Abstract
Malaria has provided a major selective pressure and has modulated the genetic diversity of the human genome. The variants of the Duffy Antigen/Receptor for Chemokines (DARC) gene have probably been selected by malaria parasites, particularly the FY*O allele, which is fixed in sub-Saharan Africa and confers resistance to Plasmodium vivax infection. Here, we showed the influence of genomic ancestry on the distribution of DARC genotypes in a highly admixed Brazilian population and confirmed the decreased susceptibility of the FY*A/FY*O genotype to clinical P. vivax malaria. FY*B/FY*O individuals were associated with a greater risk of developing clinical malaria. A remarkable difference among DARC variants concerning the susceptibility to clinical malaria was more evident for individuals who were less exposed to malaria, as measured by the time of residence in the endemic area. Additionally, we found that DARC-negative and FY*A/FY*O individuals had a greater chance of acquiring high levels of antibodies against the 19-kDa C-terminal region of the P. vivax merozoite surface protein-1. Altogether, our results provide evidence that DARC polymorphisms modulate the susceptibility to clinical P. vivax malaria and influence the naturally-acquired humoral immune response to malaria blood antigens, which may interfere with the efficacy of a future vaccine against malaria.
Similar content being viewed by others
Introduction
Plasmodium vivax is the most widespread malaria parasite species outside Africa, accounting for 64% of malaria cases in endemic regions of the Americas, greater than 30% of cases in South East Asia and 40% of cases in the Eastern Mediterranean regions1. Moreover, malaria caused by P. vivax is no longer considered a benign disease because of the reports of deadly malaria caused by this parasite2,3,4,5.
To establish erythrocyte infection in humans, P. vivax is highly dependent on the interaction of the parasite Duffy binding protein (region II, PvDBPII) and its cognate receptor in the surface of reticulocytes, Duffy antigen receptor for chemokines (DARC) (recently renamed Atypical chemokine Receptor 1)6,7,8. The DARC gene has three common alleles, FY*A, FY*B and FY*O or FY*BES (also known as Duffy negative)9,10,11. The two functional alleles, FY*A and FY*B, are the result of one non-synonymous mutation (125 G > A, rs12075) and are associated with the Fya and Fyb blood-group antigens, respectively9. The non-functional allele FY*O is defined by a mutation in the gene promoter (−33T > C, rs2814778) that abrogates its expression in the erythrocyte cell lineage10,11. An investigation of the evolutionary history of DARC has estimated the age of the FY*O allele to be 42,000 years, likely older than most other mutations associated with malaria resistance12,13. Likewise, this ancient allele seems to have undergone fixation in Africa in a scenario in which all three common DARC alleles were segregated in this continent12. Therefore, the FY*O allele has been used as a marker for African ancestry. The FY*O allele has been shown to protect against P. vivax infection6. Different susceptibilities to P. vivax have also been associated with the Duffy blood group antigens (Fya and Fyb) because polymorphisms alter the binding to the parasite’s DBP, and densities of the antigen in the erythrocyte surface14,15.
Acquired immunity against infection and clinical protection in malaria has been somehow mediated by antibodies16. Antibodies against PvDBPII showed a natural occurrence in individuals from P. vivax endemic areas17,18,19. These antibodies could block the DBP-DARC interaction, inhibit the erythrocyte invasion of P. vivax20,21 and seem to be associated with clinical protection21,22. Because the DARC-negative promoter heterozygously expresses approximately half of the total amount of DARC14,23, both the DARC antigen and its expression level may play a major role in the immunity against P. vivax24,25. We and others have shown that the frequency of naturally acquired antibodies against PvDBPII is inversely correlated with the DARC expression levels24,25. In a population-based study in the Brazilian Amazon, we showed that the naturally acquired binding-inhibitory antibodies targeting PvDBPII were more frequent in heterozygous individuals carrying the non-functional DARC allele FY*O25. Similarly, individuals expressing less DARC from Colombian malaria-endemic areas exhibited higher frequencies and magnitudes of anti-PvDBPII IgG antibodies, as evaluated by conventional serology (ELISA-detected IgG)24.
In the current study, we investigated how polymorphisms in DARC modulate the susceptibility to P. vivax malaria in individuals differing in the levels of exposure to malaria in an agricultural settlement of the Amazon region. This population showed a high genetic admixture with a significant contribution of European and Native American ancestries, which reflected the distribution of DARC genotypes. A remarkable difference among DARC variants in the susceptibility to clinical P. vivax malaria was associated with low cumulative exposure to malaria, as measured by the time of residence in the endemic area. Additionally, evidence of the association between naturally acquired immune response to the 19-kDa C-terminal region of the P. vivax merozoite surface protein-1 (PvMSP119), a leading P. vivax vaccine candidate, and DARC variants was provided, for the first time, in a community-based study.
Methods
Study Area and Population
The study was carried out in the agricultural settlement of Rio Pardo, in the Presidente Figueiredo municipality, located in the Northeast of Amazonas State. The study area and malaria transmission patterns have been described in detail elsewhere25,26. Briefly, the settlement was officially created in 1996 by the National Institute of Colonization and Agrarian Reform (INCRA) as part of a large-scale colonization project focused on agriculture and wide-ranging human settlement in the Amazon area27. The Rio Pardo population consisted of 701 inhabitants according to the census conducted between September and October 2008. Most residents were native to the Amazon region, mainly from the Amazonas State. The population lives mainly on subsistence farming and fishing along the tributaries of the Rio Pardo River.
The study area was classified as hypo- to mesoendemic based on the spleen size of the local children and parasite infection rates26. Malaria transmission occurs throughout the year. P. vivax was responsible for most cases of malaria (average of 80%) during the period of study (2003–2009) in the region. The highest number of malaria cases were in 2004 to 2006 (the average annual parasite incidence (API) was 289 cases per 1000 inhabitants), followed by a continuous decrease in the number of cases; in 2009, the API was 54.6 (Health Surveillance Secretariat of the Brazilian Ministry of Health, SVS/MS).
Study design and cross-sectional surveys
The ethical and methodological aspects of this study were approved by the Ethics Committee of Research involving Human Subjects of Institute René Rachou/Fiocruz (Reports No. 07/2006, No. 07/2009, No. 26/2013, and No. 1.395.372), according to the Resolution of the Brazilian Council on Health-CNS 466/12. All participants signed a written informed consent, including the next of kin, caretakers, or guardians, on behalf of the minors/children enrolled in the study. All the methods were carried out in accordance with the approved guidelines.
A population-based open cohort study was initiated in November 2008, followed by two cross-sectional surveys six (June 2009) and twelve months (October-November 2009) after the initial survey. The cross-sectional surveys have been described in detail elsewhere26. For each resident, the number of P. vivax malaria episodes (mono and mixed infections with P. falciparum) and information about the parasite species determined by microscopy were obtained from the Epidemiological Surveillance System for Malaria (SIVEP-Malaria) for the period residing in the Rio Pardo community between 2003 (when the data were available in the database) and 2009. In Brazil, all cases of malaria are compulsorily reported to the Ministry of Health through the national database SIVEP-Malaria28. The study area registered a median of 321 (Range = 168–391) P. vivax malaria cases, 21 (Range = 5–236) P. falciparum malaria cases and 3 (Range = 1–13) mixed cases by these two species between 2003 and 2009.
Four hundred ninety-eight subjects were enrolled at the baseline, 495 at the 2nd survey, and 495 in the 3rd survey. In total, 690 subjects were successful genotyped to DARC polymorphisms. The genomic ancestry was inferred for a subset of 339 unrelated subjects (first-degree relatives). Serological assays were carried out for 207 individuals to PvDBPII and 222 individuals to PvMSP119 who were enrolled in the three cross-sectional surveys.
