Abstract
Aldehyde dehydrogenase enzymes (ALDHs) catalyze the oxidation of aliphatic and aromatic aldehydes to their corresponding carboxylic acids using NAD+ or NADP+ as cofactors and generating NADH or NADPH. Previous studies mainly focused on the ALDH role in detoxifying toxic aldehydes but their effect on the cellular NAD(P)H contents has so far been overlooked. Here, we investigated whether the ALDHs influence the cellular redox homeostasis. We used a double T-DNA insertion mutant that is defective in representative members of Arabidopsis thaliana ALDH families 3 (ALDH3I1) and 7 (ALDH7B4), and we examined the pyridine nucleotide pools, glutathione content, and the photosynthetic capacity of the aldh mutants in comparison with the wild type. The loss of function of ALDH3I1 and ALDH7B4 led to a decrease of NAD(P)H, NAD(P)H/NAD(P) ratio, and an alteration of the glutathione pools. The aldh double mutant had higher glucose-6-phosphate dehydrogenase activity than the wild type, indicating a high demand for reduced pyridine nucleotides. Moreover, the mutant had a reduced quantum yield of photosystem II and photosynthetic capacity at relatively high light intensities compared to the wild type. Altogether, our data revealed a role of ALDHs as major contributors to the homeostasis of pyridine nucleotides in plants.
Similar content being viewed by others
Introduction
Aldehyde dehydrogenase enzymes (ALDHs) represent an evolutionary conserved protein superfamily present in nearly all organisms1,2, and up to now, 16 ALDH genes have been identified in Arabidopsis thaliana, and are classified in 10 protein families2. Several Arabidopsis ALDHs from family 3, 7 and 10 have been shown to play a role in plant stress responses and development3,4,5. For example, the overexpression of ALDH3I1 and ALDH7B4 conferred abiotic stress tolerance to Arabidopsis and tobacco plants6,7,8, and the reduced epidermal fluorescence1 (ref1) phenotype, which resulted from a mutation in the Arabidopsis ALDH2C4 gene (At3g24503), affected the synthesis of ferulic acid and sinapic acid in the phenylpropanoid pathway9,10. ALDH2B4, a member of the protein family 2, was shown to be involved in the pyruvate dehydrogenase bypass11. These examples demonstrate a role for some plant ALDHs in growth, development and response to environmental stresses (reviewed in ref.4). However, the mere transformation of the aldehyde into carboxylic acid often fails to explain how some ALDH genes are involved in complex phenotypes such as the nuclear restoration of cytoplasmic male sterility12,13, submergence tolerance14, seed maturation15,16, and cell proliferation17. Although the ALDHs use NAD+ or NADP+ as cofactors and generate NADH or NADPH, studies on both plant and animal ALDHs have only focused on their role in detoxifying toxic aldehydes, and have so far overlooked the potential contribution of the ALDH activity to the cellular NAD(P)H pools and NAD(P)H/NA(D)P ratio. In this study, we examined this hypothesis and found that the NAD(P)H pools were altered in the Arabidopsis aldh mutants particularly under conditions for high demand of reducing power.
Results
The disruption of ALDH3I1 and ALDH7B4 decreased the cellular NAD(P)H contents and altered the NAD(P)H/NAD(P) ratio
To investigate whether the ALDHs influence the cellular redox, we examined their contribution to the pyridine nucleotide NAD(P) and NAD(P)H pools by using a double T-DNA insertion mutant (KO6/62) of A. thaliana that is defective in the ALDH7B4 and ALDH3I1 genes. ALDH3I1 and ALDH7B4 were previously shown to be localized in the chloroplast and the cytosol of Arabidopsis thaliana cells, respectively, and their expression level increases in response to abiotic stresses4,18. Stress-triggered induction of ALDH7B4 and ALDH3I1 was impaired in the single T-DNA insertion lines KO6 and KO62, respectively, and both KO6 and KO62 plants accumulated higher levels of reactive oxygen species (ROS) and malondialdehyde (MDA) than the wild type7. In this study, we verified that the double mutant KO6/62 does not express the ALDH3I1 and ALDH7B4 genes. Although a transcript was still amplifiable from the KO6 mutant by using primers specific for ALDH7B4, the ALDH7B4 protein level in this line was drastically reduced, as shown for seed protein extracts (Figure S1). Next, we measured the levels of the oxidized and reduced forms of the pyridine nucleotides NAD, NADP, NADH, and NADPH in leaves of six-week-old plants (Table 1). The levels of the reduced pyridine nucleotides were lower in all aldh mutants than the wild type. Compared to the wild type, NADH and NADPH were reduced by 27% and 79% in the double mutant KO6/62, respectively. In comparison, there were above 50% increase of NAD, the preferred substrate for the plant ALDHs over NADP, in the single aldh mutant KO6 and KO62, and 27% increase in double mutant KO6/62 compared to WT. In contrast to KO6 that is defective in the cytosolic ALDH7B4, the WT and the mutants KO62 and KO6/62 showed similar levels of NADP. Moreover, the NAD(P)H/NAD(P) ratio significantly decreased by 32% (from 1.9 to 1.3) in the double mutant KO6/62 compared to WT (Fig. 1A), mainly as a result of a low concentration of the NAD(P)H pools in the ALDH single and double mutants compared to WT.
