Abstract
Methanol dehydrogenase (MDH), an NAD+-dependent oxidoreductase, reversibly converts formaldehyde to methanol. This activity is a key step for both toxic formaldehyde elimination and methanol production in bacterial methylotrophy. We mutated decameric Bacillus methanolicus MDH by directed evolution and screened mutants for increased formaldehyde reduction activity in Escherichia coli. The mutant with the highest formaldehyde reduction activity had three amino acid substitutions: F213V, F289L, and F356S. To identify the individual contributions of these residues to the increased reduction activity, the activities of mutant variants were evaluated. F213V/F289L and F213V/F289L/F356S showed 25.3- and 52.8-fold higher catalytic efficiency (kcat/Km) than wild type MDH, respectively. In addition, they converted 5.9- and 6.4-fold more formaldehyde to methanol in vitro than the wild type enzyme. Computational modelling revealed that the three substituted residues were located at MDH oligomerization interfaces, and may influence oligomerization stability: F213V aids in dimer formation, and F289L and F356S in decamer formation. The substitutions may stabilise oligomerization, thereby increasing the formaldehyde reduction activity of MDH.
Similar content being viewed by others
Introduction
Increased atmospheric CO2 is a major cause of global warming and climate change. Many in vitro studies have examined single or multiple enzyme systems to reduce CO2 and convert it to valuable chemicals and fuels1,2,3. Converting CO2 into methanol using a multi-enzyme system is one of the most promising possible routes, recycling the greenhouse gas to efficiently produce a fuel alternative. In comparison to gas-based fuels (CO, methane, etc.) produced by the single-enzyme route, the liquid methanol produced by multi-enzyme reaction has a much higher energy capacity and is easier to transport4. Converting CO2 to methanol involves the consecutive reactions of three enzymes: formate dehydrogenase reduces CO2 to formate, formaldehyde dehydrogenase reduces formate to formaldehyde, and alcohol dehydrogenase (ADH) reduces formaldehyde to methanol. This sequence has been studied in various reaction conditions: with encapsulation of the three enzymes in a porous silica sol-gel matrix5, immobilization of the enzymes and their cofactor β-nicotinamide adenine dinucleotide (NAD+) in polystyrene particles and TiO2 nanoparticles6,7, with artificial photosynthesis, by combining a photochemical cofactor reducing system with the enzymatic conversion system8, and in a thermodynamic feasibility investigation of multiple enzyme reactions9.
Methanol dehydrogenase (MDH) is an oxidoreductase that catalyses the reversible interconversion of formaldehyde and methanol10. Two classes of MDHs are expressed by methylotrophic bacteria: the pyrroloquinoline quinone-containing MDHs, which are found mostly in Gram-negative and strictly aerobic methylotrophs, and the NAD(P) + -dependent cytoplasmic MDHs, commonly found in thermophilic, Gram-positive bacteria11. Although the former has been previously used to convert CO2 to methanol12, the latter is more suitable because of their use of reduced β-nicotinamide adenine dinucleotide (NADH) as a cofactor13.
The NAD-dependent MDHs belong to ADH group III14. Three NAD-dependent MDHs are found in Bacillus methanolicus MGA3 and PB115. MDH2 and MDH3 on the chromosome of MGA3 are 96% homologous to each other and approximately 60% to MDH on the plasmid16. MDH from B. methanolicus C1, mostly studied for structures and characteristics, contains one zinc ion and two magnesium ions at its active site17,18 and acts as a decamer19. Methanol oxidation by MDH is important for methylotrophic growth. However, kinetic substrate affinity and Vmax values of MDH is higher for formaldehyde than for methanol16,20. Although the biological significance of formaldehyde reduction by MDH is not clear, MDH can be used for the reduction of formaldehyde in a sequential reaction for the conversion of carbon dioxide to methanol.
Most studies on CO2 conversion have focused on finding the optimal reaction conditions for commercial enzymes. Generally, the methanol yields reported under regular conditions by naturally evolved enzymes have been very low9, indicating the need to develop more efficient CO2 conversion enzymes. However, since the MDH protein structure at the atomic level has not been determined, it is difficult to develop novel enzymes using a rational design system.
