Abstract
Next-generation sequencing-based methods are extensively applied in studies of the human microbiota using partial 16 S rRNA gene amplicons. However, they carry drawbacks that are critical to consider when interpreting results, including differences in outcome based on the hypervariable region(s) used. Here, we show that primers spanning the V3/V4 region identify a greater number of taxa in the vaginal microbiota than those spanning the V1/V2 region. In particular, taxa such as Gardnerella vaginalis, Bifidobacterium bifidum and Chlamydia trachomatis, all species that influence vaginal health and disease, are not represented in V1/V2-based community profiles. Accordingly, missing or underestimating the frequency of these species overestimates the abundance of other taxa and fails to correctly assess the bacterial diversity in the urogenital tract. We elaborate that covering these taxa using the V3/V4 region leads to profound changes in the assignment of community state types. Altogether, we show that the choice of primers used for studying the vaginal microbiota has deep implications on the biological evaluation of the results.
Similar content being viewed by others
Introduction
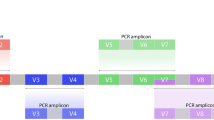
Human cavities are colonized with microorganisms and these form in their entity the human microbiome, which became intensively studied when next-generation sequencing (NGS) emerged. NGS offered the opportunity of cultivation-independent assessment of microbial communities and therefore revealed a multitude of thus far unknown bacteria. Since the first report on the analysis of the vaginal microbiota employing NGS1, amplification of partial sequences of the bacterial 16 S gene with primers spanning hypervariable regions is the common method to describe vaginal bacterial populations. In recent years, research has shown that the vagina of humans can be colonized by distinct bacterial communities, often dominated by a Lactobacillus species1. However, also diverse communities are observed1 and some communities display a significant fraction of Gardnerella vaginalis2. These different compositional states of bacterial communities were termed community state types (CSTs) by Gajer et al. subsequently3. Importantly, a rising number of reports show that the vaginal microbiota play an important role in the acquisition and prevention of sexually transmitted diseases and their long term sequels2,4,5,6,7,8. However, as it frequently appears, the amplified region used varies from study to study. While many studies use the primers 27 F and 338 R spanning the V1/V2 hypervariable region (e.g.1,3), others choose the V3/V4 region (e.g.2,9). It has been shown that the choice of the primers used for amplification can introduce bias to the results achieved when applying 16S amplicons for microbiome studies10,11,12. Thus, it is important to consider the validity of the primers for given sample type being analysed13. Despite a report of the V1/V2 region being inappropriate for targeting vaginal bacteria such as Gardnerella vaginalis14, no study to date compared the relative performance of V1/V2 to another hypervariable region for vaginal samples. To address this uncertainty, we studied a set of cervical swabs derived from different clinical sites and compared the results when using the V1/V2 spanning primers (27 F/338 R) to the same samples amplified using primers spanning the V3/V4 (319 F/806 R) hypervariable region. This analysis contributes to the standardisation of vaginal microbiota studies and provides critical insight into the comparability between studies.
