Abstract
Gestational diabetes mellitus (GDM), a common pregnancy complication, is associated with an increased risk of maternal/perinatal outcomes. We performed a prospective observational explorative study in 41 GDM patients to evaluate their microbiota changes during pregnancy and the associations between the gut microbiota and variations in nutrient intakes, anthropometric and laboratory variables. GDM patients routinely received nutritional recommendations according to guidelines. The fecal microbiota (by 16S amplicon-based sequencing), was assessed at enrolment (24–28 weeks) and at 38 weeks of gestational age. At the study end, the microbiota α-diversity significantly increased (P < 0.001), with increase of Firmicutes and reduction of Bacteroidetes and Actinobacteria. Patients who were adherent to the dietary recommendations showed a better metabolic and inflammatory pattern at the study-end and a significant decrease in Bacteroides. In multiple regression models, Faecalibacterium was significantly associated with fasting glucose; Collinsella (directly) and Blautia (inversely) with insulin, and with Homeostasis-Model Assessment Insulin-Resistance, while Sutterella with C-reactive protein levels. Consistent with this latter association, the predicted metagenomes showed a correlation between those taxa and inferred KEGG genes associated with lipopolysaccharide biosynthesis. A higher bacterial richness and strong correlations between pro-inflammatory taxa and metabolic/inflammatory variables were detected in GDM patients across pregnancy. Collectively these findings suggest that the development of strategies to modulate the gut microbiota might be a potentially useful tool to impact on maternal metabolic health.
Similar content being viewed by others
Introduction
Gestational diabetes mellitus (GDM), one of the most common pregnancy complications, is associated with a moderately increased risk of maternal and perinatal outcomes1,2. Lifestyle interventions were reported to provide benefits to the health of GDM women and their babies1. It has been hypothesized that at least some of these beneficial effects might be due to the modulation of the maternal microbiota during pregnancy3,4,5,6,7. Indeed, variations in nutrient and energy intake were associated to specific bacterial abundance8,9,10. During the course of normal pregnancy, gut microbiota has been reported to remain relatively stable11 or to change dramatically, with an increase in Proteobacteria and Actinobacteria, a decline in butyrate-producing bacteria, a reduction in bacterial richness and within-subject (α) diversity, and higher between-subject (β) diversity at the end of pregnancy3. These modifications were supposed to favor the metabolic changes which support the healthy fetal growth, such as reduced insulin sensitivity and increased nutrient absorption3,7. Only few studies have evaluated the microbiota of GDM patients, showing contrasting results: either no differences3, decreased placental abundance of the Pseudomonadales order and Acinetobacter genus11, or increased placental Proteobacteria and reduced placental Bacteroidetes and Firmicutes12 have been reported in comparison with normoglycemic mothers. In addition, dysbiosis among GDM patients was reported to be associated with a few genus belonging to Firmicutes, Bacteroides and Actinobacteria phyla of the gut microbiota and the observed main differences in comparison to healthy women are relative to the gene contents of the gut microbes13. Two studies performed in women with previous GDM showed a relatively higher stool abundance of the Prevotellaceae family and a reduced abundance of the Firmicutes phylum14, or no differences in the gut microbiota composition15 in comparison to normoglycemic post-partum controls. Furthermore, increased gut relative abundance of the Ruminococcaceae family was associated with higher odds of developing GDM16. Finally, the cross-sectional design did not allow to draw conclusions about the causal relationships of the associations found.
In consideration of these highly divergent results, mostly derived from cross-sectional studies, we aimed to perform a prospective observational study evaluating the dynamic changes of the microbiota occurring during pregnancies of GDM women. All patients, after the diagnosis of GDM, underwent an educational dietary intervention, according to guidelines17, but the compliance with the provided recommendations is variable between women.
Therefore, the aims of our study were evaluating: i) whether the within-patient gut microbiota composition varied from the second to the third trimester of pregnancy; ii) whether patients with greater adherence to dietary recommendations presented a different microbial pattern than the less adherent ones; iii) whether changes in microbiota composition were associated with variations in nutrient intakes, anthropometric and laboratory variables; iv) whether specific microbiota oligotypes were implicates in these associations.
Results
Characteristics of the participants
Nine women did not return stool samples and were lost at follow-up. Data of 41 patients were therefore analyzed. The clinical characteristics of the participants did not differ from those of the 9 women who dropped out (data not shown).
Seven women (17.1%) gave birth before the 38th week. These patients provided the fecal and blood samples and the food questionnaire about a week before all the others (37th week); they did not differ with regard to nutritional, anthropometric, or metabolic characteristics when compared to the others.
Most participants were overweight women, with excessive fat intake and lower than recommended fiber consumption. From enrolment (24–28 weeks of gestational age) to the study end (38 weeks), weight and Body Mass Index (BMI) increased, and metabolic and inflammatory patterns of participants worsened, as usually occurs during the third trimester of pregnancy (Table 1).
Adherence to the dietary recommendations
After the dietary counselling, 34.1% (14/41) of the participants declared to be adherent to the given dietary recommendations. Characteristics at enrolment did not significantly differ between adherents and non-adherents, even if adherents showed increased values of weight and BMI (Table 2). Adherent women showed reduced intakes of sugars, and increased consumption of fiber, oligosaccharides, polyunsaturated fatty acids (PUFA) than non-adherents (Table 2). All participants had abolished alcohol consumption. Adherents had a better metabolic and inflammatory pattern, with a significantly greater reduction in fasting glucose and Homeostasis Model Assessment-Insulin Resistance (HOMA-IR) levels at the end of the study.
