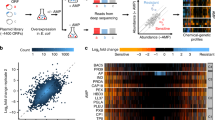
Abstract
Antimicrobial peptides (AMP) are highly conserved immune effectors across the tree of life and are employed as combinations. In the beetle Tenebrio molitor, a defensin and a coleoptericin are highly expressed in vivo after inoculation with S. aureus. The defensin displays strong in vitro activity but no survival benefit in vivo. The coleoptericin provides a survival benefit in vivo, but no activity in vitro. This suggests a potentiating effect in vivo, and here we wanted to investigate the effects of this combination on resistance evolution using a bottom-approach in vitro starting with a combination of two abundant AMPs only. We experimentally evolved S. aureus in the presence of the defensin and a combination of the defensin and coleoptericin. Genome re-sequencing showed that resistance was associated with mutations in either the pmt or nsa operons. Strains with these mutations show longer lag phases, slower Vmax, and nsa mutants reach lower final population sizes. Mutations in the rpo operon showed a further increase in the lag phase in nsa mutants but not in pmt mutants. In contrast, final MICs (minimum inhibitory concentrations) do not differ according to mutation. All resistant lines display AMP but not antibiotic cross-resistance. Costly resistance against AMPs readily evolves for an individual AMP as well as a naturally occurring combination in vitro and provides broad protection against AMPs. Such non-specific resistance could result in strong selection on host immune systems that rely on cocktails of AMPs.
Similar content being viewed by others
Introduction
In antibiotic treatments, often single drugs are successfully used to clear infections. Yet, in innate immune systems, infections typically result in the expression and release of cocktails of antimicrobial peptides (AMP)1,2,3, even though individual antimicrobial peptides can be very potent4. Possible evolutionary explanations are physiological cost savings, for example, if immune effectors synergize or potentiate and hence reduce the total amount of effectors required for clearance5,6. Other possible explanations are that resistance evolution has a lower probability if bacteria are under selection from multiple antimicrobials7 or a high probability of mixed infections.
An interesting observation is that during an infection antimicrobial peptides are expressed that have no known activity against the agent of infection. This is surprising, given that insects for example have different receptors that can distinguish between classes of infectious microbes such as fungi or bacteria8 and energetic costs of protein synthesis are considered to be high3. In the mealworm beetle Tenebrio molitor experimental infection with S. aureus induces the expression of at least ten antimicrobial peptides for a week3. A proteomic analysis shows that the majority of these inducible antimicrobial peptides remain elevated in the haemolymph after three weeks9. While some of them such as Tenecin 1, a defensin, show high activity against S. aureus in vitro, other abundant AMPs such as Tenecin 2, a coleoptericin, displays no activity against S. aureus10. The knock-down, however, of tenecin 2 led to highly increased mortality of T. molitor 3 days after S. aureus infection11. This clearly indicates that Tenecin 2 has an unknown role or activity in combination with other AMPs, of which Tenecin 1 is highly abundant9.
Here we want to use the simplest approach of understanding combinations of AMPs as observed in hosts, by using in vitro experimental evolution of bacteria to start with the minimum number in a combination: two. We explore if this combination of two highly abundant AMPs influences the evolution of bacterial resistance. We focus on the level and costs of resistance, possible underlying mutations and cross-resistance against other AMPs.
In a recent experimental evolution study investigating S. aureus resistance evolution against three different AMPs from different organisms (melittin, pexiganan and iseganan)12, we found a range of mutations associated with resistance13. These mutations were not specific for different AMPs. These results are in line with a growing number of studies that show a high probability of cross-resistance against AMPs14,15.
Bacteria have evolved a number of resistance mechanisms that provide protection against ubiquitous AMPs. As most AMPs target the cell envelope and are often cationic, most described resistance mechanisms are related to changes in the cell wall such as altering the net cell surface charge16. In the case of S. aureus, the antimicrobial peptide sensing system GraRS17 regulates the dlt operon, which controls D-alanylation of the wall teichoic acids18 and mprF expression, which is responsible for peptidoglycan lysinylation16 as well as the Bce-type ABC transporter vreFG which confers broad-spectrum AMP resistance. Two additional Bce-type ABC transporters, BraDE and vraDE, are under the control of the nisin susceptibility-associated two-component system NsaSR (also known as BceS/BceR19and BraS/BraR20). In short, the number of available resistance mechanisms against AMPs seems to be limited providing one possible explanation for the findings on cross-resistance.
