Abstract
Plac1 is an X-linked trophoblast gene expressed at high levels in the placenta, but not in adult somatic tissues other than the testis. Plac1 however is re-expressed in several solid tumors and in most human cancer cell lines. To explore the role of Plac1 in cancer progression, Plac1 was reduced by RNA interference in EO771 mammary carcinoma cells. EO771 “knockdown” (KD) resulted in 50% reduction in proliferation in vitro and impaired tumor growth in syngeneic mice; however, tumor growth in SCID mice was equivalent to tumor cells expressing a non-silencing control RNA, suggesting that Plac1 regulated adaptive immunity. Gene expression profiling of Plac1 KD cells indicated reduction in several inflammatory and immune factors, including Cxcl1, Ccl5, Ly6a/Sca-1, Ly6c and Lif. Treatment of mice engrafted with wild-type EO771 cells with a Cxcr2 antagonist impaired tumor growth, reduced myeloid-derived suppressor cells and regulatory T cells, while increasing macrophages, dendritic cells, NK cells and the penetration of CD8+ T cells into the tumor bed. Cxcl1 KD phenocopied the effects of Plac1 KD on tumor growth, and overexpression of Cxcl1 partially rescued Plac1 KD cells. These results reveal that Plac1 modulates a tolerogenic tumor microenvironment in part by modulating the chemokine axis.
Similar content being viewed by others
Introduction
Placental-specific protein 1 (Plac1) is an Xq26-linked gene that encodes a microvillous membrane protein expressed primarily in trophoblasts, at low levels in the testis, but not in other adult somatic tissues1, and has the most restricted normal tissue expression pattern in comparison to other cancer/testis antigens2. Silva first reported that Plac1 RNA was expressed over a 4-log range in >50% of human cancer cell lines covering 17 different malignancies2, suggesting that some cancers mirror an onco-placental disease or a “somatic cell pregnancy”3. This hypothesis has been confirmed by the detection of Plac1 in malignancies of the breast4,5,6, endometrium7, ovary7, lung2,8, liver9, colon6,10,11, stomach12 and prostate13. In colorectal cancer biopsies, higher levels of Plac1 were detected in 50% of stage III/IV disease in comparison to early stage disease9,10, and Plac1-dependent cytotoxic T cell (CTL) activity correlated with overall survival11.
In the MMTV-PPARd transgenic model of luminal B breast cancer, Plac1 expression was highly elevated at the onset and throughout mammary tumorigenesis14, suggesting that it might have a role in the initiation and progression of tumor development. Previous studies found that Plac1 transcription in human breast cancer cells was regulated by many of the same co-activators associated with PPARd and other nuclear receptors15,16,17, including C/EBPβ and NCOA318,19, both of which have been implicated in breast cancer progression16,20,21,22. Despite these findings, little is known about the oncogenic processes downstream of Plac1. To address this question, EO771 mammary carcinoma cells, which express high levels of Plac1, were used to examine gene expression and signaling pathways under the control of Plac1. Our findings reveal that Plac1 regulates a chemokine and immune tolerogenic signaling network necessary for sustaining tumor growth, which suggests potential therapeutic strategies that could alter the tumor microenvironment to make it more amenable to therapy.
Results
Reduction of Plac1 inhibits EO771 cell growth and tumor formation
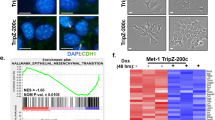
To characterize the functional role of Plac1, several mouse mammary tumor cell lines were screened by qRT-PCR for Plac1 RNA expression; among these, EO771 cells expressed the highest level, which was substantial in comparison to mouse placenta (Fig. 1a). EO771 cells were then transduced with recombinant lentiviruses expressing shRNAs targeting four regions of Plac1 mRNA (Fig. 1b). shRNA490 produced >98% reduction of Plac1 expression, and EO771 cells transduced with this shRNA (EO771/shPlac1) were used for further studies. EO771/shPlac1 cells grew in monolayer culture at 50% of the rate of control cells expressing a non-silencing RNA (Fig. 1c). Gene expression profiling revealed that Plac1 markedly suppressed several chemokine genes, including Cxcl1, Ccl7, Ccl2, Ccl5 and Cxcl10, as well as immune-related factors Lif, Ly6a/Sca-1, Ly6c and CD274 (Table 1, Fig. 1d, Supplementary Table 2). Changes in the expression of several of these genes were confirmed by qRT-PCR and most were consistent with the array profile (Fig. 1e).
Plac1 expression and lentivirus-mediated reduction of Plac1 in EO771 cells. (a) EO771 mouse mammary tumor cells expressed high levels of Plac1 in comparison to mouse placenta. (b) EO771 cells were transduced with lentiviruses expressing crambled RNA (Scr) or four Plac1 shRNAs designated sh81, sh187, sh300 and sh490; sh490 inhibited RNA expression >98%, and these cells were designated EO771/shPlac1. (c) EO771/Scr and EO771/shPlac1 cells were grown as monolayers, and the number of viable cells were quantified by sulforhodamine B staining. Shown is the mean ± S.D. of triplicate analysis of three samples. The growth of EO771/shPlac1 cells differed significantly (P < 0.001) from EO771/Scr cells by the two-sided Student’s t test. (d) qRT-PCR analysis of immune cell-related gene expression downregulated in EO771/shPlac1 cells. Shown is the mean ± S.D. of triplicate analysis of three samples. Significant differences between EO771/Scr and EO771/shPlac1 cells were obtained for CD274 (P < 0.01), Plac1 (P < 0.01), Cxcl1 (P < 0.001), Ccl5 (P < 0.001) and Lif (P < 0.001) using the two-tailed Student’s t test; values for Ccl7 were not significantly different (P > 0.05). (e) Heatmap of gene expression as determined by Affymetrix microarray analysis of EO771/Scr (Ctl) and EO771/shPlac1 (sh) cells. Shown are immune cell-related transcripts (Table 1) representing ≥3.0-fold change in expression.