Laboratory diagnosis of malaria
For participants at the cross-sectional surveys, malaria infection was diagnosed by microscopy of Giemsa solution-stained thick smears and real-time PCR amplification of a species specific segment of the 18S rRNA gene of human malaria parasites as previously described29. Briefly, real-time PCR was performed using a consensus pair of oligonucleotides (PL1473F18 [5′-TAACGAACGAGATCTTAA-3′] and PL1679R18 [5′- GTTCCTCTAAGAAGCTTT-3′]), and each 20 µL reaction mix contained 2 μL of genomic DNA (approximately 100 ng), 10 µL of Sybr Green PCR master mix (Applied Biosystems, Foster City, CA, USA), 2.5 mM MgCl2 and 0.5 μM of each oligonucleotide. The PCR conditions consisted of an initial denaturation step at 95 °C for 10 min, followed by 40 cycles of 20 sec at 90 °C, 30 sec at 50 °C and 30 sec at 60 °C. After amplification, the melting curves were observed from the dissociation curves resulting from continuous measurements of fluorescence at 530 nm during which the temperature was gradually increased from 60 °C to 95 °C. The amplification and fluorescence detection were performed using the ABI PRISM 7500 Sequence Detection System (Applied Biosystems).
The Giemsa solution-stained smears were evaluated by experienced microscopists, according to the malaria diagnosis guidelines of the Brazilian Ministry of Health. For real-time PCR, genomic DNA was extracted from either EDTA whole-blood samples (adults and children ≥ 5 years in age) or dried blood spots on filter paper (children < 5 years in age) using the Puregene blood core kit B (Qiagen, Minneapolis, MN, USA) or QIAmp DNA mini kit (Qiagen), respectively, according to the manufacturers’ instructions.
Genotyping of Ancestry-Informative Markers (AIMs)
The subjects were typed for a set of 48 single-nucleotide polymorphisms (SNPs) previously validated as a useful tool for ancestry estimation in population samples from diverse origin30 (Supplementary Table S1). This subset of markers was chosen using the ln algorithm31 and was defined as the most ancestry-informative markers (AIMs) distinguishing one or more of the four continental populations (European, African, Native American, and East Asian). The AIMs were genotyped by real-time PCR with specific hydrolysis probes for each SNP (Applied Biosystems). The assays were performed in a total volume of 5 µL and in the presence of 2.5 µL of Taqman® Universal PCR Master Mix 2 × (Applied Biosystems), 0.25 µL of Genotyping Assay Mix (Applied Biosystems), 1.25 µL of water and 1 µL of DNA (approximately 10 ng). The cycling parameters for the PCR were as follows: initial denaturation at 95 °C for 10 min, 40 cycles of 15 sec at 95 °C and 1 min at 60 °C. Amplification and fluorescence detection were carried out using the ViiA 7 Real-Time PCR System (Applied Biosystems).
Ancestry estimates
We estimated the individual ancestry using the method implemented in the program Structure 2.3.432,33. The program implements a Bayesian clustering method to infer population structure and an admixture using a Markov Chain Monte Carlo (MCMC) procedure. The burn-in was set at 50,000 steps followed by 500,000 MCMC steps, and a model of correlated allele frequencies was specified. Thirty replicates were performed assuming three clusters (K = 3) because the Brazilian population was formed by an admixture from three parental populations: Native Americans, Europeans and Africans. For all runs, lambda was set to 1.0, α parameter was estimated from the data, and the priori information for the individuals from parental populations to assist the clustering was not used (USEPOPINFO = 0). All replicates were evaluated using CLUMPP34, and specific solutions were plotted using DISTRUCT 1.135. To avoid the bias caused by the family structure in our population structure analyses, we excluded related samples that were identified using individual data of family name and household information.
The data of parental populations (European, African and Native American) were a public dataset obtained from the HGDP-CEPH Diversity Panel (http://www.hagsc.org/hgdp/files.html) and Hapmap Project (http://hapmap.ncbi.nlm.nih.gov/) and Kosoy et al.30. We downloaded all genotypes for 611 unrelated individuals, which included the following: 60 CEPH Europeans (CEU, Utah Residents with Northern and Western European ancestry), 128 European Americans (NYCPEA), 80 Yoruban Africans (Nigeria), 19 Bini West Africans (Edo State, Nigeria), 23 Kanuri West Africans (Northern Nigeria), 12 Bantus (Kenya), 31 Biaka Pygmies (Central African Republic), 15 Mbuti Pygmies (Democratic Republic of Congo), 24 Mandenka individuals (Senegal), 6 San individuals (Namibia), 75 Mayan Amerindians (Mexico and Guatemala), 25 Pimas (Mexico), 13 Colombians (Colombia), 24 Karitiana Amerindians (Brazil), 21 Surui Amerindians (Brazil), 29 Nahua Amerindians (Central Mexico), and 26 Quechuan Amerindians (Peru). The NYCPEA individuals were from the New York City, and their classification was based on the self-classification of ethnicity36.
Additionally, to represent the genetic structure of our samples, we performed principal component analysis (PCA) of individual genotypes, as implemented in Adegenet and Ade4 for the R environment37. The non-linear iterative partial least squares (NIPALS) algorithm implemented in Ade4 was applied to consider the missing values.
DARC Genotyping
We genotyped two SNPs in the FY gene (−33T > C and 125 G > A) that defines the three common DARC alleles: FY*A, FY*B, and FY*O. SNPs were genotyped by real-time PCR with allele-specific oligonucleotides, as previously described25,38. Briefly, the 20-µL reaction mix contained 50–100 ng of genomic DNA, 10 µL of SYBR Green PCR master mix (Applied Biosystems) and 0.1–1.0 µM of each oligonucleotide (Supplementary Table S2). The amplification and fluorescence were detected using the ABI Prism 7000 Sequence Detection System (Applied Biosystems). The cycling parameters for the real-time PCR comprised a cycle of 95 °C for 10 min, followed by 35 cycles of 95 °C for 15 s and 60 °C for 1 min. After amplification, melting curves were observed from the dissociation curve. Genotyping results were confirmed for approximately 10% of samples, including all FY*O/FY*O individuals, by sequencing a 942-bp fragment of the DARC gene, comprising polymorphic positions −33T > C and 125 G > A, as previously described25.
Serological Assays
We carried out enzyme-linked immunosorbent assay (ELISA) for total IgG antibodies against PvDBPII and PvMSP119. The recombinant protein PvDBPII included amino acids 243–573 (region II) of the Sal-1 PvDBPII variant; the recombinant protein was expressed as a 39-kDa 6 × His fusion protein, as previously described39. The oligonucleotides used to amplify the fragment of PvDBPII by PCR were as follows: sense (5′-TTTGGATCCACGATCTCTAGTGCTATTATAAATC-3′) and anti-sense (5′-AAACTCGAGTGTCACAACTTCCTGAGTATTTTT-3′). The recombinant protein representing PvMSP119, which contains amino acids 1616–1704 of the full-length MSP1 protein, was expressed as a 6× His fusion protein in Escherichia coli, as described elsewhere40,41,42. The PCR was performed using the sense (5′-ACCAGCCATATGACTATGAGCTCCGAGCACACA-3′) and anti-sense (5′-CGGATCCTCGCTACAGAAAACTCCCTCAAA-3′) oligonucleotides41. ELISA was carried out as previously described18 with serum samples diluted at 1:100 and PvDBPII and PvMSP119 at final concentrations of 5 μg/mL and 1 μg/mL, respectively. The sera from individuals naturally exposed to P. falciparum failed to cross-react in the ELISA assay with the same construction of the recombinant protein PvMSP119 used in the present study42. The results were expressed as the reactivity index (RI), calculated by dividing the reading values of the test (OD values) by the cut-off (mean reading for the unexposed group plus 3 SD, n = 30). Values of RI > 1.0 were considered positive.