Here we explore for the first time resistance evolution against a pair of naturally co-expressed AMPs rather than artificial combinations as used previously. We ask if this combination changes the outcome of resistance evolution with respect to level and costs of resistance, possible underlying mutations and cross-resistance against other AMPs. Using an experimental evolution protocol, we selected S. aureus for resistance against the T. molitor defensin Tenecin 1, either alone or in combination with Tenecin 2. We then investigated if resistance evolution results in fitness costs and if the presence of Tenecin 2 changes the outcome of resistance evolution against the potent Tenecin 1. The resulting strains were re-sequenced to identify mutations associated with AMP resistance and to study the degree of parallel evolution.
Methods
Bacterial strains and culturing conditions
Staphylococcus aureus strain SH1000 and Escherichia coli strain MG1655 were used in the experiments. Bacterial cultures were grown in non-cation adjusted Mueller Hinton Broth (MHB) (Panreac Applichem GmbH) at 37 °C with mild shaking and plated on Mueller Hinton Agar (MHA), unless stated otherwise.
Antimicrobial peptides
Mealworm Tenebrio molitor antimicrobial peptides Tenecin 1 (VTCDILSVEAKGVKLNDAACAAHCLFRGRSGGYCNGKRVCVCR) and Tenecin 2 (SLQPGAPSFPGAPQQNGGWSVNPSVGRDERGNTRTNVEVQHKGQDHDFNAGWGKVIKGKEKGSPTWHVGGSFRF) were chemically synthesised by Peptide Protein Research Ltd (Funtley, UK). To avoid multiple freeze-thaw cycles and prevent binding of the peptides to the vials during storage, peptides were re-suspended in sterile water to the final peptide concentration of 5 mg/ml and glycerol concentration 50% and stored at −20 °C in sterile glass vials, which were pre-treated with “Piranha” solution (3 parts of concentrated sulfuric acid and 1 part of 30% hydrogen peroxide solution).
Selection experiment
Prior to selection, bacteria were pre-adapted to the experimental conditions (following;21). For this, three randomly selected clones of S. aureus strain SH1000 were picked from a Tryptic Soy agar plate, inoculated individually into 10 ml MHB and incubated overnight with shaking at 37 °C. To mimic experimental conditions, the cultures were then diluted 1:1000 and incubated at 37 °C without agitation in 50 ml polypropylene Falcon tubes containing 3.7 ml MHB. The specific volume of MHB used for pre-adaptation was calculated to ensure the same surface-area-to-volume ratio as in 96-well plates that were used in the evolution experiment. Pre-adaptation was carried out as described above by serial passage every 24 hours (to allow for approximately 36 doublings) for 8 days, with daily measurements of optical density at 600 nm, contamination checks by plating out on MHA and cryopreservation of culture aliquots at −80 °C in 12% glycerol solution. One “ancestor” line was used to establish the initial MIC value for Tenecin 1 (see below).
