Abstract
Many hemorheologic Traditional Chinese Medicines (TCMs) that are widely-used clinically lack molecular mechanisms of action. We hypothesized that some of the active components of hemorheologic TCMs may function through targeting prothrombotic P2Y1 and/or P2Y12 receptors. The interactions between 253 antithrombotic compounds from TCM and these two G protein-coupled P2Y receptors were evaluated using virtual screening. Eleven highly ranked hits were further tested in radioligand binding and functional assays. Among these compounds, salvianolic acid A and C antagonized the activity of both P2Y1 and P2Y12 receptors in the low µM range, while salvianolic acid B antagonized the P2Y12 receptor. These three salvianolic acids are the major active components of the broadly-used hemorheologic TCM Danshen (Salvia militorrhiza), the antithrombotic molecular mechanisms of which were largely unknown. Thus, the combination of virtual screening and experimental validation identified potential mechanisms of action of multicomponent drugs that are already employed clinically.
Similar content being viewed by others
Introduction
Many hemorheologic agents in Traditional Chinese Medicine (TCM) are still widely used, including Chuanqiong (Ligusticum wallichii)1, Danshen (Salvia militorrhiza)2, and Honghua (Carthamus tinctorius)3. These hemorheologic agents are highly efficacious, although they often manifest side effects. A common strategy for developing safer drugs includes identifying and extracting the active components of TCMs. However, the molecular mechanisms including targets of their active components remain unknown in most cases. For example, Danshen depside salt, containing only several water-soluble components of Danshen4,5, has fewer side effects and is one of the common hemorheologic treatments used in China4,6,7. The principal water-soluble component of Danshen is salvianolic acid B (SAB), however, several other salvianolic acid variations, including salvianolic acid A (SAA) and salvianolic acid C (SAC), are also major components5. While SAA, SAB and SAC block adenosine 5′-diphosphate (ADP)-induced platelet aggregation8,9,10 more potently than other known inducers, such as thrombin and collagen11, the molecular mechanism of the antithrombotic action of these salvianolic acids, including their target receptors, remains largely unknown11. A recent study using atomic-force microscopy concluded that SAB can bind to the purinergic G protein-coupled receptor (GPCR) P2Y12; however, only a single dose of 280 μM was tested10.
We hypothesized that the active components of hemorheologic TCMs may function through targeting the purinergic GPCRs P2Y1 and/or P2Y12 due to their critical roles in platelet aggregation12. Both P2Y receptors can be activated by the same endogenous ligand, ADP13. Coactivation of the two receptors is essential for ADP-induced platelet aggregation12. Specifically, activation of the Gq-coupled P2Y1 receptor leads to intracellular calcium mobilization, platelet shape change and formation of small, reversible aggregates. Activation of the Gi-coupled P2Y12 receptor leads to a decrease in the level of cyclic adenosine 3′,5′-monophosphate (cAMP) that drives the reversible platelet aggregates to an irreversible state14,15. As a result, both P2Y receptors are antithrombotic drug targets; inhibition of both receptors has a synergistic effect16,17,18. Several antithrombotic drugs, such as clopidogrel, prasugrel19 and ticagrelor20, antagonize the P2Y12 receptor. However, no drugs on market targets the P2Y1 receptor. Since the structures of the P2Y1 and P2Y12 receptors were solved recently using X-ray crystallography21,22,23, we were able to take a structure-based virtual screening approach to predict ligands of the P2Y1 and P2Y12 receptors from antithrombotic components of TCMs.
Results
Eleven hits for P2Y receptors were found by ensemble docking
Ensemble docking, a method that has recently been widely used for GPCR ligand discovery24,25,26 since GPCRs tend to have flexible binding pockets21,27,28, was adopted for this study. In order to get multiple receptor conformations, 100 ns MD simulations were performed starting with the crystal structures of the P2Y1 (PDB: 4XNW) and P2Y12 receptors (PDB: 4PXZ and 4NTJ) (Figure S1). The virtual screening library consists of 253 antiplatelet aggregatory TCM components. Following the ensemble docking, the top fifty ligands ranked by Glide XP scores were further re-ranked by Induced Fit Docking29 that allows residues within 5 Å of the ligand to move. Finally, eleven hits were manually picked for experimental testing in part based on their commercial availability, resulting in the identification of SAA and SAC as antagonists of the P2Y1 and P2Y12 receptors at low µM level, while SAB inhibits only the P2Y12 receptor (Fig. 1a).
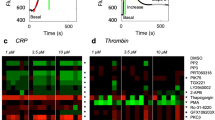
Summary of structure-based TCM ligand discovery strategy for P2Y1 and P2Y12 receptors. (a) Flowchart of ligand discovery for P2Y1 and P2Y12 receptors from TCM components with antiplatelet aggregation function. (b) Percentage inhibition of the binding of [3H]2MeSADP to the P2Y12 receptor by 11 hits (at 100 μM) selected from virtual screening. (c) 2D structures of three validated hits, SAA, SAB and SAC. The shared parts in the three compounds are in black, the different linker regions are in red, and the unique extra moiety in SAB is in blue.