In this study, we generated mutants of MDH of B. methanolicus C1 by directed evolution and screened them for improved formaldehyde reduction activity. The influences of the mutant residues on the increased reduction activity were simulated by computational structure modelling.
Results and Discussion
Screening of MDH mutants with increased formaldehyde reduction activity
To develop a screening system for MDH mutants with increased formaldehyde reduction activity, we exploited the cytotoxicity differences between formaldehyde and methanol in Escherichia coli. The minimal inhibitory concentration (MIC) of formaldehyde in E. coli is approximately 5 mM21, while that of methanol is >3 M22. When we treated E. coli BL21 (DE3) cells expressing wild type MDH with various concentration of formaldehyde, cell growth was arrested at concentrations >2 mM (data not shown). Therefore, we expected that if mutant MDH variants efficiently converted formaldehyde to methanol, the host cells would be able to survive in a medium containing 3 mM formaldehyde and would dominate the mutant library population. To identify MDH mutants with higher reduction activity, error-prone polymerase chain reaction (PCR) was performed using mdh from B. methanolicus C1, with 2–8 bp substitutions per kb. The PCR products were cloned into the pET21b plasmid under the control of a T7 promoter, and approximately 3 × 104 BL21 (DE3) transformants were grown in media containing 3 mM formaldehyde. After 24 h of culture, cells were streaked on agar plates and plasmids in single colonies were isolated and sequenced to identify mutations in mdh. We selected 20 colonies for analysis, and 18 harboured the same three mutations (F213V, F289L, and F356S). This mutant was designated MDHmt1. The two other identified mutants had six and 18 amino acid substitutions, and were named MDHmt4 and MDHmt20, respectively. Their amino acid sequence changes are shown in Fig. 1.
Tolerance to formaldehyde by mutant MDHs
To analyse their tolerance to formaldehyde, E. coli BL21 (DE3) cells expressing MDHmt1, MDHmt4, and MDHmt20 were cultivated in 2YT media containing 3 mM formaldehyde, respectively. The cells displayed growth arrest until 15 h after formaldehyde addition; however, cells expressing MDHmt1, MDHmt4, and MDHmt20 started to grow after 17, 21, and 23 h, respectively (Fig. 2). The delayed growth of the cells may be due to formaldehyde evaporation and concentration decrease by MDH mutants. However, wild type MDH cells did not grow until after 27 h, at which time MDHmt1 had already reached the stationary phase. This observation of the cell growth at 3 mM formaldehyde concentration was due to formaldehyde reduction by MDH mutants. To confirm that the formaldehyde tolerance was indeed caused by the mutations, the experiments were repeated independently in triplicate, and the same observations were made. We concluded that the MDH mutants reduced formaldehyde to methanol faster than the wild type. Therefore, host cells expressing MDH mutants escaped from the growth arrest faster than those expressing the wild type.
Growth rates of E. coli cells expressing mutant MDH in 3 mM formaldehyde. Isopropyl β-D-1-thiogalactopyranoside (IPTG) was added at an OD600 of 0.6 to induce MDH expression, and after 1 h, 3 mM formaldehyde was added. Vector, negative control with no mdh gene; WT, wild type mdh gene; mt1, MDHmt1 gene; mt4, MDHmt4 gene; mt20, MDHmt20 gene.
Site-directed mutagenesis of MDH
MDHmt1 contained three amino acid substitutions: F213V, F289L, and F356S. Interestingly, all substitutions changed phenylalanine residues, which contain bulky benzyl side groups, to smaller residues. Primary sequence alignment of the MDH protein from B. methanolicus C1 revealed that it is 97% identical to MDH of MGA316. When three substituted sites were compared with those of MDHs of B. methanolicus MGA3 strain, it was observed that Phe289 and Phe213 were conserved in MDH. However, the Phe356 residue was changed to tyrosine in MDH of MGA3 and methionine in MDH2 and MDH316. To analyse which amino acid changes contributed to increased formaldehyde reduction activity, the three mutations were introduced separately and in combination to wild type MDH to form F213V, F289L, F356S, F213V/F289L, F289L/F356S, F213V/F356S, and F213V/F289L/F356S variants.
