Abstract
An increasing number of disorders have been identified for which two or more distinct alleles in two or more genes are required to either cause the disease or to significantly modify its onset, severity or phenotype. It is difficult to discover such interactions using existing approaches. The purpose of our work is to develop and evaluate a system that can identify combinations of alleles underlying digenic and oligogenic diseases in individual whole exome or whole genome sequences. Information that links patient phenotypes to databases of gene–phenotype associations observed in clinical or non-human model organism research can provide useful information and improve variant prioritization for genetic diseases. Additional background knowledge about interactions between genes can be utilized to identify sets of variants in different genes in the same individual which may then contribute to the overall disease phenotype. We have developed OligoPVP, an algorithm that can be used to prioritize causative combinations of variants in digenic and oligogenic diseases, using whole exome or whole genome sequences together with patient phenotypes as input. We demonstrate that OligoPVP has significantly improved performance when compared to state of the art pathogenicity detection methods in the case of digenic diseases. Our results show that OligoPVP can efficiently prioritize sets of variants in digenic diseases using a phenotype-driven approach and identify etiologically important variants in whole genomes. OligoPVP naturally extends to oligogenic disease involving interactions between variants in two or more genes. It can be applied to the identification of multiple interacting candidate variants contributing to phenotype, where the action of modifier genes is suspected from pedigree analysis or failure of traditional causative variant identification.
Similar content being viewed by others
Introduction
Discrimination of causative genetic variants responsible for disease is a major challenge. An increasingly large family of algorithms and strategies has been developed to aid in identification of such variants1. These methods use properties of variants such as evolutionary conservation, predicted structural changes, allele frequency and function to predict pathogenicity. For variants in non-coding sequence regions, additional information used by computational models includes predicted regulatory function and recognized DNA–protein or DNA–RNA interactions1,2,3. Furthermore, phenotype annotations to human and model organism genes can be added to provide another layer of discrimination between involved pathogenic and non-pathogenic variants4,5,6. Phenotype-based methods can identify the likelihood that a particular gene or gene product may give rise to phenotypes observed in an individual7,8.
The increasing availability of patient sequence information coupled with resources that provide a detailed phenotypic characterization of diseases, as well as the wealth of gene-to-phenotype associations from non-human disease models9, are now enabling new approaches to the prioritization of causative variants and facilitating our ability to dissect the genetic underpinnings of disease5. PhenomeNET10, developed in 2011, is a computational framework that utilizes pan-phenomic data from human and non-human model organisms to prioritize candidate genes in genetically-based diseases10. We have combined PhenomeNET with genome-wide pathogenicity predictions to develop the PhenomeNET Variant Predictor (PVP)4 as a system that combines information about pathogenicity of variants with known gene–phenotype associations to predict causative variants. We recently developed the PVP system to classify variants into those likely to be causative or non-causative4.
While PVP has a significantly better performance in the prioritization of single variants in monogenic diseases than competing algorithms4, the phenotypes of many diseases with a recognized genetic origin show a range of severity, phenotypic spectrum, age of onset and prognosis11. While characteristic phenotypic variability can be associated with different alleles in single disease-causing genes or their mode of inheritance, it has been known for some time that in many diseases there is variable expressivity, and in some cases variable penetrance, associated with the same primary mutation in different individuals or pedigrees12. This implies that in those cases the phenotypic variability observed must be due to the presence of other modifier variants or environmental influences. Increasingly, several diseases are being understood within the context of complex inheritance and multifactorial disease phenotypes where multiple independent variants modify each other’s effect on phenotype13, or, in some cases, render the disease di- or oligogenic where variants in two or more different genes are needed for its clinical manifestation14. Evidence for digenic inheritance is available for around 50 diseases15 and details are gathered in the Digenic disease database (DIDA)16.
Epistatic interactions have been postulated to explain the missing heritability in many types of common and rare disease12, and with the increasing clinical use of next generation sequencing, further evidence is accumulating for a spectrum of types of interactions between genes. These interactions are manifest in different ways. For example, there is evidence from population genetics for phenotypic modifier genes such as the modifiers of the age of onset in Huntington’s disease17,18, and from a candidate gene approach in Parkinson’s disease19. In a similar candidate approach for amyotrophic lateral sclerosis (ALS), affected individuals with proven or potentially pathogenic mutations in two or three known ALS genes are associated again with lower age of disease onset.