Plac1 reduction blocks tumor growth in syngeneic mice, but not in SCID mice
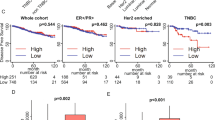
To determine the influence of Plac1 on tumor development in a host, EO771/shPlac1 cells were implanted in syngeneic C57BL/6 mice, and growth monitored by caliper measurement (Fig. 2a). EO771/shPlac1 isografts grew transiently in syngeneic mice, but when transplanted into SCID mice, their growth was similar to EO771/Scr control cells (Fig. 2b). Although all control isografts growing in syngeneic mice after 21 days expressed Plac1, there was little residual staining in mammary tissue from EO771/shPlac1-engrafted mice (Fig. 2c). Although Plac1 reduced growth in vitro, the lack of sustained growth of EO771/shPlac1 cells in syngeneic mice suggested that interactions with the tumor microenvironment may have contributed to this effect.
Growth of EO771/Scr and EO771/shPlac1 cells in syngeneic and SCID mice. (a) Syngeneic C57BL/6 mice or (b) SCID mice at five weeks of age, were inoculated in the mammary gland with 1 × 106 cells. Tumor size was measured by calipers in two dimensions. Tumor growth for EO771/Scr and EO771/shPlac1 cells in syngeneic mice differed significantly (P = 0.040) by the unpaired Student’s t test. There was no significant difference (P > 0.05) in tumor growth between the two cell lines in SCID mice. Shown is the mean ± SD, N = 5 per group. (c) H&E staining and Plac1 IHC in isografts of EO771/Scr and EO771/shPlac1 cells. Magnification 400X.
Cxcr2 antagonist inhibits tumor growth
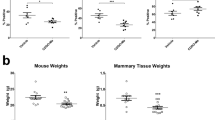
Since Cxcl1 exhibited the greatest change in expression among all chemokine genes after Plac1 downregulation (Table 1), mice were treated with a Cxcr2 antagonist to determine if it would affect tumor growth to a similar extent. Animals engrafted with EO771 cells were treated three times per week with vehicle or 2 or 20 mg/kg SB225002 beginning 10 days after transplantation (Fig. 3a). Whereas, the lower dose partially inhibited tumor growth, 20 mg/kg SB225002 completely blocked growth after a lag of two weeks. Tumor stasis at 17 days following cell inoculation was associated with reduction in expression of immune and chemokine genes, many of which were downregulated in EO771/shPlac1 cells (Fig. 3b). Cell sorting of tumor immune infiltrates indicated that SB225002 treatment reduced myeloid-derived suppressor cells (MDSC) and Treg cells, and increased CD8+/CD4+ T cells, NK cells, macrophages and dendritic cells (Fig. 3c,d). Especially noteworthy was the greater infiltration of CD8+ T cells into the tumor bed following SB25002 treatment (Fig. 3e), which was accompanied by increased macrophage and Treg cell infiltration, reduction of Plac1 and increased apoptosis (Fig. 3f). Since we did not determine tissue levels of SB25002, we determined its cytotoxicity in EO771 cell culture (Supplementary Fig. 2). SB25002 at concentration less than 100 nM were not cytotoxic, but produced cytotoxicity at concentrations exceeding 1000 nM. Since we have not carried out pharmacokinetics, the contribution of SB25002 cytotoxicity to its antitumor effect cannot be ascertained.
Growth of EO771 cells in syngeneic mice following treatment with a Cxcr2 antagonist. (a) Syngeneic 57BL/6 mice were inoculated in the mammary gland with 1 × 106 at five weeks of age, and injected i.p. daily with vehicle (blue) or 2 mg/kg (red) or 20 mg/kg (green) SB225002 beginning 11 days after cell inoculation. SB225002 completely suppressed tumor growth after 14 days. Differences between vehicle- and 2 mg/kg SB225002-treated mice were not significantly different (P = 0.145); differences between vehicle- and 20 mg/kg SB225002-treated mice were significantly different (P = 0.005) by the unpaired two-tailed Student’s t test. Shown is the mean ± SD, N = 5 per group. (b) Immune gene expression in tumors 17 days after treatment with 20 mg/kg SB225002. Shown is the relative expression in control and SB225002-treated mice in comparison to their changes in EO771/shPlac1 cells (Table 1). (c) FACS analysis of immune cell tumor infiltrates in isografts after treatment with vehicle or SB225002 as in (b). SB225002 treatment reduced the percentage of immune cell tumor infiltrates of CD11b+/Gr-1+ myeloid-derived suppressor cells (MDSC) and Foxp3+/CD25+ T cells (Treg), and increased the percentages of CD8+/CD4+ T cells (T), CD3+/NK1.1+ NK cells (NK) and F4/80+/CD80/86+ macrophages (Mϕ) and CD11c+/CD80/86+ dendritic cells (DC). Numbers in parentheses ( ) represent the percentages of each cell population. (d) Bar graph represents the mean±SD of the percent distribution of immune cell tumor infiltrates as in (c); P values were determined by the unpaired two-tailed Student’s t test, N = 4 per group. (e) CD8+ T cell infiltration determined by IHC in tumor isografts from vehicle-treated (EO771/Ctl) and SB225002-treated (EO771/SB) mice. Infiltration of CD8+ T cells increased after treatment with 20 mg/kg SB225002. Magnification 600X. (f) Macrophage (F4/80) and Treg cell (Foxp3) infiltration, Plac1 expression and apoptosis by cleaved caspase-3 expression (Caspase) in tumor isografts from vehicle-treated (EO771/Ctl) and SB225002-treated (EO771/SB) mice. Infiltration of macrophages and Treg cells were reduced and apoptosis was increased after treatment with 20 mg/kg SB225002. Magnification 400X
