding coefficient (FIS) and to test linkage disequilibrium between pairs of loci. Significance level was adjusted applying the sequential Bonferroni correction for multiple testing. Null allele frequency was estimated using FreeNA36 and the EM algorithm49.
Accession codes
Sequences containing the microsatellite loci were deposited in GenBank under accession numbers MH078079-MH078099.
Data Availability Statement
The datasets generated and/or analyzed during the current study are available in the GenBank repository (accession numbers MH078079-MH078099) but restrictions apply to the availability of these data, and so they are not publicly available until article publication. Data are however included as Supplementary Information files (Supplementary Material S1).
References
Udekem d’Acoz, C. d’. Inventaire et distribution des crustacés décapodes de l’Atlantique nord-orienal, de la Méditerranée et des eaux continentals adjacentes au nord de 25°N. Collection Patrimoines Naturels (Muséum National d’Histoire Naturelle (S.P.N.)) 40 (1999).
Grabowski, M. Rapid colonization of the Polish Baltic coast by an Atlantic palaemonid shrimp Palaemon elegans Rathke, 1837. Aquat. Invasions. 1, 116–123 (2006).
Katajisto, T., Kotta, J., Lehtiniemi, M., Malavin, S. A. & Panov, V. E. Palaemon elegans Rathke, 1837 (Caridea: Palaemonoidea: Palaemonidae) established in the Gulf of Finland. Bioinvasions Rec. 2, 125–132 (2013).
González-Ortegón, E., Sargent, P., Pohle, G. & Martinez-Lage, A. The Baltic prawn Palaemon adspersus Rathke, 1837 (Decapoda, Caridea, Palaemonidae): First record, possible establishment, and illustrated key of the subfamily Palaemoninae in northwest Atlantic waters. Aquat. Invasions. 10, 299–312 (2015).
González-Ortegón, E. & Cuesta, J. A. An illustrated key to species of Palaemon and Palaemonetes (Crustacea: Decapoda: Caridea) from European waters, including the alien species Palaemon macrodactylus. J. Mar. Biol. Assoc. U.K. 86, 93–102 (2006).
Reuschel, S., Cuesta, J. A. & Schubart, C. D. Marine biogeographic boundaries and human introduction along the european coast revealed by phylogeography of the prawn Palaemon elegans. Mol. Phylogenetics Evol. 55, 765–775 (2010).
Bilgin, R., Utkan, M. A., Kalkan, E., Karhan, S. & Bekbolet, M. DNA barcoding of twelve shrimp species (Crustacea: Decapoda) from Turkish seas reveals cryptic diversity. Mediterr. Mar. Sci. 16, 36–45 (2014).
Deli, T., Pfaller, M. & Schubart, C.D. Phylogeography of the littoral prawn species Palaemon elegans (Crustacea: Caridea: Palaemonidae) across the Mediterranean Sea unveils disparate patterns of population genetic structure and demographic history in the two sympatric genetic types II and III. Mar. Biodiv. 1–23 (2017).
Goldstein, D. B. & Schlotterer, C. Microsatellites—Evolution and Applications. Oxford University Press, Oxford (1999).
De Barba et al. High-throughput microsatellite genotyping in ecology: improved accuracy, efficiency, standardization and success with low-quantity and degradedDNA. Mol. Ecol. Resour. 17, 492–507 (2017).
Wright, J. M. & Bentzen, P. Microsatellites: genetic markers for the future. Mol. Genet. Fish. 1, 117–121 (1995).
Song, K. H., Jung, J. & Kim, W. Polymorphic microsatellite markers of freshwater prawn Palaemon paucidens (De Haan, 1844) (Crustacea: Palaemonidae). Mol. Ecol. Resour. 9, 418–420 (2009).
Jia, S. W., Liu, P., Li, J., Li, J. T. & Pan, L. Q. Isolation and characterization of polymorphic microsatellite loci in the ridgetail white prawn Exopalaemon carinicauda. Genet. Mol. Res. 12, 2816–2820 (2013).
Perina, A., González-Tizón, A. M., Vizcaíno, A., González-Ortegón, E. & Martínez-Lage, A. Isolation and characterization of 20 polymorphic microsatellite loci in Palaemon serratus and cross-amplification in Palaemon species by 454 pyrosequencing. Conserv. Genet. Resour. 8, 169–196 (2016).
