Abstract
Bacteria within the genus Alcaligenes, exhibit diverse properties but remain largely unexplored at genome scale. To shed light on the genome structure, heterogeneity and traits of Alcaligenes species, the genome of a tannery effluent isolated Alcaligenes faecalis subsp. phenolicus MB207 was sequenced and assembled. The genome was compared to the whole genome sequences of genus Alcaligenes present in the National Centre for Biotechnology Information database. Core, pan and species specific gene sequences i.e. singletons were identified. Members of this genus did not portray exceptional genetic heterogeneity or conservation and out of 5,166 protein coding genes from pooled genome dataset, 2429 (47.01%) contributed to the core, 1193 (23.09%) to singletons and 1544 (29.88%) to accessory genome. Secondary metabolite forming apparatus, antibiotic production and resistance was also profiled. Alcaligenes faecalis subsp. phenolicus MB207 genome consisted of a copious amount of bioremediation genes i.e. metal tolerance and xenobiotic degrading genes. This study marks this strain as a prospective eco-friendly bacterium with numerous benefits for the environment related research. Availability of the whole genome sequence heralds an opportunity for researchers to explore enzymes and apparatus for sustainable environmental clean-up as well as important compounds/substance production.
Similar content being viewed by others
Introduction
Alcaligenes specie strains exist in soil, water, and environment, as well as in association with humans. The bacteria of this genus are usually non-pathogenic but occasional opportunistic infections could occur in humans. Bacterial species belonging to the genus Alcaligenes have demonstrated versatile pollutant bioremediation capability, including phenols1,2, phenanthrene3, polyaromatic hydrocarbon4,5, pesticides6,7 and azo dye degradation8. Alcaligenes faecalis has been reported to convert the most toxic form of arsenic, arsenite to its less dangerous form, arsenate. Tolerance to heavy metals has been reported as well9,10. Nanoparticle production11, nematicidal12 and biocontrol activity13 has been reported in addition to production of chemicals14, detergent15, gum16, and bioplastics17. Despite such high applicability of Alcaligenes species in major spheres of research and prospective benefits in industry, agricultural and environmental domain, it remains underrepresented and understudied at whole genome level. No major comparative analysis or pan-genomic analysis has been published related to this bacterium till to date.
Here, we report the genome features of an Alcaligenes specie as well as comparison of its pan-genome and microsatellite i.e. simple sequence repeat (SSR)/ compound microsatellite (cSSR) profile with other Alcaligenes specie genomes. Alcaligenes faecalis subsp. phenolicus MB207 was isolated in 2010 from the effluent of a tannery in Multan, located in the southern zone of the province of Punjab in Pakistan and its genome was sequenced as a part of the on-going project on understanding and applying micro-remediation for environmental clean-up. Our group is further working on the various molecular aspects of this bacterium both in vitro and in silico, to further shed light on the mechanisms beneath its bioremediation capability.
Results
Overview of the sequenced genome
Total length of the genome was 4,156,248 bp with a GC content of 56.4%. Genome assembly through IDBA-UD approach resulted in 9 scaffolds. The sequence reads have been deposited in the NCBI SRA database and allocated the accession number: SRR5809679. OrthoANI value of 96.0 was obtained after comparison with the type strain Alcaligenes faecalis subsp. phenolicus DSM 16503. PGAP annotation revealed that the genome comprised of 3,812 genes, out of which 3,749 were coding DNA sequences and 63 were RNA genes. A total of 63 RNA genes were detected, including 5 S rRNA, 16 s rRNA and 23S rRNA copies. 53 tRNAs coding for all 20 amino acids and 4 ncRNAs were predicted. Pseudogenes with ambiguities like frameshift error, internal stop as well as incomplete pseudo gene sequences were predicted along with origin of replication (Fig. 1).
Incomplete prophage regions were predicted and apart from cell proliferation, chemotaxis, type I, II, VI secretion system, chaperones for cold shock, colicin V, lipase, patatin, siderophore production and antibiotic resistance proteins, biphenyl mineralization, phenol degradation, metal resistance, azo dye and Ibuprofen degradation proteins were found. Key features of some of the resistance genes are shown in Table 1. Genome sequence of Alcaligenes faecalis subsp. phenolicus MB207 harbours multiple xenobiotic degrading enzymes, enabling it to tolerate and thrive in the presence of anthropogenic, toxic compounds. All these genes were most probably encoded by chromosome as plasmid could not be detected. The presence of a repertoire of genes with bioremediation capability provides a genomic foundation for micropollutant tolerance ability of this versatile bacterium.
