Abstract
The Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES) is a multilateral environmental agreement to ensure that the international trade of threatened species is either prohibited (Appendix I listed species) or being conducted legally, sustainably, and transparently (Appendix II listed species). Twelve threatened shark species exploited for their fins, meat, and other products have been listed under CITES Appendix II. Sharks are often traded in high volumes, some of their products are visually indistinguishable, and most importing/exporting nations have limited capacity to detect illicit trade and enforce the regulations. High volume shipments often must be screened after only a short period of detainment (e.g., a maximum of 24 hours), which together with costs and capacity issues have limited the use of DNA approaches to identify illicit trade. Here, we present a reliable, field-based, fast (<4 hours), and cost effective ($0.94 USD per sample) multiplex real-time PCR protocol capable of detecting nine of the twelve sharks listed under CITES in a single reaction. This approach facilitates detection of illicit trade, with positive results providing probable cause to detain shipments for more robust forensic analysis. We also provide evidence of its application in real law enforcement scenarios in Hong Kong. Adoption of this approach can help parties meet their CITES requirements, avoiding potential international trade sanctions in the future.
Similar content being viewed by others
Introduction
The Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES) is a binding and multilateral environmental agreement with the objective of prohibiting trade of wildlife species in immediate danger of extinction (Appendix I listing) and preventing additional threatened species reaching this critical stage by regulating trade so that it is traceable, legal, and sustainable (Appendix II listing). CITES is the first line of defense against illegal wildlife trafficking, with 183 parties (i.e. signatory nations), and lists over 35,000 animal and plant species1. The vast majority of these species (96%) are listed in Appendix II and exporting parties have the obligation to document that traded specimens are traceable through the supply chain, were legally obtained, and that trade it is not detrimental for the survival of the species2. Importing parties are required to monitor imports and certify that incoming specimens are accompanied by the appropriate documentation2. Failure to properly implement CITES for any listed species can eventually lead to international trade sanctions being placed on offending parties2.
Sharks are a group of marine fish many of which are threatened by overexploitation to satisfy demand for internationally traded products, primarily fins (for use in luxury Asian soup dishes) and meat3,4. Twelve shark species have been listed in CITES Appendix II to date. The first round of shark listings took place from 2001–2004, with the whale shark Rhincodon typus (2001), basking shark Cetorhinus maximus (2001), and great white shark Carcharodon carcharias (2004). These species all share the characteristics of being iconic, extremely large bodied (>4.0 m total length at maturity), and some products from these species (e.g., fins, dressed carcasses) can be readily identified by their very large sizes (except when procured from juvenile stages) or other morphological features (e.g., jaws and teeth for great whites). These species also share the characteristic that Appendix II listing is not as stringent as the domestic protection these species receive in many jurisdictions (i.e., landing and trade prohibition4,5). Products of these species have generally not been detected in post-listing studies of major shark fin and meat markets, suggesting they are rare in trade6,7,8,9,10.
The second round of shark listings took place from 2013–2016, where the porbeagle shark Lamna nasus (2013), scalloped hammerhead Sphyrna lewini (2013), great hammerhead S. mokarran (2013), smooth hammerhead S. zygaena (2013), the oceanic whitetip shark Carcharhinus longimanus (2013), silky shark C. falciformis (2016), bigeye thresher shark Alopias superciliosus (2016), pelagic thresher shark A. pelagicus (2016), and common thresher shark A. vulpinus (2016) were added to Appendix II. This group of species all share the characteristics of being smaller at maturity than white, whale, or basking sharks, and being much more common in trade both pre and post-CITES listing9,10. The silky, scalloped hammerhead, and smooth hammerhead represent the second, fourth, and fifth most common species in the contemporary fin trade9,10. Some of the dried unprocessed fins (e.g., first dorsal, pectoral) and dressed carcasses of these sharks can be identified using morphological characters11,12. Trade in meat, other processed products, and some, less distinctive fin types (e.g., lower caudal) are more challenging to detect without the aid of genetic testing. The Secretariat of CITES recently recommended that parties “share experiences with, and knowledge of, forensic means to efficiently, reliably and cost- effectively identify shark products in trade” (SC69 Doc. 50), including genetic methods.
