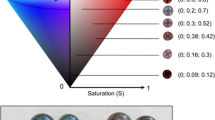
Abstract
The bivalve Pinctada margaritifera exhibits three main transplant phenotypes derived from the donor (from which a mantle graft tissue, the saibo, is excised), the recipient (into which the saibo is implanted with a nucleus, leading to the formation of a pearl sac “chimera”) and the cultured pearls themselves. This first phenome study on the species derived from a large experimental graft. Transplant phenotype was assessed at three scales: 1) macro, pearl size, colour, grade, 2) micro, pearl surface microstructure, and 3) molecular, biomineralisation gene expression level in saibo and pearl sac tissues. From donor to pearl, the phenome revealed fine variations of quality traits dependent on the position on the mantle where the saibo was cut, whose variation could overlap with inter-individual donor phenotype differences. A single donor phenotype could therefore produce multiple pearl phenotypes at the scale of the saibo position, mirroring its original activity at the mantle position level and the colour and shape of the shell. This phenome study provides essential information on phenotypic trait architecture enabling us to explore and explain the main biological functions and pave the way for a phenomic project on P. margaritifera that could benefit the pearl industry.
Similar content being viewed by others
Introduction
The phenotype is the result of the interplay of genetics with developmental, environmental and stochastic influences, where the intensity, frequency, order and interaction of these influences all affect the outcome. In the era of next generation sequencing with continued decreases in cost and increasing availability of high-throughput genotyping platforms, genomic data acquisition and associated bioinformatics treatments have become common even for non-model organisms. The complexity of plant and animal genomes, constructed from a pool of four nucleic acids and organized in a one-dimensional sequence, pales in comparison to their corresponding phenome. Phenome serves an unknown number of functions, many of which show enormous inter-individual variation that is at best only partially understood and for which the dimensionality remains unknown. This is not only due to the recent advances in genomics but also the complex multidimensional nature of phenotypes1. The vast number of phenotypic states of a genotype can be viewed as its phenotypic space, which is often referred to as its phenome. In practice, the phenome is a theoretical entity which can never be fully characterized2. Phenomics, operationally defined as the systematic study of phenotypes, is critically important to provide essential information for advances in genetic improvement for many cultured plant and animal species in the post-genomic era.
Species from the Pinctada genus are regularly used for the production of valuable free round cultured pearls, the only gems produced by a living organism3. The production of cultured pearls is both unique and biologically complex compared to that of other aquaculture industries. These are nucleated pearls produced in the gonad of a recipient pearl oyster following surgical implantation of a spherical shell-based bead (the nucleus) together with a piece of mantle (the saibo) cut from a particular section (located between posterior and anterior zone, without considering the junction of the mantle with the oyster gills) of a selected donor oyster. The mantle is clearly a metabolically and transcriptionally active tissue, indispensable for mollusc shell formation, with prominent transcriptional activity of biomineralisation genes4. The biomineralisation process is responsible for both pearl and shell formations. A few weeks after the graft operation, the pearl sac develops as a result of the epithelial cells of the mantle epidermis growing around the nucleus to completely cover it5. The pearl formation process then starts, by the deposition of successive biomineral layers onto the nucleus6. P. margaritifera is an ideal model animal for the study of biomineralisation because of the intriguing microstructure of: 1) its shells, which consist of outer calcitic prismatic layers and inner aragonitic nacreous layers, and 2) the pearls, which display mostly aragonitic structures, similar to the inner layer of the shell in both appearance and structure, through the chimeric pearl sac activity. Cultured pearl value is based on four main quality traits: size (nacre weight and thickness), shape, colour (including darkness level) and grade (the combination of lustre and surface quality)3,7.
Phenotype transmission in the Pinctada transplant model has been mostly studied from an applied point of view, in relation to its economic importance in the pearl industry. Indeed, much research effort has been focused on size and colour determination in relation, to the donor oyster, the recipient oyster, their interplay, and their interaction with the environment8,9,10,11. In P. margaritifera, pearl size is known to be mostly driven by recipient oyster growth performance, and the donor oyster to be responsible for pearl qualitative trait determination, including colour12. The genotype of the donor therefore persists within the recipient in the form of the pearl sac. This chimeric organ displays more complex interactions, particularly when spatial (geographic origin of the population, depth of culture practices), temporal (age of hatchery-produced or wild collected spat), and environmental (grow-out site culture, season of graft, experimental temperature or pH variation) factors are introduced into the equation of the pearl quality trait determination3. Previous studies have all considered the transplant model at the individual scale: i.e., at the level of individual pearl oysters13,14,15. The associated phenotypic variations recorded were then systematically related to inter-individual/ family variation among donors or recipients.