Statistical Methods
A database was created using Epidata software (http://www.epidata.dk). The Kruskal–Wallis test with Dunn’s post-hoc test and the Mann-Whitney U test were performed to compare the differences in the antibody response during the follow-up period. Fisher’s exact test was performed to compare the distribution of the frequency of DARC genotypes among areas of residence in the Rio Pardo community. Associations among age, number of malaria episodes, and time of residence in the Amazon region were estimated using the Spearman’s rank correlation test and the R cor.test function. Because of the high correlation between age and the time of residence in the Amazon region, we used the last predictor as a measure of exposure to malaria in further association analysis. All statistical analyses were performed using R software (version 3.4). A P-value < 0.05 was considered significant in all analyses.
Genetic Population Analysis
The Hardy-Weinberg equilibrium for each SNP was calculated using the R package SNPassoc43. To investigate how the ancestry-based population subdivision influenced the Rio Pardo population structure, the association between homozygosity excess (as measured by FIT for each SNP) and ancestry (estimated by FST for each SNP between the European and Native American population, the two main ancestry sources of the Rio Pardo population) was estimated using the Spearman’s rank correlation test44. The population FIT was estimated by averaging FIT across SNPs. We used Arlequin 3.545 to calculate the observed and expected heterozygosities under Hardy–Weinberg equilibrium (HO and HE, respectively) and then estimated FIT = (HE − HO)/HE for each SNP. FST was estimated using R package hierfstat46. The FIS statistic for the Rio Pardo population was estimated using Arlequin 3.5.
Regression Analysis
Multinomial logistic regression was applied to describe the association between individual genomic ancestry and the frequency of DARC genotypes. The models were fitted using the multinom function available in the R package nnet47.
The logistic regression model with stepwise backward deletion was applied to describe the association between the antibody to P. vivax blood antigens and covariates: DARC genotype, number of slide-positive malaria episodes recorded between 2003 and 2009, recent P. vivax malaria infection (determined from SIVEP-Malaria and self-reported malaria episodes), time of residence in the Amazon region and place of residence in Rio Pardo community. Covariates were selected for inclusion in the logistic models if they were associated with the outcome at the 15% level of significance in exploratory unadjusted analysis. The models were fitted using the R glm function. Only variables associated with statistical significance at the 5% level were maintained in the final model. The goodness of fit was assessed by comparing the deviance of candidate models and by the Akaike information criteria (AIC) statistics48. Because of missing values, 195 of 196 observations remained in the final logistic regression model.
Zero-inflated Poisson Regression (ZIP) was carried out to assess the relationship between the number of clinical P. vivax malaria that each subject experienced in a period of seven years (2003–2009) and covariates: DARC genotype, time of residence in the Amazon region and place of residence in Rio Pardo community. The offset of the ZIP regression was defined as the time that an individual lived in the Rio Pardo community during the study period. ZIP regression analyses were performed using zeroinfl function in the R package pscl49. The goodness of fit was assessed by comparing the deviance of candidate models and by Akaike information criteria (AIC) statistics48. The candidate models were tested by leave-one-out cross-validation and we chose the model with the least cross-validation error (the generalized Pearson statistic50 and the sum of squared error) that indicated the best predictive model. Because of missing values, 633 of 690 observations remained in the final ZIP regression model.
Clustering Analysis
K-means cluster analyses were performed on the IgG total antibodies against blood-stage antigens (PvMSP119 and PvDBPII) data using the R cluster function. We used the antibody response against each antigen measured in three periods (baseline, six and twelve months after the initial survey) to generate k clusters. Cluster numbers between 2 and 15 were evaluated and the k clusters with the lowest total within groups sum of squares was determined as the appropriate number of clusters. Clustering data were visualized using the clusplot function in the R package cluster51.
Results
Population Structure in the Amazonian Settlement of Rio Pardo
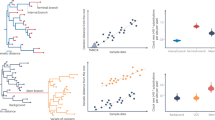
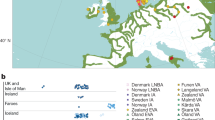
The genomic ancestry of 339 unrelated subjects was estimated by genotyping a validated set of 48 AIMs dispersed in the human genome. The allele frequencies at these loci are shown in Supplementary Table S3. One locus showed departure from Hardy–Weinberg equilibrium (HWE, P < 0.01) in the Rio Pardo population (Table S2). Population-based analyses indicated that the Rio Pardo population has strong departure of Hardy–Weinberg equilibrium (FIT = 0.04, 95% CI 0.02–0.06), which could be the result of the high level of inbreeding estimated for this population (FIS = 0.03, P = 0.015) and ancestry-based population subdivision (ρFIT-FST = 0.33, P = 0.022). The Bayesian cluster approach to inferring the genetic population structure showed that the AIMs used discriminated among European, African and Native American parental populations (Fig. 1a). The mean estimates of ancestry for the individuals from the Rio Pardo settlement pointed to a highly admixed population with significant contribution of European (44.1%, 95% CI 41.8–46.5) and Native American (37.6%, 95% CI 35.4–39.8) ancestries, but with low African ancestry (18.3%, 95% CI 16.9–19.7). Individual ancestry inference was confirmed by PCA analysis, which placed the Rio Pardo population between Europeans and Native Americans (Fig. 1b). The first two principal components (PCs) account for 95% of the variance, where PC1 clearly separates the Africans from all other groups, and PC2 clearly separates the Europeans from Native Americans. Because this set of AIMs was originally selected to maximize the differences among European Americans, Africans and Native Americans, it was not surprising that both approaches clearly distinguished the three parental populations30.
Genomic ancestry of the Rio Pardo population. (a) Genomic individual ancestry bar plot, including worldwide populations and the Rio Pardo population. Each individual is represented by a thin vertical line that is partitioned into three colored segments corresponding to the individual’s estimated membership proportion in the European (blue), African (green) and Native American (red) clusters. (b) PCA representation of individual ancestry. EURO, European; AMER, Native American; AFRI, African and BRAZ, Brazilian (Rio Pardo population).
Distribution of DARC Genotypes Among Subjects from the Rio Pardo Settlement According to Genomic Ancestry
Initially, we determined the DARC genotypes of 690 subjects from the Rio Pardo population and found a high prevalence of FY*A/FY*B (29.6%), FY*A/FY*A (23.2%) and FY*A/FY*O (20.1%) genotypes (Table 1). On the other hand, the FY*O/FY*O genotype was observed at a very low frequency (3.0%). In general, there was a predominance of the functional DARC alleles FY*A (48.0%) and FY*B (33.6%) in the study population. This locus was not found to be in HWE in the Rio Pardo population (P = 0.035), with a deficiency in the heterozygous DARC genotypes (60.3%) compared with what was expected (62.3%). Similarly, analysis with a subset of unrelated subjects also showed that this locus was not in HWE (HO = 57.0% and HE = 63.0%, P = 0.0005).