Congenital hypothyroidism has both rare monogenic recessive loss-of-function, and common, apparently sporadic, forms. The recent description of patients carrying biallelic and triallelic digenic combinations of variants in known thyroid development and function genes20 has lead to the suggestion that a frequent oligogenic origin might explain sporadic hypothyroid cases. Evidence has also been provided for rare trigenic involvement of variants, such as in TSHR, SLC26A4 and GLIS321. In addition to these examples of genetic interaction, the observations suggesting digenic/triallelic inheritance in Bardet-Biedel syndrome (BBS)22 continue to provoke interest and further research, and illustrate the challenges in establishing formal digenicity23. One of the best characterized cases of digenic inheritance is a form of Usher syndrome where digenic heterozygous mutations in CDH23 and PCDH15 have been shown to interact in both humans and mice24. Other examples are critically discussed elsewhere15.
Digenic disease can be divided into two classes: strict digenic disease where variants at both loci are required for the disease, and composite disease which is either the result of the epistatic relationship between alleles of two independent genes modifying the phenotypes of the individual mutations alone, or the phenotypic overlay of two monogenic Mendelian diseases present in the same patient25,26. Identification of the genes involved in all of these types of digenic disease usually requires pedigree information or the use of existing knowledge about candidate genes. For example, the selection of candidate genes may rely on the availability of additional information about molecular or functional connections between the entities (genes or gene products) bearing the variants20. The difficulties in establishing strong evidence for digenic inheritance are discussed elsewhere15,27.
Computational identification of likely causative alleles that are involved in digenic or genetically more complex diseases, in particular for genes not previously associated with the disease, is particularly challenging; such methods would have to be able to incorporate and utilize a large amount of background information about molecular and (patho-)physiological interactions within an organism to determine how combinations of variants jointly result in an observed phenotype. The observation that disease-implicated proteins often interact with each other has stimulated the development of network-based approaches to identification of disease modules28,29,30,31. However, relevant interactions may occur across much larger distances within pathways and networks, or at the whole organism physiological level where knowledge about biological systems and multi-scale interactions is critical for understanding pathobiology32,33. Phenotypes provide a readout for all of these disease-relevant interactions and offer insights into the underlying pathobiological mechanisms34. Phenotype data can be a powerful source of information for variant prioritization and is complementary to pathogenicity prediction methods based on molecular information4,35,36,37,38.
Here, we first evaluate the ability of the PVP system to identify combinations of variants in digenic diseases obtained from a database of digenic diseases. We then present OligoPVP, a novel algorithm for prioritizing digenic or higher order combinations of variants in personal genomes. While the OligoPVP algorithm will not identify whether a phenotype or disease has a digenic or oligogenic inheritance, it identifies and ranks potential causative variants in the same way as PVP but then prioritizes pairs or sets of higher cardinality of potentially interacting variants present in the same genome, as specified by the user, on the basis of prior knowledge about genetic, regulatory and biochemical interactions between them. It is therefore mainly useful to identify candidate causative sets of variants in cases in which digenic or oligogenic inheritance is already suspected, for example due to variable penetrance or expressivity in family studies, or as means of exclusion because established methods for variant prioritization failed.
We apply OligoPVP to the identification of variants in pairs of genes where mutations in two separate genes present in a single individual and lead to a particular phenotypic profile that is not apparent in individuals carrying only one of these variants. We demonstrate that OligoPVP is able to identify gene variant sets in digenic diseases using a set of synthetic whole genome sequences into which we insert multiple known causative gene variants. OligoPVP is freely available at https://github.com/bio-ontology-research-group/phenomenet-vp.
Materials and Methods
Digenic disease
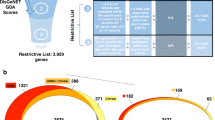
The Digenic Disease Database (DIDA) v216 consists of 258 curated digenic combinations representing 54 diseases, with 448 variants in 169 genes. Of the 258 digenic combinations, 189 have Human Phenotype Ontology (HPO) annotations, representing 52 diseases, 153 distinct genes, and 337 unique variants. We use the 189 digenic combinations with HPO annotations in our experiments. 25 of these combinations are triallelic and exhibit compound heterozygosity in one gene while the remaining 164 combinations are biallelic.