Primmer, R. C., Painter, N. J., Koskinen, T. M., Palo, U. J. & Merilä, J. Factors affecting avian cross-species microsatellite amplification. J. Avian Biol. 36, 348–60 (2005).
Barbará, T. et al. Cross-species transfer of nuclear microsatellite markers: Potential and limitations. Mol. Ecol. 16, 3759–3767 (2007).
Rutkowski, R., Sielezniew, M. & Szostak, A. Contrasting levels of polymorphism in cross-amplified microsatellites in two endangered xerothermophilous, obligatorily myrmecophilous, butterflies of the genus Phengaris (Maculinea) (Lepidoptera: Lycaenidae). Eur. J. Entomol. 106, 457–69 (2009).
Yue, G. H., Balazs, K. & Laszlo, O. A new problem with cross-species amplification of microsatellites: generation of non-homologous products. Zool. Res. 31, 131–140 (2010).
Zane, L., Bargelloni, L. & Patarnello, T. Strategies for microsatellite isolation: A review. Mol. Ecol. 11, 1–16 (2002).
Guichoux, E. et al. Current trends in microsatellite genotyping. Mol. Ecol. Res. 11, 591–611 (2011).
Abdelkrim, J., Robertson, B. C., Stanton, J. A. L. & Gemmell, N. J. Fast, cost-effective development of species-specific microsatellite markers by genomic sequencing. BioTechniques. 46, 185–192 (2009).
Schoebel, C. N. et al. Lessons learned from microsatellite development for nonmodel organisms using 454 pyrosequencing. J. Evol. Biol. 26, 600 (2013).
Castoe, T. A. et al. Rapid microsatellite identification from illumina paired-end genomic sequencing in two birds and a snake. PLoS ONE. 7, e30953 (2012).
Lance, S. L. et al. 32 species validation of a new Illumina paired-end approach for the development of microsatellites. PLoS ONE. 8, e81853 (2013).
Duan, C. X., Li, D. D., Sun, S. L., Wang, X. M. & Zhu, Z. D. Rapid development of microsatellite markers for Callosobruchus chinensis using illumina paired-end sequencing. PLoS ONE. 9, e95458 (2014).
Davey, J. W. et al. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 12, 499–510 (2011).
Wei, N., Bemmels, J. B. & Dick, C. W. The effects of read length, quality and quantity on microsatellite discovery and primer development: from Illumina to PacBio. Mol. Ecol. Resour. 14, 953–965 (2014).
Ewers-Saucedo, C., Zardus, J. D. & Wares, J. P. Microsatellite loci discovery from next-generation sequencing data and loci characterization in the epizoic barnacle Chelonibia testudinaria (Linnaeus, 1758). PeerJ. 4, e2019 (2016).
Landínez-García, R. M. & Márquez, E. J. Development and characterization of 24 polymorphic microsatellite loci for the freshwater fish Ichthyoelephas longirostris (Characiformes: Prochilodontidae). PeerJ. 4, e2419 (2016).
Gaeta, J., Acevedo, I. & Machordom, A. Characterization of 17 novel microsatellite loci for the brown spiny lobster Panulirus echinatus Smith, 1869 using MiSeq. Crustaceana. 91, 413–424 (2018).
Pan, Y. W., Chou, H. H., You, E. M. & Yu, H. T. Isolation and characterization of 23 polymorphic microsatellite markers for diversity and stock analysis in tiger shrimp (Penaeus monodon). Mol. Ecol. Notes. 4, 345–347 (2004).
Pereyra, R. T. et al. Isolation and characterization of nuclear microsatellite loci in the northern shrimp, Pandalus borealis. Conserv. Genet. Resour. 4, 109–112 (2012).
Heras, S. et al. Development and characterization of novel microsatellite markers by Next Generation Sequencing for the blue and red shrimp Aristeus antennatus. PeerJ. 4, e2200 (2016).
Hare, M. P., Karl, S. A. & Avise, J. C. Anonymous nuclear DNA markers in the American oyster and their implications for the heterozygote deficiency phenomenon in marine bivalves. Mol. Biol. Evol. 13, 334–345 (1996).
Dailianis, T., Tsigenopoulos, C. S., Dounas, C. & Voultsiadou, E. Genetic diversity of the imperilled bath sponge Spongia officinalis Linnaeus, 1759 across the Mediterranean Sea: Patterns of population differentiation and implications for taxonomy and conservation. Mol. Ecol. 20, 3757–3772 (2011).