To examine the kinetic properties of formaldehyde reduction for the wild type and mutant forms of MDH, we determined the Michaelis-Menten kinetic parameters (Table 1) in vitro using the recombinantly produced and purified enzymes. The kinetic parameters (KM and Vmax) of wild type MDH from strain C1 was similar to that of MDH from MGA316. The F213V mutant showed a 3.4-fold higher catalytic efficiency (kcat/KM) than wild type MDH. All mutants with two amino acid substitutions exhibited higher catalytic efficiencies than wild type MDH. The mutant containing all three amino acid substitutions exhibited the highest catalytic efficiency, with a 52.8-fold increase compared with wild type MDH.
Next, we measured the NADH reduction rate for F213V/F289L and F213V/F289L/F356S (Fig. 3a), which showed the highest catalytic efficiency for formaldehyde reduction. The time required for NADH to decrease by half in the presence of the mutant MDHs (100 μg) was shorter than that for the wild type enzyme, taking 15, 29, and 53 min for the F213V/F289L/F356S mutant, the F213V/F289L mutant, and wild type MDH, respectively. When we measured the conversion ratio of formaldehyde to methanol for wild type MDH and the two mutants by high-performance liquid chromatography (HPLC), 0.26 ± 0.03, 0.79 ± 0.16, 1.48 ± 0.06, and 1.63 ± 0.02 mM of methanol was produced by wild type MDH, F213V, F213V/F289L, and F213V/F289L/F356S, respectively, from 30 mM formaldehyde in a 4 h reaction at room temperature (Fig. 3b). Also, to confirm the correlation between the concentration change of formaldehyde and methanol, the consumption of formaldehyde and the production of methanol by wile type MDH and F213V/F289L/F356S was measured in a time-dependent manner (Fig. S1). The conversion rates of formaldehyde under the tested conditions were low compared to the kinetic parameters, and the reason for this is unclear. However, one possible reason is that the reduction reaction in Fig. 3b was performed at room temperature to prevent formaldehyde and methanol evaporation, although the optimal temperature for B. methanolicus MDH activity is 50 °C23 and kinetic parameters were determined at this temperature. Increase in formaldehyde reduction further could be achieved by an additional screening of mutant MDHs with increased formaldehyde reduction activity, continuous supply of the substrate, and removal of products to prevent reversible methanol oxidation11.
Formaldehyde reduction activity of MDH variants. (a) Concentration of the cofactor NADH during the reduction reaction. WT, wild type MDH. (b) After reduction by each MDH variant for 1 h at 50 °C, methanol levels were measured by HPLC. The data represent the mean value of three independent experiments.
Homology modelling of MDH
To examine the roles of the amino acid residues changed in MDH mutants, protein structure analysis was performed. However, because the experimental structure for MDH has not been characterized, we built a monomeric structure of MDH using Pseudo Quadratic Restraints with Simulated Annealing (PQR-SA) homology modelling (Fig. S2a). The generated structure was validated using various protein quality scores, including the radius of gyration (Fig. S2b) and a score radar chart (Fig. S2c). B. methanolicus MDH shares its highest sequence identity (0.43) with Zymomonas mobilis MDH (Protein Data Bank (PDB): 3OX4.A). Maximum potential sequence identity with the template combination used was 0.77. The radius of gyration was slightly smaller than that of similarly sized proteins (Fig. S2b), indicating a more compact protein structure. The MDH homology model had pack1, clash, WhatRama, MolRama, ProRama, and rotamer protein quality scores within 50% (Fig. S2c). Therefore, the generated homology model was valid for use in further structural analyses.
Since the developed homology model contains no information on interacting ligands, we performed binding site prediction for the cofactor NAD+ using the Dockable Pocket Site Prediction (DPSP) method (Fig. S3). In total, 40 residues were identified as potential NAD+ interacting residues (residue numbers 37, 39, 40, 43, 45, 68, 69, 70, 94, 95, 96, 97, 98, 100, 136, 137, 140, 142, 145, 147, 148, 149, 158, 161, 177, 180, 181, 182, 185, 188, 189, 192, 196, 252, 257, 261, 265, 274, 275, and 276). Two analogue ligands, nicotinamide-adenine-dinucleotide phosphate and 5,6-dihydroxy-NADP, shared interacting residues in the binding pocket with NAD+. The three mutation sites had no direct connection to the NAD+ ligand-interacting residues. Thus, these mutated sites do not directly influence ligand binding during reduction. Therefore, we next focused on the structural changes caused by the MDH mutants.