The objective of the present study was to explore for the first time the phenome of cultured pearl quality traits at the scale of the saibo, i.e., at an intra-individual donor scale. For this, two fixed hatchery-produced pearl oyster phenotypes of P. margaritifera, exhibiting predominately green or yellow inner shell coloration (twelve of each) were used as donors. The entire mantle was taken for saibo production, including all the following four sections: posterior, connection (with gills), middle (used in commercial production) and anterior (Fig. 1). Experimental grafts (N = 1798) with traceability between donor line and saibo mantle position has been designed and performed in a single culture site. These grafts provided the biomineralised materials for the phenome study: the saibo tissue, the chimeric pearl sac tissue, and the associated pearls. Phenotypic data were collected at the macroscopic level (pearl quality traits), microscopic level (pearl surface ultrastructure) and molecular scale (expression level of a panel of biomineralisation genes representative of the nacreous aragonite and/or prismatic calcite synthesis in both saibo and pearl sac, derived from each of the mantle sections). This first phenome study, initiated with a set of easy-to-use tools, will provide: 1) basic knowledge to help us to understand phenotypic transmission, range of variation in an animal transplant model, and 2) useful information for the improvement of pearl quality for the industry.
Donor Pinctada margaritifera of green (a) and yellow (b) phenotypes, each with two shell valves showing, on the right, the four sections of the entire dilated mantle tissue in (a), and contracted mantle tissue in (b): posterior position (P), connection with the gill (C), middle corresponding to the zone usually used for commercial saibo production (M) and anterior (A). In each case, on the left, the correspondence is shown with the zones of contact with the inner shell zone that exhibits the colourful band characteristic of donor oysters. The dotted lines indicate the areas of mantle tissue excised for saibo production from the four positions A, M, C and P. General anatomy: am, adductor muscle; gi, gills; by, byssus; cn, coloured nacreous zone.
Results
Experimental graft
The nucleus retention rate was 75.1% (N = 1350) at 45 days post-grafting (the remainder, 24.9%, N = 448, correspond to nucleus rejection, oyster mortalities and/or predation). Between the two phenotypes used as donors (green and yellow), no significant difference was detected for overall retention rate, even at the mantle section scale (i.e., pairwise comparison between oysters grafted with the same mantle section between the two phenotypes). By contrast, comparison between the mantle sections revealed that the middle position had significantly a higher retention rate than the posterior (+7.0%; p = 0.0315) and connection positions (+10.7%; p = 0.0315) (Fig. 2A).
Graft and cultured pearl quality trait variation in P. margaritifera according to both mantle graft position (posterior, connection, middle and anterior) and the two phenotypes of donor (green C and yellow L). The five variables are the: (A) experimental graft retention rate; (B) cultured pearl nacre weight in g (boxplot); (C) pearl colour category percentages (dark, green, light dark and light), (D) pearl grade rate (A–C, D+, D, D1, D2 grade and rebuts) and (E) pearl shape rate (baroque, circle, oval or round). Letters indicate significance between the mantle graft positions within each variable. Asterisks indicate within each mantle graft position the significant differences between the proportions of the categories of each cultured pearl trait, between the C (green) and L (yellow) phenotypes. The shell valve corresponded to a green C phenotype donor oyster.
Cultured pearl quality traits
The overall mean cultured pearl nacre weight was 0.67 g (±0.32 g). A significant difference (p = 0.031) was detected between the two donor phenotypes, with grafts from the yellow donor leading to heavier nacre (+6.2%), in comparison with the green phenotype: 0.65 (±0.31 g) vs. 0.69 (±0.33 g). This was due, at the mantle position scale, to the anterior position on the yellow phenotype, which led to nacre 18.8% heavier (p = 0.019) than the green phenotype (Fig. 2B). Differences between the mantle positions were observed within the donor phenotypes. For the green phenotype, the middle section was 17.3% heavier than the anterior one (0.68 ± 0.31 g vs. 0.58 ± 0.30 g; p = 0.001). For the yellow phenotype, the two extremities of the mantle gave the most disparate results: the posterior position was 20.6% significantly (p < 0.001), heavier than the anterior one (0.76 ± 0.39 g vs. 0.63 ± 0.27 g).