Chapuis, M. P. & Estoup, A. Microsatellite null alleles and estimation of population differentiation. Mol. Biol. Evol. 24, 621–631 (2007).
Ball, A. O. & Chapman, R. W. Population genetic analysis of white shrimp, Litopenaeus setiferus, using microsatellite genetic markers. Mol. Ecol. 120, 2319–2330 (2003).
Valles-Jiménez, R., Cruz, P. & Pérez-Enríquez, R. Population genetic structure of pacific white shrimp (Litopenaus vannamei) from Mexico to Panama: microsatellite DNA variation. Mar. Biotechnol. 6, 475–484 (2004).
Waqairatu, S. S. et al. Genetic analysis of black tiger shrimp (Penaeus monodon) across its natural distribution range reveals more recent colonization of fiji and other south pacific islands. Ecol. Evol. 2, 2057–2071 (2012).
Cannas, R. et al. Genetic variability of the blue and red shrimp Aristeus antennatus in the Western Mediterranean Sea inferred by DNA microsatellite loci. Mar. Ecol. 33, 350–363 (2012).
Holland, B. S. Invasion without a bottleneck: microsatellite variation in natural and invasive populations of the brown mussel Perna perna (L). Mar. Biotechnol. 3, 407–415 (2001).
Guillemaud, T., Beaumont, M. A., Ciosi, M., Cornuet, J. M. & Estoup, A. Inferring introduction routes of invasive species using approximate Bayesian computation on microsatellite data. Heredity. 104, 88–99 (2010).
Hoffman, J. R., Morris, T. J. & Zanatta, D. T. Genetic evidence for canal-mediated dispersal of Mapleleaf, Quadrula quadrula (Bivalvia: Unionidae) on the Niagara Peninsula, Canada. Freshw. Sci. 37, 82–95 (2018).
Kearse, M. et al. Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 28, 1647–1649 (2012).
Untergasser, A. et al. Primer3—New capabilities and interfaces. Nucleic. Acids. Res. 40, e115 (2012).
Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 18, 233–234 (2000).
Peakall, R. & Smouse, P. E. Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Resour. 6, 288–295 (2006).
Rousset, F. Genepop’007: a complete re-implementation of the Genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106 (2008).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Series B (methodological). 1–38 (1977).
Acknowledgements
This work was funded by a CTM2014-53838-R grant from the Spanish Government (Ministerio de Economía y Competitividad) and by a GRC2014/050 grant from Xunta de Galicia (Programa de Investigación Competitiva do Sistema Universitario Galego, Modalidade de grupos de referencia competitiva: Grupo de Investigación en Biología Evolutiva). I. González-Castellano and Z. Torrecilla were both supported by a FPU scholarship from Ministerio de Educación, Cultura y Deporte (Spain), FPU15/05265 and FPU13/05673 respectively. A. Perina was supported by a FPI scholarship from Ministerio de Economía y Competitividad (Spain), grant BES-2012-053288. We also thank the anonymous reviewers for their improvements in the manuscript.
Author information
Authors and Affiliations
Contributions
I.G.C., A.P., A.M.G.T. and A.M.L. conceived and designed the experiments. I.G.C. and Z.T. performed the experiments. I.G.C. and A.P. analyzed data. I.G.C. wrote the manuscript. All authors reviewed drafts of the manuscript and approved its final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
González-Castellano, I., Perina, A., González-Tizón, A.M. et al. Isolation and characterization of 21 polymorphic microsatellite loci for the rockpool shrimp Palaemon elegans using Illumina MiSeq sequencing. Sci Rep 8, 17197 (2018). https://doi.org/10.1038/s41598-018-35408-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35408-1
Keywords
This article is cited by
-
A review of the endangered mollusks transcriptome under the threatened species initiative of Korea
Genes & Genomics (2023)
-
Detection and application of novel SSR markers from transcriptome data for Astronium fraxinifolium Schott, a threatened Brazilian tree species
Molecular Biology Reports (2021)
-
Development and characterization of a duplex PCR assay in Chinese sturgeon (Acipenser sinensis) for genetic analysis
Scientific Reports (2020)
-
Genetic diversity and population structure of the rockpool shrimp Palaemon elegans based on microsatellites: evidence for a cryptic species and differentiation across the Atlantic–Mediterranean transition
Scientific Reports (2020)
-
Isolation and characterization of 20 polymorphic microsatellites loci for Xenocypris davidi based on high-throughput sequencing
Molecular Biology Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.