Molecular dynamics simulations of wild type MDH, F213V/F289L, and F213V/F289L/F356S were performed to examine changes to the secondary and tertiary structure. The secondary structure compositions among the three MDH variants were very similar. Alpha-helix compositions were 54, 52, and 53%, respectively, and beta-sheet compositions were all 10%. The three structures were superimposed to measure their backbone similarities, and the root mean square deviations (RMSDs) were 2.7 Å for the wild type-F213V/F289L and F213V/F289L-F213V/F289L/F356S pairs, and 2.5 Å for the wild type-F213V/F289L pair. Therefore, the three structures were very similar, and the mutations did not dramatically change the tertiary structure of MDH.
Decameric structure of the MDH mutants
Because similar MDH proteins from thermotolerant Bacillus exist as decamers in electron microscopic images24, we prepared various oligomeric structures, including dimers, pentamers, and decamers. An oligomerization model was built from the monomer by PQR-SA, and was processed using the Chemistry at HARvard Macromolecular Mechanics (CHARMM) program25. Energy-minimized structures of various oligomers were analysed to determine how the mutant residues influenced oligomer stability and activity. Two decamer oligomerization pathways are possible: (1) two monomers form a dimer, and five dimers form a decamer (1 → 2 → 10 pathway), or (2) five monomers form a pentamer, and two pentamers form a decamer (1 → 5 → 10 pathway). Gel permeation chromatography suggested that most MDH structures existed as decamers and dimers (Fig. S4), suggesting the first pathway. Accordingly, first, a dimer structure was generated from the monomer homology structure. Two monomers were superimposed and structurally aligned based on known MDH dimer structure (PDB: 3OX4.A). Among mutants with only one amino acid change, only F213V showed higher catalytic activity than wild type MDH (Table 1). As shown in Fig. 4a–c, F213 was located in the interface of two monomers. Thus, the F213V substitution may increase reduction activity by influencing dimer formation. However, the other two mutations (F289 and F356) were not located on the dimer interface and therefore do not directly influence dimerization. F356 is located on the dimer surface, while F289 is buried in the structure (Fig. 4d–f). Pentamer and decamer structures were generated by five-fold symmetry rotation of five MDH monomers and dimers, by changing two geometrical variables, the inter-dimer distance of the centre of geometries of two neighbouring monomers, ranging from 40–50 Å in 2 Å increments, and the rotation angle of one of principal axes of the MDH monomer, ranging from 2–360° in 2° increments. The potential energy changes of the pentamer and decamer were calculated as functions of the inter-dimer distance and rotation angle to find the lowest energy conformation.
MDH dimer structure. Cartoon image (a–c) and molecular surface model (d–f) of the dimer. Chain A, red; chain B, blue. Mutation positions are shown in green space-fill models accompanied by the residue name and number. The dimer structures are oriented in the XY (a,d), XZ (b,e), and YZ (c,f) planes, respectively.
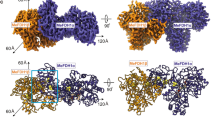
The generated model matched the electron microscopy data of Vonck et al.24 As shown in Fig. 5a, the measured energy profile indicated that two stable energy conformations existed, at rotation angles of 180° and 360° with 50 Å of inter-dimer distance. The decamer structures had unfavourable positive energy around 90° and 270° because neighbouring dimers collided when the inter-dimer distance was close, particularly at 40 Å. The side view of the decamer structure showed that the F289 and F356 residues were located in the proximity of the dimer-dimer interface. F289 was buried in the decamer model with the rotation angle of 180° and slightly exposed in the 360° rotation angle model. F289 and F356 were both located in the valley between dimers, and may influence decamer formation.