We use the combinations of variants from DIDA to generate 189 synthetic whole genome sequences by randomly inserting the causative variants in a randomly selected whole genome sequence from the 1000 Genomes Project39.
Interaction data
We downloaded all interactions occurring in humans from the STRING database version 10.\(5\)40. Then, we mapped all interactions to their respective genes using the mapping file provided by STRING to generate 989,998 interactions between genes, representing 13,770 unique genes. We use these interactions between genes to prioritize combinations of variants in OligoPVP.
PhenomeNET Variant Predictor
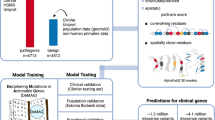
In our work, we use the PhenomeNET Variant Predictor (PVP) version 2.0 (“DeepPVP”)41. This version of PVP is based on the Human Phenotype Ontology (HPO)35 and the Mammalian Phenotype Ontology (MP)42 ontologies obtained from the AberOWL repository43 on Feb 7th, 2017. PVP uses gene-to-phenotype associations in humans as well as from the mouse and zebrafish model organisms downloaded on Feb 7th, 2017 from the HPO website35, the Mouse Genome Informatics website42, and the Zebrafish Information Network website44, respectively. The PVP system was trained on data generated from the ClinVar database45 accessed on Feb 7th, 2017.
PVP combines features that score the pathogenicity of variants with a phenotype similarity measure that aims to identify whether a variant is likely to cause the phenotypes observed in a patient. Using the cross-species phenotype ontology PhenomeNET46, phenotype similarity also can be computed between patient phenotypes and phenotypes in model organisms.
The PVP system used in our analysis, the synthetic genome sequences we generated for the evaluation of our system, and our analysis results can be found at https://github.com/bio-ontology-research-group/phenomenet-vp.
Evaluation and comparison
We compare PVP and OligoPVP to several state of the art variant prioritization method. Specifically, we compare the PVP and OligoPVP scores to variant pathogenicity prediction scores obtained from CADD v1.347, DANN v1.048, and GWAVA v1.049. Furthermore, we compare our results to the phenotype-based tool Genomiser version 7.2.150 using its default parameters.
Results
Prediction of biallelic and triallelic disease variants
We analyze each WGS using the phenotypes provided for the combination of variants in DIDA. We do not filter any variants by minor allele frequency to avoid missing potentially important interacting variants that might have medium to common frequencies in the background population. On average, each WGS in our experiments contains 2,192,967 variants.
We use the phenotypes associated with the combination of variants in DIDA as phenotypes associated with the synthetic WGS, and we use PVP4 to prioritize variants, using an “unknown” mode of inheritance model. Out of 164 whole genome sequences where two variants were inserted, we find both causative variants (i.e., the two variants we inserted) as the highest ranked variants in 88 cases (53.7%) and within the top ten ranks in 107 cases (65.2%) (see Table 1). For the 25 cases of triallelic diseases, we find all three causative variants within the first three ranks in 10 cases (40.0%) and we find all three causative variants within the top ten variants in 14 cases (56.0%) (see Table 2). Tables 1 and 2 also compare PVP to established variant prioritization methods, including CADD47, DANN48, the phenotype-based Exomiser/Genomiser system50, and GWAVA49. Out of these systems, CADD performs the best in prioritizing combinations of variants; however, PVP can rank variant involved in bi- or triallelic diseases significantly higher than CADD (Mann-Whitney U test, p = 6.8 × 10−5).
Individually, the performance of our approach differs between diseases, depending on the availability of gene–phenotype associations and high quality and informative disease–phenotype associations in DIDA. Table 3 provides an overview of the performance of PVP for individual diseases, and we provide the full analysis results on our website.
In particular, for the case of hypodontia, PVP identifies all the causative variant pairs in the top 3 ranks in all synthetic patients, and in Familial long QT syndrome, the causitive variant pairs can be found in the top 3 ranks in over 90% of the synthetic patients. Similarly, for the case of Bardet-Biedl syndrome (BBS), PVP ranks 84.21% of causative variant pairs in the top 10, and identifies digenic causative variants in 9 of the 16–20 genes now implicated in BBS23,51.
