Abstract
Accumulating evidence has revealed that aberrant Circular RNAs (circRNAs) expression plays important roles in carcinogenesis and tumor progression. However, their role in non-small cell lung cancer (NSCLC) remains unclear. In this study, we first used circRNA microarrays to screen for tumour-specific circRNA candidates in between NSCLC (n = 3) and adjacent lung (n = 3) tissue. Among the circRNA expression profile, two circRNAs (hsa_circ_0014130 and hsa_circ_0016760) were selected for validation in ten pairs of NSCLC and adjacent non-cancerous tissues by real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR). Only hsa_circ_0014130 exhibited significantly overexpressed in NSCLC tissues (P < 0.001), which were further confirmed in another 36 matched tissue samples using qRT-PCR. Hsa_circ_0014130 expression significantly correlated with TNM stage (P = 0.001) and lymphatic metastasis (P = 0.004). The area under the receiver operating characteristic curve was 0.878 (95% confidence interval = 0.804–0.951; P < 0.001), which showed good diagnostic potential. Bioinformatics platforms predicted that hsa_circ_0014130 might interact with five miRNAs and their corresponding mRNAs. Gene oncology analysis and pathway analysis revealed that hsa_circ_0014130 could participate in NSCLC development. In summary, our findings indicated that hsa_circ_0014130 could be used as a potential NSCLC biomarker and might be closely related to the carcinogenesis of NSCLC.
Similar content being viewed by others
Introduction
Lung cancer is the leading cause of cancer death among males in both more and less developed countries, and has surpassed breast cancer as the leading cause of cancer death among females in more developed countries1. Non-small cell lung cancer (NSCLC) accounts for more than 80% of all lung cancer cases, consists of squamous cell carcinoma, large cell carcinomas and adenocarcinoma, and most patients with NSCLC have advanced local invasion and/or distant metastases at the time of diagnosis2. Therefore, better understanding of the molecular mechanisms associated with NSCLC pathogenesis is critical to the development of effective diagnostic and therapeutic approaches, which in turn contributes to identify novel biomarkers for NSCLC detection and treatment targets, and even to develop personalized therapies for individual patients NSCLC with in the future.
Circular RNAs (circRNAs) are a unique class of endogenous noncoding RNAs (ncRNAs) that, unlike linear RNA, form a closed continuous loop by back-splicing with covalently joined 3′- and 5′-ends3. As a result of this closed structure, circRNAs have been shown to be highly stable and largely resistant to RNA degradative pathways4, which highlights clear advantages in using circRNAs as novel molecular biomarkers for many diseases. Many studies have reported that circRNAs have numerous biologic functions, which are involved in forming RNA-protein complexes, acting as microRNA (miRNA) sponges, and regulating targeted gene transcription and splicing5. Therefore, it has been hypothesized that circRNAs potentially regulate disease progression by sequestering a miRNA associated with a particular disease6. Recent studies indicated that circRNAs might play an important role in carcinogenesis and tumor progression7,8. Some studies have revealed that circRNAs exhibit dysregulated expression in human various cancers including esophageal squamous cell carcinoma, gastric cancer, hepatocellular carcinoma, colorectal cancer, laryngeal cancer, gliomas9,10,11,12,13,14,15 and so on. It has been reported that circRNA might serve as a novel potential biomarker for cancer diagnosis16. However, the role of circRNAs in NSCLC has not been well studied. A report has showed that circular RNA-ITCH may compete with ITCH to bind to miR-7 and miR-214 and may be involved in lung cancer development17. Hsa_circ_100876 has been suggested as a potential prognostic biomarker for NSCLC18. Hsa_circ_0013958 has been indicated to be used as a potential non-invasive biomarker for the early detection and screening of lung adenocarcinoma19. Despite this potential link with circRNAs, the global circRNA expression profile in NSCLC has not been fully uncovered.
Hence, in the present study we first performed circRNA microarray to investigate the differential expression profiles of circRNAs between NSCLC tissues and paired adjacent non-cancerous tissues. Subsequently, from these differentially expressed circRNAs detected in microarray, two circRNAs (hsa_circ_0014130 and hsa_circ_0016760) were confirmed by real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR). We then selected hsa_circ_0014130, with significantly upregulated expression in the validation test above, as a targeted circRNA to further explore its clinical significance and application in NSCLC. In addition, a bioinformatics analysis was used to predict the miRNA binding sites and the related mRNAs of hsa_circ_0014130. These results indicate that the upregulated expression of hsa_circ_0014130 significantly associates with some clinicopathological factors of NSCLC patients, and may serve as a novel potential tumor marker and therapeutic target for NSCLC.
Results
Validation of RNA quality
Total RNA was extracted from each sample, and the results of RNA quality and RNA integrity were displayed in Supplementary Figure 1.