MDH decamer structure (a) Energy profile changes by rotation angle (R: x-axis) and inter-dimer distance (D: lines with different colours). (b) Top and side views of the decamer structure with (R, D) = (180°, 50 Å). (c) Top and side views of the decamer structure with (R, D) = (360°, 50 Å). The exposed mutation positions are shown in green and accompanied by residue numbers.
Conclusions
We have successfully improved the formaldehyde reduction activity of MDH by directed evolution and screening, which might potentially be useful for the conversion of CO2 to methanol. We could suggest the positional influence of the mutant with computational modelling. The three substituted residues were located at MDH oligomerization interfaces. The substitutions may stabilise oligomerization, thereby increasing the formaldehyde reduction activity of MDH.
Methods
Strains, media, and materials
E. coli DH5α cells were used for general DNA recombinant techniques and BL21 (DE3) cells were used for MDH expression. DH5α and BL21 (DE3) competent cells were purchased from Real Biotech Corporation (Taipei, Taiwan). Cells were grown in 2YT medium (16 g yeast extract, 10 g tryptone, and 5 g NaCl per litre) supplemented with appropriate antibiotics. Restriction enzymes were purchased from New England Biolabs (Ipswich, MA, USA). DNA polymerase and ligase were from Takara (Shiga, Japan). Ni-affinity chromatography column was from GE Healthcare (Little Chalfont, UK). All chemicals except formaldehyde used in enzymatic reactions, including the cofactors NAD+ and NADH, were purchased from Sigma-Aldrich (St. Louis, MO, USA). And methanol-free 16% formaldehyde was from Thermo Scientific (Waltham, MA, USA).
Directed evolution and mutant screening
The E. coli codon-optimized mdh (GenBank accession number M65004) from B. methanolicus C126 with a C-terminal histidine tag was synthesized by Bioneer (Daejeon, Korea) and randomly mutated using a Diversify® PCR Random Mutagenesis Kit (Clontech, Mountain View, CA, USA) with the primers mdh-forward (5′-GACATATGACAAACTTTTTCATTCC-3′) and mdh-reverse (5′-AACTCGAGCAGAGCGTTTTTGATG-3′) to produce 2–8 bp substitutions per kb, according to the manufacturer’s instructions. Underlines indicate restriction enzyme sites: CATATG (NdeI) and CTCGAG (XhoI). The wild type and mutated genes were digested with NdeI/XhoI, and ligated into pET-21b (Novagen, Billerica, MA, USA) digested with the same enzymes. E. coli DH5α was transformed with ligation mix and spread on 2YT agar plate supplemented with ampicillin. The size and quality of the constructed library were analysed as described previously27. BL21 (DE3) cells were transformed with the vectors, which were prepared from DH5α, by electroporation and cultivated on agar plate or in 2YT broth containing 100 μg/mL ampicillin at 37 °C. The number of transformants was calculated from the number of colonies on plates. To express MDH, 0.1 mM of isopropyl β-D-1-thiogalactopyranoside (IPTG) was added when the optical density at 600 nm (OD600) reached 0.6. After MDH expression for 1 h, 3 mM formaldehyde was added. After an additional 24 h incubation, the medium was streaked on 2YT agar plates containing 100 μg/mL ampicillin. Plasmids were isolated from single colonies and sequenced.
To analyse the effects of the mutant MDHs on formaldehyde reduction, the growth of cells harbouring wild type and mutant MDH variants was monitored in media containing formaldehyde. After induction of MDH expression for 1 h, 3 mM formaldehyde was added, and the OD600 was measured every 2 h, until formaldehyde was detoxified to levels enabling the resumption of cell growth. To exclude additional mutations within the gene, the mutant mdh genes were sequenced at every step.