Cxcl1 reduction inhibits tumor growth and immune cell-related transcription
To evaluate the role of Cxcl1/Cxcr2 signaling in tumor growth, EO771 cells were transduced with scrambled shRNA (Scr) or four Cxcl1 shRNAs (Fig. 4a). EO771 cells expressing sh174 (shCxcl1) exhibited >95% reduction of Cxcl1 RNA expression. After 48 hr in monolayer culture, EO771/shCxcl1 cells grew at approximately 30% of the rate of control cells (Scr) (Fig. 4b). Comparison of the gene expression profile of EO771/shCxcl1 cells with EO771/Scr cells revealed a small subset of genes with ≥3-fold changes, including Ly6a, IL23a, C3, Cxcl1 and CD68 (Table 2, Supplementary Table 3). Transplantation of EO771/shCxcl1 cells into syngeneic mice resulted in impaired tumor growth in comparison to control cells (Fig. 4c), a result that was similar to EO771/shPlac1 cells (Fig. 2a). Changes in immune-related gene expression (Table 2) were confirmed by qRT-PCR, with the exception of CD68, which did not change significantly (Fig. 4d). Measurement of Cxcl1 in tumors or mammary tissue by IHC after 21days indicated the presence of Cxcl1 in EO771/Scr tumors, but not in EO771/shCxcl1 engrafted mammary tissue (Fig. 4e). Comparison of gene expression in EO771/shPlac1 vs. EO771/shCxcl1 cells indicated reduced expression in five genes in common, viz. CD68, Cxcl1, Ly6a, Plau and Rgs16 (Table 3), although CD68 was not changed significantly in EO771/shCxcl1 cells as measured by qRT-PCR (Fig. 4d).
Lentivirus-mediated reduction of Cxcl1 in EO771 cells. (a) EO771 cells were transduced with lentiviruses expressing scrambled RNA (Scr) or three Cxcl1 shRNAs designated sh118, sh174, sh218; sh174 inhibited RNA expression >99% (EO771/shCxcl1). (b) EO771/Scr and EO771/shCxcl1 cells were grown as monolayers, and the number of viable cells were determined by sulforhodamine B staining. Shown is the mean ± S.D. of triplicate analysis from three samples, which were significantly different (P < 0.001) by the two-tailed Student’s t test. (c) Growth of EO771/Scr and EO771/shCxcl1 cells in syngeneic mice. Mice at five weeks of age were inoculated in the mammary gland with 1x106 cells, and tumor size was measured by calipers in two dimensions. Differences in tumor growth between EO771/Scr and EO771/shCxcl1 cells were significantly different (P = 0.006) by the unpaired two-tailed Student’s t test; N = 5. (d) qRT-PCR analysis of genes downregulated in EO771/shCxcl1 cells. Shown is the mean ± SD of triplicate analysis of 3 samples. Significant differences between EO771/Scr and EO771/shCxcl1 cells were obtained for Plau (P < 0.02), C3 (P < 0.01), Ly6a (P < 0.01), Ccl7 (P < 0.001) and Il23a (P < 0.01) by the two-sided Student’s t test; differences for CD68 were not significantly different (P > 0.05).
Cxcl1 partially rescues Plac1 reduction in EO771 cells
To determine the contribution of Cxcl1 to the effects of Plac1 downregulation on tumor growth, EOI771/sh490 cells were transduced with a retrovirus expressing Cxcl1-mCherry and EO771/Scr and EO771/sh490 were transduced with mCherry alone (Fig. 5). After selection in G418, a significant percentage of cells co- expressed GFP and mCherry (Fig. 5a) and Cxcl1 mRNA (Fig. 5b). The growth of EO771/sh490 cells in vitro was slower rate than control cells as shown in Fig. 1c, but cells expressing Cxcl1 largely rescued this effect (Fig. 5c). Isografts of these cell lines in syngeneic mice confirmed the poor growth of EO771/sh490 cells, and further showed that Cxcl1 could partially rescue their poor tumorigenicity (Fig. 5d).
Cxcl1 rescue of EO771/sh490 cells. (a) EO771/Scr and EO771/sh490 cells expressing eGFP were transduced with a lentivirus expressing Cxcl1 and mCherry, and selected for 35 days in 3.5 mg/ml G418. The merged photo shows cells co-expressing eGFP and mCherry (yellow). Magnification 200X. (b) qRT-PCR for Plac1 and Cxcl1 in EO771/Scr, EO771/sh490 and EO771/sh490/Cxcl1 cells. Shown is the mean ± S.D. of triplicate determinations.(c) EO771/sh490/Cxcl1 cells were grown in 96-well plates at an initial density of 5,000 cells per well in media supplemented with 3.5 mg/ml G418. Cell density was determined by sulforhodamine B staining. Shown is the mean ± SD of triplicate determinations. (d) Syngeneic C57BL/6 mice were inoculated in the mammary gland with 1 × 106 at five weeks of age. There was a significant difference in the growth EO771/sh490 cells (P = 0.021) and EO771/sh490/Cxcl1 cells (P = 0.034) vs. EO771/Scr cells by the unpaired two-tailed Student’s t test. Shown is the mean ± SD, N=6 per group.