To ensure that newer versions of ontologies and our training data do not already contain, implicitly, information about associations between variants and disease, we perform a semi-prospective experiment; the PVP system we used is based on ontology versions (HPO and MP) and training data obtained on Feb 7th, 2017. We separately test the performance of our system on the digenic cases added to the DIDA database after Feb 7th, 2017. In total, 45 digenic combinations with HPO annotations are newly added to DIDA after the date our PVP system was built. Among these newly added combinations, 42 are biallelic and 3 are triallelic. Table 4 shows the performance of PVP on these cases. We find that the performance on predicting the new cases drops somewhat in comparison to cases before the PVP build date.
OligoPVP: Use of background knowledge to find causative combinations of variants
Our results demonstrate that PVP can identify combinations of variants implicated in a disease, outperforming current state-of-the-art gene prioritization approaches. The variants found by PVP are commonly in genes that form a disease module, i.e., a set of interacting genes that are jointly associated with a disease or phenotype52. For example, out of the 164 biallelic combinations used in our study, we can find evidence of interactions for 71 biallelic combinations and 16 triallelic combinations using the interaction database STRING40. The STRING resource contains background knowledge about the interaction between genes based on protein-protein interactions, co-expression, pathway involvement, or co-mention in literature, and therefore provides a wide range of distinct interaction types which may underlie a phenotype. We have now exploited this background knowledge to further improve prioritization of variants in oligogenic diseases which involve interactions between alleles of two or more genes.
OligoPVP is an algorithm that uses background knowledge from interaction networks to prioritize variants in oligogenic diseases. It identifies likely causative variants in interacting genes and ranks tuples of n variants in genes that are connected through an interaction network. OligoPVP will first rank all variants in a set of variants (such as those found in a VCF file) independently using PVP and assign each variant ν a prediction score σ(ν). When ranking combinations of n variants, OligoPVP will then evaluate all n-tuples of variants \({v}_{1},\,\mathrm{...,}\,{v}_{n}\) and assign a score \(\bar{\sigma }\) to the n-tuple \(({v}_{1},\,\mathrm{...,}\,{v}_{n})\), given an interaction network ϒ:
Algorithm 1 illustrates the procedure to find oligogenic disease modules in more detail. OligoPVP can identify combinations of variants both in exonic and non-exonic regions. For non-exonic variants, we assign the gene that is located closest to the variant as the variant’s gene.
The OligoPVP algorithm strictly relies on an interaction network as background knowledge and will not prioritize any combinations of variants if they are not connected in a known network. For digenic or oligogenic disease we assume that the interactions between alleles are mediated directly or indirectly through physical or regulatory mechanisms at any level, e.g., through DNA (for example transcriptional regulation), RNA (for example processing of RNA or half life modulation), or protein (e.g., direct functional interaction or co-participation in the same pathway or physiological processes). Data relevant to all these levels of interaction is available in STRING. OligoPVP is of course limited by the completeness of the interaction data available and until more complete high level physiological modeling is achieved, interactions at the level of the whole organism physiome will be difficult to capture.
OligoPVP utilizes beam search53 to optimize memory usage. We can simply extend OligoPVP to also consider compound heterozygote combinations of variants by adding self-loops to each node in ϒ. The main advantage of OligoPVP is its ability to identify and rank connected sets of variants higher than individual variants. Table 5 lists several cases in which OligoPVP prioritizes pairs of variants higher than PVP would prioritize them on their own. On the other hand, OligoPVP will not prioritize combinations of variants if they are in genes that are not connected in the background network ϒ. Supplementary Table 1 lists some of the cases which can be prioritized with PVP but not OligoPVP.