For the selection protocol, five independent parallel selection lines (numbered 1 to 5) were founded by plating one pre-adapted “ancestor” line on MHA and isolating five random colonies. The experiment was performed at 37 °C without shaking in a microplate reader (Synergy 2, Biotek). We used flat bottom polypropylene non-binding 96-well plates (Greiner Bio-One GmbH, Germany) to avoid attachment of the peptides to the plastic surfaces covered with clear polystyrene lids with condensation rings (Greiner Bio-One GmbH, Germany). The plates were filled with MHB, with the total volume of 200 µl per well. Growth curves were generated by taking measurements of OD600 every 30 minutes (preceded by a brief shaking for 10 seconds) for 23 hours. For each of the five replicate lines there were two experimental conditions – Tenecin 1 or a combination of Tenecin 1 + Tenecin 2, as well as four controls - negative control (culture medium control), two glycerol controls - for each AMP treatment - to account for the increasing concentrations of glycerol at higher concentrations of peptides, in which they were stored, and a non-selected control. The serial passage started at 1/2 × MIC, which corresponded to 4 µg/ml for Tenecin 1 and 4 µg/ml Tenecin 1 and 8 µg/ml Tenecin 2 for the combined treatment. To inoculate the treatment wells and the respective control wells, overnight cultures of the five replicate lines were diluted 1:100 and sub-cultured until OD600 0.5 (corresponding to 1 × 108 cfu/ml), then 10 µl of these cultures were inoculated into each treatment and control (except negative control) wells resulting in the final total volume of 200 µl and bacterial density of approximately 5 × 106 cfu per well. Two percent of the culture were transferred every 24 hours to a fresh 96 well plate and the concentration of AMPs was doubled. Twenty µl of the remaining cultures were added to 180 µl of sterile 0.9% NaCl and then serially diluted (typically from 10−1 to 10−5) and checked for contamination and viable counts by plating 5 µl of each dilution on MH-agar using the drop plate method22 and incubating the plates overnight at 30 °C. Glycerol was added to the rest of the cultures to the final concentration of 12% and the plates were stored at −80 °C. The selection experiment continued for 7 daily passages, in each of which the concentration of AMPs was doubled reaching 256 µg/ml for Tenecin 1 and 512 µg/ml Tenecin 2 on the last day of the experiment (day 7). When the volume of added peptide started to exceed 5% of the total volume of medium per well, 2-fold concentrated MHB was used to prepare stock solutions of the desired concentration to alleviate the possible effects of nutrients depletion. The resulting resistant lines were used for subsequent assays (MIC, growth curves) and sequencing both as populations and colonies.
Antimicrobial susceptibility testing
Minimal inhibitory concentration (MIC) was determined using a broth micro-dilution method23. Briefly, 5 µl (1 × 105 cfu/ml) of the mid-exponential phase bacterial culture diluted 1:100 were inoculated into the wells of polypropylene V-bottom 96-well plates (lids with condensation rings 656171, both from Greiner Bio-One GmbH, Germany, Germany) containing two-fold serial dilutions of AMPs or antibiotics in the total volume of 100 µl MHB per well. Each assay was performed in triplicate. The plates were incubated at 37 °C in a humidity chamber. The MIC was defined as the lowest concentration that inhibited visible bacterial growth after 24 hours of incubation. This standard method was used to determine the initial MIC of Tenecin 1 for the pre-adapted “ancestor” line and mutant lines and antibiotics. Because Tenecin 2 is mostly active against gram-negative bacteria, Escherichia coli strain MG1655 was used to determine the activity of this peptide.
To determine the MICs of Tenecins immediately after the selection experiment, we scaled-down the assays to the total volume of 40 µl per well because of the high number of bacterial lines and the limited amount of the antimicrobial peptide available. For this, we used flat-bottom 384-well polypropylene plates (Greiner Bio-One GmbH, Germany) and clear polystyrene lids (Greiner Bio-One GmbH, Germany). We determined MIC as the lowest concentration at which OD600 readings were indistinguishable from those of the NTC wells. As a method control, we compared the MIC determined using the standard and the small-volume protocol, and found no differences. The MICs were determined for populations and individual colonies derived from the selection lines.
Growth curves
Growth curve assays were performed by monitoring the changes in turbidity at OD600 of the selected mutant lines, non-selected controls and ancestor in un-supplemented MHB using a microtitre plate reader. For this, bacterial lines (populations and colonies) were grown until OD600 0.5, diluted 1:10 and 20 µl of the resulting cell suspension were inoculated into 180 µl MHB. Each assay had four replicates. The measurements were taken at 20 minutes intervals during 16 hours of incubation at 37 °C inside the microtitre plate reader, with 10 seconds shaking before each reading. Growth parameters such as final and maximum OD, the maximum growth rate (Vmax) and lag time (for a rationale using these)24,25,26 were calculated with Gen5 software (Biotek).
DNA isolation
Genomic DNA for whole genome sequencing was isolated using GeneMATRIX Bacterial and Yeast genomic DNA purification kit (Roboklon, Germany) following manufacturer’s instructions. Four µl of 10 mg/ml freshly prepared lysozyme and lysostaphin (both from Sigma) each were added into bacterial lysate. The DNA quantity and quality were estimated by measuring the optical density at A260/280 using the Nanodrop spectrophotometer (Thermo Scientific) and agarose gel electrophoresis.
