Abstract
Parasitic chytrid fungi have emerged as a significant threat to amphibian species worldwide, necessitating the development of techniques to isolate these pathogens into culture for research purposes. However, early methods of isolating chytrids from their hosts relied on killing amphibians. We modified a pre-existing protocol for isolating chytrids from infected animals to use toe clips and biopsies from toe webbing rather than euthanizing hosts, and distributed the protocol to researchers as part of the BiodivERsA project RACE; here called the RML protocol. In tandem, we developed a lethal procedure for isolating chytrids from tadpole mouthparts. Reviewing a database of use a decade after their inception, we find that these methods have been applied across 5 continents, 23 countries and in 62 amphibian species. Isolation of chytrids by the non-lethal RML protocol occured in 18% of attempts with 207 fungal isolates and three species of chytrid being recovered. Isolation of chytrids from tadpoles occured in 43% of attempts with 334 fungal isolates of one species (Batrachochytrium dendrobatidis) being recovered. Together, these methods have resulted in a significant reduction and refinement of our use of threatened amphibian species and have improved our ability to work with this group of emerging pathogens.
Similar content being viewed by others
Introduction
A major consequence of globalisation has been the increase of invasive species owing to trade in live animals and plants. A further outcome of this process is the concomitant rise of novel emerging fungal pathogens (EFPs1) as these infections are moved within trade networks and establish in uninfected regions – an example of fungal ‘pathogen pollution’2. Whilst EFPs can affect humans, they have also been broadly detrimental to natural populations of plants and animals, leading to worldwide losses of biodiversity. This dynamic has been most apparent across amphibians, where EFPs leading to population extirpation and species extinctions have contributed to amphibians now being the most endangered class of vertebrate3,4. In particular, emergence of parasitic fungi in the genus Batrachochytrium (phylum Chytridiomycota, order Rhizophydiales) have played a major role in driving amphibian population and species declines worldwide5,6.
While a single species, Batrachochytrium dendrobatidis (Bd), was originally thought to have caused the ongoing panzootic7, we now know that amphibian chytridiomycosis is caused by a much broader swathe of phylogenetic diversity than was previously thought8,9. Next-generation sequencing and phylogenomic analyses have shown that Bd sensu stricto is composed of deep genetic lineages which are emerging through international trade in amphibians10,11,12. Superimposed upon this background of trade-associated lineages of Bd has come the recent discovery of a new species of pathogenic chytrid, also within the Rhizophydiales, B. salamandrivorans13. This pathogen has rapidly extirpated European fire salamanders (Salamandra salamandra) in the Netherlands and a broad screening of urodeles has shown that Bsal occurs naturally in southeast Asia where it appears to asymptomatically infect salamander and newt species14.
The ability to isolate and culture both Bd and Bsal has played a key role in their discovery and by catalysing research into their pathogenesis and virulence15,16,17, phenotypic characteristics18,19,20 and a wealth of experimental studies on epidemiologically relevant parameters21,22,23. Longcore et al.24 first isolated Bd from infected amphibians by modifying techniques used to isolate other chytrids25. Longcore cleaned small (<0.5 mm dia) pieces of Bd-infected leg and foot skin by wiping them through agar and then placed skin pieces onto a clean plate of nutrient agar containing penicillin G and streptomycin. This method worked well for isolating from dead animals sent by courier from North and Central America. The method, however, requires euthanizing potentially healthy animals if their infection status was unknown. Further, it is difficult to perform these techniques in remote regions that lack suitable laboratory facilities, and the lethal sampling of amphibians may be contraindicated if the species is endangered, protected or located in protected areas.
We confronted this issue in a 2008–2014 project funded by BiodivERsA (http://www.biodiversa.org) – RACE: Risk Assessment of Chytridiomycosis to European amphibian biodiversity26. One of the objectives of this project was to adjust the original protocol of Longcore et al.24 to (i) reduce the need to kill adult amphibians, (ii) improve rates of chytrid isolation by allowing the use of more animals, (iii) develop protocols that enabled isolation in a field setting, and, (iv) integrate the data into the GPS-smartphone enabled epidemiological software application Epicollect27,28. Furthermore, ‘forewarned is forearmed’ and we wished to determine whether the protocol was able to isolate other species of chytrid that are members of the amphibian skin microbiota, and that may present a biosecurity risk. This need to more broadly characterise global chytrid biodiversity was met by using resources from RACE to train researchers worldwide in chytrid isolation techniques to provide opportunities to characterise novel chytrids as they were discovered.