Differential circRNA expression profiles in NSCLC
Using high-throughput human circRNA microarray, we assessed the differences of circRNA expression profiles between NSCLC tissues and paired adjacent non-cancerous tissues from 3 patients. We drew a Box plot to quickly visualize the distribution of the intensities from all the datasets after normalization and found that the distribution of log2 ratios was similar in all the tested samples (Fig. 1A). Cluster analysis segregated samples into groups based on differences in their expression levels, and hypothetic relationships among the samples. The results of hierarchical clustering showed distinguishable circRNA expression profiling among 6 samples (Fig. 1B). These data indicated that circRNAs have a different expression pattern in NSCLC compared with that in adjacent lung tissues. The Volcano plot was performed to visualize the significant differences (fold change > 2.0, P value < 0.05) between NSCLC and non-cancerous tissues (Fig. 1C). Moreover, the distributions of differentially expressed circRNAs in human chromosomes showed that most circRNAs were transcribed from chr1, chr3, chr9, chr12, chr16 and chr19, but seldom from chr13, chr21 and chrY (Fig. 1D). The microarray data showed 171 circRNAs were found to be differentially expressed in NSCLC tissues (fold change > 2.0, P value < 0.05). Among them, 148 circRNAs were upregulated and 23 were downregulated in tumor tissues. The top 5 upregulated and downregulated circRNAs were presented in Table 1.
Differences and characterizations of circRNA expression profile between NSCLC tissues and paired adjacent non-cancerous tissues. (A) Box plots show the distribution of circRNAs for the six samples (C for NSCLC and A for adjacent non-cancerous tissues). The distributions were nearly the same after normalization. (B) Unsupervised hierarchical clustering shows a distinguishable circRNA expression profiling among the six samples (C for NSCLC and A for adjacent non-cancerous tissues). Each column represents the expression profile of a tissue sample, and each row corresponds to a circRNA. “Red” indicates higher expression level, and “green” indicates lower expression level. (C) Volcano plots are used to visualize the differential circRNA expression. The vertical green lines correspond to 2.0-fold up and down, and the horizontal green line represents a P-value of 0.05. The red points in plot represent the differentially expressed circRNAs with statistical significance. (D) Chromosomal distributions of differentially expressed circRNAs.
Amplification of hsa_circ_0014130 and hsa_circ_0016760
We used qRT-PCR assays to verify 2 typically differential expression circRNAs (hsa_circ_0014130 and hsa_circ_0016760) in 10 samples of NSCLC tissues and their paired adjacent lung tissues. The primers were specifically capable of amplifying the backsplice sites of each circRNA, and the PCR results were validated by melt-curve analysis (Supplementary Fig. 2). These tests confirmed that hsa_circ_0014130 and hsa_circ_0016760 existed in NSCLC tissues and could be specifically amplified by qRT-PCR. As shown in Fig. 2, we found that the hsa_circ_0014130 expression levels in NSCLC tissues were significantly higher than those in corresponding non-cancerous tissues, but hsa_circ_0016760 expression levels had no significant difference between the NSCLC tissues and the corresponding non-cancerous tissues. However, the expression level of hsa_circ_0014130 was not consistent with that measured by microarray analysis, in which the hsa_circ_0014130 expression was significantly downregulated (Table 1).
The expression levels of candidate circRNAs were validated by performing RT-qPCR (in triplicate) with ten pairs of NSCLC tissues and adjacent lung tissues. RNA expression levels were normalized to expression of the reference gene β-actin, only hsa_circ_0014130 was significantly differentially expressed between the 2 groups (4.06-fold, P = 0.002).
Upregulation of hsa_circ_0014130 expression in NSCLC Tissues
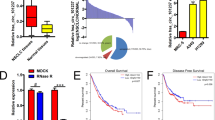
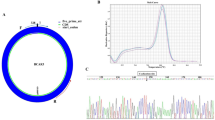
By means of inquiring from circBase (http://www.circbase.org), we knew that hsa_circ_0014130 was located at chromosome 1q21.3 and was composed of five exons (Supplementary Fig. 3A). The PCR results of hsa_circ_0014130 were validated by gel electrophoresis (Supplementary Fig. 3B) and sequencing the amplified product (Supplementary Fig. 3C), which showed that the primers were specifically capable of amplifying the back splice sites of hsa_circ_0014130. Next, to validate the previous study results, we expanded the sample size. The expression levels of hsa_circ_0014130 in 46 NSCLC tissues and their matched adjacent non-cancerous tissues were measured by qRT-PCR method, which showed that hsa_circ_0014130 expression was significantly upregulated in NSCLC tissues (P < 0.001, Fig. 3A). Moreover, the results showed that hsa_circ_0014130 expression was significantly upregulated in 82.6% (38/46) NSCLC tissues compared with the adjacent non-cancerous tissues (Fig. 3B). We then used the receiver-operating characteristic (ROC) curve to investigate the diagnostic value of hsa_circ_0014130 in distinguishing NSCLC tissues from adjacent non-cancerous tissues. When the expression level of hsa_circ_0014130 was analyzed for this purpose, the area under the ROC curve (AUC) was 0.878 (Fig. 3C), and the optimal cutoff value of hsa_circ_0014130 was 0.573, with sensitivity and specificity were 87.0% and 84.8%, respectively.
Hsa_circ_0014130 significantly upregulated in cancer tissues could serve as a biomarker for NSCLC. (A) The expression levels of hsa_circ_0014130 in the NSCLC group are significantly higher than those in corresponding non-cancerous tissues (n = 46) (P < 0.001). (B) Hsa_circ_0014130 levels were significantly upregulated in 82.6% (38/46) of NSCLC tissues. (C) Receiver operating characteristic (ROC) curve of hsa_circ_0014130 was built for differentiating NSCLC tissues from controls. The area under curve was 0.878. The cutoff value was 0.573, and the sensitivity and specificity were 87.0% and 84.8%, respectively (P < 0.001).