As an added proof of the function of Plac1 in tumorigenesis, MC cells, a cell line with low Plac1 expression (Fig. 1a), were transfected with Plac1 (Supplementary Fig. 3, Supplementary Table 5). MC/Plac1 cells grew at a higher rate than control cells and exhibited less apoptosis, and upregulated a gene expression profile that included several of the chemokines as well as CD274 that were downregulated in EO771/shPlac1 cells.
Discussion
The present study establishes the first link between the trophoblast gene, Plac1, and adaptive immunity, through its ability to modulate chemokine expression and other immune cell regulators. In retrospect, this is not too surprising since the placenta may be regarded as a foreign allograft protected from host vs. graft rejection23 in part by the presence of Treg cells in the uterine decidua24. However, the link between Plac1, chemokine signaling and immune tolerance in tumors is a novel and relevant finding since the latter processes are hallmarks of most, if not all, solid tumors25,26. The relevance of Plac1 to mammary tumorigenesis was first noted in MMTV-PPARd mice, where Plac1 was markedly upregulated at the onset and throughout tumor development14. This finding implicated nuclear receptor signaling in the transcriptional regulation of Plac1, as noted previously for its activation at alternate promoter regions in the Plac1 locus by LXR and RXRA27. However, from a mechanistic perspective, the downstream intracellular components interacting with Plac1have not been determined. Plac1 is predominantly extracellular with an N-terminal signal peptide, a small transmembrane domain and an extracellular ZP3 domain28, which promotes protein-protein interactions. By analogy, cytokine receptors lacking an intracellular signaling domain partner with co-receptors, adapter molecules and cytosolic protein tyrosine kinases to effect signaling29, and such a mechanism may also pertain to Plac1.
In the present study, the panel of immune-related genes down-regulated by Plac1 ‘knockdown’ (Table 1) suggest that one of its functions is to modulate chemokine effector pathways associated with immune evasion, such as antigen presentation, angiogenesis and myeloid cell, T cell and fibroblast activation (see scheme in Supplementary Fig. 4). One dominant downstream effector pathway was the Cxcl1/Cxcr2 axis, as shown by inhibition of tumor growth by the Cxcr2 antagonist SB25002 (Fig. 3a), and the inability of EO771/shCxcl1 cells to sustain tumor proliferation in syngeneic mice (Fig. 4c). Inhibition of Cxcr2 by SB25002 was associated with reduction of Treg cells and MDSC, and an increasing the percentage of CD8+ T cells, macrophages, NK cells and dendritic cells, which was consistent with previous studies demonstrating the ability of SB225002 to suppress MDSC infiltration in breast tumor xenografts30 and prostate tumors31, as well as metastasis via S100A8/A930. Thus, our data suggest that one role of Plac1 may be to maintain the production of inflammatory and immunoregulatory chemokines to effect changes in the stromal microenvironment conducive to immune tolerance and poor outcome30,32,33, which would explain the poor tumorigenicity of EO771/shPlac1 cells in syngeneic mice, but not in SCID mice. Although, co-expression of Plac1 and Cxcl1 in breast cancer tissue has not been reported, Cxcl1 expression in breast cancer biopsies was found to be elevated in metastases, and inversely related to ERα expression and relapse-free survival34.
Despite focusing on the immunological aspects of Plac1 function, it was apparent that it affected several signaling pathways in EO771 cells (Supplementary Table 2). Changes in gene expression common to both EO771/shPlac1 and EO771/shCxcl1 cells, apart from Cxcl1, included reduced expression of Plau, Ly6a, CD68 and Rgs16 (Table 3). Plau is a well-known marker of metastasis35, and its role in invasion has been noted in trophoblast migration36. Ly6a/Sca-1 is a mouse stem cell and tumor-initiating cell biomarker37,38 that down-regulates several tumor suppressor pathways, including PPARγ, TGF-β and PTEN39,40. CD68 is a scavenger receptor involved in phagocytosis, particularly in M2 polarized macrophages41, and has been implicated in immunotolerance by tumor-associated macrophages42. Rgs16 is a G-protein-coupled receptor associated with vascular smooth muscle cell proliferation and angiogenesis, which play prominent roles in oncogenesis43. Thus, the functions of Plac1 in placental development44 appear to phenocopy its functions in tumorigenesis, which supports the onco-placental nature of cancer3. From a therapeutic perspective, our data not only suggest that Plac1 may be a potential drug target, but that chemokine receptor antagonists developed for chronic inflammatory disorders, including COPD and psoriasis45,46, may be useful adjuvants when used in combination with other therapies to enhance the efficacy of cancer treatment47,48.