Genome re-sequencing
TruSeq DNA PCR-free libraries were constructed according to the manufacturers instructions and sequenced for 600 cycles using a MiSeq at the Berlin Center for Genomics in Biodiversity Research.
The genetic differences between strain SH1000 and other members of the 8325 lineage have been described using array-based resequencing27, and de novo genome sequencing28. The differences comprise: the excision of three prophages from 8325 (Φ11, 12, 13), 13 single-nucleotide polymorphisms (SNPs; two synonymous, 11 nonsynonymous), a 63-bp deletion in the spa-sarS intergenic region, and an 11-bp deletion in rsbU28. To account for these differences we first assembled reads from SH1000 using SPAdes29, and used the resulting contigs to correct the three phage excision sites in the 8325 reference genome. SH1000 reads were then mapped to the resulting sequence and bcftools consensus30 was used to correct the remaining 13 SNPs and two indels.
The haploid variant calling pipeline snippy31 was used to identify mutations in the selection lines. Snippy uses bwa30 to align reads to the reference genome and identifies variants in the resulting alignments using FreeBayes (Garrison and Marth 2012). All variants were independently verified using a second computational pipeline, breseq32.
Statistical analyses
Statistical analyses were performed using R version 3.4.1. Growth parameters (Vmax, duration of lag phase, final OD600 and log2 (MIC)) were analysed by using the nlme package33 to fit linear mixed-effects models specifying line as a random effect. Differences between groups were analysed with contrasts. We also used visreg34 and Ismeans35. Permutational analyses of Jaccard distance were performed using the vegan package36.
Results
Resistance evolves readily at a cost
Over the course of selection all replicates of both treatments (single and combination of AMPs) readily evolved a high level of resistance. All lines were able to grow in the presence of 256 µg/ml Tenecin 1 and 512 µg/ml Tenecin 2, resulting in an increase in MIC for Tenecin 1 (after selection with either Tenecin 1 or Tenecin 1 + 2, T = −9.98, df = 35, p = < 0.0001, Fig. 1). MICs did not differ between Tenecin 1 and Tenecin 1 + 2 lines (T = 0.894, df =35, p =0.3776). When comparing bacterial growth curves of strains isolated either from the selected lines or non-selected controls, we found consistently slower growth rates in the exponential phase for both Tenecin 1 (T = −6.8, df = 56, p = < 0.0001, Fig. 1) and Tenecin 1 + Tenecin 2-selected strains (T = −7.17, df = 56, p = < 0.0001, Fig. 1) and also extended lag phases (Tenecin 1: T = 9.4, df = 56, p = < 0.0001, Tenecin 1 + Tenecin 2: T = 9.1, df = 56, p = < 0.0001, Fig. 1). Tenecin 1 and Tenecin 1 + Tenecin 2 selected strains did not differ (vmax: T = −0.67, df = 56, p = 0.5042, lag: T = 0.012087, df = 56, p = 0.9904, Fig. 1). Final population sizes measured as OD did not differ (Tenecin 1: T = −0.57419 df = 54, p = 0.5682 Tenecin 1 + 2: T = −1.70549, df = 54, p = 0.0938). This suggests that AMP resistance incurs fitness costs irrespective of the type of selection.
Cross-Resistance to other AMPs but not antibiotics
We tested for cross-resistance against commercially available AMPs, colistin (T = −3.826, df = 27.94, p = 0.0007), and melittin (T = −7.669, df = 27.94, p < 0.0001). We found 2–8-fold cross-resistance of all AMP-selected strains against the AMPs Mellitin and Colistin (Table S1). The selected strains showed no cross-resistance against Vancomycin, a drug of last resort for staphylococcal infections with multiple resistances. There was no relationship between the AMP cross-resistance and the resistance mutations (see Table S2). We also tested the evolved strains against a panel of eight antibiotics (see Table S2) but detected no cross-resistance.