Alcaligenes spp have been used for bioplastic production with fatty acid supplementation18 and in presence of sugar beet/cane-molasses as sugar and urea as carbon source19. Our genome also encompassed enzymes for polyhydroxyalkanoate (PHA) synthesis, repression and depolymerization. Our strain carries genes for PHA (linear polyesters) production, which is usually a product of lipid and sugar fermentation by bacteria in nature. The gene is in immediate vicinity of acetyl-CoA enzyme which aids condensation of two acetyl-CoA molecules to acetoacetyl-CoA which is further reduced to the monomer hydroxybutyryl-CoA, the building block of PHA20. Intracellular stockpiling of PHA is usually carried out in the nutrient limiting and excess carbon setting, where PHA is hoarded inside the cell as energy-reserve granules. The biodegradable PHA polymer resembles petrochemical based polymer, which is unfortunately non-degradable. This heralds good news for eco-friendly bioplastic synthesis and reduce plastic related pollution. The presence of a PHA repressor protein shows a fine-tuning of the mechanism through negative regulation. Negative feedback loop could support homeostasis during environmental flux. A PHA depolymerizing enzyme occurrence indicates its possible role in plastic degradation and in turn aiding environment clean-up.
Antibiotic resistance is a pressing issue with natural history as well as human use. Many bacteria are avid antibiotic producers and resist them as well, for survival21. Genes from these are transferred horizontally to the antibiotic susceptible strains, resulting in acquiring resistant genes. In Alcaligenes faecalis subsp. phenolicus MB207, we catalogued the antibiotic resistance genes to understand the tannery polluted environmental reservoir of such genes. Only one sequence i.e. Undecaprenyl pyrophosphate phosphatase, involved in the sequestration of undecaprenyl pyrophosphate had a similarity cut-off above threshold and showed resistance to bacitracin. Other sequences had a lower cut-off value but high BLAST similarity and depicted resistance to tetracycline, chloramphenicol, fluoroquinolone, aminoglycoside, macrolide, trimethoprimlincosamide, macrolide, streptogramin_b, tigecycline, beta-lactam, carbenicillin, penicillin, erythromycin, glycylcycline, roxithromycin, kasugamycin, streptomycin, acriflavine, puromycin and t_chloride. Antibiotic resistance was also observed for mupirocin and anticoumarin via CARD (Fig. 2).
Pollutant tolerance
Alcaligenes faecalis subsp. phenolicus MB207 demonstrated tolerance to micropollutants including heavy metals22 (up to 250 µg/ml for nickel, cadmium, copper, zinc, lead and chromium in LB medium; pH:7; Temperature: 37 °C) and pharmaceutical Ibuprofen (our unpublished data). Genes for metal tolerance were searched and multiple copies of bacterioferritin, porins, ABC transporters, ATPases etc were found which are key regulators of metal transport in and out of the cell, involved in metal detoxification and survival in metal-stressed environment. Protein families for sensing and regulation for specific metals like arsenic and copper were also present.
Alcaligenes faecalis subsp. phenolicus MB207 has also shown azo dye (sulphonated mono and di-azo dye methyl orange and Congo red respectively) degradation capability23 (Supplementary Fig. 1). A degradation percentage of 64.18, 68.97 and 77.97 was achieved in LB medium (pH:7; Temperature:37 °C; Concentration: 100 µg/ml) for Congo red after 24, 48 and 72 hours respectively while a degradation percentage of 19.48, 36.8, and 40 was achieved for Methyl orange in LB medium after 24, 48 and 72 hours respectively. This is comparatively low as compared to other bacterial strains isolated from polluted environment, such as Pseudomonas24 with degradation as high as 97% achieved in 12 hours in similar conditions. A decolourization of 96 and 87% was achieved in 12 hrs for Methyl orange and double the quantity of Congo red (i.e. 200 µg/ml) respectively, at 30 °C for Shewanella xiamenensis BC0125. However, no data for degradation of both these sulphonated azo dyes is available for comparison with other Alcaligenes faecalis species. Although degradation percentage seemed low but it exhibited a remarkable capability of growth with both these dyes as sole carbon source. Genes responsible for the degradation of these dyes previously reported in literature such as azoreductase and peroxidase (GenBank accession: OQV32989.1 and OQV32923.1) were present in the genome sequence. BLAST hits showed closely related (up to 99% similar) NAD(P)H quinone dehydrogenase and azoreductase sequences in other Alcaligenes sp. genomes. A similarity of only 48% with azoreductase structure (of Pseudomonas putida, PDB ID: 4C14) in the Protein Data Bank was obtained. For peroxidase, high similarity was obtained for sequences in related genomes and only a 53% similarity was obtained with human peroxiredoxin structure in the Protein Data Bank. (PDB ID: 5B6M). There is a strong need for the prediction/solving of structures for these proteins.