Discussion
With the increasing appreciation of the relationship between complex and Mendelian diseases54, the ability to discover multiple variants contributing to disease phenotype in the same genome provides a powerful tool to help understand the genetic architecture of diseases and the underlying physiological pathways. With the advent of whole exome and whole genome sequencing, advances have been made using existing approaches to prioritize causative variants. However, use of standard criteria for the identification of rare disease variants, e.g., a low minor allele frequency (MAF) of, for example, less than 1%, are designed to detect de novo, homozygous, or compound heterozygous variants, and may not give sufficient priority to variants of low apparent pathogenicity, haploinsufficiency, or low to medium MAF, although these variants may still be important in the pathogenesis of a disease. Because the approach we take with OligoPVP and PVP makes no assumptions about allele frequency or mode of inheritance, and balances estimates of pathogenicity with phenotypic relatedness, a wider net is cast and candidate genes affecting the penetrance, expressivity or spectrum of the phenotype are more readily identified.
Genes whose variants contribute to a disease phenotype are considered likely to be situated within the same pathway or network55,56,57,58. In addition to well established studies of genes involved in, for example the ciliopathies23,59, newer studies are now identifying network relations between genes implicated in the oligogenic origins of diseases60,61. Consequently, we can exploit background knowledge on the interactions of gene products in OligoPVP and improve the ranking of candidate pairs of variants over that assigned through pathogenicity and phenotypic relatedness scores alone.
Currently, identification of multiple variants contributing to the characteristics of a disease in a cohort or individual patient rely either on a candidate gene approach or the assumption that contributing alleles are likely to be rare in the population. The contribution of rare alleles of low effect, i.e., which by themselves generate sub-clinical phenotypes, for example hypomorphs, may be missed in this way, and rare to medium frequency alleles which modify the penetrance or expressivity of a second remain difficult to identify (the former because of low potential pathogenicity and the latter because of high frequency and lack of association with a phenotype when occurring alone). An alternative strategy for identification of candidate genes for highly heterogeneous human diseases is to use mouse genetics to identify phenotypic modifier genes. For example, neural tube defects are believed to involve more than 300 genes in the mouse, mutations in many of which need to be digenic or trigenic for expression of the phenotype62. Similarly, mouse double heterozygous mutants in Nkx2-1 and Pax8 show strain-specific thyroid dysgenesis phenotypes not seen in the individual mutations63. The scale of genetic interactions becoming apparent from mouse studies strongly supports the suggestion that in the human, we are only seeing the tip of a very important iceberg64.
The OligoPVP algorithm offers a generic framework for using background knowledge about any form of interaction between genes and gene products to guide the identification of combinations of variants. In its generic form, the worst case complexity of the algorithm is \({\mathscr{O}}({n}^{k})\) where n is the number of variants and k the size of the module (the size of the module is a parameter of OligoPVP). It is clear that our algorithm, in its basic form, will not yet scale to large disease modules (i.e., large k); however, in the future, several methods can be used to further improve the average case complexity to find larger disease modules.
Furthermore, background knowledge about interactions between genes and gene products is far from complete. In particular, information about coarse scale physiological interactions, i.e., those that occur based on whole organism physiology, are significantly underrepresented in interaction databases32. Additionally, interaction networks may have biases such as over-representation of commonly studied genes65,66, and these biases will likely effect the performance of our algorithm. As more genomic data related to complex diseases becomes available, more work will be required to identify and remove these biases in the identification of phenotype modules from personal genomic data.
The function of OligoPVP is not to determine whether a disease is formally di- or oligogenic but to assess whether sets of variants may be jointly responsible for phenotype when an individual contains multiple, potentially pathogenic, variants in two or more genes that might, from background knowledge, be expected to interact to generate the phenotypic profile of the patient. The user is able to specify the cardinality of the set of variants that should be prioritized, and the rank scores provide a relative measure of the likelihood that two, three or more specific allelic variants might be involved.
While this limitation restricts OligoPVP’s applicability, we nevertheless believe our algorithm to have important applications. The most likely scenario in which OligoPVP can be used successfully is to generate hypotheses for combinations of specific variants after other variant prioritization approaches have failed to yield significant results. Alternatively, knowledge of family history or ethnic background might suggest the contribution of two or more genes to the disease phenotype, and OligoPVP can then be used to identify plausible combinations of genes and allelic variants.