Results
The V3/V4 region identifies more taxa and displays higher diversity than the V1/V2 region
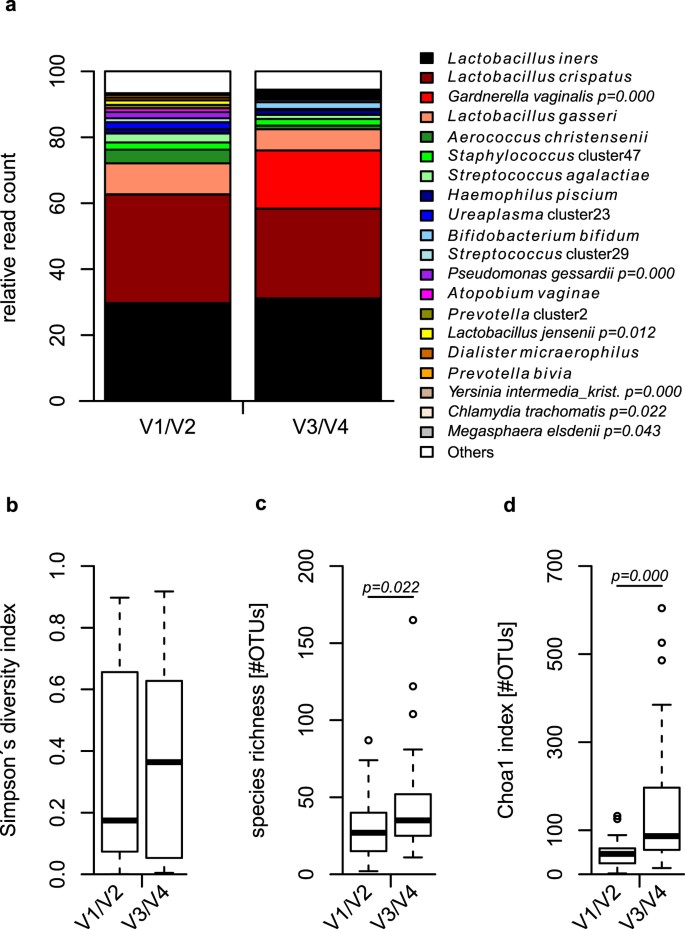
In total we sequenced the V1/V2 and V3/V4 regions for 38 samples, with a normalized sequencing coverage of 5000 reads per sample after data processing. Sequence classification was performed using the same approach15 for both data sets and we subsequently compared the resulting taxa identified. Among the most abundant taxa, we observed Gardnerella vaginalis, and Chlamydia trachomatis to be significantly enhanced in the V3/V4 data set compared to the V1/V2 region, while other taxa, such as Lactobacillus jensenii, Pseudomonas gessardii, and Megasphaera elsdenii are correspondingly decreased (Fig. 1a and Supplementary Table S1).
Relative read count of several bacterial taxa and bacterial alpha diversity differ according to the region used. The V3/V4 region shows significantly enhanced relative read count of taxa such as Gardnerella vaginalis and Chlamydia trachomatis in comparison to the V1/V2 region. Inversely, Lactobacillus jensenii, Pseudomonas gessardii and Megasphaera elsdenii are significantly reduced in their relative read count in the V3/V4 data set (a). Observed and estimated total number of OTUs within the samples are significantly higher when comparing the V3/V4 region with the V1/V2 data set using operational taxonomic units clustered with a 97% global identity threshold (b–d). Statistical comparisons were performed using Wilcoxon rank-sum test.
We further applied several methods for estimation of alpha diversity using OTUs clustered with a similarity threshold of 97%. While Simpson’s diversity index reveals a non-significant trend to be enhanced (Fig. 1b), observed (Fig. 1c) and estimated total (Fig. 1d) number of OTUs show significant higher values for the V3/V4 in comparison to the V1/V2 region.
The V3/V4 region identifies community state types with characteristic species lacking in the V1/V2 region
We observe a total of six community state types (CSTs) present in the data set using the V1/V2 region. Five of them are characterized by a dominant species, which are either Lactobacillus iners (Lin CST), L. crispatus (Lcr CST), L. gasseri (Lga CST), Aerococcus christensenii (Ach CST) or Streptococcus agalactiae (Sag CST), respectively. One CST is more diverse (div CST) (Fig. 2a) and not dominated by a single species. In contrast, the number of CSTs dominated by L. crispatus and L. gasseri appears lower based on the V3/V4 region (Fig. 3). While the div CST and the CSTs dominated by A. christensenii and S. agalactiae are either less frequent or not present in the V3/V4 data set, two CSTs with characteristic G. vaginalis abundance (Gva1 and Gva2 CST) and one CST dominated by Bifidobacterium bifidum (Bbi CST) are appearing (Fig. 2b). The differences between the two regions appear to be significant (Fig. 3, p < 0.01, chi-square test).