Furthermore, we aimed to determine whether a high level of hsa_circ_0014130 in patients was associated with clinicopathological parameters. As shown in Table 2, we found that the level of hsa_circ_0014130 expression showed no significant differences between groups of different genders (P = 0.513), ages (P = 0.239), smoking/ nonsmoking (P = 0.748), tumor sizes (P = 0.074) or pathologic types (P = 0.381). However, there were significant associations between the level of hsa_circ_0014130 expression and TNM stage (P = 0.001) or lymphatic metastasis (P = 0.004).
CircRNA-microRNA-mRNA co-expression network for the hsa_circ_0014130
As circRNA and miRNA sequences were aligned, and circRNAs interacted with miRNAs via miRNA response elements (MREs), 5 miRNAs with the highest mirSVR scores were identified for each differentially expressed circRNA using miRNA target-prediction software. Predicted Top-5 miRNAs regarding the top 5 upregulated and downregulated circRNAs including hsa_circ_0014130 were shown in Table 3. We assumed that hsa_circ_0014130 act as a miRNA sponge to regulate its circRNA-miRNA-mRNA network. We then predicted the target genes of Top-5 miRNAs by means of targetscan7.1 and mirdbV5. We generally accepted the overlapping results of two databases, and there were 333 target genes for hsa_circ_0014130 (Supplementary Fig. 4). Based on these analysis results, a total of 5 miRNAs and 333 mRNAs were predicted to have an interaction with hsa_circ_0014130 in this study. Cytoscape analysis of the circRNA-miRNA-mRNA interaction network of hsa_circ_0014130 indicated that miR-216a-3p exhibited the largest interaction network followed by miR-302a-3p, miR-892a, miR-493-5p, and miR-200c-5p (Fig. 4A). Since predicted target miRNAs including miR-216a-3p, miR-493-5p and miR-200c-5p, proved to be downregulated in cancer progression in previous research results, we utilized the public databases [circBase (http://www.circbase.org)] to screen for targeted miRNAs and the results based on seed sequence matching and specific base pairing displayed the three cancer-related miRNAs: miR-216a-3p, miR-493-5p and miR-200c-5p, had a binding site for hsa_circ_0014130 (Fig. 4B).
The predicted hsa_circ_0014130 targeted circRNA-miRNA-mRNA gene co-expression network based on sequence-pairing prediction. (A) The co-expression network was drawn with the cytoscape software. Five miRNAs and their mRNA target genes were found with overlapping results. As shown in this figure, miR-216a-3p exhibited the largest interaction network followed by miR-302a-3p, miR-892a, miR-493-5p, and miR-200c-5p. (B) Seed sequence matching predicted the direct interaction of hsa_circRNA_0014130 with the three cancer-related miRNAs: miR-216a-3p, miR-493-5p and miR-200c-5p.
Bioinformatics analysis of the predicted network genes for the hsa_circ_0014130
Gene Ontology (GO) analysis for hsa_circ_0014130 was performed to explore the functional roles of the top 10 significantly enriched target genes in terms of biological processes (BP) (Fig. 5), cellular components (CC) and molecular functions (MF) (Supplementary Fig. 5). From these results, we found that hsa_circ_0014130 showed a strong relationship with the regulation of metabolic process, the intracellular membrane-bounded organelle, DNA binding and its regulation of transcription, and so on. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis for hsa_circ_0014130 showed that there were the top 10 significantly enriched pathways including Transcriptional dysregulation in cancer, MAPK signaling pathway, Ubiquitin mediated proteolysis, TGF−beta signaling pathway and so on (Fig. 6). Especially, there were 12 target genes enrichment in MAPK signaling pathway (Supplementary Fig. 6). These processes and pathways were associated with human tumorigenesis and metastasis.
Gene ontology (GO) enrichment analysis for hsa_circ_0014130 in terms of biological processes (BP). The top 10 significantly enriched target genes and their scores (negative logarithm of P value) are listed as the x-axis and the y-axis, respectively. The horizontal axis represents the significant level of GOs.
KEGG pathway analysis for hsa_circ_0014130*. The top 10 significantly enriched pathways and their scores (negative logarithm of P value) were listed as the x-axis and the y-axis, respectively. The horizontal axis represented the significant level of pathways. *This image was obtained from KEGG (http://www.kegg.jp/kegg/kegg1.html).
Discussion
In recent years, with the rapid development and widespread application of RNA sequencing, researchers have found that many exonic transcripts can form circRNAs through non-linear reverse splicing or gene rearrangement. Moreover, they account for a large proportion of all spliced transcripts20. CircRNAs may arise from exons or introns21. An increasing number of researchers have begun to study potential functions of circRNAs in all kinds of diseases such as nervous system, cardiovascular system diseases and cancers7,8,22,23. Some altered circRNAs have been demonstrated to associate with tumor development, invasion, metastasis or patient prognosis9,10,11,12,13,14,15. Compared with other ncRNAs such as miRNAs and long noncoding RNAs (lncRNAs), circRNAs are highly conserved sequences and high degree of stability in mammalian cells4, these properties provide circRNAs with the potential to become ideal biomarkers for diagnosis of disease, including cancers16 and potential treatment target. Currently, although three circRNAs including cir-ITCH, hsa_circ_100876, hsa_circ_0013958 were studied in lung cancer17,18,19, the clinical value and role of circRNAs in NSCLC remained largely unknown.