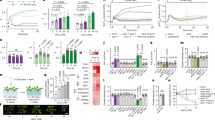
Levels of pyridine nucleotides and glucose-6-phosphate dehydrogenase (G6PDH) activity in WT and aldh mutants. (A) Ratios of total reduced to oxidised pyridine nucleotides in 6-week-old plants. (B) Total G6PDH activity from 6-week-old plants. Data in A and B represent mean values ± SE (n = 9 plants). (C) Ratios of total reduced to oxidised pyridine nucleotides in 2-week-old seedlings. (D) Total G6PDH activity from 2-week-old seedlings extracts. Data in (C and D) represent mean values ± SE (n = 4 pools of seedlings). For the measurements, seedlings were grown on half-MS agar plates for 7 days then transferred on fresh plates supplemented with 0 mM (control) or 30 mM of either potassium nitrate (shown as Nitrate) or ammonium chloride (shown as Ammonium). Seedlings were harvested after 7 days of additional growth. Single asterisks in (A and B) show significant differences between control and treatments whereas the double asterisk in (C) indicates significant differences between the accumulation of reduced pyridine nucleotides in WT and KO6/62 on nitrate (P < 0.05; Student’s t test).
Several studies reported about the sensitivity of the plant glucose-6-phosphate dehydrogenases (G6PDH) to the intracellular level of reduced pyridine nucleotides NADH and NADPH, which manifests by an increase of the G6PDH activity in response to a low cellular level of reduced pyridine nucleotides19,20,21. To verify the alteration of NAD(P)H contents found in the aldh mutant, we then measured the total G6PDH activity in the mutant and WT leaf protein extracts. We found that the G6PDH activity was about 20% higher in the double mutants KO6/62 mutants than in WT (Fig. 1B), confirming the well-known positive regulation of the G6PDH enzyme by low NAPH/NADP ratio22. In another experiment, we leveraged the fact that nitrate assimilation by the plant requires substantial amounts of NAD(P)H to further verify the alteration of NAD(P)H contents in the aldh mutant. We thus analyzed the pools of NAD(P) and NAD(P)H in wild-type and KO6/62 seedlings grown on a medium supplemented with nitrate. The rationale was that growing the seeds on a medium supplemented with nitrate would increase the demand of NAD(P)H, and allow any defect in the NAD(P)H pools in the seedlings to become visible. In agreement with this, the NAD(P)H/NAD(P) ratio increased in the WT and in the KO6/62 mutant when grown on agar plate supplemented with nitrate or ammonium. However, the increase was 18% less in the aldh double mutant KO6/62 (from 0.83 to 0.94–13%) than in WT (from 1.22 to 0.84–31%) in medium supplemented with nitrate (Fig. 1C), which shows that the supply or the consumption of NAD(P)H are altered in the aldh mutant. No difference was seen between the aldh mutant and WT when they were grown in a medium supplemented with ammonium, suggesting that the supply of NAD(P)H by the ALDH could be more relevant to nitrate assimilation than ammonium assimilation. As in the adult plants, we measured the G6PDH activity in the seedlings grown on the medium supplemented with nitrate. The G6PDH activity was 39% higher in the mutant than in WT, indicating a high demand for NAD(P)H in the aldh double mutant compared to WT (Fig. 1D). Similarly to NAD(P)/NAD(P), there were no difference in the G6PDH activity between the aldh mutant and WT when grown in a medium supplemented with ammonium. These results, altogether, indicate an alteration of NAD(P)H pool in the aldh mutant compared to WT. Nevertheless, the change in the NAD(P)H content was only observable in the leaves of adult mutant plants and not in the seedlings under normal growth conditions, as it can be seen by comparing Fig. 1A and B with the control condition shown in Fig. 1C and D, respectively.
Disruption of ALDH3I1 and ALDH7B4 affects Glutathione Metabolism
We hypothesized that the low level of reduced pyridine nucleotides in the ALDH double mutant might also affect the redox state of the glutathione pools given that the turnover of antioxidant pools such as glutathione and ascorbate depends on NAD(P)H availability23,24. To verify this hypothesis the glutathione pools were compared in WT and KO6/62 plants. The levels of the total glutathione (reduced + oxidized) and of the reduced to oxidized glutathione ratio (GSH/GSSG) were reduced by 20% and 33% in KO6/62 compared to WT, respectively (Fig. 2). This would reflect a change in the cellular redox homeostasis following the inactivation of the ALDH3I1 and ALDH7B4 genes.
Glutathione pools in WT and KO6/62 plants. (A) Total glutathione denotes reduced plus oxidised nanomole (nmol) glutathione per gram leaf fresh weight (FW). (B) GSH/GSSG denotes the ratio of reduced to oxidised glutathione pools. Shown are the mean values ± SE (n = 4 pools of leaves each from 3 plants). Asterisks indicate significant differences between WT and KO6/62 plants (P < 0.05; Student’s t test).