In addition to the non-lethal isolation protocol, a lethal method was developed in parallel to isolate chytrids from the mouthparts of larval amphibians. We describe this method as a refinement to the main isolation protocol.
Methods
Non-lethal field isolation of chytrids
Animals were captured and held in separate plastic bags or suitable containers until ready for processing (Supp. Info. RML Protocol 1 and Supp. Info. Swabbing Protocol 2). Using clean gloves and sterilized dissection scissors or scalpel blades, the terminal 1–2 mm of the phalanges of the 4th hind toe (counting from the proximal toe) was clipped and laid onto the surface of an mTGhL + antibiotic (200 mg/L penicillin-G and 400 mg/L streptomycin sulphate) agar plate. Alternatively, ~1 mm toe-webbing biopsy punches were taken (Sklar instruments, PA, USA) then laid on a plate. This allowed multiple animals to be processed rapidly in the field. Subsequently, each tissue sample was transferred to a second plate with a sterile needle or forceps then cleaned (as far as possible) of surface-contaminating bacteria and fungi by dragging it through the agar-medium. The needle or forceps was then used to place the tissue sample into a sterile 2 ml screw-cap microtube containing liquid mTGhL medium with antibiotics (200 mg/L penicillin-G and 400 mg/L streptomycin sulphate), then stored in a cool, dry place. While 4 °C appears optimal, we have successfully used shaded regions of streams to cool cultures when refrigeration was not immediately available and have even held tubes and plates for several days at >10 °C until suitable storage conditions were available.
Once back in the laboratory, samples in tubes were visually screened for evidence of yeast or bacterial contamination (when the media takes on a ‘cloudy’ appearance), or mycelial ‘balls’ around the toe that are evidence of non-chytrid fungal contaminants. Visibly clear samples were decanted into a single well of a sterile 12-well lidded culture plate then incubated at 18 °C for up to 4 weeks, topping up with extra medium to counter evaporation as necessary. Depending on the size of the initial tissue sample, toe clips and webbing were divided into several smaller samples before transferring to liquid culture media.
Isolating chytrids from tadpoles
Tadpoles often have higher burdens of infection than adults, especially long-lived tadpoles29, and have higher densities and encounter rates than adults. As incomplete data exists as to which amphibian species raise tadpoles that are susceptible to Bd, in practice tadpoles from a range of species should be tested. Where tadpoles are large and infections heavy, tadpoles were microscopically prescreened with a dissecting microscope or hand lens in order to detect areas of depigmentation and hyperkeratosis within mouth parts, especially the jaw sheaths, that are associated with infection30,31,32,33 (Fig. 1). Tadpoles are euthanized using a humane method that does not affect the fungi (e.g., overdose of MS-222) before excising their mouthparts and these preliminary microscopic screens enabled us to use only a small number of animals to isolate chytrids. Additionally, uninfected and naïve tadpoles that were reared in captivity were used as live substrates to bait chytrids from adult amphibians with low levels of Bd infection34.
When using tadpoles to bait Bd from infected adult amphibians, tadpoles from a susceptible species such as Bombina orientalis34 were co-housed with an infected animal. Susceptible tadpoles were reared until gills were resorbed and animals were free-swimming and feeding (developmental Gosner stage 25), because at earlier stages they are still developing the keratinized mouthparts. Each tadpole container was then immersed within a similar but larger container that held at least one chytrid-infected animal. Water exchange between the infected and bait animal containers occurred through small holes (<0.3 mm) drilled into the bottom of the walls of the smaller internal containers. Animals were held in these conditions for between 2 and 4 weeks at species-appropriate conditions. Tadpoles were periodically examined every fourth day for the presence of the depigmented areas in the jaw sheaths that have been associated with chytrid infection.
Isolating chytrids from tadpoles first required killing by immersion in a 5 g/L solution of MS-22235 or other approved method. Note that anaesthetics that contain ethanol, such as phenoxyethanol36, should be avoided as these kill chytrids while MS222 is not toxic37. We then dissected out keratinized jaw sheaths and cleaned the entire sheath, or sections, as above using an agar plate with antibiotics (24; Supp. Info. RML Protocol 1). Cleaned sections were then placed singly into sterile 12-well culture plates with 1 mL liquid media + antibiotics, or onto agar plates with 6–10 sections per plate, and incubated at 10–20 °C.