In this study, we first investigated circRNA expression profile in human NSCLC by high-throughput circRNA microarray. Our results showed that circRNA expression in NSCLC tissues(n = 3) is significantly different from that in adjacent non-cancerous tissues (n = 3) (Fig. 1). Compared with non-cancerous tissues, our microarray data showed a total of 148 circRNAs were significantly upregulated and 23 were significantly downregulated in NSCLC tissues. Moreover, the distributions of differentially expressed circRNAs in human chromosomes showed that most circRNAs were transcribed from chr1, chr3, chr9, chr12, chr16 and chr19 (Fig. 1D). After validating the expression of 2 circRNAs (hsa_circ_0014130 and hsa_circ_0016760) in additional tissue samples (n = 10), only hsa_circ_0014130 was confirmed to be significantly upregulated in NSCLC (P = 0.002) (Fig. 2). However, the expression level of hsa_circ_0014130 was not consistent with that measured by microarray analysis, in which the hsa_circ_0014130 expression was significantly downregulated (Table 1). These results indicated that the validation of differentially expressed circRNAs in microarray analysis was an essential step of this screening study. Furthermore, to validate the study results, we expanded the sample size to detect the expression of hsa_circ_0014130 in additional tissue samples (n = 36). Our results showed that hsa_circ_0014130 was significantly upregulated in 82.6% (38/46) NSCLC tissues with average 3.34 fold compared with their adjacent non-cancerous tissues, and the AUC of hsa_circ_0014130 was up to 0.878 (Fig. 3). The ROC analyses confirmed that hsa_circ_0014130 had a high degree of specificity and sensitivity. More importantly, considering clinicopathological factors, we found that high expression levels of hsa_circ_0014130 in NSCLC were significantly associated with TNM stage and lymph node metastasis (Table 2), which are crucial factors in the evaluation of the prognosis of NSCLC. Taken together, these findings indicated that the circRNA 0014130 might be involved in carcinogenesis, progress and metastasis of NSCLC, might be used as a potential non-invasive biomarker of NSCLC and a new target for treatment of NSCLC.
CircRNAs represent a class of widespread ncRNAs, since they are capable of negatively regulating the effects of miRNAs upon their target mRNAs by directly competing for binding to shared miRNAs via MREs, thereby essentially functioning as ‘miRNA sponges’ within the cellular RNA network24,25. Compared with linear miRNA sponges, circRNAs have more miRNA binding sites and higher expression levels, and may be more effective in sequestering miRNAs26,27. Increasing evidence has indicated that the aberrant expression of circRNAs may promote cancer pathogenesis by absorbing cancer-associated miRNAs7. In the present study, to further understand the biological function of hsa_circ_0014130, the top 5 miRNAs with the highest mirSVRs were identified (including miR-216a-3p, miR-302a-3p, miR-892a, miR-493-5p and miR-200c-5p) (Table 3), and the hsa_circ_0014130-miRNA-gene network was predicted using TargetScan and miRanda. In essence, this network map diagramed a cellular RNA network consisting of hsa_circ_0014130 interacting with 5 miRNA nodes and 333 target genes (Fig. 4A). By searching the literatures from PubMed (https://www.ncbi.nlm.nih.gov/pubmed/), before Octomber 26, 2017, miR-216a-3p, miR-493-5p and miR-200c-5p were proved to be downregulated in cancer progression in previous research results. According to the prediction results of circBase28, we found that miR-216a-3p, miR-493-5p and miR-200c-5p might interact with hsa_circ_0014130 (Fig. 4B). As hsa_circ_0014130 might function as a potent miRNA sponge, we considered the regulatory network in which the three cancer-related miRNAs: miR-216a-3p, miR-493-5p and miR-200c-5p could participate.
It have been found that miR-216a-3p was significantly reduced in pancreatic ductal adenocarcinoma and could affect colorectal cancer cell proliferation by inhibiting COX-2 and ALOX5 expression29,30. miR-493-5p has been found to be deregulated in several tumours, such as breast cancer and myeloid leukemia31,32. It has been reported that the expression level of miR-493-5p was down-regulated in NSCLC patients and the overexpression of miR-493-5p significantly inhibited NSCLC cell proliferative capacity by suppressing the expression of oncogene ITGB133. Similar to miR-216a-3p and miR-493-5p, miR-200c-5p was reportedly downregulated in human hepatocellular carcinoma (HCC) and replenishing of miR-200c-5p suppressed the proliferation, migration and invasion of HCC cells by suppressing MAD2L134. Therefore, the decreased expression and inhibited function of miR-216a-3p, miR-493-5p and miR-200c-5p in cancer further support our hypothesis that hsa_circ_0014130 functions as a miRNA sponge to regulate the more comprehensive hsa_circ_0014130-miRNA-mRNA network.