Heatmaps based on relative read counts of the major bacterial taxa including clustering of the urogenital microbial communities into several CSTs above the heatmaps. While the samples cluster into 6 CSTs within the V1/V2 data set (a), the V3/V4 region displays partially different CSTs with reduction of the proportion or elimination of the CSTs present in the V1/V2 region (b). The color coding for the CSTs is given in (c). The clusters are named based on characteristic taxa: Lin: Lactobacillus iners, Lcr: L. crispatus, Lga: L. gasseri, div: diverse, Ach: Aerococcus christensenii, Sag: Staphylococcus agalactiae, Gva: Gardnerella vaginalis, Bbi: Bifidobacterium bifidum.
Proportion and presence of assigned CSTs change depending on the 16S region used for amplification. The two regions reveal different CSTs. The proportion of Lactobacillus-dominated and diverse CST appears lower while CSTs characterized by the presence of Gardnerella vaginalis are prominent when using the V3/V4 region in comparison to the V1/V2 data set (Pearson’s chi square test: p = 0.002). The CSTs are named according to characteristic species of each cluster: Lin: Lactobacillus iners, Lcr: L. crispatus, Lga: L. gasseri, div: diverse, Ach: Aerococcus christensenii, Sag: Staphylococcus agalactiae, Gva: Gardnerella vaginalis, Bbi: Bifidobacterium bifidum.
We next analysed to which CST a sample belongs according to the different regions. Following the constructed migration plot (Fig. 4), we observe that parts of the samples belonging to the Lga CST in the V1/V2 region cluster to the Gva1/2 and div CST using the V3/V4 data set, while for the Lcr CST (V1/V2) we observe a transition to the Gva2 CST (V3/V4). The diverse CST from the V1/V2 data set mostly switches to the Gva1 CST in the V3/V4 region. The samples assigned to the singletons Ach CST and Sag CST from the V1/V2 data set appear in different CSTs using the V3/V4 region.
A considerable amount of samples display a CST when the V1/V2 region is used that appears as a different CST when using the V3/V4 region. The migration plot shows the number of samples belonging to each CST within both data sets and displays the connections between samples with different CSTs between the regions. Thus, the plot shows e.g. that a certain number of samples the V1/V2 region displays as Lcr, Lga and div CST are shown to belong to the Gva1 and Gva2 CST when using the V3/V4 region. The CSTs are named according to characteristic species of each cluster: Lin: Lactobacillus iners, Lcr: L. crispatus, Lga: L. gasseri, Gva: Gardnerella vaginalis, div: diverse, Bbi: Bifidobacterium bifidum, Ach: Aerococcus christensenii, Sag: Staphylococcus agalactiae.
We additionally performed qPCR detection of G. vaginalis in a subset of samples. Thus, we could prove the presence of G. vaginalis in all samples with a Gva CST. We further tested two samples which were negative for G. vaginalis reads in the sequencing and they were negative in the qPCR as well (Supplementary Fig. S1).
Only the V3/V4 region identifies the major sexually transmitted pathogen Chlamydia trachomatis
C. trachomatis is the most prevalent sexually transmitted disease worldwide and the major reason for severe pathological consequences such as pelvic inflammatory disease. We specifically analysed read numbers classified as C. trachomatis. While we observe no C. trachomatis reads using the V1/V2 region, C. trachomatis is clearly identified using the V3/V4 region (Fig. 1 and Supplementary Table S1), with four samples displaying C. trachomatis read numbers ranging from 22 to 526 (0.44% to 10.52% relative read count) (Table 1). We additionally confirmed the presence of C. trachomatis in these samples using conventional qPCR (Table 1). Three of the C. trachomatis positive samples appear with a change in their CST between the two regions (Supplementary Table S2). However, both regions identify certain bacterial taxa associated with C. trachomatis infection in an indicator species analysis using infection status as grouping variable (Supplementary Tables S3 and S4).
Discussion
Numerous combinations of primer pairs targeting different regions of the bacterial 16S gene have been employed to identify the composition of bacterial communities by NGS. Previous research shows that different regions yield contrasting results when applied on the same samples10,12,16. However, prior to this study the representativeness of commonly used 16S rRNA gene primers pairs was not assessed for the lower urogenital tract.