The overall distribution of the cultured pearls among the colour categories was as follows: 53.1% (N = 632) for dark, 5.8% (N = 70) for green, 5.4% (N = 64) for light dark and 35.7% (N = 425) for light. The anterior section was significantly different from the three others and showed the highest rate of light pearls whatever the donor phenotype considered. For the yellow phenotype, 73.5% light pearls were found with anterior section grafts compared with an average of 30.0% for the other sections (Fig. 2C). The same tendency was observed in the green phenotype, with 66.9% of light pearls from the anterior section grafts and an average of 15.3% for the three other sections. The posterior and middle sections differentiated the two donor phenotypes, whereas the connection and anterior sections showed no difference between the colour rates. The yellow phenotype posterior position grafts produced 18.1% more light pearls than those cut from the same position on green phenotype donors (p = 0.001), which showed a significantly higher proportion of green pearls (+8.9%; p = 0.014). The middle position also had a significantly higher proportion of light pearls for the yellow phenotype (+21.0%; p < 0.001), while the green phenotype showed more dark pearls for this position (+15.7%; p < 0.001) (Fig. 2C).
For the cultured pearl quality grade, no significant differences between positions were detected for the green phenotype (Fig. 2D). For the yellow phenotype, the anterior position was different, with only 10.5% top quality pearls (A–C grade) compared with 22.9% on average for the three other positions (p = 0.002). Comparison between the two phenotypes showed that: 1) the green phenotype produced significantly (p < 0.001) more pearls of grades A–C, with +23.1% and +20.3% for the posterior and middle positions, respectively, 2) the yellow phenotype produced significantly more rebut (reject) grade peals, with +9.5% (p = 0.012) and +12.1% (p < 0.001) for the posterior and the middle positions, respectively, and also more D2 grade pearls with + 16.0% (p < 0.001) and +9.8% (p = 0.041) for the posterior and middle positions, respectively (Fig. 2D). A significant difference was also observed between the two phenotypes when comparing the results of anterior position grafts, with the yellow donors producing more rebuts (+12.1%, p < 0.001), D2 (+13.3%, p < 0.001) and D1 (+7.1%, p < 0.001) pearls and less grade D pearls (−10.5%, p = 0.01), than the green phenotype (Fig. 2D).
For cultured pearl shapes (Fig. 2E), no significant differences were detected within donor phenotype groups among the four different mantle positions from which saibo was cut. By contrast, significant differences were observed between the two donor colour lines. For posterior and anterior positions, yellow phenotype donors produced significantly more baroque pearls than green phenotype donors with +16.2% (p = 0.01) and +16.6% (p = 0.006), respectively. For these two positions pearl circles were, on the contrary, more frequent in pearls from green phenotype donors than yellow ones, with +13.4% (p = 0.034) for posterior position saibo and +12.5% (p = 0.038) for the anterior position. For the middle position, round pearls were significantly more frequent with the green phenotype (+8.9%, p = 0.02).
Cultured pearl surface ultrastructure observation
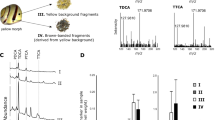
The mineralised portion of the nacre was observed on the cultured pearl surface by scanning electron microscopy and was seen to consist of aragonite tablets organized into growth fronts (Fig. 3). Observation of the corresponding microscopic patterns showed clear differences in the distance between the parallel growth fronts on the pearl surface, between the different mantle sections. The growth fronts of pearls produced with middle position saibo were significantly larger (35.1 ± 4.5 µm; Fig. 3C,G), than those of the connection zone (30.8 ± 6.4 µm; p = 0.045; Fig. 3B,F) and the posterior and anterior sections, which showed similar growth front distances (mean of both: 26.0 ± 3.5 µm; p < 0.0001; Fig. 3A,E,D,H).