We further described the association between DARC genotypes and the estimated individual genomic ancestry by fitting a multinomial logistic regression. The odds of having the FY*B/FY*B [relative risk ratio (RRR): 20.7; 95% CI 3.7–115.9; P = 0.001] and FY*A/FY*A (RRR 4.2; 95% CI 0.9–19.1; P = 0.06) genotypes increased continuously with the increase in the European and Native American ancestry, respectively, compared with the FY*A/FY*B genotype (Fig. 2a,b). An opposite trend was observed for FY*O allele carriers (RRR 0.11; 95% CI 0.02–0.64; P = 0.013 for FY*A/FY*O; RRR 0.06; 95% CI 0.01–0.43; P = 0.005 for FY*B/FY*O) and the FY*B/FY*B (RRR 0.08; 95% CI 0.01–0.47; P = 0.006) genotype to Native American ancestry. As expected, the probability of having the FY*O/FY*O genotype increased as the individual proportion of African ancestry increased (RRR > 1000; P = 0.00) (Fig. 2c).
Frequency distribution of DARC genotypes according to the estimated genomic ancestry for the Rio Pardo population. Fitted multinomial logistic regression models showing the association between the distribution of DARC genotypes and (a) European, (b) Native American and (c) African ancestry. The individual proportion of genomic ancestry is shown in the x-axis. The y-axis is labeled on the probability scale and shows the variation in the probability of a genotype along with the variation in ancestry proportions.
DARC Affects the Susceptibility to Plasmodium vivax malaria, as influenced by the Time of Malaria Exposure
To evaluate whether DARC has influenced the susceptibility to clinical P. vivax malaria the number of malaria episodes was recorded for each of the 690 participants during a period of seven years (2003–2009). The median number of episodes was zero but showed great variation (Range = 0–24) (Table 1). During the cross-sectional surveys (2008–2009), the overall prevalence of malaria (as detected by both microscopy and real-time PCR) was 7.0% (35 of 498) at baseline, with 89% of infections caused by P. vivax. At the 2nd and 3rd surveys, the overall frequencies of malaria infection were 1.6% (8 of 495) and 7.5% (37 of 495), respectively. All infections were caused by P. vivax at the 2nd survey, while 89% of infections were caused by P. vivax at the 3rd survey. No P. malariae or mixed Plasmodium infections were diagnosed by either microscopy or real-time PCR in the three cross-sectional surveys. The individuals had an average age of 25 years (IQR = 11–45) and lived for 22 years (IQR = 11–38) in the malaria endemic area (Brazilian Amazon region). In the Rio Pardo community, the time of residence was shorter (median = 6 years), partly reflecting the recent establishment of the settlement (approximately 20 years).
Next, we sought to assess the potential influence of demographic, epidemiologic and genetic factors on the P. vivax malaria incidence expressed as the total number of malaria episodes experienced per individual in a period of seven years. Three hundred twenty-two (47% of 690) subjects had at least one symptomatic P. vivax malaria episode between 2003 and 2009 (Supplementary Table S4). The number of malaria episodes was inversely and weakly correlated with age (ρ = −0.10, P = 0.010) and the time of residence in the Amazon region (ρ = −0.09, P = 0.018). As expected, because this was a stable population established mainly by migrants from the Amazon region, age was highly correlated with the time of residence in the Amazon region (ρ = 0.94, P < 0.001). Furthermore, a previous study in the community suggested spatial clustering of transmission, with a higher risk of malaria infection among people living along and around the local streams (riverine population)26.
Zero-inflated Poisson Regression (ZIP) analysis adjusted for the place of residence and time of residence in the endemic area showed a reduction in the risk of clinical P. vivax malaria of 19% (95% CI 2–32; P = 0.029) and 91% (95% CI 67–97; P = 0.0003), respectively, for subjects with the FY*A/FY*O and FY*O/FY*O genotypes compared with those with the FY*A/FY*B genotype (Table 2). By contrast, FY*B/FY*O individuals showed a greater risk (26%, 95% CI 3–53, P = 0.023) of clinical malaria compared with those with the reference genotype FY*A/FY*B. Of importance, the difference in susceptibility to malaria among DARC genotypes was decreased according to the increase in residence time in the endemic area (Fig. 3a,b). We observed a reduction of 3% (95% CI 2.5–3.4; P < 0.0001) in the risk for P. vivax malaria per additional year of residence in the endemic area and a variable risk depending on the transmission level in the place of residence (Fig. 3a,b; Supplementary Table S5). For instance, the riverine population was at an increased risk of P. vivax malaria (Fig. 3a).
Analysis of the risk of clinical P. vivax malaria for the Rio Pardo population. The predicted number of clinical malaria episodes according to the DARC genotype, time of residence in the Amazon region and place of residence in the Rio Pardo community were derived from the Zero-inflated Poisson regression model. (a) Area along or close to the Rio Pardo stream with a higher risk of malaria infection (Riverine area) and (b) Non-riverine area with a lower risk of malaria infection.
Antibody response varies with the DARC genotype
We next investigated whether the association between DARC variants and risk of P. vivax malaria was reflected in the acquisition of IgG total antibodies against blood-stage antigens. Here, to obtain a more comprehensive understanding of naturally acquired P. vivax immunity, we evaluated the antibody response against two leading blood-stage P. vivax vaccine candidates spanning a range of immunogenicity, the PvDBPII and PvMSP119 antigens. Antigen-specific total IgG levels were measured for each subject at baseline and after 6 and 12 months. The anti-PvDBPII response was evaluated for 30% (207/690) of subjects who participated in the three cross-sectional surveys. The median antibody response measured by RI at baseline, 6 months and 12 months were 0.50 (IQR = 0.21–2.12), 0.47 (IQR = 0.22–1.78) and 0.57 (IQR = 0.24–1.96), respectively (Kruskal–Wallis test, P = 0,695). We found no association between the antibody response against PvDBPII and DARC variants in this population (Supplementary Table S6).
Overall, the anti-PvMSP119 response was evaluated for 32% (222/690) of participants in the three cross-sectional surveys. The median antibody responses measured by RI at baseline, 6 months and 12 months were 1.35 (IQR = 0.80–2.50), 1.37 (IQR = 0.79–2.30) and 1.03 (IQR = 0.67–2.03), respectively (Kruskal–Wallis test, P = 0.007). An association between the antibody response against PvMSP119 and DARC variants was found at 6 months (Supplementary Table S7). We performed clustering analysis using the anti-PvMSP119 antibody levels measured in the three cross-sectional surveys to allow more consistent identification of the different profiles of antibody responders considering that the levels of malaria transmission fluctuated in the study area. Individuals were clustered into two groups, the low (group 1, N = 166) and high (group 2, N = 30) antibody level group (Fig. 4a). The frequency of non-responders to PvMSP119 (RI < 1.0) in the low-response group varied between 42% and 57% across surveys. Both the frequency and magnitude of the antibody response to PvDBPII was lower than those of the anti-PvMSP119 response in the three cross-sectional surveys (Table 3 and Fig. 4b). Due the differences in the immunogenicity of these merozoite proteins, clustering analysis was performed for both antigens separately. The high-response group included older individuals who have lived for a longer time in the malaria-endemic area and have experienced a higher number of malaria episodes than the low-response group (Table 3). In addition, 50% of subjects from the high-response group have experienced a P. vivax malaria episode recently (Table 3). We found a higher proportion of individuals from the high-response group living close to the river (Riverine area) (Table 3). The frequency and levels of anti-PvDBPII antibodies were significantly different between the two groups, probably due to differences in malaria exposure between the groups (Table 3). We previously demonstrated that the subject’s age, which was correlated with cumulative exposure to malaria in the study population, was a strong predictor of seropositivity to PvDBPII26. Additionally, the distribution of the frequency of the DARC genotypes differed significantly between the low- and high-response groups (Table 3).