Microbiota composition at enrolment and at the study end
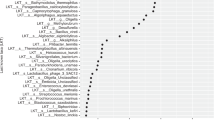
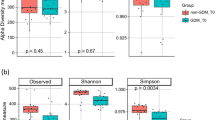
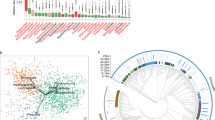
The microbiota α-diversity values were significantly different between subjects at enrolment when compared to subjects at the end of the study (P < 0.001). In details, species richness, number of different species and the Shannon index were significantly higher at the end of the study (P < 0.001) (Fig. 1). The analysis of microbial taxa abundance at phylum level showed an increase of Firmicutes at the study end, and a reduction of Actinobacteria and Bacteroidetes (Fig. 2). Going more deeply in the microbial composition, the level of diversity of the subjects based on the structure of their microbiota was clearly different across time (Supplementary Fig. S1). Moreover, Principal Component Analysis (PCA) based on microbiota composition (Fig. 3) revealed a significant relationship between genus-level microbiota composition and sampling time confirmed by ADONIS and ANOSIM statistical test (P < 0.001). Boxplot at genus level (Fig. 4) showed a significant reduction in the abundance of Bacteroides, Collinsella and Rikenellaceae, and a significant increase of Blautia, Butyricicoccus, Clostridium, Coprococcus, Dorea, Faecalibacterium, L−Ruminococcus (Ruminococcus genus assigned to Lachnospiraceae family), and Lachnospiraceae at the study end when compared to enrolment (Fig. 4).
Boxplots showing the relative abundance of Actinobacteria, Bacteroidetes, Proteobacteria and Firmicutes phyla in fecal samples of GDM patients at enrolment (green bars) and study end (blue bars). Boxes represent the interquartile range (IQR) between the first and third quartiles, and the line inside represents the median (2nd quartile). Whiskers denote the lowest and the highest values within 1.56 IQR from the first and third quartiles, respectively. Circles represent outliers beyond the whiskers.
Boxplots showing the relative abundance at genus or family level of the OTUs differentially abundant based on Wilcoxon matched pairs test (P ≤ 0.002) in fecal samples between: GDM patients at enrolment (green bars) and at the study end (blue bars); adherents to the dietary recommendations at enrolment (yellow bars) and at the study end (red bars); non-adherents to the dietary recommendations at enrolment (cyan bars) and at the study end (orange bars).
Microbiota signature between dietary adherences
The microbiota α- and β-diversity values were not significantly different between adherent and non-adherent subjects (data not shown). Similarly, there was no significant separation of the microbiota composition. We performed a number of analyses investigating the shift in microbiota as a function of the adherence to diet. Taking into account the shift in the microbiota between enrolment and study end in adherents and non-adherents (Fig. 4) a common microbiota signature was observed. Blautia, Coprococcus, Dorea and Lachnospiraceae significantly increased in both groups during the progression of pregnancy while Rikenellaceae decreased. We observed that the delta (study-end minus baseline) values of those OTUs was significantly higher in adherent patients. Between the two groups, we detected a specific microbiota shift at the study end: an impressive decrease in Bacteroides in adherents, and higher abundance of Faecalibacterium and L-Ruminococcus together with minor OTUs in non-adherents (Fig. 4).
Associations between microbiota and nutrient intakes and metabolic variables
Several different associations between nutrients/metabolic variables and microbiota could be detected both at enrolment (Supplementary Fig. S2A) and at the study end (Supplementary Fig. S2A).
At enrolment, Alistipes was found positively related with fat intakes (β = 0.10; 95% CI 0.06 0.14; P < 0.001) in a regression model, after adjusting for age and weight values. Furthermore, glycated hemoglobin (HbA1c) levels correlated with both Bacteroidales (β = 1.43; 95% CI 0.67 2.19; P < 0.001) and Prevotella (β = 0.11; 95% CI 0.06 0.16; P < 0.001).
At the end of the study, many associations among specific microbiota relative abundance and nutrient intakes, metabolic and inflammatory variables and their changes across pregnancy were detected in multiple regression analyses, after adjusting for age, weight change, and adherence to the given recommendations (Table 3). Among the relationships, we underline the direct associations between Roseburia and fiber intake (β = 0.09; 95% CI 0.02 0.16; P = 0.01), and that between L-Ruminococcus and oligosaccharides (β = 0.02; 95% CI 0.01 0.03; P = 0.006), that however did not reach the established statistical cut-offs. Furthermore, Faecalibacterium resulted inversely correlated with fasting glucose; Collinsella and Blautia were respectively directly and inversely associated with insulin and HOMA-IR values; Blautia was inversely correlated with HbA1c levels while Sutterella directly with CRP values (Table 3). Results did not change significantly, after adjusting for pre-pregnancy BMI, educational level and exercise (data not shown).
Gut microbiota signature at sub-genus level
In order to explore the possible effects at sub-genus level, we carried out oligotyping on sequences of Blautia and Roseburia since these were the only genera changing over time that showed a Shannon entropy index sufficient to identify all nucleotide positions that would resolve the oligotypes. Specific Blautia oligotypes (identified as Blautia wexlerae by BLASTn match): B1, B2, B4, B9, B11, B18, B27, B32, B36 B42, B49, B51 and B53, identified as Blautia luti were more abundant at the study end, while B41 and B59 decreased with the progress of pregnancy (Supplementary Fig. S3). When plotting the correlation between those oligotypes and dietary intake and blood variables (Supplementary Fig. S4), different correlations were found. In a multiple regression model, B42 was associated directly with total cholesterol (β = 16.6; 95% CI 7.10 26.1; P = 0.002), and B42 inversely with diastolic blood pressure (β = −3.06; 95% CI −4.75 −1.37; P = 0.001).
Roseburia oligotypes R10, R24, R50, and R70 increased at the study end when compared to enrolment (data not shown). Among the different correlations found by Spearman’s nonparametric correlations, none resulted significantly different in the regression model.
Shift in predicted metagenomes
The pathway enrichment analysis of the predicted metagenomes showed an enrichment of KEGG orthologues, at study end when compared with baseline, of glycolysis/gluconeogenesis (ko00010), fructose and mannose metabolism (ko00051), galactose metabolism (ko00052), starch and sucrose metabolism (ko005009), biosynthesis of amino acids (ko01230), and a reduction of fatty acid metabolism (ko01212), biotin metabolism (ko00780) and folate biosynthesis (ko00790). When plotting the correlations between OTUs and inferred metabolic pathways, we observed a positive correlation between Lipopolysaccaride (LPS) biosynthesis (ko00540) with Sutterella, Bacteroides and Phascolartobacterium (Supplementary Fig. S5).