P. margaritifera cultured pearl surface, assessed by electronic microscopy (magnification: x1000) according to mantle graft position (posterior, connection, middle or anterior) based on grafts with tissue from two donor oysters. The first column (A,B,C and D) corresponds to sample of pearl surfaces from grafts made with the same yellow phenotype donor, whereas the second column (E,F,G and H) corresponded to pearl surface from another donor, which had the green phenotype. Cultured pearl grades were all D grade for those pearls produced with the yellow phenotype donor (first column) and (A–C) grade for pearls produced with the green phenotype donor (second column). The black bars at the bottom of each picture correspond to 100 µm. Distances between the fronts of aragonites were expressed in µm and illustrated with white bars and arrows.
Biomineralisation gene expression levels in saibo
Concerning the genes involved in the nacreous layer produced from the mantle, all four corresponding genes (MRNP34, MSI60, Pearlin and Pif177) showed the same expression profile, which distinguished the yellow and the green donor phenotypes, whatever the mantle position considered (Fig. 4). The saibo originating from the green donor phenotype systematically showed overexpression of these four genes in comparison to the saibo from the yellow variant (Fig. 4A–D). For example, in saibo from the green phenotype the Pif177 gene was overexpressed by 4.90 (p = 0.01), 3.98 (p < 0.001) and 1.96 (p = 0.023) times for the posterior, middle and anterior positions respectively. Comparison among mantle sections within the green phenotype revealed systematic overexpression of genes in the posterior section compared with the middle and anterior sections, except for MRNP34.
Relative expression of nine biomineralization genes in the mantle graft tissue (mixed of 3 saibo per position) of P. margaritifera obtained from the green C (Posterior N = 6; Middle N = 10; Anterior N = 11) and yellow L (Posterior N = 6; Middle N = 12; Anterior N = 6) donor phenotypes. Y axes are in the logarithmic scale. Error bars correspond to standard deviations. Letters indicate significance between mantle graft positions within each phenotype.
For the four genes involved in prismatic layer formation (Aspein, Shem9, KRMP7 and Prism14), near identical expression profiles were observed among donor phenotypes and mantle positions (Fig. 4). The same tendencies were observed between the green and yellow donor phenotypes for all prismatic genes, except for the KRMP7 gene expression in the middle section, where there was significant overexpression (×4) in the yellow phenotype compared with the green one (Fig. 4H). Expression of Shem9 showed significant differences between: 1) phenotypes for the anterior position only (p = 0.048), with nearly twice the expression level in the green phenotype compared with the yellow, and 2) mantle section for both phenotypes only for the anterior position which was different from the two others (Fig. 4G). For Aspein gene expression, no differences were observed between the donor phenotypes for any of the mantle positions.
Expression levels of the Nacrein A1 gene (involved in both nacreous and prismatic formation), were not significantly different between mantle positions for the yellow donor phenotype (Fig. 4E). By contrast, the middle section of the green phenotype was significantly different from the anterior and posterior positions. Inter-phenotype comparison revealed significant differences for the posterior and middle sections, with respectively overexpression factors of 1.77 for the green and 3.02 for the yellow phenotype.
Biomineralisation gene expression levels in the pearl sac
For the genes involved in the nacreous layer produced by the pearl sac, the expression of MSI60, Pearlin and Pif177 were not different between either phenotypes or mantle positions (Fig. 5B–D). For MNRP34 gene expression, differences were detected only for the yellow phenotype, where the posterior (p = 0.002) and anterior positions (p = 0.006) were overexpressed 47 and 20 times, respectively, in comparison to the middle (Fig. 5A).
Relative expression of nine biomineralization genes in the pearl sac of P. margaritifera originated from the green C (Posterior N = 13; Middle N = 20; Anterior N = 32) and yellow L (Posterior N = 13; Middle N = 20; Anterior N = 16) donor phenotypes. Y axes are in the logarithmic scale. Error bars correspond to standard deviations. Letters indicate significance between the mantle graft positions within each phenotype.
For the genes implicated in cultured pearl prismatic layer formation, the expression of the Aspein gene showed no significant difference between phenotypes or positions (Fig. 5F). Shem9 was expressed 4.7 times more in pearl sacs from anterior position saibo from green donor phenotypes than in those from the same position in the yellow donor phenotype (p = 0.04) (Fig. 5G). For KRMP7 gene expression, the two phenotypes were highly different for all mantle sections: in pearl sacs from the green phenotype donors, KRMP7 was overexpressed 5.4 times for the anterior position, 4.5 times for the middle section and 3.8 for the posterior position, compared with the yellow phenotype (Fig. 5H). For Prism14 gene expression, only the posterior position showed a significant difference between phenotypes, with the green donor phenotype showing significantly higher expression (5.9 times, p = 0.014). Within phenotype, no significant differences were revealed between mantle positions (Fig. 5I).