In this study, we found that a large number of mRNAs might take part in this hsa_circ_0014130-miRNA-gene network described above, such as interleukin 7, insulin-like growth factor binding protein 1, transcription factor E2F7 and so on. So we functionally examined the target genes using GO and KEGG pathway analysis. GO enrichment analysis (Fig. 5) revealed that target genes were involved in the regulation of crucial metabolic processes and the transcription regulation of DNA binding, indicating that regulating these genes in the cellular response is of great importance during the development of NSCLC. Among the KEGG pathways (Fig. 6) found in this study, MAPK signaling pathway has been reported to be a key mediator of the growth, invasion, and migration of NSCLC cells35. Ubiquitin mediated proteolysis pathway has been found to promote its ubiquitin-proteasome-dependent degradation and consequently lead to radiation-induced apoptosis in NSCLC cells36. Since MAPK signaling pathway and ubiquitin mediated proteolysis pathway are strongly associated with NSCLC cell proliferation, invasion, and metastasis, we speculate that the hsa_circ_0014130-microRNA- mRNA axis may be the possible mechanism promoting the development of NSCLC, and it is worthwhile to further investigate the over-expressed hsa_circ_0014130 as an inhibitor of miRNA and its potential mechanism.
In conclusion, the present study revealed the expression profile of circRNAs in NSCLC tissues and demonstrated that circRNAs were aberrantly expressed in NSCLC. Our study validated the significant upregulation of hsa_circ_0014130 and analyzed the relationship of between this circRNA and clinical features in NSCLC patients, suggesting its potential involvement in NSCLC tumorigenesis and its potential use as a novel biomarker for NSCLC diagnosis. In the future, it’s necessary to explore the detailed molecular mechanisms by which hsa_circ_0014130 functions as miRNA sponges to regulate NSCLC occurrence and development.
Methods
Patients and specimens
The study included patients with NSCLC who underwent partial or total pulmonary lobectomy at the Department of Thoracic Surgery of Beijing Anzhen Hospital of Capital Medical University (China) between September 2016 and September 2017. NSCLC tissues were collected from the tumor surface, and the corresponding adjacent non-cancerous tissues were taken 5 cm from the edge of the cancer and contained no obvious tumor cells. After removal from the body, the specimens were preserved in liquid nitrogen within 5 minutes of excision. They were transported frozen to the laboratory for storage at −80 °C until use. For excluding confounding factors affecting the steady of patients’ circRNA profiling, none of the enrolled patients received chemotherapy, radiotherapy, or target therapy before they underwent surgery. All cases were diagnosed as NSCLC by two independent pathologists. Tumor histological grading and staging were based on the World Health Organization classification criteria and the seventh edition of the International Union against Cancer Tumor Node Metastasis (TNM) staging system. In total, 46 NSCLC tissues and 46 adjacent lung tissues were obtained from 46 patients (11 females and 35 males). For circRNA microarray analysis, 3 NSCLC tissues and corresponding 3 adjacent lung tissues were randomly selected. And other 46 NSCLC tissues and corresponding 46 adjacent tissues were prepared for qRT-PCR validation experiments. We obtained written informed consent from all patients. The study was approved by the Medical Ethics Commission of Capital Medical University. The study was in accordance with the provisions of Medical Ethics Commission of Capital Medical University. I confirmed that all methods were performed in accordance with the relevant guidelines and regulations.
Total RNA isolation and quality control (QC)
Total RNA was isolated from tumor and adjacent tissues using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. The purity and concentration of RNA samples were examined spectrophotometrically by absorbance measurements at 260 nm, 280 nm and 230 nm using the NanoDrop ND-1000 (Thermo Fisher Scientific, Wilmington, DE, USA). Specifically, OD260/OD280 ratios between 1.8 and 2.1 were deemed acceptable, while OD260/OD230 ratios of greater than 1.8 were deemed acceptable. RNA integrity and contamination was tested by denaturing agarose gel electrophoresis. RNA was prepared and stored at −80 °C for the validation experiments.
Sample Preparation and Microarray Detection
Total RNA from 6 samples (3 NSCLC samples and 3 paired adjacent non-cancerous samples) was treated with Rnase R (Epicentre, Madison, WI, USA) to remove linear RNAs and to enrich circRNAs. Then, the enriched circRNAs were amplified and transcribed into fluorescent circRNA utilizing a random priming method according to Arraystar Super RNA Labeling Kit’s instructions (Arraystar, Rockville, MD, USA). The labeled circRNAs were purified by RNeasy Mini Kit (Qiagen, Germany). The concentration and specific activity of the labeled circRNAs (pmol Cy3/μg circRNAs) were measured by NanoDrop ND-1000. The labeled circRNAs were hybridized onto the Arraystar Human circRNA Arrays (8 × 15 K, Arraystar). The slides were incubated for 17 h at 65 °C in an Agilent Hybridization Oven (Agilent Technologies, Santa Clara, CA, USA). The hybridized microarrays were washed, fixed, and scanned with an Agilent G2505C Scanner (Agilent Technologies, Santa Clara, CA, USA).