Discussion
Our results showed a shift in the microbiota composition from the second to the third trimester of pregnancy, with higher α-diversity, Firmicutes increment, and Bacteroidetes and Actinobacteria reduction. Furthermore, associations between specific bacterial abundance and dietary and laboratory variables were detected.
The reduced insulin sensitivity of late pregnancy is considered beneficial to support fetal growth and increased nutrient absorption, even if it is associated with metabolic impairment and inflammation3. Women who developed GDM have greater reduction in insulin sensitivity and their insulin secretion is not sufficient to maintain euglycemia, leading to glucose intolerance17. This is counterintuitive owing to the progressive weight gain and increase in circulating levels of insulin, lipids, and inflammatory markers in the patients and the well-known association of low bacterial richness and adiposity, insulin resistance, dyslipidemia, and inflammatory phenotypes18. Going more deeply in the microbiota composition of our patients, we observed that the higher bacterial richness was related to Firmicutes. In a Finnish study, it has been reported that glycated hemoglobin values were positively associated with microbiota richness19 and mice transplanted with feces from obese and lean individuals showed a positive correlation of OTUs richness with both fasting insulin and HOMA-IR level20. The increase in Firmicutes abundance (and the reduction in Bacteroidetes) can be justified by the patients’ gestational weight gain, already overweight before pregnancy, not dissimilarly to what happens in obese patients21,22. Accordingly, the inferred metagenomic showed an increase in pathways involved in carbohydrate metabolism (in details, glycolysis/gluconeogenesis, fructose and mannose metabolism, galactose metabolism, and starch and sucrose metabolism) with the release of simply sugars due to the higher abundance of Firmicutes that harvested more energy from the diet23. It can be hypothesized that those enriched function could be related with the progressive weight gain and could be a feature in hyperglycemic phenomena.
Literature data are controversial: in normoglycemic pregnancy, weight gain was reported to be associated with Escherichia coli24 or Bacteroides abundance25 and both an increment in Proteobacteria and Actinobacteria and a decline in butyrate-producing bacteria (such as Faecalibacterium) were found3.
A higher Bacteroides-to-Firmicutes ratio has been found to correlate with elevated plasma glucose levels26. In our patients, however, the most relevant change was the weight gain at the end of pregnancy (Table 1). We detected a reduction in Bacteroidetes across pregnancy, but we found significant direct associations between Bacteroidales and Prevotella and HbA1c levels at enrolment.
Indeed, studies are difficult to compare due to the different ethnicity and food habits of the analyzed cohorts27 leading to inter-individual variations in the gut microbiota composition and the various methods used to analyze the microbiota, both causing results sometimes contradictory.
Further aspects of previous studies make the comparison with our results difficult, such as the fact that both normoglycemic and GDM women were combined together3,7, participants taking probiotics or antibiotics were not excluded3, dietary intakes did not change during pregnancy3, early pregnancy only was evaluated3,7,28.
Overall, our patients consumed a low-fiber and high-fat diet, an unhealthy dietary pattern which has been associated with GDM1. Most of them (about 2/3) did not change substantially their dietary habits after having received nutritional recommendations and showed a worse metabolic and inflammatory pattern than the adherent women. Overall, patients increased their intake of oligosaccharides; consistent with this, we observed with the pregnancy progression an enrichment in inferred metabolic pathways related with polysaccharide degradation, which in turn could be linked to the increased insulin resistance.
We found few associations between nutrient intake and microbial abundance. Fat intake was associated with Alistipes among Bacteroidetes, while Roseburia and L-Ruminococcus among the Firmicutes appeared related, though not significantly, with nutrients related to vegetable foods (oligosaccharides and fiber). This observation is in agreement with DNA-based studies evaluating the fecal microbiota during pregnancy in healthy overweight Finnish women at early pregnancy stage (17 week)29 as well as in normal-weight Norwegian women during the second trimester of pregnancy30. On the opposite, we observed a positive association between protein intake and Faecalibacterium which is in disagreement with previous studies29,30.
Research about gut microbiota composition and dietary intakes during pregnancy showed controversial results. Either no relationships between bacterial groups and dietary intakes3,31 or association between dietary fat and vitamin D with Proteobacteria increase29 and higher gut microbiota richness and lower abundance of Bacteroidaceae with increased dietary fiber intake have been reported31. Those findings confirm the great heterogeneity of results on this topic and highlight difficulties in the comparison of results from the studies, probably due to the different dietary habits, microbiota remodeling during pregnancy owing to hormonal changes and the additional insulin resistance determined by the presence of GDM.
Short-term changes in dietary pattern have been demonstrated to modulate quickly the microbiota composition. Rapid, but transient changes occur following dietary variations8,10, although longer and persistent modifications are needed to shape the human gut microbiota. In addition, the effects of diet on gut microbiota, rather than being direct, are hypothesized to be the consequence of the weight change and the subsequent variation in white adipose tissue inflammation and insulin resistance26.
In this study, we have assessed whether the modification of the dietary habits by dietary counselling during pregnancy can affect the gut microbiota composition. Even if arbitrary, the subdivision by dietary adherence distinguished women with greater increments of fasting glucose, insulin resistance and CRP values. During normal pregnancy a low grade of inflammation develops and GDM is a pro-inflammatory state32. Accordingly, we observed higher values of CRP at the pregnancy end. An imbalance of pro- and anti-inflammatory bacterial species have been proposed to trigger low-grade inflammation and insulin resistance in humans18. In particular Faecalibacterium, an anti-inflammatory commensal bacteria, significantly increased with pregnancy progression, but also increased in non-adherents, and is consistently reported to be more prevalent in individuals with higher bacterial richness18. It could be hypothesized that this increase could be a compensatory mechanism to counterbalance the pro-inflammatory state, potentially harmful for the fetus.