Genome re-sequencing reveals mutations in a limited number of loci
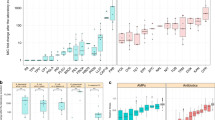
Whole genome sequencing of the selected mutants and the respective controls (at the population and single colony levels) showed differences both between treatments and between replicate lines within treatments. In each resistant strain one mutation was identified in either pmtR, pmtA, nsaS, or nsaR gene, all of which are known to be involved in envelope stress tolerance (see also Tables S3 and S4 for a full list of mutations). In case of the pmtR mutations, there were examples of missense, frameshifts, and stop gains. For pmtA, nsaS, nsaR, rpoB, and rpoC mostly missense mutations were found. Interestingly, we found the same pmtR stop-gain mutation (c.14 T > A p.Leu5*) in at least 11 strains, which has previously been described for melittin-selected S. aureus lines (Johnston et al. 2016). Additionally we also identified an nsaS missense mutation (A208E) in 4 strains which has been shown to be responsible for nisin resistance in SH100037, the same strain as used here. The final MICs achieved did not differ by mutation (T = 0.25, df = 32, p = 0.8, Fig. 2).
Reduced fitness in relation to resistance mutations
Fitness reduction, as measured by the increased lag phase, did not differ by mutation (T = 0.93, df = 56, p = 0.3576, Fig. 3A–C). Vmax differed between nsa and pmt mutations, with nsa mutants showing a lower growth rate than pmt mutants (T = 2.7, df = 56, p = 0.0092) and both mutants grow significantly slower than the controls (pmt: T = −6.46, df = 56, p = < 0.0001, nsa: T = −8.16, df = 56, p = < 0.0001, Fig. 3A–F). In the presence of a second mutation rpo, nsa mutants show a further extension of the lag phase (T = 4.21, df = 54, p = 1e-04, Fig. 3G–I). Final OD differed by mutation (T = −3.36 df = 56 p = 0.0014), final OD is lower for nsa mutants than pmt mutants or controls.
Parallel evolution
Using the Jaccard distance to calculate the degree of parallel evolution (see Wong et al. 2012), we did not find evidence for parallel evolution at the operon level. Selection treatment (T1 or T1T2) did not affect the mean proportion of shared mutated operons (permutational analysis of multivariate homogeneity of group dispersion F = 0.1925, p = 0.625) or the mean number of shared mutations (permutational multivariate analysis of Jaccard distance matrix, F = 1.063, p = 0.347, Fig. 4). Yet, as reported above, all AMP selected lines showed either a mutation in nsa or in pmt operons, but never in both (Table S1).
Discussion
Our study was one of the first to explore the evolution of resistance to AMPs that are part of the same immune system. We find that all populations under antimicrobial peptide selection evolve resistance quickly and to a similar degree irrespective of the presence of one or two AMPs. In all cases resistant strains possessed a mutation in either in the pmt or nsa operons. Mutations in these operons were not found together.
We have previously shown that evolution of resistance towards the AMP melittin in S. aureus JLA513 is associated with nonsense mutations in pmtR (previously referred to as ytrA13. Here we find that the most frequent pmtR mutation is identical to the stop-gain mutation described in S. aureus JLA51313. In S. aureus, the pmt operon encodes an essential export system for Phenol-soluble modulin peptide toxins (PSM)38. pmt expression is negatively regulated by the GntR-type transcriptional repressor PmtR which binds to an operator site within the pmt promoter16. PSMs enable pmt expression by disrupting the binding of PmtR to the operator16. pmt is the only known regulatory target of PmtR and pmtR deletion mutants show constitutive upregulation of only pmtA-D16. pmt expression is also induced by exposure to cationic AMPs39 and has recently been shown to be responsible for defending S. aureus against killing by human AMPs40. This raises the possibility that the pmtR mutations observed here may mediate Tenecin 1 resistance via repression of pmt.
The nisin susceptibility-associated (nsa) two-component system was independently discovered three times19,20,37 as being responsible for resistance to nisin and bacitracin. In the presence of it’s substrate NsaSR activates transcription of the Bce-AB type ABC transporters BraDE and VraDE which are involved in sensing and detoxification respectively20. Strikingly, in 4 strains we identified the same nsaS mutation (A208E) which was shown to confer increased nisin resistance in SH100037.