Pan-genomic analysis
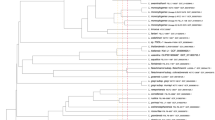
This type of analysis when applied to the sub-groups of organisms helps differentiate the serovars and pathovars, through niche and virulence-specific gene segregation. Distinct ecological and pathogenic traits could thus, be sieved out through this approach. Pan-genomic information been used as an aid to therapeutic design in bacteria26,27 as well as for heterogeneity study28. A total of eleven genomes of genus Alcaligenes were subjected to pan-genomic analysis (Table 2). The cluster map was significantly altered for the pan and core genome centred phylogeny for the studied bacterial species, apart from our bacterium Alcaligenes faecalis subsp. phenolicus MB207 and Alcaligenes faecalis M0R2, which remained grouped together (Fig. 3). BPGA tool also determined pan and core genome curve (Fig. 4) and its extrapolation through power law, to assess closing or openness of the pan-genome. The expected size of the pan-genome was calculated as 6087 while size estimated was 6032. The parameter ‘b’ was calculated to be 0.221095 and the pan-genome curve showed a plateau formation, indicating that pan-genome of this genus is yet open but may be closed soon. We have previously postulated that a pan-genome should be permanently open in bacteria due to natural evolution and horizontal gene transfer and we believe it should be the same case in genus Alcaligenes.
Phylogenetic analysis of the genus Alcaligenes based on (a) core and (b) pan-genome similarity. Clusters have been coloured separately to distinguish groups. Serial numbers referring to the genome names are same as the ones allocated in Table 2.
The highest number of new genes which contributed to the pan-genome were observed for Alcaligenes sp. EGD-AK7 (Fig. 5a). Bulk of the core genome of this genus was composed of genes having metabolic related functionality (Fig. 5b) as previously observed for the genus Serratia29, whereas hypothetical/poorly characterized genes contrived most of the unique or specie specific genome.
Secondary metabolite producing gene cluster analysis
Secondary metabolites serve as a rich source of bioactive compounds with pharmaceutical and other important properties. The compendium of genes encoding for important metabolites was detected through blast search against genes with similar architecture and composition. The genus has been underexplored for secondary metabolite genes which usually exist in clusters. Our genome was compared against gene cluster database (with information for ~3000 clusters) and predicted to encompass six such metabolite producing clusters.
The first cluster (Location: 169171–180010 nt) consisting of eleven genes, encoded butyrolactone. Homologous cluster were mined from Alcaligenes sp. EGD-AK7 and Alcaligenes sp. HPC1271 with a 28 percent similarity. Components include the major biosynthetic gene (Afsa) with A-factor biosynthesis hotdog domain, vital to streptomycin production and resistance. Major facilitator transporter was located upstream as well as downstream of biosynthetic gene. Siderophore receptor, damage inducible protein and MarR family transcriptional regulator coding genes were located upstream while downstream region comprised of TetR transcritional regulator which regulates processes like antibiotic production, resistance, efflux pump expression and osmotic stress response regulation. Second cluster (Location: 458937–469329 nt) codes for ectoine product. Out of thirteen genes, product synthesizing ones include diaminobutyrate-2-oxoglutarate-transaminase and L-ectoine synthase. Ectoine hydrolase and transporters were located downstream while metalloprotease, diaminobutyrate acetyltransferase, transcriptional regulator of class MarR and EamA family transporter were present upstream. More than 20 genes were mined upstream and downstream of terpene synthesis gene cluster with a total length of 21730 nucleotides. Ribose-5-phosphate isomerase, arsenate reductase, putrescine transporter components, sythases for squalene, phosphate starvation induced proteins, 3-deoxy-D-manno-octulosonic acid kinase, alanine racemase, alanine transaminase, homoserine dehydrogenase, periredoxin, zeta-carotene desaturase and DNA repair protein made up this cluster. Resorcinol producing cluster (Location: 920642–962552 nt) had homologs in four Alcaligenes species with similarity ranging from 24 to 17 percent. ABC transporters, dehydrogenases, TetR and LysR family transcriptional regulators, eneterobactin esterase, quercetin 2,3-dioxygenase, amidohydrolase and decarboxylating condensing enzymes coding for type III polyketide synthases, made up this cluster.