It is evident that enforcing CITES regulations is a major challenge, in part due to limited availability of rapid, reliable, and cost-effective tools to identify traded products3, especially for the species listed from 2013–201610. Wildlife forensics tools, such as DNA barcoding13,14,15 and PCR-based species-specific assays16,17,18, are the most reliable and widely applicable species identification approaches available to assess and detect illicit activities associated with wildlife trade19,20,21. However, the application of these tools to detect CITES listed species is difficult when inspections must be conducted in a short time frame (e.g. less than 24 hours as is the legal requirement in some locations, such as the United States of America, Hong Kong Special Administrative Region of China), with limited scope for randomly sampling unknown products, transferring tissue samples to a laboratory, and going through all of the steps required for DNA barcoding (i.e., DNA isolation, PCR, product purification, sequencing19) or using available species-specific PCR tests (i.e., DNA isolation, testing for CITES listed species contained in up to four separate multiplex PCR assays, gel electrophoresis19). This is especially problematic given that individual shipments of shark products can be quite large (i.e., 102–103 kg20). Real-time PCR (rtPCR) techniques are an alternative tool that reduce the number of steps to species diagnosis and offers the advantage over DNA barcoding of being able to be conducted outside of the laboratory. Typically, rtPCR techniques have been developed for the quantification of nucleic acids21, and for the detection of pathogen microorganisms22,23,24. This technique uses target-specific primers and fluorescent dyes (e.g. SYBR Green, TaqMan probes) to detect if PCR amplification occurred due to the presence of the targeted nucleic acid template, eliminating the need for sequencing and agarose gels, and enabling a reliable identification of targeted species in real time. In addition, the last stage of rtPCR protocols usually generates melt curves and melt curve temperatures, that can further be used for the identification of the target species/locus25,26. Melt curves are generated by the dissociation of double-stranded DNA during heating, and melt curve temperatures typically vary with the GC/AT ratio of the generated amplicon and its size26.
Here, we present a rtPCR protocol capable of rapidly detecting the presence of nine of the twelve CITES-listed shark species in a single multiplex PCR that could be used as a field-based tool to detect the illegal exportation/importation of these species (i.e. cases of import/export without the correct documentation). The multiplex includes all of the commercially important sharks listed from 2013–2016, with the exception of the oceanic whitetip shark. While the test itself does not discriminate which of the species is present (with a few exceptions based on melt curve analysis) it rapidly and cost effectively (~4 hours for 95 samples; $0.94 USD per sample) provides sufficient information to justify holding shipments based on the illegal presence of CITES listed species. This would provide time for more rigorous forensic testing to verify species-of-origin to meet typical evidentiary standards required for prosecution. We also present three field tests conducted in Hong Kong to inspect the contents of a shark fin consignment in collaboration with local authorities that provide proof-of-concept of the portability of this approach.
Methods
Previously published species-specific primers for CITES-listed species16,17,27,28 were used as a starting point for our multiplex rtPCR assay. The original published studies validated these primers against large, geographically broad samples of the target species, and demonstrated that the primer failed to amplify a large number of non-target species including all or most of the target species’ close relatives (i.e. congeners) through agarose gel electrophoresis18,29.Originally, published primers for hammerhead and thresher shark species were tested on 56 species17,28, great white shark primers on 68 species16, and porbeagle and silky shark primers on 34 species27. Initially, all published primers for these CITES listed species (Table 1) were tested individually with one universal reverse primer against 48 target and non-target species comprising six orders and twelve families (Table 2) to ensure complete specificity of each individual primer when adapted to SYBR Green chemistry. Each 20 μL reaction included, 10 μL of PowerUp SYBR Green Master Mix (Applied Biosystems), 5 μL of primer mix (equal mixture of primers from 10 μM stock solutions), and 5 μL of extracted DNA. DNA extractions were either conducted from dried unprocessed fin, meat, or wet unprocessed fins using Chelex DNA extraction protocols30,31, where a small sample (∼2 mm2) was added to a PCR tube containing 200 μL of 10% Chelex solution and heated for 20 minutes at 60 °C and for 25 minutes at 99 °C, followed by a brief centrifugation and storage at 4 °C. A no-template negative control was included in every extraction and rtPCR batch to control for reagent contamination. Species-specific amplifications were tested against a wide range of annealing temperatures (60°–70 °C) on the QuantStudio5 System (Thermofisher) to determine the most stringent temperature that could be used to amplify the target species and an annealing temperature of 69 °C was selected for further trials. Early trials showed that the silky shark primer, described by27, did not amplify well in a rtPCR format. It also frequently amplified genomic DNA from the spinner shark (C. brevipinna). Thus, we designed a novel species-specific primer for silky sharks (Table 1). We aligned all reference sequences for the internal transcribed spacer 2 (ITS2) deposited in GeneBank by27 in SeaView v432, and designed a putative species-specific primer based on nucleotide differences in this locus. The ITS2 locus was chosen as the target locus for silky sharks because (i) it is highly conserved within shark species, (ii) contains enough nucleotide polymorphisms to achieve species-specific diagnosis and sequence differences between distantly related species are sufficiently large to preclude meaningful alignments, and (iii) all other species-specific primers in our multiplex were designed for this particular locus, thus allowing us to use the same universal reverse primer to amplify the novel primer16,17,27,28. This novel silky shark primer was individually tested for amplification performance in the rtPCR format with silky shark tissues collected over a wide geographic area (Fig. 1) and against all test samples in Table 2. Once we had validated this silky primer, all species-specific primers and one universal reverse primer were combined into a single multiplex of 20 μL including 10 μL of PowerUp SYBR Green Master Mix (Applied Biosystems), 5 μL of primer mix (equal mixture of all primers from 10 μM stock solutions), and 5 μL of extracted DNA. This final multiplex was tested against all test species (Table 2) to detect possible primer interaction that could obstruct the amplification. The thermal cycle profile is shown in Supplementary Fig. S1. Our expected result was that genomic DNA from the nine CITES listed species would amplify and enable SYBR green to fluoresce, while DNA from any other species would not.