It is important to note that in general partial 16S sequences are critical when it comes to classification as different taxa can be classified to different levels. While we are not able to classify all taxa to species level, this is possible for many species within the urogenital tract, enabling us to discuss the detailed taxonomic analysis we provide here.
Our study indicates that the V1/V2 region fails to identify several important species inhabiting the lower urogenital tract (e.g. B. bifidum, G. vaginalis and C. trachomatis), which is in line with a previous report14. Thus, the usage of the 27 F/338 R primer pair fails to detect bacteria important to vaginal health and disease. In contrast, the primers commonly used for amplifying the V3/V4 hypervariable region do not have this limitation17.
The most drastic changes we observe between the two regions are due to the abundance of G. vaginalis. The V3/V4 region determines this species as an characteristic component of many bacterial communities in this data set, which is additionally confirmed by qPCR quantification. In contrast, the V1/V2 region lacks the G. vaginalis detection, which may lead many samples to deviate from their true underlying community structure. This is noticeably shown by a considerable amount of samples having a Lcr or Lga CST based on V1/V2 which are clustering into a CST with high relative read counts of G. vaginalis (Gva1 and Gva2 CST) based on V3/V4. These differences in CST assignment are important to consider, as the biological function in defense to pathogens largely depends on microbial composition. Within the commensal vaginal microbiome, lactobacilli are a major maintainer of vaginal balance and health, as they are capable of producing lactic acid and therefore lowering the pH18. Indeed, it was recently shown that a CST with high relative amounts of L. crispatus is associated with a decreased risk of C. trachomatis transmission from infected partners6 and that L. crispatus shows inhibitory capacity on the infectivity of C. trachomatis in vitro5. Further, the ability of the bacterial community to trap HIV is dependent on the CST present in the vagina2 and a protective role of L. crispatus-dominated CST against Escherichia coli in the vagina was shown19. The interpretation of such studies is largely driven by the primers used, as the number of Lactobacillus-dominated samples seems to be overestimated with the V1/V2 region as our data show. Thus, a recent study by Callahan et al. focusing on the presence of lactobacilli vs. Gardnerella in the preterm birth microbiota20, would have yielded severely erroneous results had they used the V1/V2 region primers. The missing C. trachomatis reads by using the V1/V2 primers further emphasizes the importance of using primers for V3/V4 or another alternative for urogenital microbiota studies.
However, while the V1/V2 region appears to be characterized by under-representing vaginal pathogens such as G. vaginalis and C. trachomatis on the one hand, it is also seems to present other important vaginal pathogens more prominent as they are in reality. As of example, Ureaplasma urealyticum is an often detected pathogen in the vagina21 and seems to be over-estimated using the V1/V2 region. In combination, the skewed representation of important vaginal bacteria might shift the focus of pathogens in a misleading direction in microbiota analysis determined by the V1/V2 region.
Considering the above-mentioned importance of CST composition in the defense against infectious diseases, a particular focus needs to be set on a complete and comparable assignment of CSTs. While Robinson et al. guide researchers towards standardization of their protocols in microbiota studies using 16S genes in general, and point out the importance of the primer choice13, we here show that the V3/V4 region seems to perform much better than the V1/V2 region for urogenital studies in females when using the common primer pairs as outlined in the method section. Of note, special formulations of primers have also been developed to improve mismatch-based inadequacy14 and have been used for microbiota studies22,23. However, this coincides with reduced efficiency and specificity during amplification14, and the validity of such mixtures has been called into question24. With our study we therefore aim to contribute to the awareness of accurate study design in microbiota studies of the urogenital tract of females.
Methods
Sample collection
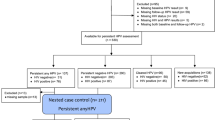
Cervical swabs from healthy females were collected at different clinical sites in Germany, avoiding contamination by other parts of the vagina or the body. All study participants were informed about the purpose of the study and usage of the data and signed a declaration of consent for participation. The study was approved by the ethics committee of the University of Lübeck with the reference number 11–185. All methods were performed in accordance with the relevant guidelines and regulations. Swabs were transported on dry ice and stored in Universal Transport Medium (UTM, COPAN Diagnostics) at −80 °C.