All of the genes for type I non-ribosomal peptide synthesis cluster (Location: 597271–644863 nt) genes showed some similarity with Alcaligenes sp. EGD-AK7 and Alcaligenes faecalis subsp. faecalis NCIB 8687 non-ribosomal peptide synthesis cluster (Fig. 6). This cluster constituted biotin metabolizing enzyme 8-amino-7-oxononanoate synthase, permeases, hypothetical proteins, polysaccharide deacetylase, glycosyl transferase, ABC transporter machinery, spore coat forming protein and capsular biosynthesis protein.
SSR and cSSR analysis
Alcaligenes sp. dataset consisted of 30,610 SSRs and 455 cSSRs ranging from a minimum of 2501 SSRs to a maximum of 3224. cSSRs ranged from 31 to 65 (Table 3). SSR density ranged from ~661 to 775 (Mean = 2782.73, S.D. = 204.07), while cSSR density ranged from 7.68 to 15.64 (Mean = 41.36, S.D. = 10.72). Strain MB207 had the highest number of SSR and cSSRs among the studied genomes. Strong correlation between SSR and cSSR density (R2 = 0.73, P < 0.01) was observed. This was almost similar to the previously analysed Lactobacillus genomes but dissimilar to the Escherichia coli genomes. This is in accordance with the hypothesis that the correlation of SSR and cSSR density is specie specific and dependent upon recombination of SSR motifs instead of the replication phenomenon28. Correlation between GC content and cSSR density (R2 = 0.13, P = 0.27) as well as genome size and cSSR density (R2 = 0.17, P = 0.19) was weak and non-significant contradictory to the results from other analyzed bacterial genomes28,30. Increment in cSSR formation is usually observed upon increase in maximum allowed distance between two adjoining SSRs. Number of cSSRs in all Alcaligenes specie genomes also increased with increased in dMAX values (Table 3).
To determine the organization and imperfection in SSR motif arrangement, giving rise to cSSRs in Alcaligenes genomes, we explored the complexity and structural make-up of cSSRs. cSSR coupled motifs (e.g. TTAAGT-CTTGTT) were unique i.e. distinct for each Alcaligenes specie and most probably arose by defective duplication.
Motif duplication i.e. similar motifs on both ends of spacer sequence existed once in Alcaligenes faecalis subsp. phenolicus DSM 16503 and Alcaligenes sp. EGD-AK7, at both dMAX = 10 and dMAX = 50. In Alcaligenes faecalis NBIB-017, Alcaligenes faecalis subsp. phenolicus IITR89, Alcaligenes sp. HPC1271, Alcaligenes faecalis MOR02, Alcaligenes faecalis strain NCIB 8687, no duplication was observed at dMAX = 10 but at dMAX = 50, adjacent duplications of 1, 1, 2, 2 and 2 motifs were mined respectively. Presence of similar duplicated motifs advocates that these Alcaligenes species have analogous array of motif duplication in their genomes. For Alcaligenes faecalis subsp. phenolicus MB207, 3 duplications at dMAX = 10 and 2 duplications at dMAX = 50 were observed. Alcaligenes faecalis NBIB-017 showed 4 and 3 pair of duplications at dMAX = 10 and dMAX = 50 respectively. In Alcaligenes faecalis ZD02, presence of 1 and 2 duplicated motifs were detected at dMAX = 10 and dMAX = 50 respectively. Alcaligenes faecalis strain P156 did not show any duplication at either dMAX = 10 or dMAX = 50.
Moreover, a cSSR is either categorized as perfect (e.g. [(TG)n(TA)n]) or overlapping (intersection of final base of the precedent motif with first base of the next motif (e.g [(ATT)n(TG)n]) (Kumar et al., 2014). Overlapping cSSR motifs were present in all Alcaligenes genomes (Table 2). Inspection of cSSR complexity indicated that cSSR assembly was very complex and intricate in the Alcaligenes genome sequences. A complexity of up to ‘32-microsatellite’ cSSRs was reached in our sequenced genome specie at dMAX = 50, which is even greater than that of eukaryote ‘24-microsatellite’ complexity31.