Disruption of ALDH3I1 and ALDH7B4 affects Photosynthesis
The low ratio of NAD(P)H/NAD(P) in the leaves of the double mutant of ALDHs led us to examine their photosynthetic capacity in comparison to that of WT. At light intensities above 200 µmol m−2 s−1, we found that the quantum yield of photosystem II (PSII) and the electron transport rate were low in KO6/62 plants compared to WT (Fig. 3A,B), but there was no difference in the maximum quantum yield of PSII (Fv/Fm) of the two genotypes (Fig. 3C). The transport of electrons is coupled with that of protons and results in the formation of a trans-thylakoid proton gradient25,26. An increase in the proton gradient in turn downregulates the electron transport and upregulates the non-photochemical quenching (NPQ) mechanism of the photosystem II27. We found that the NPQ was similar between the WT and the mutant at light intensities less than 500 µmol m−2 s−1. At higher light intensities, the NPQ increased more slowly in the KO6/62 mutant than in WT (Fig. 3D), suggesting a dysfunction in the non-photochemical mechanism, likely the xanthophyll cycle, in the ALDH double mutant.
Chlorophyll fluorescence analysis in WT and the aldh mutant KO6/62. (A) Quantum yield of photosystem II (ΦPSII); (B) Electron transport rate (ETR); (C) Maximum quantum efficiency of the photosystem II (Fv/Fm); (D) Non-photochemical quenching (NPQ). Shown are mean values ± SE (n = 9 plants). Asterisks show significant differences according to Student’s t test (P < 0.05) between WT and the mutant KO6/62.
Discussion
ALDHs oxidize aliphatic and aromatic aldehydes to their corresponding carboxylic acids using NAD+ or NADP+ as cofactors4. Given that the oxidation of aldehydes by ALDHs generates reducing equivalents in the form of NADH or NADPH, we investigated how the ALDHs may influence the cellular redox homeostasis. For this purpose, we analyzed knockout mutants of ALDHs and found that the suppression of ALDHs resulted in the accumulation of the oxidized pyridine nucleotides and a decrease of the reduced forms. This was associated with a decrease in the NAD(P)H/NAD(P) ratio and an increase of the G6PDH activity in KO6/62 adult leaves. The NAD(P)H/NAD(P) ratio and the G6PDH activity were however similar in the WT and KO6/62 seedlings under control conditions (Fig. 1C,D), which indicates a contribution of ALDH-derived NAD(P)H to the homeostasis of pyridine nucleotides in the adult leaves of A. thaliana. In the plant cell cytosol, enzymes including the NADP-dependent Isocitrate Dehydrogenase (NADP-ICDH), Glucose 6-Phosphate Dehydrogenase (G6PDH), non-phosphorylating glyceraldehyde-3-phosphate dehydrogenase (GAPN), and the NADP-Malic Enzyme2 (NADP-ME2) may contribute to the NAD(P)H content. However, the NADP(H) pools were practically not affected in the Arabidopsis single mutants defective in either NADP-ICDH or NADP-ME2 enzymes28,29. With regards to this, the alteration of NAD(P)H in the aldh mutant thus indicates that the ALDHs show less redundancy with other enzymes producing NADPH for the cell redox homeostasis.
Pyridine nucleotides are important redox sensors, since they are associated with most reactive oxygen species (ROS)-detoxifying enzymes. ROS production has been demonstrated to be influenced by the NAD(P)H/NAD(P) ratio in the chloroplast or the mitochondria30,31,32,33. Indeed, the turnover of oxidized antioxidant pools such as glutathione and ascorbate depends on availability of NAD(P)H23,24. Our observation that the NADP(H)/NAD(P) ratio decreased in parallel with the reduced to oxidised glutathione ratio indicate that ALDHs would significantly contribute to the reduced glutathione cellular content by providing NADPH. It is noteworthy that both the total and the reduction state of glutathione decreased in the aldh mutant whereas a decrease in the reduction state of glutathione led to an increased total pool in the CATALASE2 mutant (cat2) defective in oxidant processing and cat2gr1 double mutant defective in both oxidant processing and glutathione reduction by GLUTATHIONE REDUCTASE1 (GR1) in Arabidopsis34. Given that the level of lipid-peroxidation increased in the aldh mutants4,7, a rapid consumption of glutathione via Michael adduct formation with aldehydes would affect both the glutathione content and the cell redox status. Indeed, aldehydes including 4-hydroxynonenal (HNE) can be conjugated to glutathione via GLUTATHIONE S-TRANSFERASE (GST, EC 2.5.1.18) to form HNE-GST conjugates, and several GST isoforms were found to be induced in pumpkin (Cucurbita maxima Duch.) by aldehydes35,36. Thus, the decrease of total glutathione could well be due to both the altered NADPH and increased GST conjugation with the aldehydes that accumulate in the aldh mutant. Unfortunately, the