Methods
Cell lines
EO771 cells were originally isolated from a spontaneous mammary carcinoma in C57BL/6 mice49, and were provided by Dr. Louis M. Weiner, Georgetown University. EO771 cells tested negative against the IMPACT II panel of infectious agents (IDEXX). Plac1 and Cxcl1 expression were reduced with the piLenti-sRNA-GFP lentiviral vector targeting the sequence 5′-CCACCTTATGTCTACAATCAAAAGAGCAT-3′ of Plac11 mRNA (cat #i034429, ABM, Vancouver, Canada) or the sequence 5′-CTGCACCCAAACCGAAGTCATAGCCACAC-3′ of Cxcl1 mRNA (cat # i042697, ABM); a scrambled sequence was used as a control. Lentivirus expression of Cxcl1 utilized Lenti ORF clone Cxcl1 (MR220966L1) from Origene and lenti vector CMV-mCxcl1-IRES-mCherry (VB170524-1062) from VectorBuilder or the vector lacking Cxcl1 as a control. HEK293T cells were co-transfected at 50% confluence with the lentiviral shRNA plasmid, psPAX2 packaging plasmid and the VSV-G/pMD2 envelope plasmid at a ratio of 2:1:0.1 using Fugene 6 (Promega). After 18 hr, medium was replaced with fresh growth medium, and after 24–48 hr, the virus-containing supernatant was collected, filtered through a 0.45 um filter, mixed with fresh cell culture medium at a ratio of 7:1, and added to EO771 cells with 8 µg/ml polybrene. A lentivirus expressing a scrambled non-silencing control shRNA (shRNAmir, ABM) served as a negative control. Cells were selected for stable integration of the virus by incubation with 7.5μg/ml puromycin (Sigma-Aldrich Corp.) for 10 days. The efficiency of integration was monitored by GFP expression by the lentivirus. Cell lines 34T, 105T, 437T, MC and NeuT were described previously50,51.
Animals
EO771 cells at an inoculum of 1 × 106 cells/0.1 ml were injected into the no. 4 mammary gland of C57BL/6 or SCID mice (Taconic), and tumor growth was monitored daily. Cxcr2 antagonist SB225002 (Sigma-Aldrich) was dissolved in a diluent containing 8% DMSO, 10% PEG-400 and 1.75% Tween-20 in water at a concentration of 0.4 or 4.0 mg/ml, and administered i.p. Monday through Friday at a dose of 2 or 20 mg/kg, respectively52,53. Other mouse mammary tumor cells lines tested for Plac1 expression were 34T, 105T, 437T and MC40,51. Animal studies were conducted under protocols approved by the Georgetown University Animal Care and Use Committee (protocol 2016-1143) in accordance with NIH guidelines for the ethical treatment of animals.
Cell growth and cytotoxicity assays
EO771 or MC cells were grown in 96-well plates at an initial density of 5,000 cells per well. The cytotoxicity of SB225002 was determined in EO771 cells by dissolving the drug in DMSO and diluting in medium to a final DMSO concentration of 0.001%. Cell density was determined after incubation for 24, 48 and 72 hr by sulforhodamine B staining and measuring optical density at 570 nm54. Cytotoxicity assay data are shown in Supplementary Fig. 3.
Histopathology and immunohistochemistry (IHC)
Mammary tumors were excised, and formalin-fixed, paraffin-embedded sections were prepared for H&E staining and IHC by the Tissue and Histopathology Shared Resource, LCCC. Antigen retrieval was carried out by incubation of tissue sections in 10 mM sodium citrate buffer (pH 6.0) for 20 min at a sub-boiling temperature in an electric steamer as previously described14,51,55. Endogenous peroxidase activity was quenched with 3% hydrogen peroxide for 10 min, and incubated for 30 min with blocking solution (10% goat serum in Tris-buffered saline), followed by incubation overnight at 4 °C with the appropriate primary antibody diluted in blocking solution. Biotin-conjugated secondary antibodies were diluted in TBS containing 0.1% Tween-20 and incubated for 30 min at room temperature using the ABC Vectastain (Vector Laboratories) detection system and diaminobenzidine (Pierce), and slides were counterstained with Harris-modified hematoxylin (Thermo-Fisher, Inc.), dehydrated and mounted in Permount (Thermo-Fisher, Inc.). Apoptosis was determined with the SignalStain Apoptosis IHC Detection kit for cleaved caspase-3 (Cell Signaling Technology). Antibodies and their dilutions for IHC and FACS are listed in Supplementary Table 1.
Fluorescence-Activated Cell Sorting (FACS)
Tumor immune infiltrates were obtained by excising tumors, mincing them into small pieces and digestion with collagenase D (Roche) at a ratio of 15 ml collagenase solution per 2 g of tissue for 1 hr at 37 °C with shaking. The cell suspension was filtered through a 70 μm cell strainer (Falcon), washed, erythrocytes lysed and 1 × 106 cells were analyzed by FACS. Cells were stained using the Live/Dead Fixable Dead Cell Stain Kit (Invitrogen) and excluded from analysis, and non-specific binding was blocked with Fc antibody CD16/32 (Biolegend). Cells were first sorted for CD45 (macrophages, MDSC, Treg cells, NK cells and dendritic cells), CD45+CD3+ (T cells) or CD45+CD4+ (Treg cells). Cells were further sorted for: macrophages: F4/80+/MHCII+, MDSC: CD11b+/Gr-1+, dendritic cells: CD11c+/MHCII+, T cells: CD4+/CD8+, NK cells: CD45+/NK1.1+ and Treg cells: CD25+/Foxp3+. The fluorescent-conjugated monoclonal antibodies and their dilutions are listed in Supplementary Table 1. Cells were stained for Foxp3 after fixation in 1% paraformaldehyde and permeabilization (Permeabilization Buffer, eBioscience). Flow cytometry data was acquired by the Flow Cytometry & Cell Sorting Shared Resource, LCCC, with a BD LSRFortessa analyzer (BD Biosciences) and FCS Express 4 software (De Novo Software) to determine mean fluorescence intensity.