The costs of antibiotic resistance are variable and examples of cost-free resistance exist25. The issue of cost of resistance is much less researched for AMPresistance costs14,21. Here we find clear evidence for costly resistance as measured in growth rate and lag phase for both AMPtreatments. When looking at the level of the most frequent mutated genes, nsa and pmt, we find that all three measured costs, reduction in growth rate and population density (final OD) and extended lag phase are increased. The cost of resistance by mutations in nsa seems to be higher as indicated by both, growth rate (Vmax) and population density (final OD). With the exception of the double mutation nsa/rpo, which displays the longest lag phase the costs in our experiments do not differ by mutation. rpoB mutations in AMP-resistant S. aureus have been found before41. At the moment it is not clear why the more costly double-mutation evolves, given that there is no difference in MICs between nsa and nsa/rpo mutants. Limited costs in S. aureus selected for resistance against the human AMP LL-37 were reported41, but the same study found reduced growth rates in AMP-resistant strains. Antibiotic resistance evolution resulting in extended lag phases has been frequently reported (e.g.)42 and also has been proposed as a mechanism explaining antibiotic tolerance25 even in the absence of bona fide resistance.
Lack of parallel evolution
While we do find strong evidence for parallel evolution at the level of the resistance phenotype, all populations under selection evolve resistance at a comparable speed to a similar level (MIC), we do not find evidence for parallel evolution at the level of the operons and treatments. This contrasts with findings reported in Pseudomonas fluorescens under antibiotic selection43, which found a higher degree of parallel evolution, albeit in more complex environments. A low level of parallelism has been observed in other studies of bacterial antibiotic resistance evolution42,44. While our assessment is based on the Jaccard-Distance, it is noteworthy that all strains have one of two resistance-associated mutations. This is much lower than the variation observed in a previous study in S. aureus against a panel of antimicrobial peptides originating from different organisms13.
Cross-resistance
Despite the limited number of resistance mutations, both our experimentally evolved Tenecin 1 and Tenecin 1 + Tenecin 2-selected strains, display cross-resistance against colistin and melittin. All of these antimicrobial peptides have a different origin (from Paenibacillus polymixa, the African clawed frog and the honey bee, respectively) and belong to distinct AMP families. This underlines that AMP resistance mechanisms can be relatively nonspecific. In gram-positive bacteria resistance is mostly achieved by modifications to the physico-chemical properties of the cell envelope16.
Cross-resistance between AMPs has been reported before14,21 and might constitute a risk for the application of AMPs in medical treatments45. In the context of natural immune defenses it is likely that the resistance evolution to one AMP possibly results in resistance also against AMP cocktails. This could constitute a strong selection pressure by AMP resistant bacteria for those innate immune systems that are strongly dependent on AMPs, though combinations of AMPs can still be beneficial in relation to resistance as they change the probability of resistance evolution by changing the pharmacodynamics (Yu et al. 2018). Finally, while we did not find indications for cross-resistance against antibiotics, cross-resistance against human AMPs and the peptide antibiotic daptomycin, which targets the cell membrane, have been reported in MRSA46. It hence seems possible that resistance against AMPs comes with the risk of cross-resistance against new potential antibiotics such as teixobactin47 that target the cell surface.
While we found differences in costs of resistance depending on mutation, we did not find differences in the costs of resistance nor the resistance mutations acquired dependent on treatment with either Tenecin 1 alone or Tenecin 1 and 2 combined. In this respect our simplified in vitro approach did not allow us to understand the more complex interactions observed in vivo. Whether this is caused by the strong reduction in interacting immune effectors to only two, or by the physiological environment, i.e. differences between the artificial growth medium and haemolypmph remains to be investigated.
References
Westerhoff, H. et al. Functional synergism of the magainins PGLa and magainin-2 in Escherichia coli, tumor cells and liposomes. Eur. J. Biochem. 228, 257–64 (1995).
Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 415, 389–395 (2002).
Johnston, P. R., Makarova, O. & Rolff, J. Inducible Defenses Stay Up Late: Temporal Patterns of Immune Gene Expression in Tenebrio molitor. G3 (Bethesda). 4, 1–9 (2013).
Fjell, C., Hiss, J., Hancock, R. & Schneider, G. Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discov. 11, 37–51 (2012).
Yu, G., Baeder, D. Y., Regoes, R. R. & Rolff, J. The More The Better? Combination Effects of Antimicrobial Peptides. Antimicrob. Agents Chemother. 60, 1717–1724 (2016).
Rahnamaeian, M. et al. Insect antimicrobial peptides show potentiating functional interactions against Gram-negative bacteria. Proc. R. Soc. B Biol. Sci. 282, 20150293 (2015).