IgG total antibody levels against PvMSP119 during the 12-month follow-up study in the Rio Pardo Settlement. (a) Individuals were clustered according to the anti-PvMSP119 antibody levels evaluated in three cross-sectional surveys (baseline, 6 months and 12 months): Group 1: low level of response; Group 2: high level of response. The two components explained 90% of the point variability. (b) Anti-PvMSP119 antibody levels in the follow up for the low-response (light blue) and high-response (dark blue) groups. The results were expressed as the reactivity index (RI).
The low- and high-response groups showed an anti-PvMSP119 response significantly different in all three periods (Mann-Whitney U test, P < 0.001) (Fig. 4b). A logistic regression model was adjusted for the covariates: DARC genotype, number of malaria episodes, recent P. vivax malaria infection, time of residence in the Amazon region and place of residence in the Rio Pardo community. Nevertheless, the only variables significantly associated with anti-the PvMSP119 antibody levels were the DARC genotype, recent P. vivax infection and time of residence in the endemic area. By fitting the logistic regression, individuals with the FY*A/FY*O genotype showed a higher chance (OR = 4.8; 95% CI 1.2–24; P = 0.032) to acquire high anti-PvMSP119 antibody levels than the reference genotype FY*A/FY*B (Fig. 5). These individuals frequently showed high levels of the anti-PvMSP119 antibody (Fig. 6). The FY*O/FY*O genotype was strongly associated with a high-response status (OR = 24, 95% CI 3–207; P = 0.003). This pattern of high antibody levels against PvMSP119 was observed for all individuals with the FY*O/FY*O genotype, including individuals not included in logistic regression analysis due to missing data (Supplementary Fig. S1). Here, we observed that a longer period living in the malaria endemic area is associated with a slightly increased chance (OR = 1.04; 95% CI 1.02–1.07; P = 0.001) to acquire antibodies against PvMSP119. In addition, individuals who had malaria recently (in the previous 6 months) showed an increased chance to develop anti-PvMSP119 antibodies (OR = 4.8; 95% CI 1.9–13; P = 0.0008).
Association between the DARC genotype and antibody levels against PvMSP119 during the 12-month follow-up study in the Rio Pardo Settlement. The logistic regression model was adjusted for the time of residence in the endemic area and recent P. vivax infection. Only variables associated with anti-PvMSP119 antibody levels at a significance level of 0.05 (P < 0.05) were maintained in the final model. The odds ratio (OR, 95% CI) was adjusted using FY*A/FY*B as the reference group. The text in bold indicates that the OR is statistically significant.
Individual anti-PvMSP119 response evaluated in three cross-sectional surveys (baseline, 6 months and 12 months) according to the DARC genotype. The results were expressed as the reactivity index (RI). The antibody response data for low- and high-response groups are colored by light blue and dark blue, respectively.
Discussion
Malaria has provided a major selective pressure and has modulated the genetic diversity in the recent history of the human genome52. Growing evidence has pointed to erythrocyte variants that have resulted from evolutionary selection by malaria53,54,55. The variants of the DARC gene that apparently do not result in disease were probably selected by malaria parasites, particularly the FY*O allele56. Hence, most of the population of sub-Saharan Africa, where this allele is highly frequent, show the remarkable absence of P. vivax infection6. The ethnic differences in susceptibility and diverse genetic adaptations to P. vivax malaria for the other DARC variants are less clear. Here, we showed the influence of genomic ancestry on the distribution of DARC genotypes in the highly admixed Brazilian population and reinforced the recent finding that the FY*A/FY*O genotype is associated with decreased susceptibility to P. vivax malaria in a population-based follow-up. Conversely, the FY*B/FY*O individuals showed a greater risk of clinical P. vivax malaria. Of note, these differences in the susceptibility to clinical P. vivax malaria become more evident depending on the time of malaria exposure.
The distribution of DARC alleles shows a pattern of striking global divergence, whereas the FY*O allele is close to fixation (i.e., frequencies of 100%) in sub-Saharan Africa, and the most prevalent FY*A allele has become dominant across south-east Asia57. On the other hand, the ancestral FY*B allele is the least prevalent with a more limited distribution, being more frequent in Europe and parts of the Americas57. In the Americas, including Brazil, all three alleles are present, reflecting the great allelic heterogeneity. The highly heterogeneous Brazilian population was formed by the admixture between Native Amerindians, Europeans colonizers or immigrants, and African slaves58. Although the European component is preponderant in the Brazilian population (≈70%), with a slightly increasing trend from North to South of Brazil59,60,61, in the small settlement of Rio Pardo, a much lesser contribution of the European component (≈44%) was found. A significant Native American contribution (≈38%) was also estimated for this population, which comprises mostly natives of the Amazon region, mainly from the Amazonas State. Our data show that the high level of inbreeding and ancestry-based population subdivision are responsible to shape the genetic structure of this native population. Similar estimates of Native American ancestry were observed in individuals from the capital Manaus, which is located 160 km from the Rio Pardo settlement62. This genetic ancestry profile is consistent with the demographic history of northern Brazil, where land colonization and the genetic admixture process were different from other geographic regions of the country63. This process was characterized by the presence of numerous original indigenous populations64, social policies encouraging mating between European men and indigenous women during centuries of Brazilian colonization as a strategy for the occupation of the region, and the restricted use of African slave labor in the Amazon region63,65,66. Only after the 1960s occurred the frontier expansion in the Amazon as a result of the introduction of large-scale colonization projects and wide-ranging human settlement sponsored by the government67.
We further investigated the impact of ancestry on the distribution of DARC variants in the highly admixed population from the Rio Pardo community. A higher frequency of FY*A allele carriers and, consequently, the Fya+b− phenotype (i.e., FY*A/FY*A and FY*A/FY*O, 43%) was observed compared with the Fya-b+ phenotype (i.e., FY*B/FY*B and FY*B/FY*O, 24%). A consistent finding is that the probability of having the FY*B/FY*B genotype or heterozygosity for the FY*O allele decreases with the increase in the Native American component. On the other hand, the probability of having the FY*B/FY*B genotype increases as the individual proportion of European ancestry increases. Considering that similar proportions of the European and Native American components together account for 82% of the diversity in individual genetic ancestry in the Rio Pardo population, a complex pattern of admixture with the predominant carriers of functional alleles FY*A and FY*B was found. Noticeably, this complexity is increased by the percentage of African ancestry present in individuals who are associated with a higher probability of carrying the FY*O allele. The frequency distribution of DARC genotypes described here differs from that reported in other populations in the Amazon region, except for the FY*A/FY*B and FY*O/FY*O genotypes, which are, respectively, the most frequent (≈30%) and rare genotypes (≈3%) in northern Brazil15,68,69,70. The main differences were found for the FY*A/FY*O and FY*B/FY*B genotypes (20% and 13% in the Rio Pardo population, respectively), which showed median frequencies of 12% (Range = 7–15) and 18% (Range = 14–22), respectively, in the Amazon region15,68,69,70. The differences in the genotype frequencies observed for the Amazon population reflects the global scenario of the ethnic heterogeneity of the Brazilian population. Interestingly, individuals from the Rio Pardo community did not fit Hardy–Weinberg expectation. Here, we considered two possibilities to explain this result: demographic disequilibrium due to inbreeding and natural selection acting on the DARC locus. Indeed, a high level of inbreeding was estimated for this population.