CircRNA Microarray Data Analysis
Scanned images were imported into Agilent Feature Extraction software (version 11.0.1.1, Agilent) for raw data extraction. Following quantile normalization of the data using a log2 ratio (R software package, version 3.1.2), we performed a low-intensity filtering. The circRNAs with at least 3 out of 6 samples flagged in “P” or “M” (“all targets value”) were retained for further analysis. The Box Plot was drawn to visualize the distribution of the intensities from all datasets after normalization. To determine whether circRNA profiles were informative with regards to tissue type, an unsupervised, hierarchical cluster analysis was conducted based on the circRNA expression levels in 3 NSCLC specimens and adjacent lung tissues. Then, fold-change filtering and Student’s t-testing were applied to identify differential circRNA between NSCLC tumors versus matching adjacent lung tissues. Specifically, only the circRNAs that exhibited fold-changes of greater than 2.0 and Student’s t-test p-values of less than 0.05 were identified as significantly differential circRNAs. Volcano plot was used to visualize the significantly differential circRNAs between between NSCLC and non-cancerous tissues.
qRT-PCR validation of Candidate circRNAs
To validate the expression profiles of circRNAs in three matched pairs of NSCLC and adjacent specimens, considering the group raw signal intensity (>500.0), the significant difference of raw signal intensity (>100.0), the fold change of expression(FC >2.0), and that circRNAs whose target miRNAs predicted in the microarray analysis proved to be in related with cancer progression in previous research results, 2 circRNAs (hsa_circ_0014130 and hsa_circ_0016760) were selected for validation study by qRT-PCR as follows. After RNA extraction from NSCLC tumors and matching adjacent lung tissues, cDNA was synthesized with a reverse transcriptase according to the kit’s instructions (SuperScript First-Strand Synthesis System for RT-PCR, Invitrogen, Carlsbad, CA, USA). Divergent primers of hsa_circ_0014130 and hsa_circ_0016760 were designed with Primer Premier software version 5.0 (Premier Biosoft International, Palo Alto, CA, USA), ensuring that circRNAs were amplified through head-to-tail splicing4. β-actin, a housekeeping gene, was used as internal standard for normalization. Primers were synthesized by Bioligo Biotech (Shanghai, China). The sequences of hsa_circ_0014130, hsa_circ_0016760 and β-actin primers were as follows: 5′ AGATTCCCTAACCTCAACCAGA3′ (forward) and 5′ CGAATGTTCTTGCCACCTGC3′ (reverse) for hsa_circ_0014130; 5′ TGCATTGGTGCTCAGAAGCG3′ (forward) and 5′ TCTGTTCCTGGGTCTGTGTGC3′ (reverse) for hsa_circ_0016760; and 5′ GTGGCCGAGGACTTTGATTG3′ (forward) and 5′ CCTGTAACAACGCATCTCATATT3′ (reverse) for β-actin. PCR was conducted in a 10-μl reaction volume consisting of the following: 2.0 μl cDNA, 5 μl 2 × PCR Master mix (PCR Master Mix, Arraystar, Rockville, MD, USA), 0.5 μl primer forward (5 μM), 0.5 μl primer reverse (5 μM), and 2.0 μl H2O. The qPCR reaction was performed on an ABI QuantStudioTM 5 System (Applied Biosystems, DE, USA) as follows: initial denaturation at 95 °C for 10 min, 40 cycles of amplification at 95 °C for 10 sec, annealing and extension at 60 °C for 1 min. Amplification products were analyzed by 1.5% agarose gel electrophoresis, stained with 1∶10000 dilution of GelRed Nucleic Acid gel stain (Biotium, CA, USA) and visualized under ultraviolet illumination for band size consistency. All the experiments were conducted in triplicate. The data were analyzed by using the comparative cycle threshold (2−ΔΔCt) 19 to represent a relative expression level of circRNAs.
Cloning and sequencing of qRT-PCR products
Two qRT-PCR products of hsa_circ_0014130 in NSCLC tumor and adjacent lung tissue were purified using PCR Product Purification Kit (Ensurebio Biotech Co., Ltd., Shanghai, China), and then cloned using PMD18-T Vector Cloning Kit (Takara Bio Inc., Dalian, China) following the manufacturer’s instructions. DNA sequencing was performed by Life Technologies (Thermo Fisher Scientific, Wilmington, DE, USA).
Prediction of circRNA-miRNA-mRNA co-expression network for the hsa_circ_0014130
The homemade computer program of Arraystar (Rockville, MD, USA) based on TargetScan37 and miRanda38 was applied to predict miRNA targets of circRNAs and the circRNA/miRNA interaction. To focus the targeted miRNA profile, the miRNA support vector regression (mirSVR) algorithm was used to score and rank the efficiency of the predicted miRNA targets39. Accordingly, for each circRNA, we identified 5 miRNAs with the highest mirSVR score to establish a “Top-5” circRNA-miRNA network (1 circRNA connecting to 5 miRNAs). To further elucidate correlations between circRNAs and miRNA, Cytoscape 3.01 was then used to diagram the potential map of the circRNA/miRNA/mRNA interaction network of hsa_circ_0014130. The size of each node in the network map represents the number of putative miRNA functionally connected to hsa_circ_0014130.