Three ligands have antagonist activity against P2Y1 and/or P2Y12 receptors
The eleven selected components were further examined in radioligand binding and functional assays. We initially tested the eleven compounds by measuring the inhibition of [3H]2-methylthioadenosine 5′-diphosphate (2MeSADP) binding at the human P2Y12 receptor (Fig. 1b). The three salvianolic acids from Danshen, SAA, SAB and SAC (Fig. 1c), were found to have significant inhibitory effect on P2Y12 receptor binding at a concentration of 100 µM. The remaining eight tested components, including salvianolic acid D (33.5 ± 8.6% inhibition), were weakly inhibiting or inactive. Another Danshen component, rosmarinic acid, was previously shown to be more potent in antagonizing thrombin-induced than ADP-induced platelet aggregation30. Our results confirmed that it is indeed a weak ligand of the P2Y12 receptor (31.6 ± 6.2% inhibition at 100 µM).
The binding affinities of SAA, SAB and SAC for the P2Y12 receptor were determined by full inhibition curves resulting in Ki values of 20.3 ± 1.5, 36.4 ± 20.3, 15.6 ± 6.5 µM, respectively (Fig. 2a). The functional antagonism of these three salvianolic acids against the P2Y12 receptor was measured by their effect on agonist-induced inhibition of forskolin-stimulated cyclic AMP accumulation in P2Y12 receptor-expressing U2OS cells. For example, in a concentration-dependent manner, SAA shifted the agonist response curve to the right without affecting its maximum agonist effect (Fig. 2b). This observation is consistent with competitive antagonism, but allosteric antagonism cannot be excluded31. Schild analysis of the antagonism yielded KB values of 24.2 ± 5.9, 33.6 ± 6.4 and 23.7 ± 4.5 µM for SAA, SAB and SAC, respectively (Fig. 2c).
Radioligand binding assays and functional assays showed that SAA and SAC can bind and antagonize both P2Y1 and P2Y12 receptors, while SAB can bind and antagonize only the P2Y12 receptor. (a) Displacement curves of SAA, SAB and SAC against [3H]2MeSADP binding to the P2Y12 receptor. (b) Functional antagonism by SAA of 2MeSADP-induced inhibition of foskolin-stimulated cAMP accumulation in U2OS cells expressing the P2Y12 receptor. (c) Schild plots of the antagonism of SAA, SAB and SAC at the P2Y12 receptors. (d) Displacement curves of SAA, SAB and SAC against [3H]2MeSADP binding to the P2Y1 receptor. (e) 2MeSADP-induced IP-1 accumulation in U2OS cells expressing the P2Y1 receptor (compared to MRS2500). (f) Schild plots of antagonism by SAA and SAC at the P2Y1 receptor. Results are expressed as mean ± SEM. The Ki values from binding experiments and KB values from Schild analyses of functional antagonism by SAA, SAB and SAC are listed in the text and are from at least three independent experiments. MRS, MRS2500; SAR, SAR216471.
Both P2Y1 and P2Y12 receptors play important roles in platelet aggregation. Therefore, we also tested the potential effects of SAA, SAB and SAC on the P2Y1 receptor. Initial binding results showed that both SAA and SAC produced a substantial inhibitory effect on [3H]2MeSADP binding to the P2Y1 receptor while the inhibition by SAB was significantly weaker (Fig. 2d); the concentration-dependent inhibitory curves for SAA and SAC resulted in Ki values of 14.7 ± 2.4 and 23.3 ± 4.5 µM, respectively. SAA and SAC antagonized 2MeSADP-induced accumulation of inositol 1-phosphate (IP-1) (Fig. 2e). KB values calculated by Schild analysis for the P2Y1 receptor were 9.1 ± 1.5 and 15.6 ± 2.0 µM for SAA and SAC, respectively (Fig. 2f). Thus, the Schild analysis of the antagonist potencies of SAA and SAC for Gq-coupled P2Y1 and Gi-coupled P2Y12 receptors was consistent with their binding affinities measured in the radioligand assays.
To examine the possibility that the observed pharmacological activity might be due to colloidal aggregation of these compounds, we tested the effect of a nonionic detergent, Tween-2032,33,34, on the inhibition of the specific binding of [3H]2MeSADP to the P2Y1 receptor by the two potent salvianolic acids (SAA and SAC). We found that the inhibition curves are not rightward-shifted by Tween-20 (Figure S2). Thus, the results suggest that colloidal aggregation is not a major contributor to the inhibition of the specific binding of [3H]2MeSADP by salvianolic acids. Additionally, the reports that SAA and SAC are more potent at ADP-induced than at thrombin-induced platelet aggregation8,35 may also support a specific effect of these compounds.
SAA, SAB and SAC may act through additional targets
We also tested if SAA, SAB and SAC are promiscuously binding compounds, due to the fact that they all have multiple phenolic groups and are thus potentially PAINS36. To find other possible targets of the three compounds, we evaluated their potential binding to 45 diverse receptors, transporters and ion channels (human, unless noted) using radioligand binding assays via the Psychoactive Drug Screening Program (PDSP) (Table S1)37. Among the three compounds, only SAB bound to one off-target site, i.e. the α1B adrenergic receptor with a Ki value of 6.26 ± 1.46 µM (n = 3) (Figure S3), however following FLIPR assays showed that SAB is neither agonist nor antagonist of α1B (Figure S4). No other significant affinity (<50% inhibition at 10 µM) was detected for any of the proteins listed in Table S1. Therefore, these natural products may have multiple targets in vivo, but do not display the indiscriminate promiscuity characteristic of PAINS compounds.