We also tested our multiplex rtPCR protocol against processed shark fin samples collected in the Hong Kong retail markets (Table 2) in order to assess its applicability on highly degraded tissue samples. Only one of the nine species present in the multiplex was not tested on processed products (i.e., great white shark), since no samples for this species have been found after more than four years of extensive sampling in Hong Kong and mainland China (Cardeñosa et al. unpublished data). DNA extraction methods, rtPCR protocols, and thermal profiles followed the ones described above.
Three field tests were conducted in Hong Kong in collaboration with the Agriculture Fisheries and Conservation Department (AFCD). During these field tests the QuantStudio5 System, reagents, and plastic consumables were transported to local inspection facilities as a mobile lab. We used the following workflow during our field tests in Hong Kong. After AFCD personnel visually inspected the fins, tissue samples were taken and the QuantStudio5 System was used for Chelex extraction of DNA (~45 minutes), 5 μL of which was then added to the reaction plate. Two wells were used for a positive control using DNA from a previously known scalloped hammerhead sample and a negative control. Once the primer mix and SYBR Green were added, 35 PCR cycles were run with a posterior melt curve analysis (Supplementary Fig. S1). At the conclusion of the PCR and melt curve stages, the melt curve analysis was used to determine which species were present. The field tests involved (i) a container that had sacks of small, unprocessed fins (i.e., <10 cm across the base), (ii) an undeclared shipment of dried seafood that, among other products, included processed shark fins, and (iii) a container that was primarily carrying large unprocessed primary fins (i.e., first dorsal, pectorals, caudal). In the first two cases (i.e., small fins, processed fins) no positive visual identifications were made and we tested our rtPCR protocol against haphazardly selected fins. For the third case (i.e., large unprocessed primary fins), we tested our rtPCR protocol against fins identified as being from CITES listed species based on visual identification protocols (N = 21) and haphazardly selected fins that were thought not to be from these species (N = 35). Since Hong Kong authorities had not yet started to enforce the 2016 shark CITES listings at the time, the species-specific primers to detect the silky and thresher sharks were removed from the multiplex for these specific law-enforcement scenarios. For all of these cases a portion of the mitochondrial COI gene was later amplified and sequenced from unprocessed fins following the protocols by31, and from processed fins following the protocols by15 to assess concordance with the multiplex results. Sequences were compared to BOLD (FISH-BOL) and BLAST (GenBank) databases to identify them to the lowest taxonomic category possible.
Results
Our putative silky shark specific primer performed well individually and in multiplex format, amplifying silky sharks from all over the world (Fig. 1). The final multiplex rtPCR assay containing this primer and the previously published primers for eight additional species correctly detected when any of these CITES-listed species were present and failed to amplify when using genomic DNA templates from all non-target species (Fig. 2). The multiplex proved to be highly reliable in this context, with a false positive rate of 0% with the test samples used (Table 2). There were no differences in the amplification profiles when primers were run individually or as a multiplex protocol. Melt curve temperatures for CITES-listed species occurred between 89° and 91 °C (Fig. 3), making the instantaneous post-PCR identification to species difficult. However, particular melt curve presented consistent shapes that were distinctive for certain species, enabling unambiguous post-PCR identification of S. lewini and A. superciliosus (Fig. 3). Our protocol was also able to consistently detect seven of the eight species tested on processed shark fin samples, with the exception of A. superciliosus, due to its large amplicon (i.e., around 1000 bps).