Indeed, we found a strong inverse relationship between Faecalibacterium abundance and fasting glucose values, supporting the well-known association between inflammation and dysmetabolism. Accordingly, Faecalibacterium prausnitzii resulted highly discriminant for the diagnosis of type 2 diabetes in metagenomic analyses33,34. Furthermore, these butyrate-producer bacteria have been found inversely linked to diabetes in human studies on fecal microbiota35,36,37,38.
In our patients, we observed a negative associations between diastolic blood pressure and Rikenellaceae and Oscillospira. Rikenellaceae is a butyrate producers, while Oscillospira is considered an enigmatic bacterial genus that has never been cultured, probably producing butyrate. Few available data support a beneficial role on human health39. Other studies found a protective role of Odoribacter (Bacteroidetes) on blood pressure in pregnant overweight women, and its capability to produce butyrate was mainly implicated in the maintenance of normal blood pressure40.
The strong direct associations that we found between the genus Collinsella and insulin/HOMA-IR values were in line with studies during pregnancy7,41 or not37 showing higher abundance of the lactate-producing Collinsella in type 2 diabetes mellitus. These bacteria can affect the metabolism by decreasing liver glycogenesis and playing pro-inflammatory effects41.
Insulin resistance was associated positively with Erysipelotrichia and negatively with Blautia in our patients. Very few human data are available on Erysipelotrichia, suggesting a relationship with inflammatory diseases for this class, which seems in a close relationship with the class of Mollicutes, which is in turn associated with many pathological human conditions like endotoxemia, obesity and insulin resistance26,42.
In our overweight GDM patients, the butyric acid-producing genus Butyricimonas was directly associated with insulin resistance. The possible role of this taxa in human diseases awaits further investigation, even if a positive association with mean arterial pressure has been detected43.
Changes in CRP values during the third trimester of pregnancy resulted directly associated with Sutterella. Even if we did not detect an overall increase in Proteobacteria during pregnancy, as other authors observed3, we found that Sutterella, a proteobacteria with known pro-inflammatory capacity, was associated with CRP increment across pregnancy. Consistent with this, the predicted metagenomes showed a correlation between Sutterella and KEGG genes associated with LPS biosynthesis. Gram-negative bacteria could produce inflammatory LPS triggering a pro-inflammatory state, a condition characterizing both type 2 diabetes and obesity44.
We also observed a correlation between LPS inferred KEGG genes and Bacteroides. In diabetic patients, LPS from a specie belonging to Bacteroides (B. fragilis) was reported to play a major pathogenic role45. Bacteroides is often associated with high fat-animal based diet9. Consistently, we found a Bacteroides reduction in adherents only, whose total and saturated fat intake, and CRP values decreased across pregnancy (Table 2). In addition, the metagenomic content of GDM patient was reported to be enriched of genes involved in LPS biosynthesis and in the regulation of blood glucose levels13.
At genus level, strong inverse relationships between Blautia and Hba1c and insulin resistance were observed. At sub-genus level, we observed a higher number of oligotypes belonging to the same species, even if only a few of them changed during the progression of the pregnancy, and controversial associations between Blautia and blood pressure and cholesterol values. A controversial role of Blautia in the human gut is reported. Several studies showed a direct association between Blautia and hyperglycemia46,47 but other studies reported that abundance of this taxon indicates a healthy gut, reduced inflammation and blood pressure values, diminished risk for type 1 diabetes and obesity, and increased survival48,49. Our results suggest a possible different strain-dependent effect on metabolism. The diversity at sub-genus-level is indeed well known to play a key role in establishing the interconnection between gut microbiome and host responses50. As recently observed by De Filippis and colleagues51, different oligotypes belonging to the same species showed different relative abundance and different correlation patterns with metabolomic data. Those authors suggested that different putative strains could have different impact on the host51.
The knowledge of the gut bacterial composition might allow the identification of subsets of women with different metabolic risks, owing to its role in the gestational pro-inflammatory status potentially contributing to the increased insulin resistance of pregnancy. This is a topic of great interest, also in consideration of the benefits of probiotic supplementation in the reduction of inflammation in women with GDM52, a condition well-known for exposing to an increased risk for chronic health conditions not only the mother but also her child.
One limitation of this study is the small sample size; nevertheless, the power of our study to detect differences in alpha diversity was 0.84 with α = 0.01. The fecal samples were used as proxies for the microbial content of the entire gastrointestinal tract; it is reasonable to consider that mouth and skin microbiota could vary too. The limitations of the food questionnaires must be recognized, even if these were widely used. Most of our patients had a very low fiber intake and consumed a high-fat diet; indeed, the dietary intakes of our patients resembled those of other pregnant women31,53, and this finding is in line with the well-known associations between GDM and unhealthy diet1. The lack of substantial difference in dietary intakes between enrolment and study end should be recognized as a limitation; we cannot exclude that a better adherence to the dietary recommendations could have resulted in greater differences between adherents and non-adherents. Nonetheless, it is noteworthy that already small dietary changes have been able to lead to statistical and clinical significant difference between groups, suggesting the importance of a healthy diet in these patients. Owing to the observational design of this study, the presence of unmeasured confounding factors cannot be excluded. Microbiota assessment through amplicon-based sequencing has several biases due to the PCR amplification step, while shotgun metagenomic sequencing identified significantly more bacterial species per read than the 16S method54. Correlations were performed by considering individual groups of bacteria independently from each other, therefore it was not possible to establish neither the causality nor the biological relevance of the reported relationships. Finally, the predictive metagenomic profiling was obtained from the bacterial abundance and was therefore a derived result.
In our overweight GDM patients, a shift in the microbiota composition with higher α-diversity, and numerous associations between the metabolic/inflammatory pattern and specific bacterial abundance were detected. If confirmed by further studies in larger sample, these results suggest that the development of strategies to modulate the gut microbiota might be the next step in order to impact on maternal and possibly fetal health and their future risk for metabolic diseases.