Gene microarray analysis
Microarray analysis was carried out as previously described39,51,55,56,57. Briefly, tissue was snap-frozen in liquid nitrogen, pulverized in a mortar and pestle and RNA was extracted using an RNeasy Mini Kit (Qiagen) according to the manufacturer’s protocol. RNA purity was assessed by the integrity of 18S and 28S rRNA using an Agilent microfluidic chip. Array analysis was carried out with cRNA prepared from equal amounts of RNA (1 μg) pooled from three replicates of cells per group. Biotin-labeled cRNA was fragmented at 94°C for 35 min and hybridized overnight to an Affymetrix mouse 430A 2.0 GeneChip®, and scanned with an Agilent Gene Array scanner. Grid alignment and raw data generation used the Affymetrix GeneChip® Operating software 1.1. A noise value (Q) based on the variance of low-intensity probe cells was used to calculate a minimum threshold for each GeneChip. Samples were averaged and data refined by eliminating genes with signal intensities <300 in both comparison groups, and heat maps were generated from ≥3-fold changes in gene expression normalized to control tissue using unsupervised hierarchical cluster analysis as previously described58. Gene expression data for EO771 control, EO881/shPlac1 and EO771/shCxcl1 cells are included in Supplementary Tables 2 and 3, respectively. Gene expression data for mice treated with vehicle or 20 mg/kg SB25001 are included in Supplementary Table 4. Gene interaction analysis utilized Ariadne Pathway Studio version 9.1 (Supplementary Fig. 4). Data sets were deposited in the GEO public database under accession no. GSE78202.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted as described above, and RNA (1 µg) from each of 3 samples per group was reverse transcribed using the Omniscript RT kit (Qiagen) as previously described39,51,55,56,57. PCR was performed in triplicate using an ABI-Prism 7700 (Applied Biosystems, Foster City, CA) with SYBRGreen I detection (Qiagen) according to the manufacturer’s protocol. Amplification using the appropriate primers (Supplementary Table 5) was confirmed by ethidium bromide staining of the PCR products on an agarose gel. The expression of each target gene was normalized to GAPDH and is presented as the ratio of the target gene to GADPH expression calculated using the formula, 2−ΔCt, where ΔCt = CtTarget − Ct18s 39.
Statistical analysis
Statistical significance of means ± S.D. were evaluated using the two-tailed Student’s t test at a significance of P < 0.05. Differences in tumor growth in vivo were determined by the unpaired two-tailed Student’s t test at a significance of P < 0.05.
References
Fant, M., Farina, A., Nagaraja, R. & Schlessinger, D. PLAC1 (Placenta-specific 1): a novel, X-linked gene with roles in reproductive and cancer biology. Prenat. Diagn. 30, 497–502 (2011).
Silva, W. A. Jr et al. PLAC1, a trophoblast-specific cell surface protein, is expressed in a range of human tumors and elicits spontaneous antibody responses. Cancer immunity 7, 18 (2007).
Old, L. J. Cancer is a somatic cell pregnancy. Cancer immunity 7, 19 (2007).
Grigoriadis, A. et al. CT-X antigen expression in human breast cancer. Proc. Natl. Acad. Sci. USA 106, 13493–13498, https://doi.org/10.1073/pnas.0906840106 (2009).
Koslowski, M. et al. A placenta-specific gene ectopically activated in many human cancers is essentially involved in malignant cell processes. Cancer Res. 67, 9528–9534 (2007).
Wang, X., Baddoo, M. C. & Yin, Q. The placental specific gene, PLAC1, is induced by the Epstein-Barr virus and is expressed in human tumor cells. Virology journal 11, 107, https://doi.org/10.1186/1743-422X-11-107 (2014).
Devor, E. J. & Leslie, K. K. The oncoplacental gene placenta-specific protein 1 is highly expressed in endometrial tumors and cell lines. Obstetrics and gynecology international 2013, 807849, https://doi.org/10.1155/2013/807849 (2013).
Cheung, H. W. et al. Systematic investigation of genetic vulnerabilities across cancer cell lines reveals lineage-specific dependencies in ovarian cancer. Proc. Natl. Acad. Sci. USA 108, 12372–12377, https://doi.org/10.1073/pnas.1109363108 (2011).
Dong, X. Y. et al. Plac1 is a tumor-specific antigen capable of eliciting spontaneous antibody responses in human cancer patients. Int. J. Cancer 122, 2038–2043 (2008).
Liu, F. et al. Identification of two new HLA-A*0201-restricted cytotoxic T lymphocyte epitopes from colorectal carcinoma-associated antigen PLAC1/CP1. J. Gastroenterol. 49, 419–426, https://doi.org/10.1007/s00535-013-0811-4 (2014).
Liu, F. F. et al. The specific immune response to tumor antigen CP1 and its correlation with improved survival in colon cancer patients. Gastroenterology 134, 998–1006 (2008).
Otsubo, T. et al. MicroRNA-126 Inhibits SOX2 Expression and Contributes to Gastric Carcinogenesis. PloS one 6, e16617 (2011).
Ghods, R. et al. High placenta-specific 1/low prostate-specific antigen expression pattern in high-grade prostate adenocarcinoma. Cancer Immunol. Immunother. 63, 1319–1327, https://doi.org/10.1007/s00262-014-1594-z (2014).
Yuan, H. et al. PPARdelta induces estrogen receptor-positive mammary neoplasia through an inflammatory and metabolic phenotype linked to mTOR activation. Cancer Res. 73, 4349–4361, https://doi.org/10.1158/0008-5472.CAN-13-0322 (2013).