Purification of MDH variants
BL21 (DE3) cells harbouring plasmids encoding wild type or mutant MDH were cultivated in 2 L 2YT broth at 37 °C and 180 rpm. When the OD600 reached 0.4, 0.1 mM IPTG was added. After 24 h, cells were harvested and resuspended in Buffer A (20 mM Tris pH 7.5, 0.5 M NaCl). Resuspended cells were disrupted by sonication (SONIFIER250, Branson, CT, USA). Then, the crude solution was centrifuged at 10,000 × g for 30 min. The supernatant was filtered through a 0.45 μm Minisart® filter (Sartorius Stedim Biotech, Gӧettingen, Germany). The filtered fraction was applied to a HisTrap™ FF Nickel affinity column (GE Healthcare) equilibrated with Buffer A. The bound proteins were eluted with 250 mM imidazole (pH 7.5). To eliminate remaining imidazole in the eluted protein solution and replace the buffer for subsequent experiments, an Amicon® Ultra-15 Centifugal Filter (Millipore, Bedford, MA, USA) was used. After purification, the apparent molecular masses of MDH were determined on a 4–12% gradient sodium dodecyl sulphate-polyacrylamide gel (NOVEX, San Diego, CA, USA). Protein concentrations were determined using the BCA™ Protein Assay Kit (Pierce, Rockford, IL, USA) using bovine serum albumin as a standard.
Gel permeation chromatography of native MDH
Purified MDHs were diluted in 100 mM potassium hydrogen phosphate buffer (PHPB; pH 7.5) and the final concentration was adjusted to 1 mg/mL for injection into a Superose 12 10/300 GL column (GE healthcare) on an AKTA fast protein liquid chromatography system at 4 °C. The column was equilibrated with PHPB. Purified MDH (0.5 mL) was loaded into the column and eluted at a flow rate of 1 mL/min. A calibration curve was prepared with a protein standard marker (#151–1901) purchased from Bio-Rad (Hercules, CA, USA).
Formaldehyde reduction assay
Formaldehyde reduction activity of MDH was assayed spectrophotometrically by detecting decreased NADH, as described previously17,24,28. Reactions were started by adding 100 μg of MDH to a reaction solution (500 μL) containing 100 mM PHPB, 0.6 mM NADH, and 3 mM formaldehyde, and the decrease in NADH was measured at 340 nm at room temperature.
The kinetic parameters of MDH with respect to formaldehyde reduction were determined in 100 mM PHPB (500 μL) containing 0.3 mM NADH for 20 min at 50 °C. The MDH content was fixed at 10 μg, while the concentration of formaldehyde varied from 0.1–10 mM. The initial rate of the reactions was determined by measuring the change in NADH. Experiments were performed in triplicate and repeated three times.
HPLC measurements of methanol and formaldehyde
The methanol produced from formaldehyde by MDH (100 μg) in a reaction solution (500 μL) containing 100 mM PHPB, 50 mM NADH, and 30 mM formaldehyde for 1 h at room temperature was measured using an Agilent HPLC station (Santa Clara, CA, USA), equipped with an Aminex HPX-87H column (300 × 7.8 mm, Bio-Rad) and a refractive index detector. HPLC-grade 0.01 N H2SO4 was used in the mobile phase, at a flow rate of 0.6 mL/min at 60 °C. A standard curve was generated with formaldehyde and methanol.
3D structure modelling
A protein structure for MDH from B. methanolicus C1 was built using homology modelling. HHblits29 was used to find suitable template structures in the PDB. The PQR-SA protocol30 was used to generate the homology structure from identified template structures by minimizing the target energy potential, consisting of the stereochemical energy, derived multiple alignment restraint energy from the PQR-SA protocol, and statistical torsion angle potential31. We predicted binding sites in the generated structure using the DPSP method (http://psb.kobic.re.kr/dpsp/).
To examine the influence of mutant residues on the structure, 10 ns molecular dynamics simulations of wild type and mutant MDHs were performed using CHARMM25. To consider the solvation effect, a generalized Born model with a simple switch function was used32. The initial structure, which was the lowest energy structure generated in homology modelling, was energy-minimized, and we performed Langevin dynamics with 2 fs intervals at room temperature. The RMSDs, secondary structure measurements, and time profiles of surface area changes were measured.
References
Du, X. L., Jiang, Z., Su, D. S. & Wang, J. Q. Research Progress on the Indirect Hydrogenation of Carbon Dioxide to Methanol. ChemSusChem 9, 322–332 (2016).