To account for a role of natural selection in the pattern of distribution of DARC variants, we investigated the possible association between DARC genotypes and P. vivax malaria epidemiology. In the study area, FY*A/FY*B was the most frequent genotype, followed by the FY*A/FY*A and FY*A/FY*O genotypes. For other areas of the Brazilian Amazon, FY*A/FY*B has also been described as the major variant15,68,69. Although the FY*O/FY*O frequency was very low in the study area, we found three of 21 individuals infected by P. vivax during the study period. Therefore, the FY*O/FY*O genotype was associated with a very low risk of clinical P. vivax malaria in the Rio Pardo community. Plasmodium vivax infection among DARC-negative individuals has been previously described in the Brazilian Amazon71. We also found a reduced risk of clinical malaria for individuals with the FY*A/FY*O genotype compared with those with the FY*A/FY*B. By contrast, FY*B/FY*O presented a greater risk to develop clinical P. vivax malaria. These results are in accordance with previous findings describing the effect of different DARC genotypes on the risk of clinical malaria in a northern Brazilian population15. In this previous study, individuals with the FY*A/FY*O genotype showed a higher magnitude of reduction regarding the risk of clinical malaria (80%) than those with the FY*A/FY*B genotype. Additionally, subjects with the FY*B/FY*B genotype showed a greater risk (270%) of clinical P. vivax malaria. The differences in the strength of association among DARC variants and clinical malaria observed between the studies may be related to epidemiological, demographic and genetic particularities of each population. In fact, we showed that the differences among DARC variants in the susceptibility to malaria become more evident according to the residence time in the endemic area and, therefore, may reflect the immune status to malaria of the subject. Accordingly, the population of the previous study was a frontier settlement constituted mainly by migrants from the malaria-free regions of Southeast and South Brazil72 who had been living for a median of 14 years in the Brazilian Amazon19. In this population, in contrast to what we observed in the Rio Pardo population, there was a higher proportion of the FY*B/FY*B and FY*B/FY*O genotypes in addition to the FY*A/FY*B genotype15. Considering the complexity of malaria epidemiology, we cannot rule out the possibility that other factors related to the parasite (e.g., P. vivax variants) or the host (e.g., immune response and genetic variants in the blood group systems) that were not considered in both studies could also be responsible for the observed differences. Furthermore, the analysis of a higher number of individuals may be necessary to evaluate the risk of clinical malaria among the DARC variants. Despite these limitations, the findings from both studies are consistent with a protective effect of the FY*A allele and increased susceptibility of FY*B.
An important point to be investigated was how susceptibility to P. vivax infection influenced by DARC has modulated the immune response to malaria antigens. We evaluated the antibody response against two leading P. vivax vaccine candidates, the PvDBPII and PvMSP119 antigens. Of importance, although both antigens are expressed during the blood-stage cycle, they show very different profiles of an antibody response that likely is influenced by the host and parasite features26. Here, we found no association between the conventional antibody response against PvDBPII, as detected by ELISA, and DARC variants. These results confirmed our previous findings in which the frequencies of ELISA-detected IgG antibodies were similar between individuals carrying one or two DARC functional alleles (FY*A and FY*B)25. However, the association between DARC variants and susceptibility to P. vivax malaria seems to be reflected in the acquisition of anti-PvMSP119 antibodies. A major finding was that DARC-negative individuals had a greater chance to acquire high levels of anti-PvMSP119 antibodies in the entire follow-up period than FY*A/FY*B individuals. This is an unexpected result because most P. vivax isolates do not proceed to the blood-stage cycle in DARC-negative individuals. Another interesting result was that FY*A/FY*O individuals had a higher chance to acquire high levels of anti-PvMSP119 antibodies than FY*A/FY*B individuals, although FY*A/FY*O individuals showed a reduced risk to develop clinical malaria. Likewise, individuals with the FY*A/FY*O genotype from the Caribbean coast of Colombia showed a higher trend to acquire anti-PvMSP-119 than those with the FY*A/FY*B genotype24. Until now, two studies have investigated the immune response to P. vivax antigens regarding the DARC variants. In the African-Colombian population from the Colombian Pacific coast, it was found that the N terminus of PvMSP1 (200 L fragment) was similarly recognized by DARC-positive and -negative individuals73. Conversely, the frequency and magnitude of anti-PvMSP119 antibodies were higher in DARC-positive individuals from the Colombia’s Caribbean coast24. However, this result should be interpreted with caution because both the frequency and levels of the antibody response to PvMSP119 were very low in the study population24.
The reasons for the association found between DARC genotypes (Fy*O/Fy*O and Fy*A/Fy*O) and the anti-PvMSP119 antibody response are not clear; however, we have raised three not mutually exclusive possibilities. First, the host immune response may be activated during the liver stage of parasite development because MSP1 is expressed during this stage, as shown for P. falciparum and P. berghei74,75. Thus, although most of the P. vivax isolates do not infect DARC-negative reticulocytes, the host immune response might be activated and/or boosted during the liver stage. Another possibility is the occurrence of P. vivax parasites adapted to use other receptors for reticulocyte invasion in the study area. Recently, DNA expansion of PvDBP was described in DARC-negative P. vivax infections, suggesting that it may be selected to allow low-affinity binding to another receptor on DARC-negative erythrocytes76. Finally, the lowest binding between PvDBP and DARC has been shown for FY*O allele carriers, who express decreased amounts of DARC compared with carriers of two functional alleles14,15,19,77. Therefore, it is possible to speculate that the low efficiency of invasion of FY*A/FY*O reticulocytes by merozoites might increase PvMSP119 exposure to the immune system and boost the anti-PvMSP119 antibody response.
In conclusion, this community-based study provides concluding remarks about modulation of the susceptibility of P. vivax malaria in a hypo to mesoendemic area such as the Amazon Brazilian region. We have shown how genomic ancestry is related to the distribution frequency of DARC genotypes in a population formed mainly by natives from the Amazon and characterized by the high contribution of Native Amerindian ancestry. In this population with an increased frequency of the FY*A allele, our results are consistent with a significant protective effect of the FY*A/FY*O genotype. Conversely, the FY*B/FY*O genotype is associated with increased susceptibility to clinical P. vivax malaria. Of importance, the differences in the susceptibility to malaria among DARC variants seem to be influenced by the time of exposure to malaria. Additionally, our findings indicate that differences in the susceptibility to malaria among DARC variants may modulate the immune response to malaria blood antigens such as PvMSP119. Together, these findings have important implications for vaccine development and indicate the need to consider the level of exposure to malaria to evaluate differences in DARC mediating P. vivax susceptibility.
Data Availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Who. World Malaria Report 2017 196 (2017).
Price, R. N. et al. Vivax malaria: Neglected and not benign. Am. J. Trop. Med. Hyg. 77, 79–87 (2007).
Genton, B. et al. Plasmodium vivax and mixed infections are associated with severe malaria in children: a prospective cohort study from Papua New Guinea. PLoS Med. 5, e127 (2008).
Tjitra, E. et al. Multidrug-resistant Plasmodium vivax associated with severe and fatal malaria: a prospective study in Papua, Indonesia. PLoS Med. 5, e128 (2008).
Lacerda, M. V. et al. Postmortem characterization of patients with clinical diagnosis of Plasmodium vivax malaria: to what extent does this parasite kill? Clin. Infect. Dis. 55, e67–74 (2012).
Miller, L., Mason, S., Clyde, D. & McGinniss, M. The resistance factor to Plasmodium vivax in blacks. The Duffy-blood-group genotype, FyFy. N. Engl. J. Med. 295, 302–304 (1976).
Barnwell, J., Nichols, M. & Rubinstein, P. In vitro evaluation of the role of the Duffy blood group in erythrocyte invasion by Plasmodium vivax. J. Exp. Med. 169, 1795–1802 (1989).
Wertheimer, S. P. & Barnwell, J. W. Plasmodium vivax interaction with the human Duffy blood group glycoprotein: identification of a parasite receptor-like protein. Exp. Parasitol. 69, 340–350 (1989).