Methods
Patients recruitment
The participants were 50 patients with GDM consecutively recruited from the “Città della Salute e della Scienza” Hospital of Turin from April 2016. Each participant gave her written informed consent to participate in the study. The study protocol was approved by the Ethics Committee of the “Città della Salute e della Scienza” Hospital of Turin (approval 707/2016). All research was performed in accordance with relevant guidelines/regulations.
Inclusion criteria were: gestational age between 24–28 weeks, Caucasian race, GDM diagnosed by a 75 g oral glucose tolerance test (OGTT). Women who had the following criteria were excluded from the study: twin pregnancy, use of prebiotics/probiotics, antibiotics or any drug during pregnancy, any pathological conditions before or during pregnancy (known diabetes mellitus, hypertension, cardiovascular, pulmonary, autoimmune, joint, liver or kidney diseases, thyroid dysfunction, cancer, any other disease/condition), no compliance to the study protocol. All women were taking folic acid supplementation.
GDM was diagnosed by OGTT performed at 24–28 gestational weeks in the morning, after at least 8h-overnight fast, when the fasting plasma glucose was ≥92 mg/dL and/or 1 h post-OGTT glycemia ≥180 mg/dL and/or 2 h post-OGTT glycemia ≥153 mg/dL, according to international criteria17. In our cohort, all the patients with GDM routinely received dietary counselling and nutritional recommendations in line with guidelines (carbohydrates 45% total energy, rapidly absorbed sugars <10% total energy, proteins 18–20% total energy, fats 35% total energy, at least 20–25 g/day fibre intake, no alcohol)17. Furthermore, 30-min daily moderate exercise was recommended (i.e. brisk walking). Patients were instructed to self-monitor finger-prick capillary blood glucose at least 4 times per day. Insulin treatment was prescribed by the physicians in the presence of hyperglycemia, in accordance with guidelines55.
Sample collection, anthropometric measurements and dietary information
Questionnaires, anthropometric values, fasting blood samples and stool samples were collected for all participants both at 24–28 weeks of gestational age at the time of GDM diagnosis (enrolment), and at 38 weeks, or before delivery, in the case of preterm delivery (study end). The researchers were in continuous contact with the patients, through weekly telephone contact. In this way, they were aware of the progress of pregnancy.
Stool samples were self-collected by the patients as previously described27. Briefly, the subjects were instructed on how to self-collect the samples, and all materials were provided in a convenient, refrigerated, specimen collection kit. Patients were provided with sterile containers to collect the feces (VWR, Milan, Italy). The fecal samples were collected at home and transferred to the sterile sampling containers using a polypropylene spoon (3 spoons of about 10 g) and immediately stored at 4 °C. The specimens were transported to the laboratory within 12 hours of collection at a refrigerated temperature. Containers were immediately stored at −80 °C for DNA extraction. No storage medium was used.
Participants completed a 3-day food record (2 weekdays and 1 weekend day) and the Minnesota-Leisure-Time-Physical Activity Questionnaire56 at enrolment and at the study end. Detailed information on how to record food and drink consumed by using common household measures was provided to all participants. Two dieticians checked all questionnaires for completeness, internal coherence and plausibility.
Data relative to pre-pregnancy weight was self-reported; weight, height, and arterial blood pressure (BP) were measured at time of enrolment, and weight and BP at the study end. Body weight was measured to the nearest 0.1 kg, and height was measured to the nearest 0.1 cm with a stadiometer (SECA model 711, Hamburg, Germany), with the participants wearing light clothes and no shoes. Arterial BP was measured from the left arm, in a sitting position, after at least 10 min of rest, with a mercury sphygmomanometer with appropriate cuff sizes (ERKA Perfect-Aneroid, Germany). Two measurements were taken by trained personnel with arm supported at heart level and the values reported were the means of the two. Glucose levels were self-measured by the patients by the BGSTAR® glucometer (Sanofi-Deutscland GmbH, Frankfurt, Germany). The average of the values measured 1-hour after each meal during the third trimester has been reported.
Babies were classed as large for gestational age (LGA) if their birthweights were >90th percentile, considering neonatal anthropometric standards for Northern Italy57.
Blood analyses
Serum glucose was measured by the glucose oxidase method (Sentinel Ch., Milan) with an intra-assay CV of 1.1% and an inter-assay CV of 2.3%. HbA1c levels were determined by a latex-based method (Sentinel, Milan, Italy). The intra-assay e inter-assay CVs were respectively 1.1–1.5% and 1.1–1.6%. Triglycerides and cholesterol were assayed by enzymatic colorimetric assays (Sentinel, Milan) with an intra-assay CV of 3.0% and an inter-assay CV of 3.5% for triglycerides and with an intra-assay CV of 2.2% and an inter-assay CV of 3.4% for cholesterol. HDL-cholesterol was determined by enzymatic colorimetric assay after precipitation of LDL and VLDL fractions using heparin-MnCl2 solution and centrifugation at 4 °C and it had an intra-assay variation CV of 2.5% and an inter-assay CV of 4.1%. Insulin was measured by a biotin labelled antibody-based sandwich enzyme immunoassay (LDN, Germany). The kit had a sensitivity of less than 1.8 U/mL and a range of 0–100 U/mL. The intra-assay and inter-assay CVs were respectively 1.8–2.6% and 2.9–6.0%. Serum CRP values were determined using a high-sensitivity-latex agglutination assay on HITACHI 911 Analyzer (Sentinel, Milan). The intra-assay and inter-assay CVs were 0.8–1.3% and 1.0–1.5%, respectively. All laboratory measurements were centralized.
BMI was calculated as weight divided for the square of height. The HOMA-IR was calculated according to the published algorithm58. Adherence to the given dietary recommendations was considered in the presence of all the following criteria: consuming at least 20 g/day fiber (or increasing fiber intake more than 50% than enrolment) and reducing sugars <10% of total energy and abolishing alcohol intake.
Fecal DNA extraction
Nucleic acid was extracted from the feces collected. Total DNA from the samples was extracted using the RNeasy Power Microbiome KIT (Qiagen, Milan, Italy) following the manufacturer’s instructions. One microliter of RNase (Illumina Inc. San Diego. CA) was added to digest RNA in the DNA samples, with an incubation of 1 h at 37 °C. DNA was quantified using the QUBIT dsDNA Assay kit (Life Technologies, Milan, Italy) and standardized at 5 ng/μL.