OligoPVP is, to the best of our knowledge, the first phenotype-based method to identify disease modules in personal genomic data. With the large (i.e., exponential) number of combinations of variants that have to be evaluated in finding disease modules, it is clear that any computational method has to make use of background knowledge to restrict the search space of potentially causative combinations of variants. OligoPVP is such a method which uses knowledge about interactions and phenotype associations to limit the search space. In the future, more background knowledge can be incorporated to improve OligoPVP’s coverage as well as accuracy. OligoPVP is freely available at https://github.com/bio-ontology-research-group/phenomenet-vp.
References
Eilbeck, K., Quinlan, A. & Yandell, M. Settling the score: variant prioritization and mendelian disease. Nat. Rev. Genet. 18, 599 (2017).
Huang, Y.-F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Preprint at https://www.biorxiv.org/content/early/2016/08/15/069682 (2016).
Flygare, S. et al. The vaast variant prioritizer (VVP): ultrafast, easy to use whole genome variant prioritization tool. BMC Bioinformatics 19, 57 (2018).
Boudellioua, I. et al. Semantic prioritization of novel causative genomic variants. PLOS Comput. Biol. 13, 1–21 (2017).
Robinson, P. N. et al. Improved exome prioritization of disease genes through cross-species phenotype comparison. Genome Res 24, 340–348 (2014).
Aerts, S. et al. Gene prioritization through genomic data fusion. Nat. Biotechnol. 24, 537–544 (2006).
Gkoutos, G. V., Schofield, P. N. & Hoehndorf, R. The anatomy of phenotype ontologies: principles, properties and applications. Brief Bioinform, bbx035 (2017).
Smedley, D. et al. Phenodigm: analyzing curated annotations to associate animal models with human diseases. Database 2013, bat025 (2013).
de Angelis, M. H. et al. Analysis of mammalian gene function through broad-based phenotypic screens across a consortium of mouse clinics. Nat. Genet, 47, 969–978 (2015).
Hoehndorf, R. et al. Phenomenet: a whole-phenome approach to disease gene discovery. Nucleic Acids Res. 39, e119 (2011).
Haldane, J. B. S. The relative importance of principal and modifying genes in determining some human diseases. J. Genet. 41, 149–157 (1941).
Cooper, D. N., Krawczak, M., Polychronakos, C., Tyler-Smith, C. & Kehrer-Sawatzki, H. Where genotype is not predictive of phenotype: towards an understanding of the molecular basis of reduced penetrance in human inherited disease. Hum Genet. 132, 1077–130 (2013).
Katsanis, N. The continuum of causality in human genetic disorders. Genome Biol 17, 233 (2016).
Kousi, M. & Katsanis, N. Genetic modifiers and oligogenic inheritance. Cold Spring Harb Perspect Med 5 (2015).
Schaffer, A. A. Digenic inheritance in medical genetics. J. Med. Genet. 50, 641–52 (2013).
Gazzo, A. M. et al. DIDA: A curated and annotated digenic diseases database. Nucleic Acids Res. 44, D900 (2016).
Lee, J.-M. et al. Identification of genetic factors that modify clinical onset of Huntington’s disease. Cell 162, 516–526 (2015).
Chao, M. J. et al. Population-specific genetic modification of Huntington’s disease in venezuela. PLOS Genet. 14, e1007274 (2018).
Lubbe, S. J. et al. Additional rare variant analysis in Parkinson’s disease cases with and without known pathogenic mutations: evidence for oligogenic inheritance. Hum Mol Genet 25, 5483–5489 (2016).
Nicholas, A. K. et al. Comprehensive screening of eight known causative genes in congenital hypothyroidism with gland-in-situ. J. Clin. Endocrinol. Metab. 101, 4521–4531 (2016).
de Filippis, T. et al. A frequent oligogenic involvement in congenital hypothyroidism. Hum. Mol. Genet. 26, 2507–2514 (2017).
Eichers, E., Lewis, R. A., Katsanis, N. & Lupski, J. Triallelic inheritance: a bridge between mendelian and multifactorial traits. Annals Medicine 36, 262–272 (2004).
Shaheen, R. et al. Characterizing the morbid genome of ciliopathies. Genome Biol 17, 242 (2016).
Zheng, Q. Y. et al. Digenic inheritance of deafness caused by mutations in genes encoding cadherin 23 and protocadherin 15 in mice and humans. Hum Mol Genet. 14, 103–11 (2005).