Binding poses of SAA, SAB and SAC are consistent with their functions
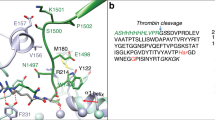
We analyzed the binding poses of SAA, SAB and SAC in the binding sites of P2Y1 and P2Y12 receptors. Their poses were consistent with their antagonistic function against these receptors. For the P2Y1 receptor, the binding poses of SAA and SAC were similar to the co-crystalized antagonist (1′R,2′S,4′S,5′S)-4-(2-iodo-6-methylamino-purin-9-yl)-1-[(phosphato)-methyl]-2-(phosphato)-bicyclo[3.1.0]hexane (MRS2500) (Fig. 3), consistent with their function as P2Y1 antagonists. However, unlike SAA and SAC, SAB had a more external binding pose (Fig. 3b), likely due to the steric bulk of SAB, consistent with SAB having little effect on the P2Y1 receptor.
Docking poses of SAA (green), SAB (purple) and SAC (yellow) in P2Y1 receptor. (a) Comparison of the docking poses of SAA and SAC with co-crystalized antagonist MRS2500 (cyan) in P2Y1 receptor shows that SAA and SAC align well with MRS2500. (b) Comparison of the binding pose of SAB with MRS2500, SAA and SAC shows that it does not occupy the same space as MRS2500/SAA/SAC. (c,d) SAA and SAC have good interactions with key residues in the pocket of P2Y1 receptor. Light blue solid lines and magenta dashed lines represent hydrogen bonds and π-π/cation-π interactions, respectively.
At the P2Y12 receptor, SAA, SAB and SAC were predicted to bind similarly, interacting with all transmembrane helices except I and II, mimicking the co-crystalized antagonist ethyl 6-(4-((benzylsulfonyl)carbamoyl)piperidin-1-yl)-5-cyano-2-methylnicotinate (AZD1283) but not the co-crystalized agonist 2MeSADP (Fig. 4). These models are consistent with the fact that all three salvianolic acids are inhibitors of the P2Y12 receptor. Each of the three compounds interacted with several key residues such as Tyr1053.33 (Ballesteros-Weinstein numbering38 scheme), Asn1594.60, His171ECL2, Arg2566.55 and Lys2807.34, albeit the detail interactions were slightly different. The binding poses of the three compounds showed that unpaired hydrogen bond donors and acceptors increased in the order of SAC, SAA, and SAB. It is uncertain whether these differences contributed to the small differences in their potencies.
Docking poses of SAA (green), SAB (purple) and SAC (yellow) at the P2Y12 receptor. (a) Comparison of the docking poses of SAA, SAB and SAC with co-crystalized antagonist AZD1283 (magenta) and agonist 2MeSADP (blue) in P2Y12 receptor show that SAA, SAB and SAC can superimpose with the co-crystalized antagonist but not the agonist. (b–d) SAA, SAB and SAC bind the P2Y12 receptor in a similar pose and show good interactions with residues lining the pocket. Light blue solid lines and magenta dashed lines represent hydrogen bonds and π-π/cation-π interactions, respectively.
Discussion
A significant finding is that while SAB only antagonizes the P2Y12 receptor, SAA and SAC are dual inhibitors of P2Y1 and P2Y12 receptors. Simultaneous inhibition of P2Y1 and P2Y12 receptors by a combination of selective inhibitors at low concentration has already been shown to synergistically enhance antithrombotic activity, especially for high shear stress-induced platelet aggregation5. A combination of P2Y1 and P2Y12 receptor antagonists at low concentrations reduced platelet–platelet cohesion and mural thrombosis12. Although these non-purine salvianolic acids do not bind to the P2Y receptors as potently as marketed P2Y12 receptor antagonist drugs, they are nevertheless effective in practice. We hypothesize that the therapeutic action of these salvianolic acids, especially when administered by injection, may benefit from the synergistic antithrombotic effect of blocking both P2Y receptors. Furthermore, it is well known that nanomolar inhibitors of the P2Y12 receptor are associated with a significantly increased bleeding risk19,20. A case in point is that clinical use often requires two antithrombotic drugs. Administration of aspirin and a P2Y12 antagonist is recommended for all patients with a clear diagnosis of acute coronary syndrome (ACS) to balance the benefits and bleeding complications39. A potential advantage of the P2Y1 receptor antagonists as antithrombotic drugs compared to the P2Y12 receptor antagonists may be the controllable bleeding time12. Thus, the finding that multiple components from Danshen have low μM antagonistic activity against both P2Y1 and P2Y12 receptors may explain the clinical efficacy and the relatively safe use of Danshen for the prevention and treatment of ACS during the past two millennia40.
The antithromobotic effects of SAA, SAB and SAC both in vitro and in vivo have been previously reported8,9,10,35, although the potential mechanism mediated via a specific P2Y receptor has been largely unexplored. The effect of SAA was found to be more potent on ADP-induced than thrombin- or arachidonic acid-induced rat platelets aggregation both in vivo and in vitro8. The antiplatelet efficacy of SAB was demonstrated in ACS patients undergoing treatment with clopidogrel and aspirin10. SAC was found to inhibit platelet aggregation in vitro and in vivo9,35. In addition, although salvianolic acids are often believed to be redox-active and function as reactive oxygen species scavengers, actually their half-lives are not very short. It was reported that in rat plasma, t1/2 of SAB is about 105 min (100 mg/kg, i.v.) and 248 min (500 mg/kg, p.o.), and t1/2 of SAA is 197 min (100 mg/kg, p.o.)11.