Because zoospore release may occur immediately, especially from tadpole mouthparts, cultures were examined with an inverted microscope for the presence of active zoospores every day for up to one week following the day that they were initiated. After that, checks every two days were sufficient.
Culture and diagnosis of chytrid isolates
Subsequent culture methods for Bd followed those of Longcore et al.24. When isolation of Bsal was anticipated an incubation temperature of 15 °C was required38 whereas a temperature of 18–22 °C is closer to the measured growth optimum of Bd23,24. Once growth of zoospores and/or zoosporangia was observed, 100–500 µL volume of the culture was transferred by pipette to a new 12-well plate with liquid medium and no antibiotics, and incubated at 15–20 °C. All successfully cultured isolates were subcultured into larger volumes, then centrifuged at 1700 rpm for 10 min before cryopreservation. A portion of the initial pellet was also be used for DNA extraction, while the remaining volume was resuspended in 10% dimethyl sulfoxide (DMSO) and 10% fetal calf serum (FCS) in liquid media and transferred into six 2 mL cryotubes for cryopreservation at −80 °C39.
We confirmed the identity of Bd and Bsal by quantitative PCR with an MGB Taqman probe assay in either single-plex or multiplex40,41. We identified non-Batrachochytrium chytrids by sequencing appropriate regions of the ribosomal RNA gene with universal fungal primers followed by comparison against OTUs held in UNITE database (Unified system for DNA-based fungal species linked to classification: https://unite.ut.ee) to establish a species-hypothesis for the chytrid isolate in question42. If further genetic data were required, then multilocus analysis or whole-genome sequencing was undertaken using chytrid-specific methods1,7,16,43.
Collation of data
To track and report chytrid isolation for the RACE project, we used a generic data collection tool that allows the collection and submission of geotagged data forms from field locations, Epicollect5 (https://five.epicollect.net). This software has the advantage that it can be used on mobile devices with or without internet connection, and allows the immediate sharing of data across the research community. Our database at https://five.epicollect.net/project/bd-global-isolation-protocol included the following data fields: Date; Continent, Country, Site name; Latitude/Longitude; Wild caught or trade?; Amphibian species; Life history stage; Number sampled; Chytrid isolated?; Number isolated; Species of chytrid isolated; Chytrid lineage; Photograph of amphibian; Name of researchers.
All field-collection and application of protocols were performed in accordance with the relevant local guidelines, regulations and licensing. Experimental protocols were approved after ethical review by the Imperial College and Institute of Zoology ethical review committees and were performed under UK Home Office Project Licences held by MC Fisher and TWJ Garner.
Data accessibility
https://five.epicollect.net/project/bd-global-isolation-protocol.
Results
The ‘RACE modified Longcore (RML) Protocol’ for the non-lethal isolation of chytrids from amphibians is detailed in Supp. Info. 1. Researchers should ensure that they have the relevant licences, permits and permissions from ethical committees to follow the RML protocol 1, swabbing protocol 2 and isolation from larval amphibians.
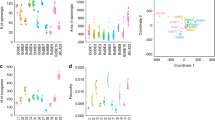
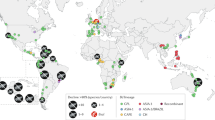
Following the formalisation and distribution of the RACE protocols, our Epicollect5 project summarised chytrid surveys from 2007 through to 2017 (Table 1). The Epicollect5 database can be spatially visualised at https://five.epicollect.net/project/bd-global-isolation-protocol/data. Figure 2 depicts the isolation of amphibian-associated chytrids using the RACE protocols from 5 continents (Africa, Asia, Australia, Europe and South America), 23 countries, 239 sampling episodes, and from latitudes spanning −44.1 S (Batrachyla antartandica, Chile) through to 55.6 N (Bufo viridis, Sweden). Chytrids have been non-lethally isolated from 1,906 animals comprising 34 amphibian species, of which 28 were anuran and 5 were caudatan species. Of the Bd isolated, 170 (80%) were determined to be BdGPL, 5 (2%) were BdCAPE, 34 (16%) were BdBRAZIL, 1 (>1%) was BdCH and 3 (1%) were hybrids. The database also contains 5 records of chytrids that were non-lethally sampled from the amphibian trade.