Gazzo, A. et al. Understanding mutational effects in digenic diseases. Nucleic Acids Res 45, e140 (2017).
Posey, J. E. et al. Resolution of disease phenotypes resulting from multilocus genomic variation. New Engl. J. Medicine 376, 21–31 (2016).
Robinson, J. F. & Katsanis, N. Oligogenic Disease, 243–262 (Springer, Berlin, Heidelberg, 2010).
Feldman, I., Rzhetsky, A. & Vitkup, D. Network properties of genes harboring inherited disease mutations. Proc Natl Acad Sci USA 105, 4323–8 (2008).
Gandhi, T. K. et al. Analysis of the human protein interactome and comparison with yeast, worm and fly interaction datasets. Nat Genet. 38, 285–93 (2006).
Bauer-Mehren, A. et al. Gene-disease network analysis reveals functional modules in mendelian, complex and environmental diseases. PLoS One 6, e20284 (2011).
Menche, J. et al. Disease networks. Uncovering disease-disease relationships through the incomplete interactome. Science 347, 1257601 (2015).
de Bono, B., Hoehndorf, R., Wimalaratne, S., Gkoutos, G. V. & Grenon, P. The ricordo approach to semantic interoperability for biomedical data and models: strategy, standards and solutions. BMC Res. Notes 4, 313 (2011).
Hoehndorf, R. et al. Integrating systems biology models and biomedical ontologies. BMC Syst. Biol. 5, 124 (2011).
Schofield, P. N., Hoehndorf, R. & Gkoutos, G. V. Mouse genetic and phenotypic resources for human genetics. Hum Mutat 33, 826–36 (2012).
Köhler, S. et al. The human phenotype ontology in 2017. Nucleic Acids Res. 45, D865–D876 (2017).
Singleton, M. V. et al. Phevor combines multiple biomedical ontologies for accurate identification of disease-causing alleles in single individuals and small nuclear families. Am J Hum Genet. 94, 599–610 (2014).
Smedley, D. & Robinson, P. N. Phenotype-driven strategies for exome prioritization of human mendelian disease genes. Genome Medicine 7, 1–11 (2015).
Sifrim, A. et al. eXtasy: variant prioritization by genomic data fusion. Nat. Methods 10, 1083–1084 (2013).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Research 45, D362–D368 (2017).
Boudellioua, I., Kulmanov, M., Schofield, P. N., Gkoutos, G. V. & Hoehndorf, R. DeepPVP: phenotype-based prioritization of causative variants using deep learning. Preprint at https://www.biorxiv.org/content/early/2018/05/02/311621 (2018).
Blake, J. A. et al. Mouse genome database (MGD)-2017: community knowledge resource for the laboratory mouse. Nucleic Acids Res. 45, D723–D729 (2017).
Hoehndorf, R., Slater, L., Schofield, P. N. & Gkoutos, G. V. Aber-OWL: a framework for ontology-based data access in biology. BMC Bioinformatics 16, 26 (2015).
Howe, D. G. et al. The zebrafish model organism database: new support for human disease models, mutation details, gene expression phenotypes and searching. Nucleic Acids Res. 45, D758–D768 (2017).
Landrum, M. J. et al. Clinvar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2013).
Rodriguez-Garcia, M. A., Gkoutos, G. V., Schofield, P. N. & Hoehndorf, R. Integrating phenotype ontologies with PhenomeNET. J. Biomed. Semant. 8, 58 (2017).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–5 (2014).
Quang, D., Chen, Y. & Xie, X. DANN: a deep learning approach for annotating the pathogenicity of genetic variants. Bioinformatics 31, 761–763 (2015).
Ritchie, G. R. S., Dunham, I., Zeggini, E. & Flicek, P. Functional annotation of noncoding sequence variants. Nat. Methods 11, 294–296 (2014).
Smedley, D. et al. A Whole-Genome analysis framework for effective identification of pathogenic regulatory variants in mendelian disease. Am J Hum Genet. 99, 595–606 (2016).
Forsythe, E. & Beales, P. L. Bardet-Biedl syndrome. Eur J Hum Genet. 21, 8–13 (2013).
Jasny, B. R. A network approach to finding disease modules. Science 347, 836–836 (2015).