SAA, SAB and SAC are polyphenolic phytochemicals, which are often thought to bind to many membrane proteins indiscriminately36. Indeed, it was reported SAB/SAA can bind MMP-9 (Ki 79.2 μM)41, SH2 domain of Src family kinases (IC50 41–90 μM)42 and CD36 (IC50 23.2 μM)43. Our study added two more members to the list: P2Y1 and P2Y12 receptors, with SAA and SAC having the highest affinities among all the determined Ki/EC50/IC50. Our broad screening of the three compounds against 45 receptors/transporters/ion channels at PDSP found no hits, thus the three compounds pass through the orthogonal target assays suggested by the recent review on PAINS44. Nevertheless, the important clinical distinction is that Danshen is proven safe and efficacious in human use. Although we realize that by the current standards of pharmaceutical development it would be highly unlikely that such a drug combination would be considered now, we have shed light on the mechanism of action of a proven TCM treatment.
Finding target(s) for multicomponent-TCM is critical for understanding their mechanisms and improving the safety. Computational methods such as network-based and machine learning-based methods have been developed to predict drug-target interactions45,46 and synergistic drug combinations47. Here we used virtual screening to predict that SAA, SAB and SAC, three components of a broadly used antithrombotic TCM Danshen, bind to two known antithrombotic targets: P2Y1 and P2Y12 receptors. This prediction was validated experimentally revealing binding affinity at low μM levels and functional antagonistic activity against these two receptors. These results suggest a plausible mechanism for the antithrombotic activity of Danshen. By blocking both P2Y receptors through the three major components, Danshen exhibits a synergistic antithrombotic effect while also keeping bleeding risk tolerable. This mechanism provides a basis for further development of a dual antagonist against the P2Y1 and P2Y12 receptors as an antithrombotic agent. We showed that virtual screening and experimental validation can identify a molecular mechanism of action of multicomponent drugs that are already in clinical use.
Methods
Protein preparation
Agonist-bound and antagonist-bound structures of human P2Y12 receptor (PDB: 4PXZ, 4NTJ) and an antagonist-bound structure of human P2Y1 receptor (PDB: 4XNW) were prepared for molecular dynamics simulation. Fusion protein, water molecules, ions, solvent molecules and ligand were removed from the three structures. Then the structures were processed using the protein preparation wizard in Schrodinger Suite 2015–448. During the process, hydrogens, disulfide bonds, missing side chains and loops were appropriately included; N-terminal and C-terminal ends were capped, and protonation states were assigned at pH 7.0 by Protassign; and a minimization with 0.3 Å RMSD constraints on heavy atoms was performed.
Molecular dynamics simulations
Molecular dynamics simulations were performed using GROMACS 5.0.649. The three prepared structures of P2Y1 and P2Y12 receptors were embedded into a POPC (1-palmytoil-2-oleoyl-sn-glycero-3-phosphatidylcholine) lipid bilayer, respectively. Membrane Builder50 of CHARMM-GUI51 was used to build the protein/membrane complex systems. Each system consisted of a receptor, a lipid bilayer of ~120 POPC molecules, ~10700 TIP3P water molecules and 0.15 M NaCl with respective counter ions. Nearly 50,000 atoms in total were contained in a periodic box ~75 × 75 × 115 Å3. Van der Waals and short-range electrostatic interactions were cut off at 10 Å. Equilibration was performed using a NPT run at 1 atm and 310 K for 10 ns with a time step of 1 fs. Molecular dynamics simulations were performed in the NPT ensemble for 100 ns using standard CHARMM 36 force field51 at 1 atm and 310 K with a time step of 2 fs.
MD trajectories clustering
The MD simulation trajectories of the three P2Y receptor systems were used to examine their potential small molecule interactions. The ligand binding site was defined by protein residues within 5 Å of the cocrystalized ligand in each X-ray structure. Based on the root-mean-square deviation (RMSD) of the selected residues, receptor snapshots taken every 1 ps in the simulation trajectory were clustered for each target site using the g_cluster tool in GROMACS package. The gromos clustering algorithm was applied. 262, 304 and 554 clusters were generated from the MD trajectories of 4XNW, 4PXZ and 4NTJ, respectively, with a 1.0 Å RMSD cutoff. The top 10 populated clusters from each trajectory were selected for ensemble docking.
Building a library of TCM components with antiplatelet aggregation function
A library of 253 TCM components with antiplatelet aggregation function was built. These compounds were filtered from the Traditional Chinese Medicine Database52 and an in-house TCM component library by using the keyword “antiplatelet aggregation”. Unspecified chiral centers were manually checked and assigned if available in literature. LigPrep was used to generate 3D structures of the ligands. During the process an OPLS3 force field53 was used for geometry optimization, pH 7.0 ± 2.0 was used to generate possible protonation states and the option “retain specified chiralities (vary other chiralities)” was used to generate stereoisomers. In total 560 compounds were generated through LigPrep processing.