Worldwide distribution of sites where the RML Longcore protocol has been used to isolate chytrids. Numbers denote the quantity of amphibian species investigated. A browseable version of this Epicollect 5 map can be accessed at https://five.epicollect.net/project/bd-global-isolation-protocol. Tiles © Esri — Source: Esri, i-cubed, USDA, USGS, AEX, GeoEye, Getmapping, Aerogrid, IGN, IGP, UPR-EGP, and the GIS User Community and OpenStreetMap © OpenStreetMap.org contributors licence CC BY-SA (https://creativecommons.org/licenses/by-sa/2.0/).
Non-lethal isolation from adult and juvenile amphibians
In total, 1,152 animals were non-lethally sampled, recovering 207 chytrid isolates and resulting in a recovery rate of 18% (~1 isolate per 5 animals sampled). Of these chytrids, 203 (98%) were Bd, 2 were Rhizophydium sp., 2 were Kappamyces sp. and none were Bsal (Table 1). Of the Bd isolated, 42 (88%) were determined to be BdGPL, 5 (10%) were BdCAPE, and 1 (2%) was BdCH.
Isolation of chytrids from larval amphibians
In total, 784 tadpoles were sampled recovering 334 chytrid isolates and resulting in a recovery rate of 43% (~1 isolate per 2–3 animals sampled) (Table 2). Isolates were recovered from 34 species of amphibian, all of which were anurans. These chytrid isolates were all Bd and, of the lineages recorded, 128 (78%) were BdGPL, 34 (20%) were BdBRAZIL and 3 (2%) were hybrids.
Baiting chytrid isolates from live adult animals using tadpoles was used successfully in South Korean Bombina orientalis as previously described34. Here, six tadpoles were co-housed with adult B. orientalis, yielding a single isolate of Bd for each attempt equating to a rate of success of ~20%.
Discussion
The RML protocol, based on the original suggestions of Joyce Longcore for the non-lethal isolation of chytrids from amphibians, has been a success with isolates of chytrids recorded from five continents. There are likely many other unrecorded uses of this method because this protocol has been widely dispersed during the 5-year span (2008–2014) of the RACE project which trained a cohort of amphibian disease researchers in these techniques.
In some circumstances chytrids could not be recovered from toe-clips when sampling populations with persistent infection despite repeated attempts. This was particularly evident when the prevalence and burden of chytrid infections in surveys was low14,34,44 or when host species occupied habitats with high bacterial, non-target fungal contaminants, or both. In these situations we isolated chytrids from tadpole mouthparts as an associated method to the RML protocol. The value of the RML protocol in propelling forward research on amphibian chytridiomycosis has been very clear: for instance, of the 59 scientific papers produced by RACE, 15 directly used isolates of Bd that were generated by this protocol for experimental trials. Further, subsequently many more studies using these isolates have extended our knowledge of the genetic diversity of Bd7,8,43,45, the development of novel diagnostics46, the genetic repertoire that underpins the virulence of these pathogens16,17 and the biogeographic distributions of Bd diversity worldwide8,45.
Clearly some uncontrolled biases and unanswered questions in these studies need attention. First, the majority of Bd isolates belong to the BdGPL lineage. This could be because this lineage is more widespread (and therefore more readily recovered) than other lineages47. Alternatively, the intensity of BdGPL infections or its rate of zoospore production may be higher than for other lineages, which would also equate to a higher rate of isolation. To achieve a true and unbiased understanding of the distribution of these lineages, a lineage-specific diagnostic will need to be developed and deployed. Second, if lineage-specific differences in the probability of successful isolation exist, then mixed infections where these lineages co-occur may not be detected. This can be controlled for by isolating and genotyping many isolates from a single host and population, although this may not fully account for this bias. A related bias is that not all infectious species of chytrid will respond equally to culturing attempts. For instance, despite known attempts to isolate Bsal from across its endemic southeast Asian range using the protocol, to date no successful isolations of Bsal have been recorded. This is likely due to a combination of the low prevalence and burden of infection in salamanders and newts combined with the low initial growth-rate of Bsal13,14. With the RML protocol, however, workers have been able to isolate non-Bd species of chytrid (e.g., Kappamyces spp. and Rhizophydium sp. Table 1). This diversity likely represents only a fraction of the diversity of amphibian-associated chytrids that occur, and non-biased estimators of this diversity by, for instance, profiling the nuclear ribosomal RNA cistron42, are sorely needed.