Concerning the gene involved in both nacreous and prismatic layer formation, Nacrein A1 showed no significant differences between donor phenotypes or mantle positions (Fig. 5F). For the two phenotypes, no significant differences were detected between positions.
Discussion
Phenotypic variations were assessed in the present study in a particular transplanted animal model, P. margaritifera, which combines a complex three-way interaction between the donor oyster, recipient oyster and the final cultured pearl product. The associated phenome, in relation to biomineralising and biomineralised tissues (graft tissue, pearl sac tissue, pearl samples) was assessed through a set of easy-to-use tools (visual and microscopic observation, RT-PCR) applied at three observation levels: macro-, micro- and molecular.
This initial cultured pearl quality phenome mirrored, to a certain extent, P. margaritifera shell morphology and characteristics. Oyster shell and cultured pearls are respectively formed in two distinct biomineralised tissues: the mantle and the pearl sac, which are derived from mantle tissue from the donor16. Around the nucleus, a pearl sac is formed by proliferation of the outer mantle epithelial cells of the mantle graft, which secretes successive nacre layers on the nucleus17,18. The pearl sac consists of mucous cells containing large acidophilic granules and epidermal cells19 that secrete proteins resulting in cultured pearl formation, a highly controlled biomineralisation process similar to the development of the inner shell regulated by the mantle20. Similarly to other bivalves, the shell of P. margaritifera consists of two polymorphs of calcium carbonate: the inner nacreous layer, which is composed of aragonite, and the outer prismatic layer, which is made of calcite21,22,23. Shell formation is a highly controlled process involving multiple matrix proteins24,25,26. In the Pinctada genus, the anterior zone of the shell is characterised by two growth-related features. First, the byssus location gives an anteriorly oblique shell conformation, with specifically subtriangular anterior auricles. Second, the concentric growth lines are closer in the anterior zone, than in the posterio-ventral part of the shell corresponding to the other mantle sections3.
At macro-scale observation, and for pearl size, results revealed that the anterior shell-growth potential was conveyed by the saibo from the donor into the chimera formed in the recipient and thereby impacted nacre weight and thickness of pearls produced, giving smaller pearls compared to those grown from saibo from the other mantle sections, whatever the donor phenotype considered. This intra-phenotype variation from the donor, at the level of mantle position, was sometimes greater than variation observed at an inter-phenotype level. This result is consistent with earlier findings where donor effect for pearl size was shown using the same middle mantle section compared among wild27 or hatchery-produced oysters28. The present study pointed the existence of a mantle effect within a same donor phenotype, which must be considered for pearl size determination to avoid any artefacts for genetic selection program aiming to improve this trait.
Colour transmission to the pearls is also dependent on the section of the mantle used at the scale of the individual donor. Here again, intra-individual variation could be greater than inter-individual variation. This was the case for example for the attractive green pearl colour, whose rate was significantly lower for the posterior section of yellow donor phenotypes compared with overall results from green donors, but similar to results from the green anterior section. Differences in pearl colour distribution also mirrored, to a certain extent, the inner shell colour profile of the donor oyster. Indeed, when looking at the interior nacreous surface of the shell, the anterior zone corresponds to the least colourful part of the donor oyster, compared with the parts adjacent to the other three sections, which show the largest and strongest intensity of the characteristic coloured band, particularly in the middle section of the mantle (Fig. 1). It is commonly known that donor tissue influences the colour of the resulting pearls and is mostly dependent on the species of Pinctada used11,29. Among pearl oysters of the genus, P. margaritifera is a good model for phenotypic colour variation studies, as it displays the largest range of pearl colours, reflected by the large diversity of inner shell colour phenotypes, in comparison to its two competing species P. maxima and P. fucata3. The green and yellow inner shell colour phenotypes have been studied recently and are known to also depend on rearing/culture site30, with a colour “signature” at the archipelago scale in French Polynesia7.