16S rRNA amplicon target sequencing
DNA directly extracted from fecal samples was used to assess the microbiota by the amplification of the V3-V4 region of the 16S rRNA gene using the primers and protocols described by Klindworth et al.59. PCR amplicons were cleaned using Agencourt AMPure kit (Beckman Coulter, Milan, Italy) and the resulting products were tagged by using the Nextera XT Index Kit (Illumina Inc. San Diego. CA) according to the manufacturer’s instructions. After the 2nd purification step, amplicons products were quantified using a QUBIT dsDNA Assay kit (Life Technologies). Subsequently, equal amounts of amplicons from different samples were pooled. The pooled sample was run on an Experion workstation (Biorad, Milan, Italy) for quality analysis prior to sequencing. The sample pool (4 nM) was denatured with 0.2 N NaOH, diluted to 12 pM, and combined with 20% (vol/vol) denatured 12 pM PhiX, prepared according to Illumina guidelines. The sequencing was performed with a MiSeq Illumina instrument (Illumina) with V3 chemistry and generated 250 bp paired-end reads according to the manufacturer’s instructions.
Bioinformatics analysis
Paired-end reads were first assembled using FLASH software60 with default parameters. Joint reads were further quality filtered (at Phred <Q20) using QIIME 1.9.0 software61 and short reads (<250 bp) were discarded through Prinseq62. Chimera filtering was performed through USEARCH software version 8.163. Operational Taxonomic Units (OTUs) were picked at 97% of similarity threshold by UCLUST algorithms64 and centroids sequences of each cluster were matched to the Greengenes 16S rRNA gene database version 2013. After sequencing, a total of 2,100,009 raw reads (2 × 250 bp) were obtained. After joining, a total of 1,919,311 reads passed the filters applied with QIIME, with an average value of 23,406 ± 31,535 reads/sample and a sequence length of 457 bp. The rarefaction analysis and Good’s coverage, expressed as percentages, indicated that there was satisfactory coverage for all the samples (Good’s coverage average, 92%). In order to avoid biases due to the different sequencing depth, OTU tables were rarefied to the lowest number of sequences per sample (4078 reads/sample). The OTU table displays the higher taxonomy resolution that was reached; when the taxonomy assignment was not able to reach the genus, family name was displayed. Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt) was used to predict abundances of KEGG orthologs (KO) based on 16S-based structure of the microbiota65. The KO abundance table was then collapsed at level 3 of the KEGG annotations in order to display the inferred metabolic pathways and the table was imported in gage Bioconductor package66 in order to carry out pathway enrichment analysis to identify biological pathways overrepresented or underrepresented between samples.
Oligotyping analysis
In order to identify sub-OTUs populations, reads assigned to genera within Ruminococcaceae and Lachnospiraceae were extracted and entropy analysis and oligotyping were carried out67. Briefly, the extracted reads were then used to identify nucleotide positions that will explain the maximum amount of biological diversity across the samples utilizes Shannon entropy in order to identify positional variation to facilitate the identification of nucleotide positions of interest67. Only Blautia and Roseburia oligotypes showed a higher level of entropy and were the only two taxa able to be differentiated in sub-OTUs. After the first round of oligotyping, high entropy positions were chosen (-C option) 8, 9, 12, 223, 225, 247, 261, 282, 432, 433 for Blautia; while position 8, 9, 12, 245, 432, 433 and 434 were chosen for Roseburia. To reduce the noise, each oligotype was required to appear in at least 10 samples, occur in more than 1.0% of the reads for at least one sample, represent a minimum of 500 reads in all samples combined, and have a most abundant unique sequence with a minimum abundance of 100. BLASTn was used to query the representative oligotype sequences against the NCBI nr database, and the top hit was considered for taxonomic assignment.
Statistical analysis
Gut microbiota α-diversity was assessed by Chao1 index, estimating the number of different taxa, and by Shannon diversity index, evaluating the taxa richness and evenness calculated using the diversity function of the vegan package68 in R environment (http://www.r-project.org). OTU table was used to build a principal-component analysis (PCA) as a function of the sampling time by using the made4 package of R. ADONIS and ANOSIM statistical test was used to detect significant differences in the overall microbial community by using the Weighted UniFrac distance matrices and the OTU table.
Not-normally distributed variables were presented as median (range interquartile). The individual differences between endof the study values minus baseline values were calculated (deltas). The delta median values were reported. Within-participant differences in bacterial richness and in the variables at enrolment compared with values at the pregnancy-end were evaluated by paired-sample t-test, or Wilcoxon matched pairs test, as appropriate. Differences between categorical variables were computed by chi-square test.
Differences in gut microbiota or oligotypes between adherents and non-adherents were calculated by t-Student test or Mann-Whitney test. Box plots represented the interquartile range between the first and the third quartile, with the error bars showing the lowest and the highest value. Pairwise Spearman’s non-parametric correlations were used to study the relationships between the relative abundance of microbial taxa abundance or oligotypes and dietary and metabolic variables, and between gut microbiota and inferred metabolic pathways. The correlation plots were visualized in R using the corrplot package of R.
Multiple regression analyses were performed to evaluate the associations between log-transformed microbial taxa abundance or oligotypes (dependent variable) and nutrient intakes after adjusting for age and weight (variables at baseline), or age and weight change (variables at the pregnancy end). The associations between blood pressure and laboratory variables (dependent variables) with OTUs were calculated by a multiple regression model, after adjusting for age and weight (variables at baseline) or age, weight change, and adherence to the given dietary recommendations (variables at the pregnancy end) (Statistica, ver. 7.0; StatSoft Inc., Tulsa, OK, USA).
Bonferroni’s correction for multiple comparisons was applied; a P value of 0.002 or lower was considered as statistically significant.
Ethics approval and consent to participate
Each participant gave her written informed consent to participate in the study. The study protocol was approved by the Ethics Committee of the “Città della Salute e della Scienza” Hospital of Turin (approval 707/2016).