Furcy, D. & Koenig, S. Limited discrepancy beam search. In Proceedings of the 19th International Joint Conference on Artificial Intelligence, IJCAI’05, 125–131 (Morgan Kaufmann Publishers Inc., San Francisco, CA, USA, 2005).
Blair, D. R. et al. A nondegenerate code of deleterious variants in mendelian loci contributes to complex disease risk. Cell 155, 70–80 (2013).
Oti, M. & Brunner, H. G. The modular nature of genetic diseases. Clin Genet. 71, 1–11 (2007).
Goh, K.-I. et al. The human disease network. Proc. Nat. Acad. Sci. 104, 8685–8690 (2007).
Khurana, V. et al. Genome-Scale networks link neurodegenerative disease genes to α-Synuclein through specific molecular pathways. Cell systems, 4, 157-170 (2017).
Marbach, D. et al. Tissue-specific regulatory circuits reveal variable modular perturbations across complex diseases. Nat. methods, 13, 366-370 (2016).
Hildebrandt, F., Benzing, T. & Katsanis, N. Ciliopathies. N Engl J Med 364, 1533–1543 (2011).
Priest, J. R. et al. De novo and rare variants at multiple loci support the oligogenic origins of atrioventricular septal heart defects. PLoS Genet. 12, e1005963 (2016).
Li, Y. et al. Against all odds: blended phenotypes of three single-gene defects. Eur J Hum Genet, 24, 1274-1279 (2016).
Leduc, R. Y., Singh, P. & McDermid, H. E. Genetic backgrounds and modifier genes of ntd mouse models: An opportunity for greater understanding of the multifactorial etiology of neural tube defects. Birth Defects Res 109, 140–152 (2017).
Amendola, E. et al. A mouse model demonstrates a multigenic origin of congenital hypothyroidism. Endocrinol. 146, 5038–47 (2005).
Nadeau, J. H. Modifier genes in mice and humans. Nat Rev Genet 2, 165–74 (2001).
Gillis, J. & Pavlidis, P. “Guilt by Association” is the exception rather than the rule in gene networks. PLoS Comput Biol 8, e1002444 (2012).
Schaefer, M. H., Serrano, L. & Andrade-Navarro, M. A. Correcting for the study bias associated with protein-protein interaction measurements reveals differences between protein degree distributions from different cancer types. Front. Genet. 6, 260 (2015).
Acknowledgements
The authors thank Dr. Nadia Schoenmakers and Professor Eamonn Maher for helpful comments on our manuscript. This work was supported by funding from King Abdullah University of Science and Technology (KAUST) Office of Sponsored Research (OSR) under Award No. URF/1/3454-01-01, FCC/1/1976-08-01, and FCS/1/3657-02-01. GVG acknowledges support from H2020-EINFRA (731075) and the National Science Foundation (IOS:1340112) as well as support from the NIHR Birmingham ECMC, NIHR Birmingham SRMRC and the NIHR Birmingham Biomedical Research Centre and the MRC HDR UK. The views expressed in this publication are those of the authors and not necessarily those of the NHS, the National Institute for Health Research, the Medical Research Council or the Department of Health.
Author information
Authors and Affiliations
Contributions
I.B. and R.H. conceived the experiments, I.B. conducted the experiments, I.B. and M.K. implemented the software and algorithms, I.B., P.N.S., G.V.G., R.H. analyzed the results. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boudellioua, I., Kulmanov, M., Schofield, P.N. et al. OligoPVP: Phenotype-driven analysis of individual genomic information to prioritize oligogenic disease variants. Sci Rep 8, 14681 (2018). https://doi.org/10.1038/s41598-018-32876-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32876-3
Keywords
This article is cited by
-
Oligogenic basis of premature ovarian insufficiency: an observational study
Journal of Ovarian Research (2024)
-
Faster and more accurate pathogenic combination predictions with VarCoPP2.0
BMC Bioinformatics (2023)
-
Linking common human diseases to their phenotypes; development of a resource for human phenomics
Journal of Biomedical Semantics (2021)
-
Recent innovations and in-depth aspects of post-genome wide association study (Post-GWAS) to understand the genetic basis of complex phenotypes
Heredity (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.