At the micro-scale observation level, assessment of the pearl surface growth fronts by electron microscopy showed the smallest distance growth front for the middle mantle section, where maximum growth was observed and is characteristic of the Pinctada shell shape. Such fronts have been already observed at the growth surface of bivalve nacre31,32 and in cultured pearls from P. margaritifera33, without looking for any direct connection between the pearl surface and the shell zone adjacent to their position of origin. The dynamics of nacre assembly therefore vary according to shell zone, and thus impact pearl quality depending on where the saibo was excised from the mantle. Change in structural assembly was related to variation in optical and mechanical properties, and was consequently connected to pearl colour, a character known to arise from light interference within the nanocomposite structure of the aragonite tablets34. This contributes to explaining the relation between microstructure and pearl colour expression.
At the molecular phenotype level, gene expression analysis, based on a panel of genes encoding proteins implicated in the shell biomineralisation process, indicated for the first time significant differences among donor phenotypes. The green and yellow donor groups revealed clearly different patterns of expression in the saibo tissue, particularly for the genes related to aragonite formation (MRNP34, MSI60, Pearlin and Pif177). The proteins corresponding to Pif177 and MSI60 genes regulate growth, nucleation and the organization of the aragonite crystal25,35,36. Pearlin is the protein equivalent to the N14 protein previously identified in P. maxima37 and seems to be specifically involved in the formation of the nacreous layer and promotion of aragonite crystal nucleation38. Whatever the mantle section, the saibo from the green donor phenotype showed systematic overexpression of these four aragonite-related genes compared with the yellow variant. These results could correspond to the higher proportion of good quality pearls (A–C and D+ grades), or difference among pearl colour distribution observed using the green phenotype compared to the yellow one. Pearl grade and colour was mainly attributed to aragonite tablets nature and assembly, which composed nacre produced from the internal regions of the mantle when this was used as saibo. The marginal area of the mantle produced the outer shell layer, constructed from densely packed calcite prisms, was associated with low pearl grade39,40. This marginal mantle area was excluded from the saibo cutting process. The different pattern of aragonite gene expression discriminating the two donor phenotype groups could not be attributed to any saibo cutting artefacts. Indeed, calcite gene expression pattern was equivalent for both donor phenotype group and followed the same tendency, such as for Prism14 gene expression.
From an aquaculture point of view, such a phenome study could be of benefit to the pearl industry. At the scale of the donor phenotype, the present study detected specific donor-expression level of biomineralisation genes, with regard to the diversity of the shell colouration29. An establishment of systematic donor-specific phenomes could then be an interesting tool for the prediction of more appropriate colour lines to be selected and/or propagated for specific pearl colour and/or quality production. At the scale of the mantle graft phenotype, the results clearly show that mantle tissue from the posterior and connection sections can be successfully used as saibo for pearl production, as the resulting pearl quality traits were comparable to those produced with the middle zone, which is the part commonly used on commercial pearl farms, as was the nucleus retention rate. Indeed, by using these sections, more saibo could be obtained from the same number of donors. As the supply of pearl oysters for producers in French Polynesia has always been wild collection, finding colourful donor oysters was always a prerequisite, and sometimes limiting, step before the grafting process. Frequency of colourful donor oysters has been studied recently29, revealing that such individuals are rare and dependent on spat collection site. It is therefore crucial to find enough donor oysters to supply grafters, who need from 400 to 700 saibo per day.
Our exploration of phenotype variation at the level of saibo position was the first conducted in the Pinctada genus. A single donor oyster individual exhibiting a particular shell colour phenotype could therefore produce multiple pearl phenotypes at the scale of the saibo unit, mirroring its original activity at the mantle position level. This intra-phenotypic variation could overlap with the inter-donor phenotype variation. Our results suggest that systematic study of multiple phenotypes across multiple biological function and scales would be important in future phenome studies in P. margaritifera. Increased sample sizes could potentially succeed in revealing robust genetic associations with pearl quality trait improvement that would benefit the pearl industry.