DNA-isolation
One swab per woman was vortexed at highest speed for 1 min. DNA was isolated from 1 ml of the remaining buffer using the MoBio PowerSoil® Kit following the instructions from the manufacturer’s protocol. We introduced a 2 h incubation with OB-Protease at 50 °C followed by homogenisation of the sample using a MoBio PowerLyzer®, both prior to the first centrifugation step. Furthermore, double amounts of solutions C2, C3, and C4 were applied due to the high sample volume. Isolated DNA was stored at −20 °C.
Sequencing
We amplified two partial 16S gene sequences from each isolated DNA in separate PCR-reactions. We used the primer pair 27 F/338 R (27 F: 5′-AGAGTTTGATCCTGGCTCAG-3′/ 338 R: 5′-TGCTGCCTCCCGTAGGAGT-3′) to amplify the V1/V2 region, and the primer pair V3F/V4R (V3F: 5′-CCTACGGGAGGCAGCAG-3′/ V4R: 5′-GGACTACHVGGGTWTCTAAT-3′) for the V3/V4 hypervariable region from the same sample. Sequencing was performed on a MiSeq sequencer (Illumina). All primers contained unique identifier sequences (barcodes) to distinguish between the samples following published methods25 with an optimized primer design26.
Data processing
The raw data were processed using mothur27 version 1.38.1 following a script we optimized based on our previous studies28. In brief, we removed all sequences with deviations from the original primer and barcode sequences. We removed all sequences with ambiguous bases, a length greater than the amplified fragment, and a homopolymer length greater than 12. The sequences were aligned with the SILVA references database29, chimera were removed using the uchime algorithm30. We clustered the processed sequences into operational taxonomic units (OTU) with a global identity threshold of 97% or performed classification based analysis. Species level classification was performed using STIRRUPS using either the V1/V2 or the V3 region15.
Statistical analysis
All statistical analysis and graphical visualisations were carried out in R version 3.2.231. Differences in relative read count of taxa and alpha diversity were tested using Wilcoxon rank sum test. Simpson’s diversity index, species richness, and chao1 index were used as a measure for alpha diversity and computed using R package vegan32 based on OTUs. Heatmaps were constructed with R package BoutrosLab.plotting.general33 using the most abundant taxa. Community state types (CSTs) were assigned by euclidean distances based on relative read counts of taxa with complete linkage clustering and CSTs were assigned on the same level of the hierarchical clustering to enable unbiased CST building. Differences in the proportion of CSTs between the two methods were tested using chi-square testing. A migration plot showing the CST changes between the regions was elaborated using R packages migest34 and circlize35.
Diagnosis of Chlamydia trachomatis
Samples with reads classified as C. trachomatis in the microbiota sequencing were additionally tested by a specific PCR test to prove infection with C. trachomatis (LightMix® Kit C. trachomatis, TIB MOLBIOL) following the manufacturer’s instructions.
Identification of bacterial taxa associated with C. trachomatis-positive samples
We performed an indicator species analysis using the R package indicspecies36 for each region separately to identify bacterial taxa which associate with C. trachomatis infection with infection status as grouping variable.
Quantification of Gardnerella vaginalis
We quantified the abundance of G. vaginalis using qPCR by calculating number of bacteria within a sample using a standard curve generated from a serial dilution with known colony forming units corresponding to DNA-content. The primer sequences used were taken from elsewhere37 (forward primer sequence: CGCATCTGCTAAGGATGTTG; reverse primer sequence: CAGCAATCTTTTCGCCAACT).
Data availability
The datasets analyzed during the current study are available at the European Nucleotide Archive (ENA) with the accession number PRJEB23508.
References
Ravel, J. et al. Vaginal microbiome of reproductive-age women. Proc Natl Acad Sci USA 108, 4680–4687 (2011).
Nunn, K. L. et al. Enhanced Trapping of HIV-1 by Human Cervicovaginal Mucus Is Associated with Lactobacillus crispatus-Dominant Microbiota. MBio 6, e01084–01015, https://doi.org/10.1128/mBio.01084-15 (2015).