Discussion
Alcaligenes faecalis subsp. phenolicus is a gram-negative rod-shaped bacterium. It has the unique ability to utilize phenol as a sole carbon source1. Here, we have sequenced and reported the characteristics of Alcaligenes faecalis subsp. phenolicus MB207. Sequencing provided a glimpse into its micropollutant tolerance capability and gene apparatus responsible for these properties was identified and being investigated further, both in vitro and in silico. Our isolate had micropollutant resistance, azo dye and ibuprofen degradation properties. This bacterium has ample genes for metal sensing and transport which enables it to create metal homeostasis system that helps it survive/thrive in polluted environment. Further analysis like varied expression profiling and proteome alteration under metal stress could provide better understanding regarding metal homeostasis in the bacterium. Genome sequencing of Alcaligenes faecalis subsp. phenolicus MB207 is an important milestone in understanding its remediation and eco-friendly properties. A lot of antibiotic resistance genes were mined and since antibiotic resistance system impacts metal homeostasis and vice versa, it would be interesting to explore this facet too.
We also elucidated the bioplastic forming and depolymerizing apparatus in addition to gene clusters responsible for butyrolactone, ectoine, resorcinol, terpene, nrps and pks. In silico inspection in Alcaligenes specie genomes revealed a wealth of SSRs and cSSRs. The most complex structured cSSR was detected in our sequenced genome. It is demonstrated that previously uncorrelated genome data can be utilized for mining of new biological information by means of available softwares, databases and high-performance computation. Democratization of genome sequencing has made bacterial genomics a mature and easy approach for researchers from interdisciplinary fields like environment, evolution and scientists working in the biomedical disciplines. Genomic data stored in repositories is available to public for comparison with their own datasets which has made the studies concurrently deeper and diffused, leading to interesting results and conclusions. A striking example is the concept of pangenome, introduced originally with pathogenic strains and now widely studied for non-pathogenic species/genus of interest. We have touched upon this analysis but for full scale comparison with other bacteria of similar sizes and to make solid conclusions, similar scale approaches need to be undertaken. The software and parameters need to be standardized for quality assessment. Secretion system protein clusters did not show significant alignment with a particular genus or specie although each protein of the system resembled a similar protein of different specie but on the whole, system level conservation was not observed, although interspecies similarity was high.
Tandem repeats of nucleotide motifs (sized 1–6 bp) are called SSRs and give rise to cSSRs upon joining. They are known to exist in all genomes and their importance ranges from use as molecular markers to studying genome evolution. cSSRs have an alleged role in the expression of gene regulation and functional dictation of proteins in numerous species30,32. Hence, it is important to study their distribution, enrichment and polymorphism in the genomes of interest. Correlation between SSR and cSSR density of our isolate was almost similar to the previously analysed Lactobacillus genomes but dissimilar to the Escherichia coli genomes. This is in accordance with the hypothesis put forward by us28 that the correlation of SSR and cSSR density is specie specific and dependent upon recombination of SSR motifs instead of the replication phenomenon. Since the correlation of GC content with cSSR density and genome size with cSSR density was weak and non-significant. This did not comply with the previous analyses on bacterial genomes28,30. cSSR coupled motif structure was consistent with Escherichia coli and Lactobacillus cSSR motifs, with distinct bases30 unlike eukaryotic cSSRs, that have similar motifs in ~90% cases31. Occurrence of ‘32-microsatellite’ cSSR complexity divulges from previously analysed Escherichia coli and Lactobacillus genome study, which did not cross the maximum complexity of more than ‘5-microsatellite’ cSSRs upon dMAX increment to 50. Complexity of prokaryotic cSSR does not seem to be dependent upon genome size as genome size of eukaryotes is colossal as compared to genome size of bacteria. This is also consistent with previous study that complexity seems to be depends on SSR abundance as it might augment the frequency of joining SSRs into cSSRs by chance28. Our analysis is computational and only an intelligent guess due to lack of absolute certainty in the GC content and genome sizes of the genomes used in the study i.e. unfinished/draft genome sequences. The actual impact needs to be proven experimentally.
Methods
Genome sequencing, assembly and annotation
DNA extraction was done with high pure PCR template preparation kit (Roche, Switzerland). Whole-genome sequencing was performed using the MiSeq PE300 sequencer with 2 × 300 bp pair-end library. A total of 794,028 reads (55× coverage of the genome) were generated, cleaned and quality filtered using Trimmomatic33. Reads were then corrected for errors through String graph assembler which utilizes a k-mer centric algorithm34. De novo assembly was attempted through IDBA-UD algorithm centred on de bruijn graph approach35. Genome annotation was carried out using the NCBI ‘Prokaryotic genome annotation pipeline’. Genomic context was visualized through GView36. Prophage regions were identified using PHAST37 and antibiotic resistance was profiled through BLAST module of ARDB38. Resistance Gene Identifier (RGI) listed at CARD site (https://card.mcmaster.ca/analyze/rgi), was used for mapping of resistome featuring homology and SNP model with strict criteria.