Adhikary, T. et al. Genomewide analyses define different modes of transcriptional regulation by peroxisome proliferator-activated receptor-beta/delta (PPARbeta/delta). PloS one 6, e16344, https://doi.org/10.1371/journal.pone.0016344 (2011).
Torres-Arzayus, M. I. et al. High tumor incidence and activation of the PI3K/AKT pathway in transgenic mice define AIB1 as an oncogene. Cancer cell 6, 263–274 (2004).
Torres-Arzayus, M. I., Zhao, J., Bronson, R. & Brown, M. Estrogen-dependent and estrogen-independent mechanisms contribute to AIB1-mediated tumor formation. Cancer Res. 70, 4102–4111, https://doi.org/10.1158/0008-5472.CAN-09-4080 (2010).
Koslowski, M. et al. Selective activation of trophoblast-specific PLAC1 in breast cancer by CCAAT/enhancer-binding protein beta (C/EBPbeta) isoform 2. J. Biol. Chem. 284, 28607–28615 (2009).
Wagner, M. et al. NCOA3 is a selective co-activator of estrogen receptor alpha-mediated transactivation of PLAC1 in MCF-7 breast cancer cells. BMC cancer 13, 570, https://doi.org/10.1186/1471-2407-13-570 (2013).
Anzick, S. L. et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science 277, 965–968 (1997).
Liao, L. et al. Molecular structure and biological function of the cancer-amplified nuclear receptor coactivator SRC-3/AIB1. J. Steroid Biochem. Mol. Biol. 83, 3–14 (2002).
Grimm, S. L. & Rosen, J. M. The role of C/EBPbeta in mammary gland development and breast cancer. J. Mammary Gland Biol. Neoplasia 8, 191–204 (2003).
Lunghi, L., Ferretti, M. E., Medici, S., Biondi, C. & Vesce, F. Control of human trophoblast function. Reproductive biology and endocrinology: RB&E 5, 6, https://doi.org/10.1186/1477-7827-5-6 (2007).
Clark, D. A., Chaput, A. & Tutton, D. Active suppression of host-vs-graft reaction in pregnant mice. VII. Spontaneous abortion of allogeneic CBA/J × DBA/2 fetuses in the uterus of CBA/J mice correlates with deficient non-T suppressor cell activity. J. Immunol. 136, 1668–1675 (1986).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer cell 27, 450–461, https://doi.org/10.1016/j.ccell.2015.03.001 (2015).
Li, B. et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome biology 17, 174, https://doi.org/10.1186/s13059-016-1028-7 (2016).
Chen, Y., Moradin, A., Schlessinger, D. & Nagaraja, R. RXRalpha and LXR activate two promoters in placenta- and tumor-specific expression of PLAC1. Placenta 32, 877–884, https://doi.org/10.1016/j.placenta.2011.08.011 (2011).
Cocchia, M. et al. PLAC1, an Xq26 gene with placenta-specific expression. Genomics 68, 305–312 (2000).
Bezbradica, J. S. & Medzhitov, R. Integration of cytokine and heterologous receptor signaling pathways. Nature immunology 10, 333–339, https://doi.org/10.1038/ni.1713 (2009).
Acharyya, S. et al. A CXCL1 paracrine network links cancer chemoresistance and metastasis. Cell 150, 165–178, https://doi.org/10.1016/j.cell.2012.04.042 (2012).
Di Mitri, D. et al. Tumour-infiltrating Gr-1+ myeloid cells antagonize senescence in cancer. Nature 515, 134–137, https://doi.org/10.1038/nature13638 (2014).
Zou, A. et al. Elevated CXCL1 expression in breast cancer stroma predicts poor prognosis and is inversely associated with expression of TGF-beta signaling proteins. BMC cancer 14, 781, https://doi.org/10.1186/1471-2407-14-781 (2014).
Finak, G. et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat. Med. 14, 518–527, https://doi.org/10.1038/nm1764 (2008).
Bieche, I. et al. CXC chemokines located in the 4q21 region are up-regulated in breast cancer. Endocrine-related cancer 14, 1039–1052, https://doi.org/10.1677/erc.1.01301 (2007).
Harbeck, N., Schmitt, M., Paepke, S., Allgayer, H. & Kates, R. E. Tumor-associated proteolytic factors uPA and PAI-1: critical appraisal of their clinical relevance in breast cancer and their integration into decision-support algorithms. Crit. Rev. Clin. Lab. Sci. 44, 179–201, https://doi.org/10.1080/10408360601040970 (2007).
Lala, P. K. & Chakraborty, C. Factors regulating trophoblast migration and invasiveness: possible derangements contributing to pre-eclampsia and fetal injury. Placenta 24, 575–587 (2003).
Grange, C., Lanzardo, S., Cavallo, F., Camussi, G. & Bussolati, B. Sca-1 identifies the tumor-initiating cells in mammary tumors of BALB-neuT transgenic mice. Neoplasia 10, 1433–1443 (2008).
Liu, J. C., Deng, T., Lehal, R. S., Kim, J. & Zacksenhaus, E. Identification of tumorsphere- and tumor-initiating cells in HER2/Neu-induced mammary tumors. Cancer Res. 67, 8671–8681 (2007).
Upadhyay, G. et al. Stem cell antigen-1 enhances tumorigenicity by disruption of growth differentiation factor-10 (GDF10)-dependent TGF-beta signaling. Proc. Natl. Acad. Sci. USA 108, 7820–7825, https://doi.org/10.1073/pnas.1103441108 (2011).