Availability of data and material
All the sequencing data were deposited at the Sequence Read Archive of the National Center for Biotechnology Information (SRA accession number: SRP135886).
References
Brown, J. et al. Lifestyle interventions for the treatment of women with gestational diabetes. Cochrane Database Syst. Rev (2017).
Billionnet, C. et al. Gestational diabetes and adverse perinatal outcomes from 716,152 births in France in 2012. Diabetologia 60, 636–644 (2017).
Koren, O. et al. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150, 470–480 (2013).
Dunlop, A., Mulle, J., Ferranti, E., Edwards, S. & Dunn, A. The Maternal microbiome and pregnancy outcomes that impact infant health: A Review. Adv Neonatal Care 15, 377–385 (2015).
Soderborg, T. K., Borengasser, S. J., Barbour, L. A. & Friedman, J. E. Microbial transmission from mothers with obesity or diabetes to infants: an innovative opportunity to interrupt a vicious cycle. Diabetologia 59, 895–906 (2016).
Garcia-Mantrana, I. & Collado, M. C. Obesity and overweight: Impact on maternal and milk microbiome and their role for infant health and nutrition. Mol. Nutr. Food Res. 60, 1865–1875 (2016).
Gomez-Arango, L. F. et al. Connections between the gut microbiome and metabolic hormones in early pregnancy in overweight and obese women. Diabetes 65, 2214–2223 (2016).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108 (2011).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Carrothers, J. M. et al. Fecal microbial community structure is stable over time and related to variation in macronutrient and micronutrient intakes in lactating women. J. Nutr. 145, 2379–2388 (2015).
Di Giulio, D. B. et al. Temporal and spatial variation of the human microbiota during pregnancy. Proc. Natl. Acad. Sci. 112, 11060–11065 (2015).
Zheng, J. et al. The placental microbiota is altered among subjects with gestational diabetes mellitus: A pilot study. Front. Physiol. 8, 1–12 (2017).
Kuang, Y.-S. et al. Connections between the human gut microbiome and gestational diabetes mellitus. Gigascience 6, 1–12 (2017).
Fugmann, M. et al. The stool microbiota of insulin resistant women with recent gestational diabetes, a high risk group for type 2diabetes. Sci. Rep. 5, 1–8 (2015).
Hasan, S. et al. Gut microbiome in gestational diabetes: a cross-sectional study of mothers and offspring 5 years postpartum. Acta Obstet. Gynecol. Scand. 97, 38–46 (2018).
Mokkala, K. et al. Gut microbiota aberrations precede diagnosis of gestational diabetes mellitus. Acta Diabetol. 54, 1147–1149 (2017).
Hod, M. et al. The international federation of gynecology and obstetrics (FIGO) Initiative on gestational diabetes mellitus: A pragmatic guide for diagnosis, management, and care. Int. J. Gynecol. Obstet. 131, S173–S211 (2015).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Org, E. et al. Relationships between gut microbiota, plasma metabolites, and metabolic syndrome traits in the METSIM cohort. Genome Biol. 18, 70 (2017).
Zhang, L. et al. Environmental spread of microbes impacts the development of metabolic phenotypes in mice transplanted with microbial communities from humans. ISME J. 11, 676–690 (2017).
Fändriks, L. Roles of the gut in the metabolic syndrome: an overview. J. Intern. Med. 281, 319–336 (2017).
Dao, M. C., Everard, A., Clément, K. & Cani, P. D. Losing weight for a better health: Role for the gut microbiota. Clin. Nutr. Exp. 6, 39–58 (2016).
Cani, P. D. Gut microbiota and obesity: Lessons from the microbiome. Brief. Funct. Genomics 12, 381–387 (2013).
Santacruz, A. et al. Gut microbiota composition is associated with body weight, weight gain and biochemical parameters in pregnant women. Br. J. Nutr. 104, 83–92 (2010).
Collado, M. C., Isolauri, E., Laitinen, K. & Salminen, S. Distinct composition of gut microbiota during pregnancy in overweight and normal-weight women. Am. J. Clin. Nutr. 88, 894–899 (2008).
Suez, J., Shapiro, H. & Elinav, E. Role of the microbiome in the normal and aberrant glycemic response. Clin. Nutr. Exp. 6, 59–73 (2016).
Ferrocino, I. et al. Fecal microbiota in healthy subjects following omnivore, vegetarian and vegan diets: culturable populations and rRNA DGGE profiling. PLoS One 10, e0128669 (2015).
Aguirre, M. & Venema, K. Does the gut microbiota contribute to obesity? Going beyond the gut feeling. Microorganisms 3, 213–235 (2015).
Mokkala, K. et al. Gut microbiota richness and composition and dietary intake of overweight pregnant women are related to serum zonulin concentration, a marker for intestinal permeability. J. Nutr. 146, 1694–1700 (2016).
Mandal, S. et al. Fat and vitamin intakes during pregnancy have stronger relations with a pro-inflammatory maternal microbiota than does carbohydrate intake. Microbiome 4, 55 (2016).
Portune, K. J., Benítez-Páez, A., Del Pulgar, E. M. G., Cerrudo, V. & Sanz, Y. Gut microbiota, diet, and obesity related disorders. The good, the bad, and the future challenges. Mol. Nutr. Food Res. 252, 1–17 (2016).
Röytiö, H., Mokkala, K., Vahlberg, T. & Laitinen, K. Dietary intake of fat and fibre according to reference values relates to higher gut microbiota richness in overweight pregnant women. Br. J. Nutr. 118, 343–352 (2017).
Lekva, T., Norwitz, E. R., Aukrust, P. & Ueland, T. Impact of systemic inflammation on the progression of gestational diabetes mellitus. Curr. Diab. Rep. 16, 26 (2016).
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012).
Tilg, H. & Moschen, A. R. Microbiota and diabetes: an evolving relationship. Gut 63, 1513–1521 (2014).
Zhang, X. et al. Human gut microbiota changes reveal the progression of glucose intolerance. PLoS One 8, e71108 (2013).