Highfield, J. Advances and recent trends in heterogeneous photo(electro)-catalysis for solar fuels and chemicals. Molecules 20, 6739–6793 (2015).
Kumar, M., Sundaram, S., Gnansounou, E., Larroche, C. & Thakur, I. S. Carbon dioxide capture, storage and production of biofuel and biomaterials by bacteria: A review. Bioresour. Technol. 247, 1059–1068 (2018).
Shi, J. et al. Enzymatic conversion of carbon dioxide. Chem. Soc. Rev. 44, 5981–6000 (2015).
Obert, R. & Dave, B. C. Enzymatic conversion of carbon dioxide to methanol: enhanced methanol production in silica sol-gel matrices. J. Am. Chem. Soc. 121, 12192–12193 (1999).
El-Zahab, B., Donnelly, D. & Wang, P. Particle-tethered NADH for production of methanol from CO2 catalyzed by coimmobilized enzymes. Biotechnol. Bioeng. 99, 508–514 (2008).
Sun, Q., Jiang, Y., Jiang, Z., Zhang, L. & Li, J. Green and Efficient Conversion of CO2 to Methanol by Biomimetic Coimmobilization of Three Dehydrogenases in Protamine-Templated Titania. Ind. Eng. Chem. Res. 48, 4210–4215 (2009).
Amao, Y. & Watanabe, T. Photochemical and enzymatic methanol synthesis from HCO 3− by dehydrogenases using water-soluble zinc porphyrin in aqueous media. Appl. Catal., B 86, 109–113 (2009).
Baskaya, F. S., Zhao, X., Flickinger, M. C. & Wang, P. Thermodynamic feasibility of enzymatic reduction of carbon dioxide to methanol. Appl. Biochem. Biotechnol. 162, 391–398 (2010).
Reid, M. F. & Fewson, C. A. Molecular characterization of microbial alcohol dehydrogenases. Crit. Rev. Microbiol. 20, 13–56 (1994).
Price, J. V., Chen, L., Whitaker, W. B., Papoutsakis, E. & Chen, W. Scaffoldless engineered enzyme assembly for enhanced methanol utilization. Proc. Natl. Acad. Sci. USA 113, 12691–12696 (2016).
Kuwabata, S., Tsda, R. & Yoneyama, H. Electrochemical conversion of carbon dioxide to methanol with the assistance of formate dehydrogenase and methanol dehydrogenase as biocatalysts. J. Am. Chem. Soc. 116, 5437–5443 (1994).
Muller, J. E., Heggeset, T. M., Wendisch, V. F., Vorholt, J. A. & Brautaset, T. Methylotrophy in the thermophilic Bacillus methanolicus, basic insights and application for commodity production from methanol. Appl. Microbiol. Biotechnol. 99, 535–551 (2015).
Arfman, N. et al. Properties of an NAD(H)-containing methanol dehydrogenase and its activator protein from Bacillus methanolicus. Eur. J. Biochem. 244, 426–433 (1997).
Heggeset, T. M. et al. Genome sequence of thermotolerant Bacillus methanolicus: features and regulation related to methylotrophy and production of L-lysine and L-glutamate from methanol. Appl. Environ. Microbiol. 78, 5170–5181 (2012).
Krog, A. et al. Methylotrophic Bacillus methanolicus encodes two chromosomal and one plasmid born NAD+ dependent methanol dehydrogenase paralogs with different catalytic and biochemical properties. PLoS One 8, e59188 (2013).
Arfman, N., Van Beeumen, J., De Vries, G. E., Harder, W. & Dijkhuizen, L. Purification and characterization of an activator protein for methanol dehydrogenase from thermotolerant Bacillus spp. J. Biol. Chem. 266, 3955–3960 (1991).
Kloosterman, H., Vrijbloed, J. W. & Dijkhuizen, L. Molecular, biochemical, and functional characterization of a Nudix hydrolase protein that stimulates the activity of a nicotinoprotein alcohol dehydrogenase. J. Biol. Chem. 277, 34785–34792 (2002).
Arfman, N. et al. Bacillus methanolicus sp. nov., a new species of thermotolerant, methanol-utilizing, endospore-forming bacteria. Int. J. Syst. Bacteriol. 42, 439–445 (1992).