Gajer, P. et al. Temporal Dynamics of the Human Vaginal Microbiota. Sci Transl Med 4, 132–152 (2012).
van Houdt, R. et al. Lactobacillus iners-dominated vaginal microbiota is associated with increased susceptibility to Chlamydia trachomatis infection in Dutch women: a case-control study. Sex Transm Infect 94, 117–123, https://doi.org/10.1136/sextrans-2017-053133 (2018).
Nardini, P. et al. Lactobacillus crispatus inhibits the infectivity of Chlamydia trachomatis elementary bodies, in vitro study. Sci Rep 6, 29024, https://doi.org/10.1038/srep29024 (2016).
van der Veer, C., Bruisten, S. M., van der Helm, J. J., de Vries, H. J. & van Houdt, R. The Cervicovaginal Microbiota in Women Notified for Chlamydia trachomatis Infection: A Case-Control Study at the Sexually Transmitted Infection Outpatient Clinic in Amsterdam, The Netherlands. Clin Infect Dis 64, 24–31, https://doi.org/10.1093/cid/ciw586 (2017).
Graspeuntner, S. et al. Microbiota-based analysis reveals specific bacterial traits and a novel strategy for the diagnosis of infectious infertility. PLoS One 13, e0191047, https://doi.org/10.1371/journal.pone.0191047 (2018).
Loeper, N., Graspeuntner, S. & Rupp, J. Microbiota changes impact on sexually transmitted infections and the development of pelvic inflammatory disease. Microbes Infect, https://doi.org/10.1016/j.micinf.2018.02.003 (2018).
Shipitsyna, E. et al. Composition of the vaginal microbiota in women of reproductive age–sensitive and specific molecular diagnosis of bacterial vaginosis is possible? PLoS One 8, e60670, https://doi.org/10.1371/journal.pone.0060670 (2013).
Meisel, J. et al. Skin Microbiome Surveys Are Strongly Influenced by Experimental Design. Journal of Investigative Dermatology 136, 947–956 (2016).
Koren, O. et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc Natl Acad Sci USA 108, 4592–4598 (2011).
Albertsen, M., Karst, S. M., Ziegler, A. S., Kirkegaard, R. H. & Nielsen, P. H. Back to Basics - The Influence of DNA Extraction and Primer Choice on Phylogenetic Analysis of Activated Sludge Communities. PLoS One 10 (2015).
Robinson, C. K., Brotman, R. M. & Ravel, J. Intricacies of assessing the human microbiome in epidemiologic studies. Ann Epidemiol 26, 311–321, https://doi.org/10.1016/j.annepidem.2016.04.005 (2016).
Frank, J. A. et al. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol 74, 2461–2470, https://doi.org/10.1128/AEM.02272-07 (2008).
Fettweis, J. M. et al. Species-level classification of the vaginal microbiome. BMC Genomics (2012).
Cai, L., Ye, L., Tong, A. H., Lok, S. & Zhang, T. Biased diversity metrics revealed by bacterial 16S pyrotags derived from different primer sets. PLoS One 8, e53649, https://doi.org/10.1371/journal.pone.0053649 (2013).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41, e1, https://doi.org/10.1093/nar/gks808 (2013).
Lamont, R. F. et al. The vaginal microbiome: new information about genital tract flora using molecular based techniques. BJOG: an international journal of obstetrics and gynaecology 118, 533–549, https://doi.org/10.1111/j.1471-0528.2010.02840.x (2011).
Ghartey, J. P. et al. Lactobacillus crispatus dominant vaginal microbiome is associated with inhibitory activity of female genital tract secretions against Escherichia coli. PLoS One 9, e96659, https://doi.org/10.1371/journal.pone.0096659 (2014).
Callahan, B. J. et al. Replication and refinement of a vaginal microbial signature of preterm birth in two racially distinct cohorts of US women. Proc Natl Acad Sci USA, https://doi.org/10.1073/pnas.1705899114 (2017).