Ensemble Docking
Glide XP48,54,55,56 was used to dock the antiplatelet aggregation library into the 30 conformations generated from clustered MD trajectories. Docking boxes were defined by the residues in the ligand binding site. Default parameters of Glide XP were used except for the inner and outer box lengths, which were set to 14.0 Å and 28.2 Å, respectively. MM-GBSA was used to optimize ligand conformation and rescore relative binding affinity. Ligand and receptor residues within 5 Å of the ligand were allowed to move in MM-GBSA calculation. Fifty top-scored ligands were selected from all docking results. Then Induced-Fit Docking (IFD)29,57,58,59 was further applied to optimize the binding poses and binding score. Finally, 11 antiaggregatory compounds were manually selected based on their docking scores and poses for experimental validation.
Membrane preparation and binding assays
U2Os cells expressing the P2Y1 or P2Y12 receptors were cultured in DMEM supplemented with 10% fetal bovine serum, 100 Units/ml penicillin, 100 µg/ml streptomycin and 2 µmol/ml glutamine. After reaching confluence, cells were detached from plates by scraping into PBS and centrifuged at 250 g for 5 min. The resulting pellets were re-suspended in ice-cold Tris-HCl buffer (50 mM, pH 7.4) and homogenized. After homogenization and suspension, cells were centrifuged at 1000 g for 10 min and the pellet was discarded. The suspension was re-centrifuged at 20,000 g for 60 min at 4 °C. The pellets were re-suspended, split into aliquots and stored at −80 °C. The protein concentration was measured using the Bradford assay. For P2Y1 and P2Y12 receptor binding assays, saturation curves were measured by addition of various concentrations of 50 µl [3H]2MeSADP (specific activity, 7.5 Ci/mmol; Movarek Biochemicals, Brea, CA) and 100 μl of membrane suspension to a total of 200 μl of total assay volume. Non-specific binding was determined using 10 µM MRS2500 (Tocris; St. Louis, MO) for the P2Y1 receptor and 10 μM SAR216471 (Tocris; St. Louis, MO) for the P2Y12 receptor. For the displacement assay, various concentrations of inhibitors were incubated with [3H]2MeSADP (1.0 nM for P2Y1; 0.4 nM for P2Y12) and membrane preparations (20 µg/sample) at 25 °C for 60 min. The separation of bound from free radioligand was accomplished by rapid vacuum filtration of the incubation mixture over GF/B filter using a Brandel cell harvester (Brandel Inc., Gaithersburg, MD). Filters were washed two times with 3 ml of ice-cold Tris-HCl buffer, pH 7.5. Radioactivity on the filters was quantified using a Tri-carb Liquid Scintillation Counter (PerkinElmer Life and Analytical Sciences). 2MeSADP was purchased from Sigma (St. Louis, MO). All other materials were from standard commercial sources and of analytical grade. For radioligand binding, calculation of apparent binding affinities, Ki = IC50/(1 + [radioligand]/Kd), was performed using the nonlinear iterative curve-fitting program of Prism (GraphPad Software Inc., San Diego, CA).
Functional assays
IP-one assay
Inositol 1-phosphate (IP-1) was measured using the IP-One Tb HTRF kit (Cisbio Bioassays, Bedford, MA) as described previously60,61. Briefly, after overnight growth, U2Os cells expressing the P2Y1 receptor were first treated with an antagonist for 20 min before the treatment with agonist and incubated for another 60 min. IP-1 detection reagents were added as instructed by the manual from the manufacturer. The assay plates were read on a Mithras LB940 reader (Berthold Technologies, Oak Ridge, TN) using a time-resolved fluorescence ratio (665/620 nm).
cAMP assay
U2Os cells expressing the P2Y12 receptor were cultured in DMEM medium containing 10% fetal bovine serum, 100 units/ml penicillin, 100 µg/ml streptomycin and 2 µmol/ml glutamine. For the assay of cAMP accumulation, cells were plated in 96-well plates in 100 µl medium overnight. Cells were then treated with assay buffer containing rolipram (10 mM) and antagonists for 20 min followed by the addition of agonists and incubate for 10 min. After 10 min incubation with agonist, Forskolin (10 μM) was added to the mixture and the incubation was continued for another 10 min. The reaction was terminated upon removal of the supernatant and addition of 100 µl Tween-20 (0.3%). Intracellular cAMP levels were measured with an ALPHAScreen cAMP assay kit as instructed by the manufacturer (PerkinElmer). Binding and functional parameters were calculated using Prism 7.0 software (GraphPAD, San Diego, CA, USA). Data were expressed as mean ± sem.
FLIPR assay
For agonist assay: cell line expressing target receptor was seeded in a 384-well black-wall, clear-bottom plate about 18 hours prior to the day of experiment and maintained at 37 °C/5% CO2. Then, 20 μl of dye-loading solution was added into the wells and the plates were subsequently placed into a 37 °C incubator for 60 min, followed by a 15 min incubation at room temperature. At last, 10 μl of compounds or control agonist were added into respective wells of the assay plate during reading in FLIPR.