amount of HNE-GST and aldehyde-GST conjugates in the aldh mutant were not quantifiable by the method of glutathione detection used in this study. This information will be very important to elucidate the fate of NAD(P)H generated by ALDHs. As shown for the cytosolic NADP-ICDH28, we speculate that this may serve as substrate either to GR1 for reducing oxidised glutathione or to the NADPH oxidase encoded by AtRbohF for the generation of oxidant under certain conditions. With regards to this, the limited increase of NAD(P)H/NAD(P) in the KO6/62 mutant compared to WT when grown on agar plate supplemented with nitrate could imply that NAD(P)H-derived from the ALDH activity partly support nitrate assimilation. In fact, the NAD(P)H/NAD(P) ratio was similar in WT and KO6/62 under control conditions and on medium supplemented with ammonium but not on medium supplemented with nitrate (Fig. 1). Although the increase of G6PDH activity in the KO6/62 seedlings could compensate for the reduced NAD(P)H supply in the mutant, this was not sufficient, which suggests that the ALDH activities may have a pivotal role, besides photosynthesis, in contributing to the reducing power required for the nitrate assimilation. We found that the efficiency of photosynthesis decreased in the aldh mutant KO6/KO62 compared to WT, which could also explain the diminution of NAD(P)H. It is however unclear how the mutation of the two ALDH genes might alter photosynthesis under control conditions. The reduced quantum yield of PSII and the increase in NPQ of the aldh mutant KO6/62 in the absence of photoinhibition (Fig. 3D) at high-light conditions suggests a dysfunction of the xanthophyll cycle, which most likely result from the altered NAD(P)H contents in the mutant plants. While an analysis of photosynthetic pigments is required to confirm this thesis, it is noteworthy that the NAD levels were higher in the aldh mutants than in WT (Table 1). Such an increase of NAD was also observed in Arabidopsis nadk2 mutant deficient in the chloroplast NAD kinase (EC 2.7.1.23) that phosphorylates NAD+ to NADP+ 37,38,39, and in which a low NAD(P)H/NAD(P) ratio was demonstrated to cause the inhibition of zeaxanthin epoxidase (EC 1.14.13.90) and the accumulation of zeaxanthin. A high level of zeaxanthin decreases the quantum yield of energy transfer from the antenna complex to the PSII reaction center, lowering thereby the photosynthetic yield. Previous reports also showed that both intrathylakoid pH and NADPH control the epoxidation reaction in the xanthophyll cycle40,41. The altered NAD(P)H/NAD(P) following an increase of NAD+ and a decrease of NAD(P)H in the aldh mutant has likely modified the redox state of the photosynthetic pigments, thereby affecting photosynthesis efficiency as shown in Fig. 3. Concurrently, several enzymes of the “dark reactions” of photosynthesis can be inactivated by carbonylation with the aldehydes that accumulate in the aldh mutant4,7,42. For instance, the Calvin cycle enzymes phosphoribulokinase, glyceraldehyde-3-phosphate dehydrogenase, fructose-1,6-bisphophatase, sedoheptulose-1,7-bisphosphatase, aldolase, and Rubisco were irreversibly inactivated by acrolein. Acrolein treatment caused a rapid drop of the glutathione pool, prior to the inactivation of photosynthesis. GSH exogenously added to chloroplasts suppressed the acrolein-induced inactivation of photosynthesis, but ascorbic acid did not show such a protective effect43,44. Thus, as a result of the mutation of ALDHs, aldehydes can accumulate and inhibit photosynthesis by depleting GSH and inactivating multiple enzymes in the Calvin cycle.
In conclusion, we have showed that the mutation of the ALDH3I1 and ALDH7B4 altered the cellular contents of NAD(P)H, the total and reduction state of glutathione, and decreased the efficiency of photosynthesis in A. thaliana. This is in agreement with the interdependency between the pyridine nucleotides, cellular redox, and photosynthesis and indicates a role of ALDHs as major contributors to the cellular redox homeostasis in A. thaliana.
Methods
Plant Materials and Growth Conditions
Arabidopsis thaliana (ecotype Col-0) was used throughout this work. Wild-type (WT) and transgenic plants were grown on half-strength-Murashige and Skoog (MS) agar plates45 or soil under white light. The soil was a 3:1 mixture of potting soil (Rolfs Gärtnereinkauf, Siegburg, Germany) and vermiculite. The intensity of the light was about 150 μmol m−2 s−1 with 8/16 h day/night-cycle at 22 to 24 °C. Leaves of fourteen-day- and six-week-old plants were immediately frozen in liquid nitrogen and used for analyses. The harvest for all samples was four hours after the light was switched on, as this should reflect stable day metabolites.