Prediction of circRNA-miRNA-target gene associations for the hsa_circ_0014130
To gain further insights into the functions of hsa_circ_0014130, putative target genes of miRNAs were predicted by two databases including targetscan7.1 (http://www.targetscan.org/vert_71/) and mirdbV5 (http://mirdb.org/miRDB/). GO analysis was performed to explore the functional roles of target genes in terms of biological processes, cellular components and molecular functions. Biological pathways defined by KEGG40, Biocarta and Reactome (http://www.genome.jp/kegg/) were identified by Database for Annotation, Visualization and Integrated Discovery (DAVID; http://www.david.abcc.ncifcrf.gov/). The predicted gene functions of the hsa_circ_0014130 in the circRNA-miRNA-target gene associations were annotated using GO and KEGG pathway analysis.
Statistical Analysis
All results are expressed as the mean ± SD. All statistical data were analyzed using Statistical Program for Social Sciences (SPSS) 19.0 software (SPSS, Chicago, IL, USA). GraphPad Prism 5.0 (GraphPad Software, La Jolla, CA) was used to plot all graphs. The significance of qRT-PCR validation between the NSCLC tissue group and the adjacent tissue group was analyzed by the Student t test for paired data. The correlations between expression levels of hsa_circ_0014130 and various clinicopathological parameters of NSCLC were further analyzed by the Student t test or one-way analysis of variance (ANOVA). All tests were 2-sided, and P < 0.05 was considered statistically significant.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA: a cancer journal for clinicians 65, 87–108, https://doi.org/10.3322/caac.21262 (2015).
Ettinger, D. S. et al. Non-Small Cell Lung Cancer, Version 5.2017, NCCN Clinical Practice Guidelines in Oncology. Journal of the National Comprehensive Cancer Network: JNCCN 15, 504–535, https://doi.org/10.6004/jnccn.2017.0050 (2017).
Hentze, M. W. & Preiss, T. Circular RNAs: splicing’s enigma variations. The EMBO journal 32, 923–925, https://doi.org/10.1038/emboj.2013.53 (2013).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338, https://doi.org/10.1038/nature11928 (2013).
Chen, L. L. The biogenesis and emerging roles of circular RNAs. Nature reviews. Molecular cell biology 17, 205–211, https://doi.org/10.1038/nrm.2015.32 (2016).
Chen, Y., Li, C., Tan, C. & Liu, X. Circular RNAs: a new frontier in the study of human diseases. Journal of medical genetics 53, 359–365, https://doi.org/10.1136/jmedgenet-2016-103758 (2016).
Li, J. et al. Circular RNAs in cancer: novel insights into origins, properties, functions and implications. American journal of cancer research 5, 472–480 (2015).
Wang, Y. et al. Circular RNAs in human cancer. Molecular cancer 16, 25, https://doi.org/10.1186/s12943-017-0598-7 (2017).
Xia, W. et al. Circular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferation. Scientific reports 6, 35576, https://doi.org/10.1038/srep35576 (2016).
Li, P. et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clinica chimica acta; international journal of clinical chemistry 444, 132–136, https://doi.org/10.1016/j.cca.2015.02.018 (2015).
Qin, M. et al. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer biomarkers: section A of Disease markers 16, 161–169, https://doi.org/10.3233/cbm-150552 (2016).
Shang, X. et al. Comprehensive Circular RNA Profiling Reveals That hsa_circ_0005075, a New Circular RNA Biomarker, Is Involved in Hepatocellular Crcinoma Development. Medicine 95, e3811, https://doi.org/10.1097/md.0000000000003811 (2016).
Wang, X. et al. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. International journal of clinical and experimental pathology 8, 16020–16025 (2015).
Xuan, L. et al. CircularRNA: a novel biomarker for progressive laryngeal cancer. American journal of translational research 8, 932–939 (2016).
Song, X. et al. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic acids research 44, e87, https://doi.org/10.1093/nar/gkw075 (2016).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell research 25, 981–984, https://doi.org/10.1038/cr.2015.82 (2015).
Wan, L. et al. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/beta-Catenin Pathway. BioMed research international 2016, 1579490, https://doi.org/10.1155/2016/1579490 (2016).
Yao, J. T. et al. Over-expression of CircRNA_100876 in non-small cell lung cancer and its prognostic value. Pathology, research and practice 213, 453–456, https://doi.org/10.1016/j.prp.2017.02.011 (2017).
Zhu, X. et al. hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. The FEBS journal 284, 2170–2182, https://doi.org/10.1111/febs.14132 (2017).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PloS one 7, e30733, https://doi.org/10.1371/journal.pone.0030733 (2012).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Molecular cell 51, 792–806, https://doi.org/10.1016/j.molcel.2013.08.017 (2013).
Lu, D. & Xu, A. D. Mini Review: Circular RNAs as Potential Clinical Biomarkers for Disorders in the Central Nervous System. Frontiers in genetics 7, 53, https://doi.org/10.3389/fgene.2016.00053 (2016).
Wang, K. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. European heart journal 37, 2602–2611, https://doi.org/10.1093/eurheartj/ehv713 (2016).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–352, https://doi.org/10.1038/nature12986 (2014).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388, https://doi.org/10.1038/nature11993 (2013).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome biology 15, 409, https://doi.org/10.1186/s13059-014-0409-z (2014).