Yuan, H., Upadhyay, G., Yin, Y., Kopelovich, L. & Glazer, R. I. Stem cell antigen-1 deficiency enhances the chemopreventive effect of peroxisome proliferator-activated receptor{gamma} activation. Cancer prevention research 5, 51–60 (2012).
Biswas, S. K. & Mantovani, A. Macrophage plasticity and interaction with lymphocyte subsets: cancer as a paradigm. Nature immunology 11, 889–896, https://doi.org/10.1038/ni.1937 (2010).
DeNardo, D. G. et al. Leukocyte complexity predicts breast cancer survival and functionally regulates response to chemotherapy. Cancer discovery 1, 54–67, https://doi.org/10.1158/2159-8274.CD-10-0028 (2011).
Hendriks-Balk, M. C. et al. Sphingosine-1-phosphate regulates RGS2 and RGS16 mRNA expression in vascular smooth muscle cells. Eur. J. Pharmacol. 606, 25–31, https://doi.org/10.1016/j.ejphar.2009.01.018 (2009).
Jackman S. K. X. & Fant M. Plac1 (Placenta-specific 1) Is Essential for Normal Placental and EmbryonicDevelopment. Mol. Reprod. Dev. in press (2012).
Stadtmann, A. & Zarbock, A. CXCR2: From Bench to Bedside. Frontiers in immunology 3, 263, https://doi.org/10.3389/fimmu.2012.00263 (2012).
Horuk, R. Chemokine receptor antagonists: overcoming developmental hurdles. Nature reviews. Drug discovery 8, 23–33, https://doi.org/10.1038/nrd2734 (2009).
Jamieson, T. et al. Inhibition of CXCR2 profoundly suppresses inflammation-driven and spontaneous tumorigenesis. J. Clin. Invest. 122, 3127–3144, https://doi.org/10.1172/JCI61067 (2012).
Highfill, S. L. et al. Disruption of CXCR2-mediated MDSC tumor trafficking enhances anti-PD1 efficacy. Science translational medicine 6, 237ra267, https://doi.org/10.1126/scitranslmed.3007974 (2014).
Dunham, L. J. & Stewart, H. L. A survey of transplantable and transmissible animal tumors. J. Natl. Cancer Inst. 13, 1299–1377 (1953).
Lindsay, J. et al. ErbB2 induces Notch1 activity and function in breast cancer cells. Clinical and translational science 1, 107–115, https://doi.org/10.1111/j.1752-8062.2008.00041.x (2008).
Yin, Y., Yuan, H., Zeng, X., Kopelovich, L. & Glazer, R. I. Inhibition of peroxisome proliferator-activated receptor gamma increases estrogen receptor-dependent tumor specification. Cancer Res. 69, 687–694, https://doi.org/10.1158/0008-5472.CAN-08-2446 (2009).
de Vasconcellos, J. F. et al. SB225002 Induces Cell Death and Cell Cycle Arrest in Acute Lymphoblastic Leukemia Cells through the Activation of GLIPR1. PloS one 10, e0134783, https://doi.org/10.1371/journal.pone.0134783 (2015).
Ijichi, H. et al. Inhibiting Cxcr2 disrupts tumor-stromal interactions and improves survival in a mouse model of pancreatic ductal adenocarcinoma. J. Clin. Invest. 121, 4106–4117, https://doi.org/10.1172/JCI42754 (2011).
Skehan, P. et al. New colorimetric cytotoxicity assay for anticancer-drug screening. J. Natl. Cancer Inst. 82, 1107–1112 (1990).
Yin, Y. et al. Peroxisome proliferator-activated receptor delta and gamma agonists differentially alter tumor differentiation and progression during mammary carcinogenesis. Cancer Res. 65, 3950–3957 (2005).
Pollock, C. B. et al. PPARdelta activation acts cooperatively with 3-phosphoinositide-dependent protein kinase-1 to enhance mammary tumorigenesis. PloS one 6, e16215, https://doi.org/10.1371/journal.pone.0016215 (2011).
Yin, Y. et al. Characterization of medroxyprogesterone and DMBA-induced multilineage mammary tumors by gene expression profiling. Mol. Carcinog. 44, 42–50, https://doi.org/10.1002/mc.20119 (2005).
Herschkowitz, J. I. et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome biology 8, R76, https://doi.org/10.1186/gb-2007-8-5-r76 (2007).
Acknowledgements
This work was supported by grants from the Nina Hyde Foundation, the Avon Foundation for Women, contract 1NO1 CN43302-WA19 from the National Cancer Institute, NIH, and award 1P30 CA051008 from the National Cancer Institute, NIH, to the Lombardi Comprehensive Cancer Center. This investigation was conducted using the Animal Research, Genomics and Epigenomics, Tissue and Histology and Microscopy and Imaging Shared Resources of the LCCC, and by an animal facilities construction grant from the NIH.
Author information
Authors and Affiliations
Contributions
R.I.G. conceived of the hypothesis. R.I.G., H.Y., L.M.W. and Y.T. designed the experiments. H.Y., X.W., C.S., L.J., J.L., J.H., N.V., V.D., and S.W., A.Z. performed the experiments. R.I.G., H.Y., L.J. and J.L. analyzed and interpreted the data. R.I.G. and H.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, H., Wang, X., Shi, C. et al. Plac1 Is a Key Regulator of the Inflammatory Response and Immune Tolerance In Mammary Tumorigenesis. Sci Rep 8, 5717 (2018). https://doi.org/10.1038/s41598-018-24022-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-24022-w
This article is cited by
-
PLAC1: biology and potential application in cancer immunotherapy
Cancer Immunology, Immunotherapy (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.