Isolation of Homozygous T-DNA Insertion Mutants
Single mutants KO6 (SALK_143309) for ALDH7B4 (At1g54100) and KO62 (nos. 16843)46 for ALDH3I1 (At4g34240) were previously described7. Similarly, the single mutant KO69 (nos. 22184)46 for ALDH3H1 (At1g44170) was described and referred as 3h1-B in Missihoun et al.47. The single mutant KO76 (SALK_091250) for ALDH3F1 (At4g36250) carries a T-DNA insertion in the fourth exon. Double mutants KO6/62, KO6/76, KO6/69, and KO69/76 were obtained by crossing the corresponding homozygous single mutants. The second seed generation derived from the crossing was screened to isolate homozygous double mutants. Homozygous T-DNA insertion mutants were identified by two consecutive PCR assays. The first PCR assay involved the use of two gene-specific primers. Gene-specific primers comprised 5′-TTGCCAAGGGTTTTTCTCCTGCCAG-3′ and 5′-TAAGATCCGCGTCCCCTGAAAAGCT-3′ for ALDH3I1; 5′-CATACGAGGATGATCGTGGCAATG-3′ and 5′-TCACCTCTCTTAGGAGCCGTAACCT-3′ for ALDH7B4; 5′-ATCGGCGGAAGCGAGTAATTTGGTG-3′ and 5′-TATGGCGGATACCTGACGGCTGAATC-3′ for ALDH3H1; and 5′-GAAGCCATGGAAGCTATGAAGGAGAC-3′ and 5′-GTCTCTGTCTCTCACTTTCCCCCTT-3′ for ALDH3F1. In the second assay, one gene-specific primer (underlined above for each gene) was combined with one T-DNA specific primer: FISH1 (5′-CTGGGAATGGCGAAATCAAGGCATC-3′)46 for ALDH3I1 and ALDH3H1 or LBa3 (5′-ACCCAACTTAATCGCCTTGCAGCAC-3′)48 for ALDH7B4 and ALDH3F1.
RNA and Protein Analyses
Isolation of total RNA and RT-PCR analyses were performed as described before3. Gene-specific primers were used to amplify first strand cDNAs. These were 5′-GAAGCAATAGCCAAAGACACACGC-3′ and 5′-GATATCTCGATTATCGTAGGCTCC-3′′ for ALDH7B4, 5′-CTACTGGATGTGCCTGAAGCATC-3′ and 5′-TAAGATCCGCGTCCCCTGAAAAGCT-3′ for ALDH3I1, 5′-CAGCTAAAGAACTGGATGGCTC-3′ and 5′-TCAACCAACTAAGTCCATGTTTGA-3′ for ALDH3H1, and 5′-GAAGCCATGGAAGCTATGAAGGAGAC-3′ and 5′-GTCTCTGTCTCTCACTTTCCCCCTT-3′ for ALDH3F1. Transcripts of the Arabidopsis ACTIN-2 (At3g18780) gene were used as reference49 and were amplified with the primers 5′-GGAATCCACGAGACAACCTATAAC-3′ and 5′-AGGAATCGTTCACAGAAAATGTTTC-3′.
Protein expression was examined by immunoblot detection. Crude protein extracts were separated on 12% SDS–polyacrylamide gels and electroblotted from the gel onto a nitrocellulose Protran BA-85 membrane (Whatman, Dassel, Germany) at 100 Volts for 1 h in pre-chilled protein-blot transfer buffer (25 mM Tris, 192 mM glycine, 20% [v/v] methanol)50. The membrane was stained with Ponceau red solution (0.2% [w/v] Ponceau S in 3% [w/v] trichloroacetic acid). Then, unspecific binding sites were blocked by incubating the membrane for 1 h at room temperature or overnight at 4 °C in blocking solution containing 0.1% (v/v) Tween-20 and 4% (w/v) non-fat dry milk powder dissolved in 1X TBS (20 mM Tris-HCl; 150 mM NaCl; pH 7.5). The membrane was probed with 5000-fold diluted ALDH7B4 antiserum7. The immunodetection assay was performed by using the ECL Plus Western Blotting detection Kit (Amersham, Braunschweig, Germany). Signals were detected under a CCD camera (Intelligent Dark Box II, Fujifilm Corporation, Tokyo, Japan).
Chlorophyll Fluorescence and Gas Exchange Measurements
Measurements of chlorophyll fluorescence were performed using GFS-3000 Portable Gas Exchange Fluorescence System (LED-Array/PAM-Fluorometer 3055FL, Heinz Walz, Effeltrich, Germany). Parameters were set as follows: Photosynthesis Photon Flux Density (PPFD), 100 to 1800 µmol m−2 s−1; chamber temperature, 25 °C; flow rate, 750 µmol s−1; relative humidity, 60–70%; 350 µl L−1 CO2; 21% oxygen. Plants were allowed to adapt to the respective conditions for 15 minutes before starting measurement.