In this age of the global amphibian crisis, research on the affects of chytrid infections is transitioning to attempts to mitigate their impacts48,49,50. Both of these research streams benefit from the availability of chytrid isolates, but the ethics behind these research programs can be improved. To that end, our data on isolation success suggest that tadpoles are a better target for isolation than metamorphosed animals. This is to some degree unfortunate, because isolation from tadpoles requires killing. However, we have outlined one refinement where captive reared tadpoles can be used to ‘bait’ infections from wild-caught amphibians to isolate chytrids without killing adult amphibians. Here, it is important to recognise that amphibians that have been co-housed in collections should not be returned to the wild owing to the danger of cross-transmission of pathogens during husbandry51. If it is necessary to isolate chytrids directly from wild tadpoles without using bait animals, we suggest that researchers focus on more fecund species with long larval periods as the focal species in aquatic amphibian communities. Removal of small numbers of tadpoles when clutch sizes are in the hundreds or thousands should ensure minimal ecological impact; for this reason sacrificing tadpoles is preferable to killing adult animals.
The extent to which toe-clipping affects the fitness of amphibians has been much debated52,53). Toe-clipping has been shown to decrease amphibian survival, but this effect, when present, is linearly related to the number of toes removed54,55. For the single toe-clip that the RML protocol requires, reduction in survival appears to be negligible53,56, and toe clipping is certainly preferred to killing the animal. Attention should be paid to this issue, however, and, where appropriate, survival estimates should be undertaken to determine the health implications of this procedure. Also, antiseptic and analgesic protocols can be considered to ensure that wounds where tissue samples are excised are at low risk of secondary infection57.
In summary, modification of Longcore’s original Bd-isolation protocol24 has enabled a broad community of scientists to engage with research on emerging chytrid pathogens of amphibians. This research has had an impact worldwide, and is contributing to the ongoing dialogue that is occurring among scientists, conservationists and policy-makers about how we might mitigate against these infections now and into the future58.
References
Farrer, R. A. & Fisher, M. C. D. Genomic and Epigenomic Traits Underpinning Emerging Fungal Pathogens. Adv Genet 100, 73–140, https://doi.org/10.1016/bs.adgen.2017.09.009 (2017).
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194, https://doi.org/10.1038/Nature10947 (2012).
Stuart, S. N. et al. Status and trends of amphibian declines and extinctions worldwide. Science 306, 1783–1786 (2004).
Mendelson, J. R. 3rd. et al. Biodiversity. Confronting amphibian declines and extinctions. Science 313, 48 (2006).
Berger, L. et al. Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America. Proceedings of the National Academy of Science, USA 95, 9031–9036 (1998).
Fisher, M. C., Garner, T. W. J. & Walker, S. F. Global Emergence of Batrachochytrium dendrobatidis and Amphibian Chytridiomycosis in Space, Time, and Host. Annual Review of Microbiology 63, 291–310 (2009).
James, T. Y. et al. Rapid expansion of an emerging fungal disease into declining and healthy amphibian populations. PLoS Pathogens 5, e1000458 (2009).
Farrer, R. A. et al. Multiple emergences of genetically diverse amphibian-infecting chytrids include a globalized hypervirulent recombinant lineage. P Natl Acad Sci USA 108, 18732–18736, https://doi.org/10.1073/Pnas.1111915108 (2011).
Schloegel, L. M. et al. Novel, panzootic and hybrid genotypes of amphibian chytridiomycosis associated with the bullfrog trade. Mol Ecol 21, 5162–5177, https://doi.org/10.1111/J.1365-294x.2012.05710.X (2012).
Schloegel, L. M. et al. The North American bullfrog as a reservoir for the spread of Batrachochytrium dendrobatidis in Brazil. Anim Conserv 13, 53–61, https://doi.org/10.1111/J.1469-1795.2009.00307.X (2010).
Schloegel, L. M. et al. Magnitude of the US trade in amphibians and presence of Batrachochytrium dendrobatidis and ranavirus infection in imported North American bullfrogs (Rana catesbeiana). Biol Conserv 142, 1420–1426, https://doi.org/10.1016/j.biocon.2009.02.007 (2009).
Fisher, M. C. & Garner, T. W. J. The relationship between the introduction of Batrachochytrium dendrobatidis, the international trade in amphibians and introduced amphibian species. Fungal Biology Reviews 21, 2–9 (2007).
Martel, A. et al. Batrachochytrium salamandrivorans sp nov causes lethal chytridiomycosis in amphibians. P Natl Acad Sci USA 110, 15325–15329, https://doi.org/10.1073/Pnas.1307356110 (2013).
Laking, A. E., Ngo, H. N., Pasmans, F., Martel, A. & Nguyen, T. T. Batrachochytrium salamandrivorans is the predominant chytrid fungus in Vietnamese salamanders. Sci Rep 7, https://doi.org/10.1038/srep44443 (2017).