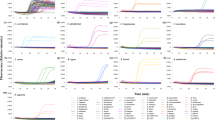
The rtPCR protocol achieved positive identification of fins from CITES listed species (N = 24) across all three field tests in Hong Kong (Fig. 4). All positive results from the rtPCR were subsequently corroborated with DNA barcoding (i.e., there were no false positives). There were four false negative results out of a total of 118 fins tested from CITES listed species, involving one processed fin and three large unprocessed fins. Three of these not only failed with rtPCR but also with DNA barcoding and were only identified as CITES listed species using morphological guides. The barcoding analysis for the first case identified 77% of the samples as milk shark (Rhizoprionodon acutus [55%]), spinner shark (C. brevipinna [16%]), silky shark (C. falciformis [3%]) and scalloped hammerhead (S. lewini [3%]), the latter of which tested positive with the rtPCR. For the second case, species identification was achieved in 92.9% of the tested samples and were identified as blue shark (P. glauca [23.8%]), brownbanded bamboo shark (Chiloscyllium punctatum [21.4%]), bigeye thresher shark (A. superciliosus [19.1%]), scalloped hammerhead (S. lewini [7.1%, all but one of which had tested positive with the rtPCR]), white-spotted guitarfish (Rhinchobatus australiae [7.1%]), spottail shark (C. sorrah [4.8%]), whitecheek shark (C. dussumieri [4.8%]), blacktip shark (C. limbatus [2.4%]), and salmon shark (L. ditropis [2.4%]). Barcoding analysis identified 71.4% of the samples from the third case. All fins that tested positive with rtPCR were CITES-listed smooth hammerhead (S. zygaena [16.1%]), great hammerhead (S. mokarran [8.9%]), and scalloped hammerhead (S. lewini [7.1%]). All of them were correctly visually identified as being from CITES listed hammerheads by AFCD personnel. Three large fins marked as CITES-listed species during the visual inspection failed to amplify with both the rtPCR protocol and the barcoding analysis. Fins that had negative test results with rt-PCR later proved to be non-CITES listed tiger shark (Galeocerdo cuvier [7.1%]), blacktip shark (C. limbatus [3.6%]), snaggletooth shark (Hemipristis elongata [3.6%]), bull shark (C. leucas [1.8%]), silvertip shark (C. albimarginatus [1.8%]), and graceful shark (C. amblyrhynchoides [1.8%]). Barcoding also verified CITES listed oceanic whitetip (C. longimanus [7.1%]) and silky sharks (C. falciformis [1.8%]) that were not part of the multiplex.
Amplification plots for the three field-based tests showing the detection of CITES listed species (red curves) among Non-CITES species (grey curves), a positive control (blue curve) and a negative control (black curve). (A) Shows the amplification plot of the small unprocessed fins case, (B) shows the amplification plot of the processed fins case, and (C) shows the amplification plot of the large unprocessed fins case.
Discussion
Here, we present an easy-to-use, reliable, and to our knowledge, the fastest molecular protocol to date to detect the majority of CITES-listed shark species that are relatively common in the international trade (at least from the perspective of fins9). The multiplex used previously published and extensively tested PCR primers, adapting them to rtPCR format with SYBR Green chemistry. The only exception was the silky shark primer presented in27, which did not work well in this format. The novel silky primer we designed produced a smaller amplicon, promoting more efficient PCR and reliable detection. It amplified 96 known silky shark samples collected from all ocean basins where they occur and failed to amplify any other species, including 22 of 28 described congeners. We did not test the silky primer against all Carcharhinus species and none of the species that we lacked tissue samples for had a sequence in the National Center of Biotechnology Information (NCBI) GeneBank (http://www.ncbi.nlm.nih.gov/genbank/) that would enable us to check for primer mismatches that could prevent amplification. However, the untested Carcharhinus species (n = 6) are phylogenetically more distant to the silky shark than others that we have examined and presented negative results33 (Table 2). Therefore, it is unlikely that the novel primer amplifies other untested Carcharhinus species, especially at an annealing temperature of 69 °C. We also tested against all of the Carcharhinus species that are common in the fin trade, with the untested species occurring only rarely or not occurring at all9. Across all species our rtPCR multiplex generated 0% false positives during the testing phase and field trials. It is likely that the false positive rate of this multiplex will be zero or near zero because of the extensive testing of the published primers16,17,27,28 and the testing we conducted of the novel silky primer. Overall, we contend that the most probable source of false positives is contamination, which can be minimized by rinsing tissues with clean water prior to sampling and by using negative controls. A positive result from this protocol is robust evidence of the presence of one of these CITES listed species, which should be sufficient probable cause to at least hold the product for DNA barcoding that is currently widely accepted in an evidentiary context.