Pyridine Nucleotides, Glutathione, and Enzyme Activity Measurements
Chemicals were obtained from Sigma-Aldrich. Approximately 50 to 100 mg fresh leaves was used for each assay, if not otherwise stated. Determination of NAD+, NADP+, NADH, and NADPH contents followed a previously described enzymatic cycling procedure51,52. Tissues were homogenized with 1 mL of 0.1 N KOH containing 50% ethanol (v/v) (for determination of NADPH and NADH) or 1 mL of 0.1 N HCl containing 50% (v/v) ethanol (for determination of NADP+ and NAD+). The suspension was heated for 5 min at 95 °C, cooled on ice and centrifuged (15,000 g) for 20 min at 4 °C. The oxidized and reduced forms of the nucleotides were determined from the supernatant or a standard solution corresponding to 0 to 500 pmol of NADP(H) in 1 mL reaction mixture containing 100 mM HEPES-KOH (pH 8.0), 0.5 mM EDTA, 2.5 mM glucose-6-phosphate (G6P), 1.66 mM phenazine ethosulfate, 0.42 mM 3-(4,5-dimethyl-2-thiazolyl)−2,5-diphenyl-2H-tetrazolium bromide. The reaction was started by adding 0.7 unit of glucose-6-phosphate dehydrogenase (G6PDH) enzyme. For the determination of NAD+ and NADH, the reaction mixture was the same except that Glc-6-P was replaced by 100 µl absolute ethanol. For the determination of NADH, the mixture contained 0.2 mM phenazine ethosulfate. The reaction was started by the addition of 0.7 unit of alcohol dehydrogenase. The rate of thiazolyl blue reduction was recorded by measuring the absorbance change at 570 nm. The pyridine nucleotides were measured in a GENESYS 10 UV spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The concentrations of each pyridine nucleotides in the extract were calculated from the corresponding standard curves obtained for pyridine nucleotide solutions per tissue fresh weight. Freshly extracted samples and standards were assayed at the same time. Glutathione measurement was performed in a Biotek Powerwave XS2 Spectrophotometer by following the procedure of Griffith53.
Total G6PDH activities were determined essentially as reported earlier54,55. Tissues were extracted in 100 mM Tris-HCl pH 8.0, 10 mM MgCl2, 5 mM EDTA pH 8.0, 1 mM phenylmethylsulphonyl fluoride, 10% glycerol, 15 µM NADP+. The assay reaction contains 50 mM Tris-HCl pH 8.0, 10 mM MgCl2, 150 µM NADP+, 3 mM Glc-6-P, and 10 to 20 µl extract. The reduction of NADP+ was measured through the increase of the absorbance at 340 nm. The activity was expressed as nmol NADPH min−1 mg−1 protein.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information file).
References
Sophos, N. A. & Vasiliou, V. Aldehyde dehydrogenase gene superfamily: the 2002 update. Chem. Biol. Interact. 143–144, 5–22 (2003).
Brocker, C. et al. Aldehyde dehydrogenase (ALDH) superfamily in plants: gene nomenclature and comparative genomics. Planta 237, 189–210 (2013).
Missihoun, T. D., Schmitz, J., Klug, R., Kirch, H.-H. & Bartels, D. Betaine aldehyde dehydrogenase genes from Arabidopsis with different sub-cellular localization affect stress responses. Planta 233, 369–82 (2011).
Stiti, N., Missihoun, T. D., Kotchoni, S. O., Kirch, H.-H. & Bartels, D. Aldehyde Dehydrogenases in Arabidopsis thaliana: Biochemical Requirements, Metabolic Pathways, and Functional Analysis. Front. Plant Sci. 2, 65 (2011).
Missihoun, T. D. et al. Overexpression of ALDH10A8 and ALDH10A9 Genes Provides Insight into Their Role in Glycine Betaine Synthesis and Affects Primary Metabolism in Arabidopsis thaliana. Plant Cell Physiol. 56, 1798–807 (2015).
Sunkar, R., Bartels, D. & Kirch, H.-H. Overexpression of a stress-inducible aldehyde dehydrogenase gene from Arabidopsis thaliana in transgenic plants improves stress tolerance. Plant J. 35, 452–464 (2003).
Kotchoni, S. O., Kuhns, C., Ditzer, A., Kirch, H.-H. & Bartels, D. Over-expression of different aldehyde dehydrogenase genes in Arabidopsis thaliana confers tolerance to abiotic stress and protects plants against lipid peroxidation and oxidative stress. Plant, Cell Environ. 29, 1033–1048 (2006).
Raza, H. Functional Characterization of Transgenic Arabidopsis Thaliana Plants Co-Overexpressing Aldehyde Dehydrogenases and Genes for Soluble Osmolytes. (University of Bonn, 2010).
Nair, R. B., Bastress, K. L., Ruegger, M. O., Denault, J. W. & Chapple, C. The Arabidopsis thaliana Reduced Epidermal Fluorescence1 gene encodes an aldehyde dehydrogenase involved in ferulic acid and sinapic acid biosynthesis. Plant Cell 16, 544–554 (2004).
Missihoun, T. D., Kotchoni, S. O. & Bartels, D. Active Sites of Reduced Epidermal Fluorescence1 (REF1) Isoforms Contain Amino Acid Substitutions That Are Different between Monocots and Dicots. PLoS One 11, e0165867 (2016).
Wei, Y., Lin, M., Oliver, D. J. & Schnable, P. S. The roles of aldehyde dehydrogenases (ALDHs) in the PDH bypass of Arabidopsis. BMC Biochem. 10, 7 (2009).
Cui, X., Wise, R. P. & Schnable, P. S. The rf2 Nuclear Restorer Gene of Male-Sterile T-Cytoplasm Maize. Science (80-). 272, 1334–1336 (1996).