Rolff, J. & Schmid-Hempel, P. Perspectives on the evolutionary ecology of arthropod antimicrobial peptides. Phil. Trans. R. Soc. B 371, 20150297 (2016).
Kounatidis, I. & Ligoxygakis, P. Drosophila as a model system to unravel the layers of innate immunity to infection. Open Biol. 2, 120075 (2012).
Makarova, O. et al. Antimicrobial defence and persistent infection in insects revisited. Philos. Trans. R. Soc. B B 371, 20150296 (2016).
Roh, K.-B. et al. Proteolytic cascade for the activation of the insect toll pathway induced by the fungal cell wall component. J. Biol. Chem. 284, 19474–19481 (2009).
Zanchi, C., Johnston, P. R. & Rolff, J. Evolution of defence cocktails: Antimicrobial peptide combinations reduce mortality and persistent infection. Mol. Ecol. 26, 5334–5343 (2017).
Dobson, A. J., Purves, J. & Rolff, J. Increased survival of experimentally evolved antimicrobial peptide-resistant Staphylococcus aureus in an animal host. Evol. Appl. 7, 905–912 (2014).
Johnston, P. R., Dobson, A. J. & Rolff, J. Genomic signatures of experimental adaptation to antimicrobial peptides in Staphylococcus aureus. G3 Genes|Genomes|Genetics 6, 1535–1539 (2015).
Habets, M. & Brockhurst, M. A. Therapeutic antimicrobial peptides may compromise natural immunity. Biol. Lett. 8, 416–418 (2012).
Lofton, H., Pränting, M., Thulin, E. & Andersson, D. I. Mechanisms and Fitness Costs of Resistance to Antimicrobial Peptides LL-37, CNY100HL and Wheat Germ Histones. PLoS One 8, e68875 (2013).
Joo, H.-S., Fu, C. & Otto, M. Bacterial Strategies of Resistance to AntimicrobialPeptides. Phil. Trans. R. Soc. B 371, 20150295 (2016).
Yang, S. et al. The Staphylococcus aureus two-component regulatory system, grars, senses and confers resistance to selected cationic antimicrobial peptides. Infect. Immun. 80, 74–81 (2012).
Koprivnjak, T. & Peschel, A. Bacterial resistance mechanisms against host defense peptides. Cell. Mol. Life Sci. 68, 2243–2254 (2011).
Yoshida, Y. et al. Bacitracin sensing and resistance in Staphylococcus aureus. FEMS Microbiol. Lett. 320, 33–39 (2011).
Hiron, A., Falord, M., Valle, J., Débarbouillé, M. & Msadek, T. Bacitracin and nisin resistance in Staphylococcus aureus: A novel pathway involving the BraS/BraR two-component system (SA2417/SA2418) and both the BraD/BraE and VraD/VraE ABC transporters. Mol. Microbiol. 81, 602–622 (2011).
Dobson, A. J., Purves, J., Kamysz, W. & Rolff, J. Comparing Selection on S. aureus between Antimicrobial Peptides and Common Antibiotics. PLoS One 8, 3–7 (2013).
Naghili, H. et al. Validation of drop plate technique for bacterial enumeration by parametric and nonparametric tests. Vet. Res. forum an Int. Q. J. 4, 179–83 (2013).
Wiegand, I., Hilpert, K. & Hancock, R. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Kassen, R. & Bataillon, T. Distribution of fitness effects among beneficial mutations before selection in experimental populations of bacteria. Nat. Genet. 38, 484–488 (2006).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Fridman, O., Goldberg, A., Ronin, I., Shoresh, N. & Balaban, N. Q. Optimization of lag time underlies antibiotic tolerance in evolved bacterial populations. Nature 513, 418–421 (2014).
O’Neill, A. Staphylococcus aureus SH1000 and 8325-4: Comparative genome sequences of key laboratory strains in staphylococcal research. Lett. Appl. Microbiol. 51, 358–361 (2010).
Bæk, K. et al. Genetic Variation in the Staphylococcus aureus 8325 Strain Lineage Revealed by Whole-Genome Sequencing. PLoS One 8, 1–16 (2013).
Bankevich, A. et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 19, 455–477 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Seemann, T. Snippy: fast bacterial variant calling from NGS reads. https://github.com/tseemann/snippy.(2015).