Specie demarcation and pan-genomic analysis
OrthoANI39 scheme was used for specie demarcation using whole genome sequence data. This type of orthologous average nucleotide identity (ANI) calculation between genomes is valid for differentiation at the species scale for microorganisms. A value of 95% and above indicates that the queried bacterium belongs to the same species as that of the reference.
BPGA (Bacterial Pan-genome Analysis tool) was used for estimation of core, pan and specie specific genome analysis40. The thresholds of the score and E-value used for BLAST were greater than 50 and less than 1e−8, respectively. Annotated genomes were taken as a seed substance for the construction of a pan-genome. Alcaligenes faecalis genomes present in the NCBI database (till the accomplishment of this study i.e. March 2017) were subjected to a pair wise homology search through BLAST. Orthologs were calculated for all possible genome pairs. In case of a partial or incomplete gene sequence, the reciprocity cannot be marked clearly due to small length. Even with a similarity of 100%, gene cannot be captured in accessory or core genome data pool and labelled as a singleton. To circumvent this problem, a length of 50% or more was considered for gene reciprocity. Initial clustering was done through Usearch algorithm and output processed into pan, core and accessory gene distribution of the genus Alcaligenes. The empirical power law equation f(x) = a.x^b and exponential equation f1(x) = c.e^(d.x) were used for extrapolation of the pan and core genome curves respectively. Exclusive presence and absence of genes/families was determined to infer specie-specific gene families. Upon addition of each new genome in the analysis pipeline, 20 random permutations of genomes were carried out to circumvent any bias. Evolutionary analysis based on concatenated gene alignments and binary (presence/absence) pan-matrix was conducted with Neighbour joining approach. COG distribution, KEGG pathway analyses and phylogeny based on core and pan proteome was then attempted.
Secondary metabolite analysis
antiSMASH41 was used for secondary metabolite gene cluster detection as well as detailed comparison to related clusters in other microorganisms. It is based on hidden Markov model profiling of genes associated with important metabolite production of all known broad chemical classes. The boundary of the gene clusters is estimated via various greedily chosen cut-off values, specified per gene cluster type and genes represented in specified colours referring to certain functionality.
SSR and cSSR analysis
SSR and cSSR information was extracted using the software IMEx42 in batch, using the parameters: Include flanking regions: 10 bp, Type of repeat: imperfect; Repeat Size: all; Minimum Repeat Number: Mono: 12, Di: 6, Tri: 4, Tetra: 3, Penta: 3, Hexa: 3, Imperfection limit/repeat unit: Mono: 1, Di: 1, Tri: 1, Tetra: 2, Penta: 2, Hexa: 3, Percent imperfection in repeat tract: 10%, Maximum distance allowed between any two adjacent SSRs forming a cSSR (i.e. dMAX in bp): 10 with complete standardization28. The obtained results were then compared to microsatellites in previously studied prokaryotic species i.e. Escherichia coli30 and Lactobacillus28. Linear regression (R2) was calculated using IBM SPSS v22, to evaluate the impact of GC content and genome size on the SSR and cSSR composition as well as correlation among SSR density (number of SSR/Mb) and cSSR density (number of cSSR/Mb). A P-value of <0.05 was considered as significant.
Nucleotide sequence accession numbers
The accession numbers of the sequences of Alcaligenes faecalis subsp. phenolicus MB207 determined in this study can be found in GenBank (http://www.ncbi.nlm.nih.gov) under the accession no. MTBI01000001-MTBI01000009.
References
Rehfuss, M. & Urban, J. Alcaligenes faecalis subsp. phenolicus subsp. nov. a phenol-degrading, denitrifying bacterium isolated from a graywater bioprocessor. Syst. Appl. Microbiol. 28, 421–429 (2005).
Kumar, A. et al. Optimization of culture condition for growth and phenol degradation by Alcaligenes faecalis JF339228 using Taguchi methodology. Desalin. Water Treat. 51, 3153–3163 (2013).
Kiyohara, H., Takizawa, N., Date, H., Torigoe, S. & Yano, K. Characterization of a phenanthrene degradation plasmid from Alcaligenes faecalis AFK2. J. Ferment. Bioeng. 69, 54–56 (1990).
Bharali, P., Das, S., Konwar, B. K. & Thakur, A. J. Crude biosurfactant from thermophilic Alcaligenes faecalis: feasibility in petro-spill bioremediation. Int. Biodeterior. Biodegrad. 65, 682–690 (2011).