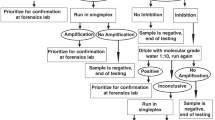
False negative results are unavoidable and no method, genetic or morphological, is entirely immune to them. With our rtPCR assay the frequency of false negative results could vary between individual runs according to the frequency of samples with low DNA quality (e.g., when tested on processed products) or the presence of PCR inhibitors. When the rtPCR multiplex was tested against processed shark fin samples, the bigeye thresher was the only CITES listed species included in the multiplex that failed to amplify, possibly due to the resulting large amplicon (i.e., >1000 bp)15. If false negatives are considered to be a major problem in law enforcement contexts we suggest that all test samples could be run twice: once with the rtPCR multiplex described and once with a protocol expected to amplify all shark products (e.g., a universal shark mini-barcode assay34 or universal ITS2 primers27). Samples that fail to amplify using both approaches could therefore be deemed inconclusive rather than registering as false negatives.
This protocol identifies nine out of twelve CITES-listed shark species, only omitting whale, basking, and oceanic whitetip sharks. The whale and basking sharks are relatively rare in the fin9,10 and meat trade and many of their fins are visually identifiable given their large size and other morphological characters12. The oceanic whitetip is more common in the contemporary shark fin trade9,10 but there are no published primers for this species. Examining the sequences for this species available on GenBank such as ITS2 and cytochrome oxidase I (COI) revealed very high sequence similarity (98%) between this and the dusky and Galapagos sharks (C. obscurus, C. galapagensis), which makes it extremely difficult to design a specific primer (see Shivji et al.27 who noted similar issues). While future efforts will explore other loci, the fins of oceanic whitetips have unique morphological characteristics (e.g. rounded apices with mottled white markings) that make them easily recognizable for law enforcement officers and inspectors12 and reduce the need for them to be in the present multiplex to monitor the fin trade. We conclude that this assay is currently ready-to-use for monitoring the fin trade given that the omitted species all have visually identifiable fins. It is ready-to-use for monitoring the trade of meat and other products, with the caveat that it does not amplify three CITES listed species, two of which are likely to be extremely rare7,10. Future work should aim to develop specific primers for these species that could be integrated into the present multiplex, although this may require surveys of additional loci.
The largest obstacle to routine screening of shark products is cost and time given large shipment volumes typical of the shark trade. The QuantStudio5 System and other rtPCR thermal cyclers have a relatively small footprint and are sufficiently portable, together with the laptop and all necessary reagents, pipettes, and disposable plastics, for this protocol to be deployed in ports or customs inspection areas as shown during our first field tests. The only on-site requisite is a source of electricity and cover from the elements. The speed at which this multiplex can detect these species is a key advance. We suggest the following workflow for sample identification using this multiplex, which can be completed in approximately 4 hours for 94 samples. After taking tissue samples (up to 94 for one run), the QuantStudio5 System can be used for Chelex extraction of DNA (~45 minutes), 5 μL of which is then added to all but one well of a 96 well reaction plate. Once the primer mix and SYBR Green are added, 35 PCR cycles should be run and positive amplifications (i.e., presence of CITES listed species) detected in wells that reach a conservative ΔRn value (i.e. the magnitude of the fluorescent signal generated by the PCR reaction) of 1.00 (Fig. 2), since no non-target species ever crossed that threshold during our testing with the optimized protocol. This threshold is normally reached in 60–70 minutes. We suggest reserving two wells, on a 96 well plate, for a positive control that includes DNA from a known CITES listed shark species in the multiplex that has previously amplified, and a negative control without a template. If the positive control sample fails to amplify it suggests the reagents are compromised. At the conclusion of PCR, the melt curve can be analyzed to determine which species is present (when possible; see Fig. 3), although the presence of any CITES listed species without the proper documentation is illegal and allows for the shipment to be seized and inspected in more detail for prosecution. Given this workflow, the concentrations of reagents and consumables used, the cost to screen one sample with the rtPCR multiplex was $0.94 USD.