Iwamoto, S., Omi, T., Kajii, E. & Ikemoto, S. Genomic organization of the glycoprotein D gene: Duffy blood group Fya/Fyb alloantigen system is associated with a polymorphism at the 44-amino acid residue. Blood 85, 622–626 (1995).
Mallinson, G., Soo, K., Schall, T., Pisacka, M. & Anstee, D. Mutations in the erythrocyte chemokine receptor (Duffy) gene: the molecular basis of the Fya/Fyb antigens and identification of a deletion in the Duffy gene of an apparently healthy individual with the Fy(a-b-) phenotype. Br. J. Haematol. 90, 823–829 (1995).
Tournamille, C., Colin, Y., Cartron, J. & Le Van Kim, C. Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nat. Genet. 10, 224–228 (1995).
McManus, K. F. et al. Population genetic analysis of the DARC locus (Duffy) reveals adaptation from standing variation associated with malaria resistance in humans. PLoS Genet. 13, e1006560 (2017).
Hedrick, P. W. Population genetics of malaria resistance in humans. Heredity 107, 283–304 (2011).
Michon, P. et al. Duffy-null promoter heterozygosity reduces DARC expression and abrogates adhesion of the P. vivax ligand required for blood-stage infection. FEBS Lett. 495, 111–114 (2001).
King, C. L. et al. Fy(a)/Fy(b) antigen polymorphism in human erythrocyte Duffy antigen affects susceptibility to Plasmodium vivax malaria. Proc. Natl. Acad. Sci. USA 108, 20113–20118 (2011).
Longley, R. J., Sattabongkot, J. & Mueller, I. Insights into the naturally acquired immune response to Plasmodium vivax malaria. Parasitology 143, 154–170 (2016).
Fraser, T. et al. Expression and serologic activity of a soluble recombinant Plasmodium vivax Duffy binding protein. Infect. Immun. 65, 2772–2777 (1997).
Ceravolo, I. P. et al. Anti-Plasmodium vivax Duffy binding protein antibodies measure exposure to malaria in the Brazilian Amazon. Am. J. Trop. Med. Hyg. 72, 675–681 (2005).
Souza-Silva, F. A. et al. Naturally acquired antibodies to Plasmodium vivax Duffy binding protein (DBP) in Brazilian Amazon. Am. J. Trop. Med. Hyg. 82, 185–193 (2010).
Grimberg, B. T. et al. Plasmodium vivax invasion of human erythrocytes inhibited by antibodies directed against the Duffy binding protein. PLoS Med. 4, 1940–1948 (2007).
King, C. et al. Naturally acquired Duffy-binding protein-specific binding inhibitory antibodies confer protection from blood-stage Plasmodium vivax infection. Proc. Natl. Acad. Sci. USA 105, 8363–8368 (2008).
Nicolete, V. C., Frischmann, S., Barbosa, S., King, C. L. & Ferreira, M. U. Naturally Acquired Binding-Inhibitory Antibodies to Plasmodium vivax Duffy Binding Protein and Clinical Immunity to Malaria in Rural Amazonians. J. Infect. Dis. 214, 1539–1546 (2016).
Zimmerman, P. et al. Emergence of FY*A(null) in a Plasmodium vivax-endemic region of Papua New Guinea. Proc. Natl. Acad. Sci. USA 96, 13973–13977 (1999).
Maestre, A. et al. Acquired antibody responses against Plasmodium vivax infection vary with host genotype for duffy antigen receptor for chemokines (DARC). PLoS One 5, e11437 (2010).
Souza-Silva, F. A. et al. Duffy Antigen Receptor for Chemokine (DARC) Polymorphisms and Its Involvement in Acquisition of Inhibitory Anti-Duffy Binding Protein II (DBPII) Immunity. PLoS One 9, e93782 (2014).
Kano, F. S. et al. Plasmodium vivax Duffy binding protein: baseline antibody responses and parasite polymorphisms in a well-consolidated settlement of the Amazon Region. Trop. Med. Int. Health 17, 989–1000 (2012).
de Castro, M. C., Monte-Mor, R. L., Sawyer, D. O. & Singer, B. H. Malaria risk on the Amazon frontier. Proc. Natl. Acad. Sci. USA 103, 2452–2457 (2006).
Oliveira-Ferreira, J. et al. Malaria in Brazil: an overview. Malar J. 9, 115 (2010).
Mangold, K. A. et al. Real-time PCR for detection and identification of Plasmodium spp. J. Clin. Microbiol. 43, 2435–2440 (2005).
Kosoy, R. et al. Ancestry informative marker sets for determining continental origin and admixture proportions in common populations in America. Human mutation 30, 69–78 (2009).
Rosenberg, N. A., Li, L. M., Ward, R. & Pritchard, J. K. Informativeness of genetic markers for inference of ancestry. Am. J. Hum. Genet. 73, 1402–1422 (2003).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332 (2009).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007).
Rosenberg, N. DISTRUCT: a program for the graphical display of population structure. Mol. Ecol. Notes 4, 137–138 (2004).
Mitchell, M. K., Gregersen, P. K., Johnson, S., Parsons, R. & Vlahov, D. The New York Cancer Project: rationale, organization, design, and baseline characteristics. J. Urban Health 81, 301–310 (2004).
Jombart, T. & Ahmed, I. Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27, 3070–3071 (2011).
Sousa, T. N., Sanchez, B. A. M., Ceravolo, I. P., Carvalho, L. H. & Brito, C. F. A. Real-time multiplex allele-specific polymerase chain reaction for genotyping of the Duffy antigen, the Plasmodium vivax invasion receptor. Vox Sanguinis 92, 373–380 (2007).
Ntumngia, F. B. et al. Conserved and variant epitopes of Plasmodium vivax Duffy binding protein as targets of inhibitory monoclonal antibodies. Infect. Immun. 80, 1203–1208 (2012).
Barbedo, M. B. et al. Comparative recognition by human IgG antibodies of recombinant proteins representing three asexual erythrocytic stage vaccine candidates of Plasmodium vivax. Mem. Inst. Oswaldo Cruz 102, 335–339 (2007).
Cunha, M. G., Rodrigues, M. M. & Soares, I. S. Comparison of the immunogenic properties of recombinant proteins representing the Plasmodium vivax vaccine candidate MSP1(19) expressed in distinct bacterial vectors. Vaccine 20, 385–396 (2001).
Rodrigues, M. H. et al. Serological detection of Plasmodium vivax malaria using recombinant proteins corresponding to the 19-kDa C-terminal region of the merozoite surface protein-1. Malar J. 2, 39 (2003).
Gonzalez, J. R. et al. SNPassoc: an R package to perform whole genome association studies. Bioinformatics 23, 644–645 (2007).
Kehdy, F. S. et al. Origin and dynamics of admixture in Brazilians and its effect on the pattern of deleterious mutations. Proc. Natl. Acad. Sci. USA 112, 8696–8701 (2015).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Goudet, J. Hierfstat, a package for r to compute and test hierarchical F-statistics. Mol. Ecol. Notes 5, 184–186 (2005).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S (Springer, NY, USA, 2002).
Sakamoto, Y., Ishiguro, M. & Kitagawa, G. Akaike Information Criterion Statistics (Boston: Reidel Publishing Company, 1986).
Achim, Z., Christian, K. & Simon, J. Regression Models for Count Data in R. J. Stat. Softw. 27 (2008).
McCullagh, P. & Nelder, J. A. Generalized Linear Models (Chapman and Hall Ltd, London, United Kingdom, 1989).