John, R. C., Essien, J. P., Akpan, S. B. & Okpokwasili, G. C. Polycyclic aromatic hydrocarbon-degrading bacteria from aviation fuel spill site at Ibeno, Nigeria. Bull. Environ. Contam. Toxicol. 88, 1014–1019 (2012).
Siripattanakul, S., Wirojanagud, W., McEvoy, J., Limpiyakorn, T. & Khan, E. Atrazine degradation by stable mixed cultures enriched from agricultural soil and their characterization. J. Appl. Microbiol. 106, 986–992 (2009).
Kong, L. et al. Biodegradation of organochlorine pesticide endosulfan by bacterial strain Alcaligenes faecalis JBW4. J. Environ. Sci. 25, 2257–64 (2013).
Shah, P. D., Dave, S. R. & Rao, M. S. Enzymatic degradation of textile dye Reactive Orange 13 by newly isolated bacterial strain Alcaligenes faecalis PMS-1. Int. Biodeterior. Biodegrad. 69, 41–50 (2012).
Gupta, S. & Nirwan, J. Evaluation of mercury biotransformation by heavy metal-tolerant Alcaligenes strain isolated from industrial sludge. Int. J. Env. Sci. Technol. 12, 995–1002 (2015).
Abbas, S. et al. A heavy-metal tolerant novel bacterium, Alcaligenes pakistanensis sp. nov., isolated from industrial effluent in Pakistan. Antonie van Leeuwenhoek 108, 859–70 (2015).
Abo-Amer, A. E., El-Shanshoury, A. E. R. R. & Alzahrani, O. M. Isolation and molecular characterization of heavy metal-resistant Alcaligenes faecalis from sewage wastewater and synthesis of silver nanoparticles. Geomicrobiol. J. 32, 836–845 (2015).
Ju, S. et al. Alcaligenes faecalis ZD02, a novel nematicidal bacterium with an extracellular serine protease virulence factor. Appl. Environ. Microbiol. 82, 2112–2120 (2016).
Liu, X. et al. The genome sequence of Alcaligenes faecalis NBIB-017 contains genes with potentially high activities against Erwinia carotovora. Genome Announc. 4, e00222–16, https://doi.org/10.1128/genomeA.00222-16 (2016).
Xu, H. M., He, C. L., Xiong, W. J., Wei, Y. C. & Xia, S. W. Preparation of (R)-o-chloromandelic acid via dynamic kinetic resolution of o-chloromandelonitrile by whole cells of Alcaligenes faecalis CGMCC 1.2006. J. Mol. Catal. 2, 174–181 (2014).
Muthuraman, M. S., Kumari, S. & Sivasubramanian, A. Production, characterization and purification of alkaline protease from Alcaligenes sp., and its application in detergent industry. Asian J. Pharm. Clin. Res. 1, 152–155 (2013).
Ai, H. et al. Improved welan gum production by Alcaligenes sp. ATCC31555 from pretreated cane molasses. Carbohydr. Polym. 129, 35–43 (2015).
Tripathi, A. D., Srivastava, S. K. & Singh, R. P. Statistical optimization of physical process variables for bio-plastic (PHB) production by Alcaligenes sp. Biomass Bioenergy 55, 243–250 (2013).
Srivastava, S. K. & Tripathi, A. D. Effect of saturated and unsaturated fatty acid supplementation on bio-plastic production under submerged fermentation. 3 Biotech 3, 389–397 (2013).
Tripathi, A. D., Yadav, A., Jha, A. & Srivastava, S. K. Utilizing of sugar refinery waste (cane molasses) for production of bio-plastic under submerged fermentation process. J. Polym. Environ. 20, 446–453 (2012).
Doi, Y. & Steinbuchel, A. Biopolymers. (Wiley-VCH, 2002).
Perry, J., Waglechner, N. & Wright, G. The prehistory of antibiotic resistance. Cold Spring Harb. Perspect. Med. 6, a025197 (2016).
Tanveer, F. Bioaccumulation of selected heavy metals by various bacterial strains from tannery effluents. Masters dissertation. Fatima Jinnah Women University, Rawalpindi (2011).
Basharat, Z. In vitro and in silico analysis of enzymes from indigenous bacteria for remediation of azo dyes. M. Phil. dissertation. Fatima Jinnah Women University, Rawalpindi (2014).
Telke, A. A., Joshi, S. M., Jadhav, S. U., Tamboli, D. P. & Govindwar, S. P. Decolorization and detoxification of Congo red and textile industry effluent by an isolated bacterium Pseudomonas sp. SU-EBT. Biodegradation 21, 283–296 (2010).