Liu, F., Cui, X., Horner, H. T., Weiner, H. & Schnable, P. S. for Male Fertility in Maize. Society 13, 1063–1078 (2001).
Tsuji, H. et al. Induction of mitochondrial aldehyde dehydrogenase by submergence facilitates oxidation of acetaldehyde during re-aeration in rice. FEBS Lett. 546, 369–373 (2003).
Shin, J.-H., Kim, S.-R. & An, G. Rice aldehyde dehydrogenase7 is needed for seed maturation and viability. Plant Physiol. 149, 905–15 (2009).
Shen, Y. et al. Mutation of OsALDH7 causes a yellow-colored endosperm associated with accumulation of oryzamutaic acid A in rice. Planta 235, 433–441 (2012).
Marcato, P., Dean, C. A., Giacomantonio, C. A. & Lee, P. W. K. Aldehyde dehydrogenase its role as a cancer stem cell marker comes down to the specific isoform. Cell Cycle 10, 1378–1384 (2011).
Missihoun, T. D., Hou, Q., Mertens, D. & Bartels, D. Sequence and functional analyses of the aldehyde dehydrogenase 7B4 gene promoter in Arabidopsis thaliana and selected Brassicaceae: regulation patterns in response to wounding and osmotic stress. Planta 239, 1281–1298 (2014).
Esposito, S., Carfagna, S., Massaro, G. & Vona, V. & Di Martino Rigano, V. Glucose-6-phosphate dehydrogenase in barley roots: Kinetic properties and localisation of the isoforms. Planta 212, 627–634 (2001).
Kruger, N. J. & Von Schaewen, A. The oxidative pentose phosphate pathway: Structure and organisation. Curr. Opin. Plant Biol. 6, 236–246 (2003).
Née, G., Zaffagnini, M., Trost, P. & Issakidis-Bourguet, E. Redox regulation of chloroplastic glucose-6-phosphate dehydrogenase: A new role for f-type thioredoxin. FEBS Lett. 583, 2827–2832 (2009).
Esposito, S. Nitrogen Assimilation, Abiotic Stress and Glucose 6-Phosphate Dehydrogenase: The Full Circle of Reductants. Plants 5, 24 (2016).
Noctor, G., Queval, G., Mhamdi, A., Chaouch, S. & Foyer, C. H. Glutathione. Arab. B. 9, 1–32 (2011).
Noctor, G. et al. Glutathione in plants: An integrated overview. Plant, Cell Environ. 35, 454–484 (2012).
Kramer, D. M., Cruz, J. A. & Kanazawa, A. Balancing the central roles of the thylakoid proton gradient. Trends Plant Sci. 8, 27–32 (2003).
Avenson, T. J. et al. Integrating the proton circuit into photosynthesis: progress and challenges. Plant, Cell Environ. 28, 97–109 (2005).
Foyer, C. H., Neukermans, J., Queval, G., Noctor, G. & Harbinson, J. Photosynthetic control of electron transport and the regulation of gene expression. J. Exp. Bot. 63, 1637–1661 (2012).
Mhamdi, A. et al. Cytosolic NADP-dependent isocitrate dehydrogenase contributes to redox homeostasis and the regulation of pathogen responses in Arabidopsis leaves. Plant. Cell Environ. 33, 1112–1123 (2010).
Li, S., Mhamdi, A., Clement, C., Jolivet, Y. & Noctor, G. Analysis of knockout mutants suggests that Arabidopsis NADP-MALIC ENZYME2 does not play an essential role in responses to oxidative stress of intracellular or extracellular origin. J. Exp. Bot. 64, 3605–3614 (2013).
Dutilleul, C. et al. Mitochondria-Driven Changes in Leaf NAD Status Exert a Crucial Influence on the Control of Nitrate Assimilation and the Integration of Carbon and Nitrogen Metabolism. PLANT Physiol. 139, 64–78 (2005).
Geigenberger, P. & Fernie, A. R. Metabolic control of redox and redox control of metabolism in plants. Antioxid. Redox Signal. 21, 1389–421 (2014).
Shen, W. et al. Involvement of a glycerol-3-phosphate dehydrogenase in modulating the NADH/NAD+ ratio provides evidence of a mitochondrial glycerol-3-phosphate shuttle in Arabidopsis. Plant Cell 18, 422–41 (2006).
Sweetlove, L. J. et al. Mitochondrial uncoupling protein is required for efficient photosynthesis. Proc. Natl. Acad. Sci. USA 103, 19587–19592 (2006).
Mhamdi, A. et al. Arabidopsis Glutathione Reductase1 plays a crucial role in leaf responses to intracellular hydrogen peroxide and in ensuring appropriate gene expression through both salicylic acid and jasmonic acid signaling pathways. Plant Physiol. 153, 1144–60 (2010).
Fujita, M. & Hossain, M. Z. Modulation of pumpkin glutathione S-transferases by aldehydes and related compounds. Plant Cell Physiol. 44, 481–490 (2003).
Curtis, J. M. et al. Protein carbonylation and metabolic control systems. Trends Endocrinol. Metab. 23, 399–406 (2012).