Ochsner, A. M., Muller, J. E., Mora, C. A. & Vorholt, J. A. In vitro activation of NAD-dependent alcohol dehydrogenases by Nudix hydrolases is more widespread than assumed. FEBS Lett. 588, 2993–2999 (2014).
Mazzola, P. G., Jozala, A. F., Novaes, L. C. L., Moriel, P. & Penna, T. C. V. Minimal inhibitory concentration (MIC) determination of disinfectant and/or sterilizing agents. Braz. J. Pharm. Sci. 45, 241–248 (2009).
Ganske, F. & Bornscheuer, U. T. Growth of Escherichia coli, Pichia pastoris and Bacillus cereus in the presence of the ionic liquids [BMIM][BF4] and [BMIM][PF6] and Organic Solvents. Biotechnol. Lett. 28, 465–469 (2006).
Hektor, H. J., Kloosterman, H. & Dijkhuizen, L. Identification of a magnesium-dependent NAD(P)(H)-binding domain in the nicotinoprotein methanol dehydrogenase from Bacillus methanolicus. J. Biol. Chem. 277, 46966–46973 (2002).
Vonck, J. et al. Electron microscopic analysis and biochemical characterization of a novel methanol dehydrogenase from the thermotolerant Bacillus sp. C1. J. Biol. Chem. 266, 3949–3954 (1991).
Brooks, B. R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614 (2009).
de Vries, G. E., Arfman, N., Terpstra, P. & Dijkhuizen, L. Cloning, expression, and sequence analysis of the Bacillus methanolicus C1 methanol dehydrogenase gene. J. Bacteriol. 174, 5346–5353 (1992).
Hanson-Manful, P. & Patrick, W. M. Construction and analysis of randomized protein-encoding libraries using error-prone PCR. Methods Mol. Biol. 996, 251–267 (2013).
Arfman, N. et al. Methanol metabolism in thermotolerant methylotrophic Bacillus strains involving a novel catabolic NAD-dependent methanol dehydrogenase as a key enzyme. Arch. Microbiol. 152, 280–288 (1989).
Remmert, M., Biegert, A., Hauser, A. & Soding, J. HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9, 173–175 (2012).
Kim, T. R. et al. A simplified homology-model builder toward highly protein-like structures: an inspection of restraining potentials. J. Comput. Chem. 33, 1927–1935 (2012).
Kim, T. R., Yang, J. S., Shin, S. & Lee, J. Statistical torsion angle potential energy functions for protein structure modeling: a bicubic interpolation approach. Proteins 81, 1156–1165 (2013).
Im, W., Lee, M. S. & Brooks, C. L. 3rd Generalized born model with a simple smoothing function. J. Comput. Chem. 24, 1691–1702 (2003).
Acknowledgements
The Intelligent Synthetic Biology Center of the Global Frontier Project (2015M3A6A8065831) and the Pioneer Research Center Program (2013M3C1A3064780) of the National Research Foundation of Korea, the New & Renewable Energy Core Technology Program (No. 20153030101580) of the Korea Institute of Energy Technology Evaluation and Planning, and the Research Initiative Program of KRIBB supported this work.
Author information
Authors and Affiliations
Contributions
J.Y., B.H.S. and J.H.S. designed the experiments. J.Y. and B.H.S. constructed the M.D.H. mutant library and analysed enzyme activity. J.L. and G.T.L. revealed protein structure by computational modelling. J.Y. and K.D.K. acquired the kinetic parameters. B.H.S., S.G.L. and J.H.B. analysed the data. J.Y., J.L., B.H.S., S.C.K. and J.H.S. wrote the paper. S.C.K. and J.H.S. supervised the research. S.C.K. and J.H.S. conceived and oversaw the project. In addition to their contributions to experimental work and analysis, all authors reviewed manuscript drafts and helped to revise the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yi, J., Lee, J., Sung, B.H. et al. Development of Bacillus methanolicus methanol dehydrogenase with improved formaldehyde reduction activity. Sci Rep 8, 12483 (2018). https://doi.org/10.1038/s41598-018-31001-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31001-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.