Ng, I. S. et al. Decolorization of textile azo dye and Congo red by an isolated strain of the dissimilatory manganese-reducing bacterium. Shewanella xiamenensis BC01. Appl. Microbiol. Biotechnol. 98, 2297–2308 (2014).
Muzzi, A., Masignani, V. & Rappuoli, R. The pan-genome: towards a knowledge-based discovery of novel targets for vaccines and antibacterials. Drug Discov. Today 12, 429–439 (2007).
Ali, A. et al. Pan-genome analysis of human gastric pathogen H. pylori: comparative genomics and pathogenomics approaches to identify regions associated with pathogenicity and prediction of potential core therapeutic targets. BioMed Res. Int. 2015, 139580, https://doi.org/10.1155/2015/139580 (2015).
Basharat, Z. & Yasmin, A. Survey of compound microsatellites in multiple Lactobacillus genomes. Can. J. Microbiol. 61, 898–902 (2015).
Basharat, Z. & Yasmin, A. Pan-genome analysis of the genus Serratia. arXiv preprint arXiv:1610.04160 (2016).
Chen, M. et al. Compound microsatellites in complete Escherichia coli genomes. FEBS Lett. 585, 1072–1076 (2011).
Kofler, R., Schlötterer, C., Luschützky, E. & Lelley, T. Survey of microsatellite clustering in eight fully sequenced species sheds light on the origin of compound microsatellites. BMC Genomics 9, 612 (2008).
Kashi, Y. & King, D. G. Simple sequence repeats as advantageous mutators in evolution. Trends Genet. 22, 253–259 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 1, 2114–20 (2014).
Simpson, J. T. & Durbin, R. Efficient de novo assembly of large genomes using compressed data structures. Genome Res. 22, 549–556 (2012).
Peng, Y., Leung, H. C., Yiu, S. M. & Chin, F. Y. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Petkau, A., Stuart-Edwards, M., Stothard, P. & Van Domselaar, G. Interactive microbial genome visualization with GView. Bioinformatics 26, 3125–3126 (2010).
Zhou, Y., Liang, Y., Lynch, K. H., Dennis, J. J. & Wishart, D. S. PHAST: a fast phage search tool. Nucleic Acids Res. 39, W347–W352 (2011).
Liu, B. & Pop, M. ARDB—antibiotic resistance genes database. Nucleic Acids Res. 37, D443–D447 (2009).
Lee, I., Kim, Y. O., Park, S. C. & Chun, J. OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103 (2016).
Chaudhari, N. M., Gupta, V. K. & Dutta, C. BPGA-an ultra-fast pan-genome analysis pipeline. Sci. Rep. 6, 24373, https://doi.org/10.1038/srep24373 (2016).
Weber, T. et al. antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res. 43, W237–W243 (2015).
Mudunuri, S. B. & Nagarajaram, H. A. IMEx: imperfect microsatellite extractor. Bioinformatics 23, 1181–1187 (2007).
Acknowledgements
The authors are grateful to all those scientists, research labs and universities who made computational biology tools, data and web servers freely available for public use.
Author information
Authors and Affiliations
Contributions
Z.B., Y.T. and A.Y. conceived and designed the experiments. A.Y. and Y.T. provided reagents/material for the experiments and administered the project. Z.B. and T.H. conducted the experiments. Z.B. and A.Y. curated the data. Z.B. drafted the original manuscript while A.Y., T.H. and Y.T. reviewed and edited the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Basharat, Z., Yasmin, A., He, T. et al. Genome sequencing and analysis of Alcaligenes faecalis subsp. phenolicus MB207. Sci Rep 8, 3616 (2018). https://doi.org/10.1038/s41598-018-21919-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21919-4
This article is cited by
-
‘A plant’s major strength in rhizosphere’: the plant growth promoting rhizobacteria
Archives of Microbiology (2023)
-
Genomic and resistome analysis of Alcaligenes faecalis strain PGB1 by Nanopore MinION and Illumina Technologies
BMC Genomics (2022)
-
Genome Mining of Three Plant Growth-Promoting Bacillus Species from Maize Rhizosphere
Applied Biochemistry and Biotechnology (2021)
-
Probiotic Properties of Alcaligenes faecalis Isolated from Argyrosomus regius in Experimental Peritonitis (Rat Model)
Probiotics and Antimicrobial Proteins (2021)
-
Whole genome sequence analysis reveals high genetic variation of newly isolated Acidithiobacillus ferrooxidans IO-2C
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.