Combining this multiplex with morphological identification guides12 enabled highly efficient and cost effective screening of imports at the border for the most common CITES listed sharks in trade from a wide range of potential fin products. The rtPCR protocol also proved to enhance the capacity of law-enforcement officers in Hong Kong to detect illicit trade. First, it achieved detection of CITES-listed species fins that otherwise went undetected because they were outside of the scope of morphological identification guides that are part of the AFCD inspection protocol (e.g., small fins and processed fins in the first and second cases respectively). In the third case it gave them near real time genetic confirmation of suspected CITES-listed shark fins that were flagged after their initial visual inspection, with at least 85.7% of their visual identifications that were tested proving to be accurate. The remainder failed to amplify with the rtPCR protocol and the barcoding analysis, probably due to degraded DNA or PCR inhibitors present in the extraction. Overall, our results highlight the need to implement this new rtPCR protocol to detect the CITES listed species in shipments of fins from small sharks, small secondary fin types (e.g., pelvic, anal, second dorsal fins), and processed fins that have been previously gone uninspected due to the lack of morphological identification protocols. All three cases contained CITES-listed shark fins that were exported without the required permits, aligning with recent evidence of CITES-listed species remaining among the most common in the contemporary shark fin trade in Hong Kong10. Also, the barcoding analysis of the COI locus allowed the confirmation of the CITES species detection by our protocol in all three cases.
The rtPCR approach described here can aid law enforcement officers in major shark trade hubs around the world meet their CITES requirements and can also serve as a model for other monitoring applications for sharks. For example, the International Commission for the Conservation of Atlantic Tunas (ICCAT) prohibits the retention, transhipment, landing, storing, or trade of the three hammerheads and bigeye thresher sharks35, which could be monitored and enforced by member states in port using a modified five-primer version of our multiplex assay. Domestic and regional prohibitions on other species await either the development of a specific multiplex or design of species-specific primers. Usually, rtPCR protocols for quantification purposes (e.g. gene expression21), or using TaqMan probes, are recommended for amplicon sizes of less that 200 bps. We demonstrated that the SYBR Green chemistry allows for amplification and detection of fragments as long as 1000 bps (Table 1), increasing the scope for finding an appropriate region to design many species-specific primers within a locus (e.g., ITS2) and tailor rtPCR assays for sharks according to the specific legal application. Given the widespread detection of illegal trade of meat and fins from sharks and their relatives8,10,36,37,38,39,40 it is imperative that we deploy all available tools, including portable rtPCR field assays, for better monitoring and enforcement of laws intended to protect them.
References
CITES. The CITES species. Available at, http:// www.cites.org/eng/disc/species.php (accessed 31 December 2017) (2012).
CITES. Appendices I, II and III: valid from 25 September 2012. Available at, http://www.cites.org/eng/app/appendices.php (accessed 31 December 2017) (2012).
Dent, F. & Clarke, S. State of the global market for shark products. FAO Fisheries and Aquaculture Technical Paper No. 590. Rome, FAO PP (2015).
Fergusson, I. K., Compagno, L. J. V. & Marks, M. ‘Carcharodon carcharias’, IUCN Red List of Threatened Species. Version 2012.2, available at, www.iucnredlist.org, accessed 17 Jan 2018 (2012).
Techera, E. J. & Klein, N. Sharks: Conservation, Governance and Management. (Abingdon, Oxon; New York: Routledge, 2014).
Jabado, R. W., Ghais, A, S. M., Hamza, W. & Henderson, A. C. The shark fishery in the United Arab Emirates: an interview based approach to assess the status of sharks. Aquatic Conserv: Mar. Freshw. Ecosyst. 25 (2014).
Sembiring, A. et al. DNA barcoding reveals targeted fisheries for endangered sharks in Indonesia. Fisheries Research 164, 130–134 (2015).
Chuang, P.-S., Hung, T.-C., Chang, H.-A., Huang, C.-K. & Shiao, J.-C. The Species and Origin of Shark Fins in Taiwan’s Fishing Ports, Markets, and Customs Detention: A DNA Barcoding Analysis. PLoS ONE 11, e0147290–13 (2016).
Fields, A. T. et al. Species composition of the international shark fin trade assessed through a retail-market survey in Hong Kong. Conservation Biology 32, 376–389 (2017).
Cardeñosa, D. et al. CITES-listed sharks remain among the top species in the contemporary fin trade. Conservation Letters 43, e12457–7 (2018).