Voyles, J. et al. Electrolyte depletion and osmotic imbalance in amphibians with chytridiomycosis. Dis Aquat Organ 77, 113–118 (2007).
Farrer, R. A. et al. Genomic innovations linked to infection strategies across emerging pathogenic chytrid fungi. Nat Commun 8, 14742, https://doi.org/10.1038/ncomms14742 (2017).
Rosenblum, E. B., Poorten, T. J., Joneson, S. & Settles, M. Substrate-Specific Gene Expression in Batrachochytrium dendrobatidis, the Chytrid Pathogen of Amphibians. Plos One 7, https://doi.org/10.1371/journal.pone.0049924 (2012).
Piotrowski, J. S., Annis, S. L. & Longcore, J. E. Physiology of Batrachochytrium dendrobatidis, a chytrid pathogen of amphibians. Mycologia 96, 9–15 (2004).
Fisher, M. C. et al. Proteomic and phenotypic profiling of the amphibian pathogen Batrachochytrium dendrobatidis shows that genotype is linked to virulence. Mol Ecol 18, 415–429 (2009).
Becker, C. G. et al. Variation in phenotype and virulence among enzootic and panzootic amphibian chytrid lineages. Fungal Ecol 26, 45–50, https://doi.org/10.1016/j.funeco.2016.11.007 (2017).
Garner, T. W. et al. Life history trade-offs influence mortality associated with the amphibian pathogen Batrachochytrium dendrobatidis. Oikos, 783–791 (2009).
Rosenblum, E. B., Poorten, T. J., Settles, M. & Murdoch, G. K. Only skin deep: shared genetic response to the deadly chytrid fungus in susceptible frog species. Mol Ecol 21, 3110–3120, https://doi.org/10.1111/j.1365-294X.2012.05481.x (2012).
Ribas, L. et al. Expression Profiling the Temperature-Dependent Amphibian Response to Infection by Batrachochytrium dendrobatidis. Plos One 4, https://doi.org/10.1371/journal.pone.0008408 (2009).
Longcore, J. E., Pessier, A. P. & Nichols, D. K. Batrachochytrium dendrobatidis gen et sp nov, a chytrid pathogenic to amphibians. Mycologia 91, 219–227 (1999).
Barr, D. J. S. In Zoosporic fungi in teaching and research (eds M.S. Fuller & A. Jaworski) p 118–120 (Southeastern Publishing Corp., 1987).
Fisher, M. C. et al. RACE: Risk Assessment of Chytridiomycosis to European Amphibian Biodiversity. Froglog, 45–47 (2012).
Aanensen, D. M., Huntley, D. M., Feil, E. J., al-Own, F. & Spratt, B. G. EpiCollect: Linking Smartphones to Web Applications for Epidemiology, Ecology and Community Data Collection. Plos One 4, https://doi.org/10.1371/journal.pone.0006968 (2009).
Aanensen, D. M., Huntley, D. M., Menegazzo, M., Powell, C. I. & Spratt, B. G. EpiCollect+: linking smartphones to web applications for complex data collection projects. F1000Res 3, 199, https://doi.org/10.12688/f1000research.4702.1 (2014).
Skerratt, L. F. et al. Survey protocol for detecting chytridiomycosis in all Australian frog populations. Dis Aquat Organ 80, 85–94 (2008).
Fellers, G. M., Green, D. E. & Longcore, J. E. Oral chytridiomycosis in the mountain yellow-legged frog (Rana muscosa). Copeia, 945–953, https://doi.org/10.1643/0045-8511(2001)001[0945:Ocitmy]2.0.Co;2 (2001).
Smith, K. G. & Weldon, C. A. Conceptual Framework for Detecting Oral Chytridiomycosis in Tadpoles. Copeia 4, 1024–1028 (2007).
Navarro-Lozano, A., Sanchez-Domene, D., Rossa-Feres, D. C., Bosch, J. & Sawaya, R. J. Are oral deformities in tadpoles accurate indicators of anuran chytridiomycosis? Plos One 13, e0190955, https://doi.org/10.1371/journal.pone.0190955 (2018).
Marantelli, G., Berger, L., Speare, R. & Keegan, L. Changes in distribution of Batrachochytrium dendrobatidis and keratin during tadpole development leading to high mortality after metamorphosis. Pacific Conservation Biology 10, 173–179 (2004).