Chai, M. F. et al. NADK2, an Arabidopsis chloroplastic NAD kinase, plays a vital role in both chlorophyll synthesis and chloroplast protection. Plant Mol. Biol. 59, 553–564 (2005).
Hashida, S. N., Takahashi, H. & Uchimiya, H. The role of NAD biosynthesis in plant development and stress responses. Ann. Bot. 103, 819–824 (2009).
Takahashi, H. et al. Chloroplast NAD Kinase is Essential for Energy Transduction Through the Xanthophyll Cycle in Photosynthesis. Plant Cell Physiol. 47, 1678–1682 (2006).
Demmig-Adams, B. Carotenoids and photoprotection in plants: A role for the xanthophyll zeaxanthin. Biochim. Biophys. Acta - Bioenerg. 1020, 1–24 (1990).
Eskling, M., Arvidsson, P.-O. & Akerlund, H.-E. The xanthophyll cycle, its regulation and components. Physiol. Plant. 100, 806–816 (1997).
Missihoun, T. D. & Kotchoni, S. O. Aldehyde dehydrogenases may modulate signaling by lipid peroxidation-derived bioactive aldehydes. Plant Signal. Behav. e1387707, https://doi.org/10.1080/15592324.2017.1387707 (2017).
Mano, J., Miyatake, F., Hiraoka, E. & Tamoi, M. Evaluation of the toxicity of stress-related aldehydes to photosynthesis in chloroplasts. Planta 230, 639–648 (2009).
Liu, X.-L., Yu, H.-D., Guan, Y., Li, J.-K. & Guo, F.-Q. Carbonylation and loss-of-function analyses of SBPase reveal its metabolic interface role in oxidative stress, carbon assimilation, and multiple aspects of growth and development in Arabidopsis. Mol. Plant 5, 1082–99 (2012).
Murashige, T. & Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiol. Plant. 15, 473–497 (1962).
Ríos, G. et al. Rapid identification of Arabidopsis insertion mutants by non-radioactive detection of T-DNA tagged genes. Plant J. 32, 243–253 (2002).
Missihoun, T. D., Kirch, H.-H. & Bartels, D. T-DNA insertion mutants reveal complex expression patterns of the aldehyde dehydrogenase 3H1 locus in Arabidopsis thaliana. J. Exp. Bot. 63, 3887–98 (2012).
Sessions, A. et al. A high-throughput Arabidopsis reverse genetics system. Plant Cell 14, 2985–94 (2002).
An, Y.-Q. et al. Strong, constitutive expression of the Arabidopsis ACT2/ACT8 actin subclass in vegetative tissues. Plant J. 10, 107–121 (1996).
Towbin, H., Staehelin, T. & Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl. Acad. Sci. USA 76, 4350–4 (1979).
Maciejewska, U. & Kacperska, A. Changes in the level of oxidized and reduced pyridine nucleotides during cold acclimation of winter rape plants. Physiol. Plant. 69, 687–691 (1987).
Tamoi, M., Miyazaki, T., Fukamizo, T. & Shigeoka, S. The Calvin cycle in cyanobacteria is regulated by CP12 via the NAD(H)/NADP(H) ratio under light/dark conditions. Plant J. 42, 504–513 (2005).
Griffith, O. W. Determination of Glutathione and Glutathione Disulfide Using Glutathione-Reductase and 2-Vinylpyridine. Anal. Biochem. 106, 207–212 (1980).
Esposito, S. et al. Glutamate synthase activities and protein changes in relation to nitrogen nutrition in barley: The dependence on different plastidic glucose-6P dehydrogenase isoforms. J. Exp. Bot. 56, 55–64 (2005).
Cardi, M. et al. Abscisic acid effects on activity and expression of barley (Hordeum vulgare) plastidial glucose-6-phosphate dehydrogenase. J. Exp. Bot. 62, 4013–4023 (2011).
Acknowledgements
DB acknowledges the financial support of the German Research Foundation (Deutsche Forschungsgemeinschaft: DFG). TDM acknowledges the financial support of DAAD for the accomplishment of this work. The authors are grateful to C. Buchholz for growing plants, Dr A. Ditzer for providing mutants, S. Höller and Dr. MJ Fryer for advice on glutathione measurement.
Author information
Authors and Affiliations
Contributions
T.D.M. designed research; T.D.M. performed research; D.B. and S.O.K. contributed new reagents/analytic tools; T.D.M., S.O.K., and D.B. analyzed data; T.D.M. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Missihoun, T.D., Kotchoni, S.O. & Bartels, D. Aldehyde Dehydrogenases Function in the Homeostasis of Pyridine Nucleotides in Arabidopsis thaliana. Sci Rep 8, 2936 (2018). https://doi.org/10.1038/s41598-018-21202-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21202-6
This article is cited by
-
TMT-based quantitative proteomic analysis reveals defense mechanism of wheat against the crown rot pathogen Fusarium pseudograminearum
BMC Plant Biology (2021)
-
Where do the electrons go? How numerous redox processes drive phytochemical diversity
Phytochemistry Reviews (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.