FAO. SharkFin Guide: identifying sharks for their fins. 1–144 (2016).
Abercrombie, D., Chapman, D. D., Gulak, S. J. B. & Carlson, J. K. Visual Identification of Fins from Common Elasmobranch in the Northwest Atlantic Ocean. NMFS-SEFSC 643, 1–57 (2013).
Ward, R. D., Zemlak, T. S., In es, B. H., Last, P. R. & Hebert, P. D. N. DNA barcoding Australia’s fish species. Philosophical Transactions of the Royal Society B: Biological Sciences 360, 1847–1857 (2005).
Naylor, G. et al. A DNA sequence-based approach to the identification of shark and ray species and its implications for global elasmobranch diversity and parasitology. Bulletin of the American Museum of Natural History 367, 1–262 (2012).
Cardeñosa, D. et al. A multiplex PCR mini-barcode assay to identify processed shark products in the global trade. PLoS ONE 12, e0185368–9 (2017).
Chapman, D. D. et al. A streamlined, bi-organelle, multiplex PCR approach to species identification: application to global conservation and trade monitoring of the great white shark. Carcharodon carcharias. Conserv Genet 4, 415–425 (2003).
Abercrombie, D. L., Clarke, S. C. & Shivji, M. S. Global-scale genetic identification of hammerhead sharks: Application to assessment of the international fin trade and law enforcement. Conserv Genet 6, 775–788 (2005).
Caballero, S., Cardeñosa, D., Soler, G. & Hyde, J. Application of multiplex PCR approaches for shark molecular identification: feasibility and applications for fisheries management and conservation in the Eastern Tropical Pacific. Molecular Ecology Resources 12, 233–237 (2011).
Abercrombie, D. L., Cardeñosa, D. & Chapman, D. D. Identification of shark fins and other shark products using genetic approaches for international trade monitoring and enforcement. Abercrombie and Fish, Marine Biological Consulting, Suffolk County, NY 1–19 (2018).
Shea, K. H. & To, A. W. L. From boat to bowl: Patterns and dynamics of shark fin trade in Hong Kong — implications for monitoring and management. Marine Policy 81, 330–339 (2017).
Klein, D. Quantification using real-time PCR technology: applications and limitations. Trends Mol Med 8, 257–260 (2002).
Boyle, D. G., Boyle, D. B., Olsen, V., Morgan, J. A. T. & Hyatt, A. D. Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real-time Taqman PCR assay. Dis. Aquat. Org. 60, 141–148 (2004).
Qvarnstrom, Y., Visvesvara, G. S., Sriram, R. & Da Silva, A. J. Multiplex Real-Time PCR Assay for Simultaneous Detection of Acanthamoeba spp., Balamuthia mandrillaris, and Naegleria fowleri. Journal of Clinical Microbiology 44, 3589–3595 (2006).
Kolby, J. E. et al. First Evidence of Amphibian Chytrid Fungus (Batrachochytrium dendrobatidis) and Ranavirus in Hong Kong Amphibian Trade. PLoS ONE 9, e90750–6 (2014).
Rouleau, E. et al. Quantitative PCR high-resolution melting (qPCR-HRM) curve analysis, a new approach to simultaneously screen point mutations and large rearrangements: application to MLH1germline mutations in Lynch syndrome. Hum. Mutat. 30, 867–875 (2009).
Ririe, K. M., Rasmussen, R. P. & Wittwer, C. T. Product differentiation by analysis of DNA melting curves during the polymerase chain reaction. Anal. Biochem. 245, 154–160 (1997).
Shivji, M. et al. Genetic identification of pelagic shark body parts for conservation and trade monitoring. Conservation Biology 16, 1036–1047 (2002).
Abercrombie, D. Efficient PCR-Based Identification of Shark Products in Global Trade: Applications for the Management and Conservation of Commercially Important Mackerel Sharks (Family Lamnidae), Thresher Sharks (Family Alopiidae) and Hammerhead Sharks (Family Sphyrnidae). Master’s thesis. Nova Southeastern University. Retrieved from NSUWorks, Oceanographic Center. (131), http://nsuworks.nova.edu/occ_stuetd/131 (2004).
Clarke, S. C., Magnussen, J. E., Abercrombie, D. L., McAllister, M. K. & Shivji, M. S. Identification of shark species composition and proportion in the Hong Kong shark fin market based on molecular genetics and trade records. Conservation Biology 20, 201–211 (2006).