Materials and Methods
Animals
First generation hatchery-produced P. margaritifera were used as donor oysters for this study. These oyster families were issued from multi-parental crosses using highly coloured broodstock with green “C” (Fig. 1a) and yellow “L” (Fig. 1b) phenotypes (inner shell coloration), carried out at the Regahiga Pearl Farm and Hatchery company, located on Mangareva Island (Gambier archipelago, French Polynesia). To discern the inner shell colour for this set, the grafter used a speculum to gently pry open the oyster valves. Broodstock breeding, larval rearing and culture of this family were done as described previously41. At the age of 30 months post-hatching, 24 donor pearl oysters from the green (N = 12) and yellow (N = 12) phenotypes, were randomly selected from a sets of healthy animals with a mean (±SD) dorso-ventral measurement of 113.9 ± 8.7 mm. The entire list of the shell length corresponding to each donor can be found in Supplementary Table S1.
Wild P. margaritifera were collected as spat in the Mangareva Island lagoon (Gambier Archipelago, French Polynesia) to serve as recipients. Passive techniques were employed for catching spat using commercial collectors. Oysters were reared from the juvenile to adult stage as previously described42. The oysters were used in the grafting procedure once they were almost 20 months old, with a mean (±SD) dorso-ventral measurement of 76.30 ± 6.5 mm.
Experimental graft
The grafting operation was conducted in October 2014, with all grafts performed by an expert from the Regahiga Pearl Farm and Hatchery company as previously43. The nuclei used for this purpose were made from the shells of freshwater mussels (1.8 BU size, equivalent to 5.45 mm diameter, 0.26 g weight; Imai Seikaku Co. Ltd., Japan). The thickness and hardness of the nacreous layers of these beads offer a specific gravity and thermal conductivity that make them particularly suitable for use as pearl nuclei44. The epithelial cells required for grafting saibo were excised from the entire mantle of the selected donor pearl oysters and include all the following sections: posterior, connection, middle and anterior. A total of 1798 grafts were performed over five days (865 grafts using the green phenotype and 933 grafts using the yellow phenotype). Supplementary Table S1 gives the number of grafts performed for each donor oyster and per mantle section. All the grafted oysters were checked for nucleus retention/rejection and mortality 45 days after the grafting operation, as previously described42. The oysters that had retained their nuclei were drilled and fixed onto chaplets (within chaplets, oysters were attached in pairs to a rope with a monofilament fishing line), which constituted the rearing system. All recipient oysters were individually labelled (attribution of a plastic label with a number) in such a way as to maintain the traceability between graft position and corresponding harvested pearls. Furthermore, the pearl oysters were regularly cleaned in order to remove biofouling (epibiota), which can hinder healthy oyster growth and pearl production.
Pearl quality trait measurements
After approximately 20 months, the cultured pearls were harvested. The pearls were cleaned by ultrasonication in soapy water (hand washing) with a LEO 801 laboratory cleaner (2-L capacity, 80 W, 46 kHz); they were then rinsed in distilled water. Pearl size was assessed by measuring nacre thickness and nacre weight as previously described28. Shape, colour and grade were evaluated for all pearls by the same professional expert from the Poe O Rikitea association, according to Tahitian pearl auction classification categories7.
Electron Microscopy
The structure of the pearl surface was observed by scanning electron microscopy (SEM) with a Hitachi Analytical Table Top SEM TM3030 at the electron microscopy facility of Université de la Polynésie Française, Tahiti. Before observation, the pearls were sawn at the point of their largest diameter so as to be positioned flat on the observation plate. Subsequently, the surfaces were metallized (Quorum Technologies, Q150R ES model) with a thin layer of gold (15 nm). This step eliminates any electromagnetic load that might interfere with observation. Observations were based on pictures taken at the pearl surface and particularly of the space between two layers of aragonite deposition (magnification 1000x, accelerating voltage 15 kV). The gap between layers of aragonite deposition was measured with post-acquisition image tools: we measured the gap between a finished deposition layer and the next or previous one (Fig. 3). Several locations on the surface of the pearl were thus measured for a statistical analysis of the averages.
Gene expression analysis
Two donors among the twelve per phenotype were randomly sampled. For each donor (N = 4), all saibo pieces prepared from 1 valve were preserved in RNAlater (Qiagen) (50 mg/mL). The saibo obtained from the other valve were used in the graft processes and the corresponding pearl sacs were all sampled (at the same time as the pearls) and likewise preserved in RNAlater (Qiagen). For the pearl sac sampling, the gonads were first cut from the recipient. The gonad tissue was then removed with a surgical blade to leave only a thin (<0.5 mm) layer of tissue surrounding the pearl. At this point, only the pearl sac and the pearl remained. Next, an incision was made in the pearl sac and the pearl removed. The pearl sac was transferred into a 5.0-ml tube with RNAlater® until RNA extraction45. The samples were stored at −80 °C until RNA extraction. A total of 170 saibo and 120 pearl sacs were sampled.