For antagonist assay: cell line expressing target receptor was seeded in a 384-well black-wall, clear-bottom plate about 18 hours prior to the day of experiment and maintained at 37 °C/5% CO2. Then the dye-loading solution and 10 μl of control antagonist and test samples were added into respectively wells in sequence. The plates were subsequently placed into a 37 °C incubator for 60 min, followed by a 15 min incubation at room temperature. At last, 12.5 μl of control agonist were added into respective wells of the assay plate during reading in FLIPR.
Change history
24 July 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Guo, M., Liu, Y. & Shi, D. Cardiovascular Actions and Therapeutic Potential of Tetramethylpyrazine (Active Component Isolated from Rhizoma Chuanxiong): Roles and Mechanisms. Biomed Res Int 2016, 2430329, https://doi.org/10.1155/2016/2430329 (2016).
Yufei, F. Analysis of Salvia miltiorrhiza (Danshen). African Journal of Microbiology Research 6, https://doi.org/10.5897/ajmr11.1256 (2012).
Duan, Z. Z. et al. Danhong injection protects cardiomyocytes against hypoxia/reoxygenation- and H2O2-induced injury by inhibiting mitochondrial permeability transition pore opening. J Ethnopharmacol 175, 617–625, https://doi.org/10.1016/j.jep.2015.08.033 (2015).
Wu, W. Y. & Wang, Y. P. Pharmacological actions and therapeutic applications of Salvia miltiorrhiza depside salt and its active components. Acta Pharmacol Sin 33, 1119–1130, https://doi.org/10.1038/aps.2012.126 (2012).
Chen, J. H., Wang, F. M., Lee, F. S. C., Wang, X. R. & Xie, M. Y. Separation and identification of water-soluble salvianolic acids from Salvia miltiorrhiza Bunge by high-speed counter-current chromatography and ESI-MS analysis. Talanta 69, 172–179, https://doi.org/10.1016/j.talanta.2005.09.041 (2006).
Yan, Y. Y. et al. Post-Marketing Safety Surveillance of the Salvia Miltiorrhiza Depside Salt for Infusion: A Real World Study. PLoS One 12, e0170182, https://doi.org/10.1371/journal.pone.0170182 (2017).
Chang, Y. et al. Postmarketing safety evaluation: depside salt injection made from Danshen (Radix Salviae Miltiorrhizae). J Tradit Chin Med 34, 749–753 (2014).
Fan, H. Y. et al. Antiplatelet and antithrombotic activities of salvianolic acid A. Thromb Res 126, e17–22, https://doi.org/10.1016/j.thromres.2010.04.006 (2010).
Chen, Y. et al. A Platelet/CMC coupled with offline UPLC-QTOF-MS/MS for screening antiplatelet activity components from aqueous extract of Danshen. J Pharm Biomed Anal 117, 178–183, https://doi.org/10.1016/j.jpba.2015.06.009 (2016).
Liu, L. et al. Salvianolic acid B inhibits platelets as a P2Y12 antagonist and PDE inhibitor: evidence from clinic to laboratory. Thromb Res 134, 866–876, https://doi.org/10.1016/j.thromres.2014.07.019 (2014).
Ho, J. H. & Hong, C. Y. Salvianolic acids: small compounds with multiple mechanisms for cardiovascular protection. J Biomed Sci 18, 30, https://doi.org/10.1186/1423-0127-18-30 (2011).
Jin, J. G. & Kunapuli, S. P. Coactivation of two different G protein-coupled receptors is essential for ADP-induced platelet aggregation. Proc. Natl. Acad. Sci. USA 95, 8070–8074, https://doi.org/10.1073/pnas.95.14.8070 (1998).
Hardy, A. R. et al. P2Y1 and P2Y12 receptors for ADP desensitize by distinct kinase-dependent mechanisms. Blood 105, 3552–3560, https://doi.org/10.1182/blood-2004-07-2893 (2005).
Offermanns, S. Activation of platelet function through G protein-coupled receptors. Circ Res 99, 1293–1304, https://doi.org/10.1161/01.RES.0000251742.71301.16 (2006).
Sangkuhl, K., Shuldiner, A. R., Klein, T. E. & Altman, R. B. Platelet aggregation pathway. Pharmacogenet Genomics 21, 516–521, https://doi.org/10.1097/FPC.0b013e3283406323 (2011).
Turner, N. A., Moake, J. L. & McIntire, L. V. Blockade of adenosine diphosphate receptors P2Y(12) and P2Y(1) is required to inhibit platelet aggregation in whole blood under flow. Blood 98, 3340–3345 (2001).
Nylander, S., Mattsson, C., Ramstrom, S. & Lindahl, T. L. Synergistic action between inhibition of P2Y12/P2Y1 and P2Y12/thrombin in ADP- and thrombin-induced human platelet activation. Br J Pharmacol 142, 1325–1331, https://doi.org/10.1038/sj.bjp.0705885 (2004).
Gremmel, T. et al. Synergistic Inhibition of Both P2Y(1) and P2Y(12) Adenosine Diphosphate Receptors As Novel Approach to Rapidly Attenuate Platelet-Mediated Thrombosis. Arterioscl Throm Vas 36, 501–509, https://doi.org/10.1161/Atvbaha.115.306885 (2016).
Wiviott, S. D. et al. Prasugrel versus clopidogrel in patients with acute coronary syndromes. N Engl J Med 357, 2001–2015, https://doi.org/10.1056/NEJMoa0706482 (2007).