Wilusz, J. E. & Sharp, P. A. Molecular biology. A circuitous route to noncoding RNA. Science (New York, N.Y.) 340, 440–441, https://doi.org/10.1126/science.1238522 (2013).
Glazar, P., Papavasileiou, P. & Rajewsky, N. circBase: a database for circular RNAs. RNA (New York, N.Y.) 20, 1666–1670, https://doi.org/10.1261/rna.043687.113 (2014).
Yonemori, K. et al. The microRNA expression signature of pancreatic ductal adenocarcinoma by RNA sequencing: anti-tumour functions of the microRNA-216 cluster. Oncotarget 8, 70097–70115, https://doi.org/10.18632/oncotarget.19591 (2017).
Wang, D., Li, Y., Zhang, C., Li, X. & Yu, J. MiR-216a-3p inhibits colorectal cancer cell proliferation through direct targeting COX-2 and ALOX5. Journal of cellular biochemistry 119, 1755–1766, https://doi.org/10.1002/jcb.26336 (2018).
Zhao, L. et al. miR-493-5p attenuates the invasiveness and tumorigenicity in human breast cancer by targeting FUT4. Oncology reports 36, 1007–1015, https://doi.org/10.3892/or.2016.4882 (2016).
Bhutra, S., Lenkala, D., LaCroix, B., Ye, M. & Huang, R. S. Identifying and validating a combined mRNA and microRNA signature in response to imatinib treatment in a chronic myeloid leukemia cell line. PloS one 9, e115003, https://doi.org/10.1371/journal.pone.0115003 (2014).
Liang, Z. et al. High expression of miR-493-5p positively correlates with clinical prognosis of non small cell lung cancer by targeting oncogene ITGB1. Oncotarget 8, 47389–47399, https://doi.org/10.18632/oncotarget.17650 (2017).
Li, Y., Bai, W. & Zhang, J. MiR-200c-5p suppresses proliferation and metastasis of human hepatocellular carcinoma (HCC) via suppressing MAD2L1. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie 92, 1038–1044, https://doi.org/10.1016/j.biopha.2017.05.092 (2017).
Wei, C. H. et al. MicroRNA-330-3p promotes cell invasion and metastasis in non-small cell lung cancer through GRIA3 by activating MAPK/ERK signaling pathway. Journal of hematology & oncology 10, 125, https://doi.org/10.1186/s13045-017-0493-0 (2017).
Kim, E. J. & Juhnn, Y. S. Cyclic AMP signaling reduces sirtuin 6 expression in non-small cell lung cancer cells by promoting ubiquitin-proteasomal degradation via inhibition of the Raf-MEK-ERK (Raf/mitogen-activated extracellular signal-regulated kinase/extracellular signal-regulated kinase) pathway. The Journal of biological chemistry 290, 9604–9613, https://doi.org/10.1074/jbc.M114.633198 (2015).
Enright, A. J. et al. MicroRNA targets in Drosophila. Genome biology 5, R1, https://doi.org/10.1186/gb-2003-5-1-r1 (2003).
Pasquinelli, A. E. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nature reviews. Genetics 13, 271–282, https://doi.org/10.1038/nrg3162 (2012).
Betel, D., Koppal, A., Agius, P., Sander, C. & Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome biology 11, R90, https://doi.org/10.1186/gb-2010-11-8-r90 (2010).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30, https://doi.org/10.1093/nar/28.1.27 (2000).
Acknowledgements
This study was supported by Talent Training Project of High Level Health Technology from Beijing Health System (NO. 2015-3-052 to Xiaoli Zeng) and National Natural Science Foundation of China (NO. 81500332 to Xiaoli Zeng). Microarray experiments and bioinformatics analysis were performed by KangChen Biotech (Shanghai, China).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments; S.Z., X.Z., S.O. and H.Y. Performed the experiments: S.Z., X.Z., T.D., L.G. and Y.L. Analyzed the data: S.Z., X.Z., T.D. and Y.L. Contributed reagents/ materials/analysis tools: S.Z., X.Z. and L.G. Wrote the paper: S.Z., X.Z. and T.D. Access to full-text articles: X.Z.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, S., Zeng, X., Ding, T. et al. Microarray profile of circular RNAs identifies hsa_circ_0014130 as a new circular RNA biomarker in non-small cell lung cancer. Sci Rep 8, 2878 (2018). https://doi.org/10.1038/s41598-018-21300-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21300-5
This article is cited by
-
Circ-10720 as a ceRNA adsorbs microRNA-1238 and modulates ZEB2 to boost NSCLC development by activating EMT
European Journal of Medical Research (2024)
-
Circulating nucleic acids as liquid biopsies for disease prediction, screening and diagnosis
Science China Chemistry (2023)
-
CircRNA hsa_circ_0014130 function as a miR-132-3p sponge for playing oncogenic roles in bladder cancer via upregulating KCNJ12 expression
Cell Biology and Toxicology (2022)
-
The role of PIP5K1A in cancer development and progression
Medical Oncology (2022)
-
Circ_0016760 Serves as a Cancer Promoter in Non-small Cell Lung Cancer Through miR-876-3p/NOVA2 Axis
Biochemical Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.