Pison, G., Struyf, A. & Rousseeuw, P. J. Displaying a Clustering with CLUSPLOT. Comput. Stat. Data Anal. 30, 381–392 (1999).
Kwiatkowski, D. P. How malaria has affected the human genome and what human genetics can teach us about malaria. Am. J. Hum. Genet. 77, 171–192 (2005).
Allen, S. J. et al. alpha+ −Thalassemia protects children against disease caused by other infections as well as malaria. Proc. Natl. Acad. Sci. USA 94, 14736–14741 (1997).
Genton, B. et al. Ovalocytosis and cerebral malaria. Nature 378, 564–565 (1995).
Ruwende, C. et al. Natural selection of hemi- and heterozygotes for G6PD deficiency in Africa by resistance to severe malaria. Nature 376, 246–249 (1995).
Hamblin, M. T. & Di Rienzo, A. Detection of the signature of natural selection in humans: Evidence from the Duffy blood group locus. Am. J. Hum. Genet. 66, 1669–1679 (2000).
Howes, R. E. et al. The global distribution of the Duffy blood group. Nat. Commun. 2, 266 (2011).
Salzano, F. M. & Freire-Maia, N. Populações Brasileiras: Aspectos Demográficos, Genéticos e Antropológicos (Companhia EditoraNacional, São Paulo, Brazil, 1967).
Callegari-Jacques, S. M. et al. Historical genetics: spatiotemporal analysis of the formation of the Brazilian population. Am. J. Hum. Biol. 15, 824–834 (2003).
Pena, S. D. et al. The genomic ancestry of individuals from different geographical regions of Brazil is more uniform than expected. PLoS One 6, e17063 (2011).
Lins, T. C., Vieira, R. G., Abreu, B. S., Grattapaglia, D. & Pereira, R. W. Genetic composition of Brazilian population samples based on a set of twenty-eight ancestry informative SNPs. Am. J. Hum. Biol. 22, 187–192 (2010).
Saloum de Neves Manta, F. et al. Revisiting the genetic ancestry of Brazilians using autosomal AIM-Indels. PLoS One 8, e75145 (2013).
Benchimol, S. Amazônia: Formação Social e Cultural (Valer, Manaus, Brazil, 1999).
Carneiro-da-Cunha, M. História dos Índios no Brasil (Companhia das letras, São Paulo, Brazil, 1992).
Batista dos Santos, S. E., Rodrigues, J. D., Ribeiro-dos-Santos, A. K. & Zago, M. A. Differential contribution of indigenous men and women to the formation of an urban population in the Amazon region as revealed by mtDNA and Y-DNA. Am. J. Phys. Anthropol. 109, 175–180 (1999).
Carneiro-da-Cunha, M. História dos Índios no Brasil 9–24 (Companhia das Letras, São Paulo, 1992).
Moran, E. F. In Change in the Amazon Basin: The Frontier after a Decade of Colonization Vol. II (ed. J. Hemming) 91–102 (Manchester University Press, 1985).
Cavasini, C. E. et al. Duffy blood group gene polymorphisms among malaria vivax patients in four areas of the Brazilian Amazon region. Malar J. 6, 167 (2007).
Albuquerque, S. R. et al. FY polymorphisms and vivax malaria in inhabitants of Amazonas State, Brazil. Parasitol. Res. 106, 1049–1053 (2010).
Carvalho, T. A. et al. Plasmodium vivax infection in Anajas, State of Para: no differential resistance profile among Duffy-negative and Duffy-positive individuals. Malar J. 11, 430 (2012).
Cavasini, C. E. et al. Plasmodium vivax infection among Duffy antigen-negative individuals from the Brazilian Amazon region: an exception? Trans. R. Soc. Trop. Med. Hyg. 101, 1042–1044 (2007).
da Silva-Nunes, M. et al. Malaria on the Amazonian frontier: transmission dynamics, risk factors, spatial distribution, and prospects for control. Am. J. Trop. Med. Hyg. 79, 624–635 (2008).
Herrera, S. et al. Antibody response to Plasmodium vivax antigens in Fy-negative individuals from the Colombian pacific coast. Am. J. Trop. Med. Hyg. 73, 44–49 (2005).
Suhrbier, A., Holder, A. A., Wiser, M. F., Nicholas, J. & Sinden, R. E. Expression of the precursor of the major merozoite surface antigens during the hepatic stage of malaria. Am. J. Trop. Med. Hyg. 40, 351–355 (1989).
Szarfman, A. et al. Allelic forms of gp195, a major blood-stage antigen of Plasmodium falciparum, are expressed in liver stages. J. Exp. Med. 167, 231–236 (1988).
Gunalan, K. et al. Role of Plasmodium vivax Duffy-binding protein 1 in invasion of Duffy-null Africans. Proc. Natl. Acad. Sci. USA 113, 6271–6276 (2016).
Woolley, I. J., Hotmire, K. A., Sramkoski, R. M., Zimmerman, P. A. & Kazura, J. W. Differential expression of the Duffy antigen receptor for chemokines according to RBC age and FY genotype. Transfusion 40, 949–953 (2000).
Acknowledgements
We thank the inhabitants of Rio Pardo for their enthusiastic participation in the study, the local malaria control team in Presidente Fiqueiredo for their logistic support, and the units of Oswaldo Cruz Foundation in Manaus, AM (Fiocruz-AM) and Belo Horizonte, MG (Fiocruz-Minas) for overall support. The authors thank the Program for Technological Development in Tools for Health-PDTIS-FIOCRUZ for use of the DNA sequencing (RPT01E) and Real-Time PCR (RPT09D) facilities. I.S.S., C.A.B., C.J.F., L.H.C., M.A.C. and T.N.S. thank Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the research scholarship support. A.M.S. thanks the Research Foundation of Minas Gerais (FAPEMIG) for scholarship support. L.M.T. thanks the Coordination for the Improvement of Higher Education Personnel (CAPES) for scholarship support. This work was supported by CNPq, PAPES VI/CNPq/FIOCRUZ and FAPEMIG.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: C.F.A.B., F.S.K., L.H.C. and T.N.S. Performed the experiments: A.M.S., B.A.M.S., C.J.F.F., F.S.K., F.A.S.S., I.S.S., L.M.T., M.A.C. and T.N.S. Analyzed the data: A.M.S., F.S.K., M.A.C. and T.N.S. Contributed reagents/materials/analysis tools: C.F.A.B., F.S.K. and L.H.C. Writing - original draft: C.F.A.B., F.S.K., L.H.C. and T.N.S. Writing - review & editing: C.F.A.B., I.S.S., F.S.K., L.H.C. and T.N.S.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kano, F.S., de Souza, A.M., de Menezes Torres, L. et al. Susceptibility to Plasmodium vivax malaria associated with DARC (Duffy antigen) polymorphisms is influenced by the time of exposure to malaria. Sci Rep 8, 13851 (2018). https://doi.org/10.1038/s41598-018-32254-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32254-z
Keywords
This article is cited by
-
Epidemiology of Plasmodium vivax in Duffy negatives and Duffy positives from community and health centre collections in Ethiopia
Malaria Journal (2024)
-
Differential transmissibility to Anopheles arabiensis of Plasmodium vivax gametocytes in patients with diverse Duffy blood group genotypes
Malaria Journal (2023)
-
Vivax malaria in Duffy-negative patients shows invariably low asexual parasitaemia: implication towards malaria control in Ethiopia
Malaria Journal (2022)
-
Duffy blood system and G6PD genetic variants in vivax malaria patients from Manaus, Amazonas, Brazil
Malaria Journal (2022)
-
Plasmodium—a brief introduction to the parasites causing human malaria and their basic biology
Journal of Physiological Anthropology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.