Barrick, D. Identification of Mutations in Laboratory-Evolved Microbes from Next-Generation Sequencing Data Using breseq. In Methods in Molecular Biology, 165–188 (2014).
Pinheiro, J., Bates, D., DebRoy, S. & Sarkar, D. NLME: Linear and nonlinear mixed effects models. R Packag. version 3.1–122, http//CRAN.R-project.org/package=nlme Version 3, 1–336 (2013).
Breheny, P. & Burchett, W. Visualization of regression models using visreg. R Packag., 1–15, http://myweb. uiowa.edu/pbreheny/publications/visreg.pdf) (2013).
Lenth, R. Least-Squares Means: The R Package lsmeans. J. Stat. Softw. 69 (2016).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.4–3 (2017).
Blake, K., Randall, C. & O’Neill, A. J. In vitro studies indicate a high resistance potential for the lantibiotic nisin in Staphylococcus aureus and define a genetic basis for nisin resistance. Antimicrob. Agents Chemother. 55, 2362–2368 (2011).
Chatterjee, S. S. et al. Essential Staphylococcus aureus toxin export system. Nat. Med. 19, 364–367 (2013).
Li, M. et al. The antimicrobial peptide-sensing system aps of Staphylococcus aureus. Mol. Microbiol. 66, 1136–1147 (2007).
Cheung, G. et al Antimicrobial peptide resistance mechanism contributes to Staphylococcus aureus infection Accepted. J., 1–34 (2018).
Kubicek-Sutherland, J. Z. et al. Antimicrobial peptide exposure selects for Staphylococcus aureus resistance to human defence peptides. J. Antimicrob. Chemother. 72, 115–127 (2017).
Barbosa, C. et al. Alternative evolutionary paths to bacterial antibiotic resistance cause distinct collateral effects. Mol. Biol. Evol. 34, 2229–2244 (2017).
Bailey, S. F., Rodrigue, N. & Kassen, R. The effect of selection environment on the probability of parallel evolution. Mol. Biol. Evol. 32, 1436–1448 (2015).
Vogwill, T., Kojadinovic, M., Furió, V. & Maclean, R. C. Testing the role of genetic background in parallel evolution using the comparative experimental evolution of antibiotic resistance. Mol. Biol. Evol. 31, 3314–3323 (2014).
Bell, G. Arming the enemy: the evolution of resistance to self-proteins. Microbiology 149, 1367–1375 (2003).
Mishra, N. N., Bayer, A. S., Moise, P., Yeaman, M. R. & Sakoulas, G. Reduced susceptibility to host-defense cationic peptides and daptomycin coemerge in methicillin-resistant Staphylococcus aureus from daptomycin-naive bacteremic patients. J. Infect. Dis. 206, 1160–7 (2012).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459 (2015).
Acknowledgements
We thank Vitali Laba for support in the lab. Accession number for Genome Data: PRJNA399645. O.M., P.R.J. and J.R. were supported by the European Research Council (EVORESIN 260986). A.R.R. and J.R. by the Deutsche Forschungsgemeinschaft (SFB 973) and BeS by the DAAD and the Egyptian Government.
Author information
Authors and Affiliations
Contributions
J.R., O.M., P.R.J. designed the study. O.M. led the experimental work supported by A.R.R., B.E.S. and J.M. P.R.J. carried out the bioinformatics and led the statistical analysis. All authors contributed to the writing of the paper. JR wrote the first draft.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Makarova, O., Johnston, P., Rodriguez-Rojas, A. et al. Genomics of experimental adaptation of Staphylococcus aureus to a natural combination of insect antimicrobial peptides. Sci Rep 8, 15359 (2018). https://doi.org/10.1038/s41598-018-33593-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33593-7
Keywords
This article is cited by
-
In Silico Characterization of a Transcript Code Based Screening of Antimicrobial Peptide from Trichogramma chilonis
International Journal of Peptide Research and Therapeutics (2021)
-
Insect antimicrobial peptides: potential weapons to counteract the antibiotic resistance
Cellular and Molecular Life Sciences (2021)
-
Probiotics Modulate a Novel Amphibian Skin Defense Peptide That Is Antifungal and Facilitates Growth of Antifungal Bacteria
Microbial Ecology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.