Bataille, A. et al. Genetic evidence for a high diversity and wide distribution of endemic strains of the pathogenic chytrid fungus Batrachochytrium dendrobatidis in wild Asian amphibians. Mol Ecol 22, 4196–4209, https://doi.org/10.1111/mec.12385 (2013).
Torreilles, S. L., McClure, D. E. & Green, S. L. Evaluation and Refinement of Euthanasia Methods for Xenopus laevis. J Am Assoc Lab Anim 48, 512–516 (2009).
Gentz, E. J. Medicine and surgery of amphibians. ILAR J 255–259 (2007).
Webb, R., Berger, L., Mendez, D. & Speare, R. MS-222 (tricaine methane sulfonate) does not kill the amphibian chytrid fungus Batrachochytrium dendrobatidis. Dis Aquat Organ 68, 89–90 (2005).
Blooi, M. et al. Treatment of urodelans based on temperature dependent infection dynamics of Batrachochytrium salamandrivorans. Sci Rep-Uk 5, https://doi.org/10.1038/srep08037 (2015).
Boyle, D. G. et al. Cryo-archiving of Batrachochytrium dendrobatidis and other chytridiomycetes. Dis Aquat Organ 56, 59–64 (2003).
Boyle, D. G., Boyle, D. B., Olsen, V., Morgan, J. A. T. & Hyatt, A. D. Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real-time Taqman PCR assay. Dis Aquat Organ 60, 141–148 (2004).
Blooi, M. et al. Duplex Real-Time PCR for rapid simultaneous detection of Batrachochytrium dendrobatidis and Batrachochytrium salamandrivorans in amphibian samples. J Clin Microbiol 51, 4173–4177, https://doi.org/10.1128/Jcm.02313-13 (2013).
Schoch, C. L. et al. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. P Natl Acad Sci USA 109, 6241–6246, https://doi.org/10.1073/pnas.1117018109 (2012).
Farrer, R. A. et al. Chromosomal Copy Number Variation, selection and uneven rates of recombination reveal cryptic genome diversity linked to pathogenicity. Plos Genet 9, https://doi.org/10.1371/Journal.Pgen.1003703 (2013).
Swei, A. et al. Is Chytridiomycosis an Emerging Infectious Disease in Asia? Plos One 6, https://doi.org/10.1371/journal.pone.0023179 (2011).
Jenkinson, T. S. et al. Amphibian-killing chytrid in Brazil comprises both locally endemic and globally expanding populations. Mol Ecol 25, 2978–2996, https://doi.org/10.1111/mec.13599 (2016).
Dillon, M. J. et al. Tracking the amphibian pathogens Batrachochytrium dendrobatidis and Batrachochytrium salamandrivorans using a highly specific monoclonal antibody and lateral- flow technology. Microbial. Biotechnology 10, 381–394, https://doi.org/10.1111/1751-7915.12464 (2017).
James, T. Y. et al. Disentangling host, pathogen, and environmental determinants of a recently emerged wildlife disease: lessons from the first 15 years of amphibian chytridiomycosis research. Ecol Evol 5, 4079–4097, https://doi.org/10.1002/ece3.1672 (2015).
Schmeller, D. S. et al. Microscopic aquatic predators strongly affect infection dynamics of a globally emerged pathogen. Curr Biol 24, 176–180, https://doi.org/10.1016/J.Cub.2013.11.032 (2014).
Garner, T. W. et al. Mitigating amphibian chytridiomycoses in nature. Philos Trans R Soc Lond B Biol Sci 371, https://doi.org/10.1098/rstb.2016.0207 (2016).
Canessa, S. et al. Decision making for mitigating emerging wildlife diseases: from theory to practice for an emerging fungal pathogen of salamanders. Journal of Applied Ecology, https://doi.org/10.1111/1365-2664.13089 (2018).
Walker, S. et al. Invasive pathogens threaten species recovery programs. Curr Biol 18, R853–R854 (2008).
May, R. M. Ecology - Ethics and amphibians. Nature 431, 403–403, https://doi.org/10.1038/431403a (2004).
Funk, W. C., Donnelly, M. A. & Lips, K. R. Alternative views of amphibian toe-clipping. Nature 433, 193–193, https://doi.org/10.1038/433193c (2005).
McCarthy, M. A. & Parris, K. M. Clarifying the effect of toe clipping on frogs with Bayesian statistics. Journal of Applied Ecology 41, 780–786, https://doi.org/10.1111/j.0021-8901.2004.00919.x (2004).