Hyde, J. R. et al. Shipboard identification of fish eggs and larvae by multiplex PCR, and description of fertilized eggs of blue marlin, shortbill spearfish, and wahoo. Marine Ecology Progress Series 286, 269–277 (2005).
Cardeñosa, D., Hyde, J. & Caballero, S. Genetic Diversity and Population Structure of the Pelagic Thresher Shark (Alopias pelagicus) in the Pacific Ocean: Evidence for Two Evolutionarily Significant Units. PLoS ONE 9, e110193–12 (2014).
Gouy, M., Guindon, S. & Gascuel, O. SeaView Version 4: A Multiplatform Graphical User Interface for Sequence Alignment and Phylogenetic Tree Building. Mol. Biol. Evol. 27, 221–224 (2010).
Naylor, G. et al. In Biology of Sharks and Their Relatives, Second Edition (eds Carrier, J. C., Musick, J. A. & Heithaus, M. R.) 20123460, 31–56 (CRC Press, 2012).
Fields, A. T., Abercrombie, D. L., Eng, R., Feldheim, K. & Chapman, D. D. A Novel Mini-DNA Barcoding Assay to Identify Processed Fins from Internationally Protected Shark Species. PLoS ONE 10, e0114844 (2015).
ICCAT. Resolutions, Recommendations and Other Decisions, available at, www.iccat.int/en/Recs-Regs.asp, accessed 26 January 2017 (2011).
Alexandre de-Franco, B., Fernandes Mendonça, F., Oliveira, C. & Foresti, F. Illegal trade of the guitarfish Rhinobatos horkelii on the coasts of central and southern Brazil: genetic identification to aid conservation. Aquatic Conserv: Mar. Freshw. Ecosyst. 22, 272–276 (2012).
Liu, S. Y. V., Chan, C. L. C., Lin, O., Hu, C. S. & Chen, C. A. DNA Barcoding of Shark Meats Identify Species Composition and CITES-Listed Species from the Markets in Taiwan. PLoS ONE 8, e79373 (2013).
Asis, A. M. J. M., Lacsamana, J. K. M. & Santos, M. D. Illegal trade of regulated and protected aquatic species in the Philippines detected by DNA barcoding. Mitochondrial DNA 27, 659–666 (2014).
Tolotti, M. T., Bach, P., Hazin, F., Travassos, P. & Dagorn, L. Vulnerability of the Oceanic Whitetip Shark to Pelagic Longline Fisheries. PLoS ONE 10, e0141396–17 (2015).
O’Malley, M. P., Townsend, K. A., Hilton, P., Heinrichs, S. & Stewart, J. D. Characterization of the trade in manta and devil ray gill plates in China and South-east Asia through trader surveys. Aquatic Conserv: Mar. Freshw. Ecosyst. 27, 394–413 (2016).
Acknowledgements
This work was supported by Paul G. Allen Philanthropies (to D.D.C. and D.C.). Thanks to COLCIENCIAS-Colombia for supporting D.C. during his PhD studies. We would also thank Daniel Leddy, Karen Hiza, and the Thermofisher Tech. Support team for their assistance and advice during the development stage of this study. We are also grateful to the Agriculture Fisheries and Conservation Department (AFCD) and Customs and Excise Department in Hong Kong for collaborating with us on the field trials of this protocol.
Author information
Authors and Affiliations
Contributions
D.D.C., conceived the experiment. D.D.C and D.C. obtained funding. D.C., J.Q., conducted the analysis in the lab and analyzed the results. D.D.C., K.H. Shea organized the field tests. D.C. conducted the field tests. D.C. and D.D.C. wrote the manuscript. All authors provided comments and edits to the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cardeñosa, D., Quinlan, J., Shea, K.H. et al. Multiplex real-time PCR assay to detect illegal trade of CITES-listed shark species. Sci Rep 8, 16313 (2018). https://doi.org/10.1038/s41598-018-34663-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-34663-6
Keywords
This article is cited by
-
A tangled web: global review of fishing interactions with rhino rays
Reviews in Fish Biology and Fisheries (2024)
-
A new multiplex qPCR assay to detect and differentiate big cat species in the illegal wildlife trade
Scientific Reports (2023)
-
DNA-based species identification of shark fins traded in thai markets
Conservation Genetics (2023)
-
Multiplex and on-site PCR detection of swine diseases based on the microfluidic chip system
BMC Veterinary Research (2021)
-
From molecule to conservation: DNA-based methods to overcome frontiers in the shark and ray fin trade
Conservation Genetics Resources (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.