Ikonomidis, A. et al. Prevalence of Chlamydia trachomatis, Ureaplasma spp., Mycoplasma genitalium and Mycoplasma hominis among outpatients in central Greece: absence of tetracycline resistance gene tet(M) over a 4-year period study. New Microbes New Infect 9, 8–10, https://doi.org/10.1016/j.nmni.2015.11.005 (2016).
Brotman, R. M. et al. Association between cigarette smoking and the vaginal microbiota: a pilot study. BMC Infect Dis 14, 471, https://doi.org/10.1186/1471-2334-14-471 (2014).
Zeeuwen, P. L. et al. Reply to Meisel et al. The Journal of investigative dermatology 137, 961–962, https://doi.org/10.1016/j.jid.2016.11.013 (2017).
Kalle, E., Kubista, M. & Rensing, C. Multi-template polymerase chain reaction. Biomol Detect Quantif 2, 11–22 (2014).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79, 5112–5120, https://doi.org/10.1128/AEM.01043-13 (2013).
Fadrosh, D. W. et al. An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2, 6, https://doi.org/10.1186/2049-2618-2-6 (2014).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75, 7537–7541, https://doi.org/10.1128/AEM.01541-09 (2009).
Giles, D. A. et al. Thermoneutral housing exacerbates nonalcoholic fatty liver disease in mice and allows for sex-independent disease modeling. Nat Med 23, 829–838, https://doi.org/10.1038/nm.4346 (2017).
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Research 35, 7188–7196 (2007).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/ (2015).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.3-2. http://CRAN.R-project.org/package=vegan (2015).
Boutros, P. C. BoutrosLab.plotting.general: Functions to Create Publication-Quality plots. R package version 5.3.4 (2015).
Abel, G. J. migest: Methods for the Indirect Estimation of BilateralMigration. R package version 1.7.3 http://CRAN.R-projekt.org/package=migest (2016).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize Implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812, https://doi.org/10.1093/bioinformatics/btu393 (2014).
De Caceres, M. & Legendre, P. Associations between species and groups of sites: indices and statistical inference. Ecology 90, 3566–3574 (2009).
Menard, J. P., Fenollar, F., Henry, M., Bretelle, F. & Raoult, D. Molecular quantification of Gardnerella vaginalis and Atopobium vaginae loads to predict bacterial vaginosis. Clin Infect Dis 47, 33–43, https://doi.org/10.1086/588661 (2008).
Acknowledgements
We are grateful to A. Gravenhorst, S. Paetzmann, A. Hellberg and K. Cloppenborg-Schmidt for excellent technical assistance. This work was supported by grants from the German Research Foundation (DFG) within the Research Training Group 1743, “Genes, Environment and Inflammation” and the Excellence Cluster 306, “Inflammation at Interfaces”.
Author information
Authors and Affiliations
Contributions
S.G., J.F.B., and J.R. designed the study. S.G. and S.K. performed the laboratory work. S.G. and N.L. performed the data processing and S.G. the statistical analysis. Results were interpreted by S.G., N.L., J.F.B., and J.R. S.G. and N.L. wrote the draft of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Graspeuntner, S., Loeper, N., Künzel, S. et al. Selection of validated hypervariable regions is crucial in 16S-based microbiota studies of the female genital tract. Sci Rep 8, 9678 (2018). https://doi.org/10.1038/s41598-018-27757-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-27757-8
This article is cited by
-
Determining the most accurate 16S rRNA hypervariable region for taxonomic identification from respiratory samples
Scientific Reports (2023)
-
Contamination source modeling with SCRuB improves cancer phenotype prediction from microbiome data
Nature Biotechnology (2023)
-
Hearing rehabilitation and microbial shift after middle ear surgery with Vibrant Soundbridge in patients with chronic otitis media
European Archives of Oto-Rhino-Laryngology (2023)
-
Analysis of the cervical microbiome in women from the German national cervical cancer screening program
Journal of Cancer Research and Clinical Oncology (2023)
-
Alterations of the vaginal microbiome in healthy pregnant women positive for group B Streptococcus colonization during the third trimester
BMC Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.