Grafe, T. U., Stewart, M. M., Lampert, K. P. & Rödel, M.-O. Putting Toe Clipping into Perspective: A Viable Method for Marking Anurans. J Herpetol 45, 28–35 (2011).
Ott, J. A. & Scott, D. E. Effects of toe-clipping and PIT-tagging on growth and survival in metamorphic Ambystoma opacum. J Herpetol 33, 344–348, https://doi.org/10.2307/1565740 (1999).
Chevalier, H. et al. Marking techniques in the Marbled Newt (Triturus marmoratus): PIT-Tag and tracking device implant protocols. Acta Herpetol 12, 79–88 (2017).
O'Hanlon, S. J. et al. Recent Asian origin of chytrid fungi causing global amphibian declines. Science, https://doi.org/10.1126/science.aar1965 (2018).
Acknowledgements
T.W.J.G., M.C.F., D.S.S., A.L., E.C., F.C.C., J.B., A.A.C., C.M., F.S., B.R.S., S.O., were supported through the Biodiversa project RACE: Risk Assessment of Chytridiomycosis to European Amphibian Biodiversity (NERC standard grant NE/K014455/1 and NE/E006701/1; ANR-08-BDVA-002-03). M.C.F., J.S., C.W., P.G. were supported by the Leverhulme Trust (RPG-2014-273), M.C.F., A.C., C.W. were supported by the Morris Animal Foundation. J.V. was supported by the Bolyai János Research Grant of the Hunagrian Academy of Sciences (BO/00597/14). F.G. and D.G. were supported by the Conservation Leadership Programme Future Conservationist Award. C.S.A. was supported by Fondecyt (No. 1181758). M.C.F. and A.C. were supported by. Mohamed bin Zayed Species Conservation Fund Project (152510704). GMR held a doctoral scholarship (SFRH/BD/69194/2010) from Fundação para a Ciência e a Tecnologia. L.F.T., C.L., L.P.R. K.R.Z., T.Y.J., T.S.J. were supported by São Paulo Research Foundation (FAPESP #2016/25358-3), the National Counsel of Technological and Scientific Development (CNPq #300896/2016–6) and a Catalyzing New International Collaborations grant from the United States NSF (OISE-1159513). C.S.A. was supported by Fondecyt (No. 1181758). T.M.D. was supported by the Royal Geographical Society and the Royal Zoological Society of Scotland. B.W. was supported by the National Research Foundation of Korea (2015R1D1A1A01057282).
Author information
Authors and Affiliations
Contributions
M.C.F. and T.W.J.G. wrote the main manuscript text. P.G., J.M.G.S., K.B., L.B., C.W., G.M.R., R.A.F., D.M.A., M.A.-R., A.B., L.B., S.B., J.B., F.C.C., E.C., A.C., A.A.C., T.M.D.-B., F.G., D.J.G., J.H., T.Y.J., T.S.J., T.A.K., C.L., A.L., C.-F.L., A.L., A.M., S.M., C.M., P.M., S.N., S.O., F.P., T.R., F.C.E.R., L.P.R., D.S.S., B.R.S., L.S., F.S., C.S.-A., G.T., L.F.T., A.V.-S., R.V., J.V., B.W., R.J.W., C.W., E.W., K.R.Z., J.E.L. contributed data and input to the manuscript text.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fisher, M.C., Ghosh, P., Shelton, J.M.G. et al. Development and worldwide use of non-lethal, and minimal population-level impact, protocols for the isolation of amphibian chytrid fungi. Sci Rep 8, 7772 (2018). https://doi.org/10.1038/s41598-018-24472-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-24472-2
This article is cited by
-
Chytridiomycosis in Asian Amphibians, a Global Resource for Batrachochytrium dendrobatidis (Bd) Research
Journal of the Indian Institute of Science (2021)
-
Early-diverging fungal phyla: taxonomy, species concept, ecology, distribution, anthropogenic impact, and novel phylogenetic proposals
Fungal Diversity (2021)
-
Presence of low virulence chytrid fungi could protect European amphibians from more deadly strains
Nature Communications (2020)
-
Global parasite trafficking: Asian Gyrodactylus (Monogenea) arrived to the U.S.A. via invasive fish Misgurnus anguillicaudatus as a threat to amphibians
Biological Invasions (2020)
-
Bullfrog farms release virulent zoospores of the frog-killing fungus into the natural environment
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.