Wallentin, L. et al. Ticagrelor versus clopidogrel in patients with acute coronary syndromes. N Engl J Med 361, 1045–1057, https://doi.org/10.1056/NEJMoa0904327 (2009).
Zhang, J. et al. Agonist-bound structure of the human P2Y12 receptor. Nature 509, 119–122, https://doi.org/10.1038/nature13288 (2014).
Zhang, K. et al. Structure of the human P2Y12 receptor in complex with an antithrombotic drug. Nature 509, 115–118, https://doi.org/10.1038/nature13083 (2014).
Zhang, D. et al. Two disparate ligand-binding sites in the human P2Y1 receptor. Nature 520, 317–321, https://doi.org/10.1038/nature14287 (2015).
Miao, Y. et al. Accelerated structure-based design of chemically diverse allosteric modulators of a muscarinic G protein-coupled receptor. Proc Natl Acad Sci USA 113, E5675–5684, https://doi.org/10.1073/pnas.1612353113 (2016).
Cavasotto, C. N. et al. Discovery of novel chemotypes to a G-protein-coupled receptor through ligand-steered homology modeling and structure-based virtual screening. Journal of Medicinal Chemistry 51, 581–588, https://doi.org/10.1021/jm070759m (2008).
Kim, S. K. et al. Predicted structures of agonist and antagonist bound complexes of adenosine A3 receptor. Proteins 79, 1878–1897, https://doi.org/10.1002/prot.23012 (2011).
Lagerstrom, M. C. & Schioth, H. B. Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat Rev Drug Discov 7, 339–357, https://doi.org/10.1038/nrd2518 (2008).
Hua, T. et al. Crystal structures of agonist-bound human cannabinoid receptor CB1. Nature. https://doi.org/10.1038/nature23272 (2017).
Schrödinger Induced Fit Docking Protocol 2015-4, Glide version 6.4, Prime version 3.7 (Schrödinger, LLC) (2015b).
Wang, Y. et al. Aqueous extract of Rabdosia rubescens leaves: forming nanoparticles, targeting P-selectin, and inhibiting thrombosis. Int J Nanomedicine 10, 6905–6918, https://doi.org/10.2147/IJN.S91316 (2015).
Gao, Z. G. & Jacobson, K. A. Distinct Signaling Patterns of Allosteric Antagonism at the P2Y1 Receptor. Mol Pharmacol 92, 613–626, https://doi.org/10.1124/mol.117.109660 (2017).
Irwin, J. J. & Shoichet, B. K. Docking Screens for Novel Ligands Conferring New Biology. J Med Chem 59, 4103–4120, https://doi.org/10.1021/acs.jmedchem.5b02008 (2016).
Duan, D., Doak, A. K., Nedyalkova, L. & Shoichet, B. K. Colloidal aggregation and the in vitro activity of traditional Chinese medicines. ACS Chem Biol 10, 978–988, https://doi.org/10.1021/cb5009487 (2015).
Ryan, A. J., Gray, N. M., Lowe, P. N. & Chung, C. W. Effect of detergent on “promiscuous” inhibitors. J Med Chem 46, 3448–3451, https://doi.org/10.1021/jm0340896 (2003).
Zhao, T. et al. Specific Combination of Salvianolic Acids As Core Active Ingredients of Danhong Injection for Treatment of Arterial Thrombosis and Its Derived Dry Gangrene. Front Pharmacol 8, 361, https://doi.org/10.3389/fphar.2017.00361 (2017).
Ingolfsson, H. I. et al. Phytochemicals perturb membranes and promiscuously alter protein function. ACS Chem Biol 9, 1788–1798, https://doi.org/10.1021/cb500086e (2014).
Besnard, J. et al. Automated design of ligands to polypharmacological profiles. Nature 492, 215–220, https://doi.org/10.1038/nature11691 (2012).
Ballesteros, J. A. & Weinstein, H. In Methods in Neurosciences Vol. 25 (ed. Sealfon Stuart, C.) 366–428 (Academic Press, 1995).
Huber, K. et al. Antiplatelet and anticoagulation agents in acute coronary syndromes: what is the current status and what does the future hold? Am Heart J 168, 611–621, https://doi.org/10.1016/j.ahj.2014.06.014 (2014).
Wang, L. et al. Salvia miltiorrhiza: A Potential Red Light to the Development of Cardiovascular Diseases. Curr Pharm Des 23, 1077–1097, https://doi.org/10.2174/1381612822666161010105242 (2017).
Jiang, B. et al. Salvianolic acid B functioned as a competitive inhibitor of matrix metalloproteinase-9 and efficiently prevented cardiac remodeling. BMC Pharmacol 10, 10, https://doi.org/10.1186/1471-2210-10-10 (2010).
Sperl, B., Seifert, M. H. & Berg, T. Natural product inhibitors of protein-protein interactions mediated by Src-family SH2 domains. Bioorg Med Chem Lett 19, 3305–3309, https://doi.org/10.1016/j.bmcl.2009.04.083 (2009).
Wang, L. et al. Discovery of antagonists for human scavenger receptor CD36 via an ELISA-like high-throughput screening assay. J Biomol Screen 15, 239–250, https://doi.org/10.1177/1087057109359686 (2010).