Furet, J. et al. Differential adaptation of human gut microbiota to bariatric surgery-induced weight loss. Diabetes 59, 3049–3057 (2010).
Del Chierico, F. et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology 65, 451–464 (2017).
Gomez-Arango, L. F. et al. Increased systolic and diastolic blood pressure is associated with altered gut microbiota composition and butyrate production in early pregnancy. Hypertension 68, 974–981 (2016).
Gomez-Arango, L. F. et al. Low dietary fiber intake increases Collinsella abundance in the gut microbiota of overweight and obese pregnant women. Gut Microbes in press, (2017).
Griffen, A. L. et al. Distinct and complex bacterial profiles in human periodontitis and health revealed by 16S pyrosequencing. ISME J. 6, 1176–1185 (2012).
Shin, J.-H., Sim, M., Lee, J.-Y. & Shin, D.-M. Lifestyle and geographic insights into the distinct gut microbiota in elderly women from two different geographic locations. J. Physiol. Anthropol. 35, 31 (2016).
Allin, K. H., Nielsen, T. & Pedersen, O. Mechanisms in endocrinology: Gut microbiota in patients with type 2 diabetes mellitus. Eur. J. Endocrinol. 172, R167–R177 (2015).
Mancuso, G. et al. Bacteroides fragilis derived lipopolysaccharide produces cell activation and lethal toxicity via toll-like receptor 4. Infect. Immun. 73, 5620–5627 (2005).
Egshatyan, L. V. et al. Gut microbiota and diet in patients with different glucose tolerance. Endocr. Connect. https://doi.org/10.1530/EC-15-0094 (2015).
Qi, C.-J. et al. Imbalance of fecal microbiota at newly diagnosed type 1 diabetes in chinese children. Chin. Med. J. (Engl). 129, 1298 (2016).
Jenq, R. R. et al. Intestinal Blautia is associated with reduced death from graft-versus-host disease. Biol. Blood Marrow Transplant. 21, 1373–1383 (2015).
Hong, P. Y., Croix, J. A., Greenberg, E., Gaskins, H. R. & Mackie, R. I. Pyrosequencing-based analysis of the mucosal microbiota in healthy individuals reveals ubiquitous bacterial groups and micro-heterogeneity. PLoS One 6, (2011).
Berni Canani, R. et al. Lactobacillus rhamnosus GG-supplemented formula expands butyrate-producing bacterial strains in food allergic infants. ISME J. 10, 1–9 (2015).
De Filippis, F., Pellegrini, N., Laghi, L., Gobbetti, M. & Ercolini, D. Unusual sub-genus associations of faecal Prevotella and Bacteroides with specific dietary patterns. Microbiome 4, 57 (2016).
Taylor, B. L. et al. Effect of probiotics on metabolic outcomes in pregnant women with gestational diabetes: A systematic review and meta-analysis of randomized controlled trials. Nutrients 9, E461 (2017).
Bo, S. et al. Simple lifestyle recommendations and the outcomes of gestational diabetes. A 2 × 2 factorial randomized trial. Diabetes Obes. Metab. 16, 1032–1035 (2014).
Ranjan, R., Rani, A., Metwally, A., McGee, H. S. & Perkins, D. L. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 469, 967–977 (2016).
Tests, D. & Diabetes, F. O. R. Classification and diagnosis of diabetes. Diabetes Care 38, S8–S16 (2015).
Taylor, H. L. et al. A questionnaire for the assessment of leisure time physical activities. J. Chronic Dis. 31, 741–755 (1978).
Bertino, E. et al. Standard antropometrici neonatali dell’Italia Nord-occidentale (Anthropometric neonatal standards based on a North-West Italian). Ital. Riv. Pedriatr. 25, 899–906 (1999).
Matthews, D. R. et al. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28, 412–419 (1985).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Magoc, T. & Salzberg, S. L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Langille, M. G. I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 10, 161 (2009).
Eren, A. M. et al. Oligotyping: Differentiating between closely related microbial taxa using 16S rRNA gene data. Methods Ecol. Evol. 4, 1111–1119 (2013).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Acknowledgements
This study was supported by a grant from the Ministry of Education, University and Research of Italy (ex-60% 2017).
Author information
Authors and Affiliations
Contributions
I.F. and S.B. designed and conducted the research. I.F., R.R. and S.B. performed the statistical analysis and analyzed the data. I.F., V.P. and S.B. wrote the paper. I.F., V.P. and Z.A. were responsible for collected biospecimens and for DNA extraction. I.F. and A.R. performed the 16S rRNA gene sequencing. I.F. carried out the bioinformatics analyses and generated the manuscript figures. R.G., C.M. and L.C. supervised the data analysis and contributed to manuscript preparation. V.P., R.G., Z.A., C.M. performed the analyses of the blood samples, L.F., M.C., G.I., G.G., B.F. enrolled and followed-up the patients, L.F., M.C., G.I., G.G., B.F. and L.C. contributed to manuscript preparation. I.F. and S.B. had primary responsibility for final content. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ferrocino, I., Ponzo, V., Gambino, R. et al. Changes in the gut microbiota composition during pregnancy in patients with gestational diabetes mellitus (GDM). Sci Rep 8, 12216 (2018). https://doi.org/10.1038/s41598-018-30735-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30735-9
This article is cited by
-
Dynamic changes in the gut microbiota during three consecutive trimesters of pregnancy and their correlation with abnormal glucose and lipid metabolism
European Journal of Medical Research (2024)
-
Diet, lifestyle and gut microbiota composition among Malaysian women with gestational diabetes mellitus: a prospective cohort study
Scientific Reports (2024)
-
Study of altered gut microbial dynamics and their association with gestational diabetes mellitus
Systems Microbiology and Biomanufacturing (2024)
-
Maternal microbiota and gestational diabetes: impact on infant health
Journal of Translational Medicine (2023)
-
Comprehensive characterization of maternal, fetal, and neonatal microbiomes supports prenatal colonization of the gastrointestinal tract
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.