Baell, J. B. & Nissink, J. W. M. Seven Year Itch: Pan-Assay Interference Compounds (PAINS) in 2017-Utility and Limitations. ACS Chem Biol 13, 36–44, https://doi.org/10.1021/acschembio.7b00903 (2018).
Chen, X., Liu, M. X. & Yan, G. Y. Drug-target interaction prediction by random walk on the heterogeneous network. Mol Biosyst 8, 1970–1978, https://doi.org/10.1039/c2mb00002d (2012).
Chen, X. et al. Drug-target interaction prediction: databases, web servers and computational models. Brief Bioinform 17, 696–712, https://doi.org/10.1093/bib/bbv066 (2016).
Chen, X. et al. NLLSS: Predicting Synergistic Drug Combinations Based on Semi-supervised Learning. PLoS Comput Biol 12, e1004975, https://doi.org/10.1371/journal.pcbi.1004975 (2016).
Schrödinger Glide, version 6.9 (Schrödinger, LLC) (2015a).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25, https://doi.org/10.1016/j.softx.2015.06.001 (2015).
Wu, E. L. et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J Comput Chem 35, 1997–2004, https://doi.org/10.1002/jcc.23702 (2014).
Lee, J. et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J Chem Theory Comput 12, 405–413, https://doi.org/10.1021/acs.jctc.5b00935 (2016).
Traditional Chinese Medicine Database (TCMdb), Neotrident Co., Ltd., Beijing, Chinese edition, 2011.
Harder, E. et al. OPLS3: A Force Field Providing Broad Coverage of Drug-like Small Molecules and Proteins. J Chem Theory Comput 12, 281–296, https://doi.org/10.1021/acs.jctc.5b00864 (2016).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47, 1739–1749, https://doi.org/10.1021/jm0306430 (2004).
Friesner, R. A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49, 6177–6196, https://doi.org/10.1021/jm051256o (2006).
Halgren, T. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem 47, 1750–1759, https://doi.org/10.1021/jm030644s (2004).
Farid, R., Day, T., Friesner, R. A. & Pearlstein, R. A. New insights about HERG blockade obtained from protein modeling, potential energy mapping, and docking studies. Bioorgan Med Chem 14, 3160–3173, https://doi.org/10.1016/j.bmc.2005.12.032 (2006).
Sherman, W., Beard, H. S. & Farid, R. Use of an induced fit receptor structure in virtual screening. Chem Biol Drug Des 67, 83–84, https://doi.org/10.1111/j.1747-0285.2005.00327.x (2006).
Sherman, W., Day, T., Jacobson, M. P., Friesner, R. A. & Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J Med Chem 49, 534–553, https://doi.org/10.1021/jm050540c (2006).
Violin, J. D. et al. Selectively engaging beta-arrestins at the angiotensin II type 1 receptor reduces blood pressure and increases cardiac performance. J Pharmacol Exp Ther 335, 572–579, https://doi.org/10.1124/jpet.110.173005 (2010).
Rajagopal, S. et al. Quantifying Ligand Bias at Seven-Transmembrane Receptors. Mol Pharmacol 80, 367–377, https://doi.org/10.1124/mol.111.072801 (2011).
Acknowledgements
We would like to thank Dr. Bryan L. Roth (Univ. North Carolina at Chapel Hill), Dr. Xi-Ping Huang (Univ. North Carolina at Chapel Hill) and the National Institute of Mental Health’s Psychoactive Drug Screening Program (Contract #HHSN-271–2008–00025-C) for screening data. We thank Xiaoyan Liu, Dr. Yueming Xu, and Qiwen Tan for their suggestion and discussion on functional assays. We also thank Dr. Andrej Sali for his critical review of the manuscript. SZ was supported by ShanghaiTech University and the Natural Science Foundation of Shanghai (16ZR1448500). KAJ and ZGG were supported by NIH Intramural Research Program (NIDDK, ZIA DK031116).
Author information
Authors and Affiliations
Contributions
S.Z. and K.A.J. designed the study. X.L. and Z.-G.G. performed the dry and wet experiments respectively. Y.W. helped analyze the docking data. R.C.S. coordinated the assays. X.L., Z.-G.G., K.A.J. and S.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
R.C.S. is a founder and board member of Bird Rock Bio, a company focuses on GPCR therapeutic antibodies. The remaining authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, X., Gao, ZG., Wu, Y. et al. Salvianolic acids from antithrombotic Traditional Chinese Medicine Danshen are antagonists of human P2Y1 and P2Y12 receptors. Sci Rep 8, 8084 (2018). https://doi.org/10.1038/s41598-018-26577-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26577-0
This article is cited by
-
Pharmacological characterization of P2Y receptor subtypes – an update
Purinergic Signalling (2023)
-
G protein-coupled receptors: structure- and function-based drug discovery
Signal Transduction and Targeted Therapy (2021)
-
Targeting P2 receptors in purinergic signaling: a new strategy of active ingredients in traditional Chinese herbals for diseases treatment
Purinergic Signalling (2021)
-
Salvianolic acid a inhibits platelet activation and aggregation in patients with type 2 diabetes mellitus
BMC Cardiovascular Disorders (2020)
-
UPLC/MS-based untargeted metabolomics reveals the changes of metabolites profile of Salvia miltiorrhiza bunge during Sweating processing
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.