Expression levels were measured by RT-PCR for a panel of nine genes involved in biomineralisation: four aragonite-related genes (MRNP34, MSI60, Pearlin and Pif177), four calcite-related genes (Aspein, Shematrin 9, Prismalin-14 and KRMP7) and one involved in the formation of both nacreous aragonite and prismatic calcite (Nacrein A1). Two housekeeping genes were also measured, chosen based on their ubiquitous and constitutive expression pattern in P. margaritifera tissue: SAGE (SAGES: AGCCTAGTGTGGGGGTTGG/ SAGER: ACAGCGATGTACCCATTTCC) (called REF36) and GAPDH (GAPDHS: AGGCTTGATGACCACTGTCC/ GAPDHR: AGCCATTCCCGTCAACTTC)46. Primer sequences of the nine biomineralisation genes can be found in Supplementary Table S2.
After removing the RNAlater by pipetting and absorption, total cellular RNA was extracted from the individual graft tissue (pooled into 55 samples with respect to the different mantle positions studied, with 3–4 saibo caming from the same individual per extraction) or pearl sac samples (N = 120), using TRIzol® reagent (Life Technologies) according to the manufacturer’s recommendations. Saibo from connection zone were not used for this part of the study as only 3 to 6 pooled sample could be obtained. RNA was quantified using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Inc.). Total RNA of each individual was then treated with DNAse I using a DNA-free Kit (Ambion). First, strand cDNA was synthesized from 500 ng total RNA using the Transcriptor First Strand cDNA Synthesis Kit (Roche) and a mix of poly (dT) and random hexamer primers. Real-Time PCR amplifications were carried out on a Roche Light Cycler® 480. The amplification reaction contained 5 μL LC 480 SYBR Green I Mast (Roche), 4 μL cDNA template, and 1 μL of primer (1 µM), in a final volume of 10 μL. Each run included a positive cDNA and a blank control for each primer pair. The run protocol was as follows: initial denaturation at 95 °C for 10 min followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 60 s. Lastly, the amplicon melting temperature curve was analysed using a melting curve program: 45–95 °C with a heating rate of 0.1 °C s−1 and continuous fluorescence measurement. All measurements were made in duplicate and all analyses were based on the Ct values of the PCR products.
Relative gene expression levels were calculated using the delta–delta method, normalized with two reference genes, to compare the relative expression results47 as follows: Relative expression(target gene, sample x) = 2−(ΔCt sample, sample x−ΔCt calibrator, sample x) = 2−ΔΔCt. Here, the ΔCt calibrator represents the mean of the ΔCt values obtained for the tested gene. The delta threshold cycle (ΔCt) is calculated by the difference in Ct for the target and reference genes. The relative stability of the GAPDH and SAGE combination was confirmed using NormFinder48. PCR efficiency (E) was estimated for each primer pair by determining the slopes of standard curves obtained from serial dilution analysis of a cDNA to ensure that E ranged from 90 to 110%.
Statistical analysis
All analyses were performed using R© version 3.2.3 software (R foundation for Statistical Computing). The significance threshold was set at p ≤ 0.05 and the tests were two-sided. All measures are given as means and variability as standard deviations.
Pearl quality and retention differences according to saibo positions were tested, for qualitative parameters, using χ² tests and Fisher’s exact tests when an expected value <5 was found. When significant differences were detected, pairwise comparisons were made for proportions. For quantitative categories, normality and homoscedasticity were tested using Shapiro–Wilk test and Bartlett’s tests, respectively. Due to non-normality, Kruskal–Wallis tests were used to test differences between saibo positions. When differences were detected, post-hoc analyses were performed with Dunn tests and Bonferroni correction. Kruskal-Wallis tests were also used to test difference of gene expression levels between the phenotypes.
Data availability statement
The authors declare that all datas are available.
References
Houle, D. Numbering the hairs on our heads: the shared challenge and promise of phenomics. Proc Natl Acad Sci. 07, 1793–9 (2010).
Houle, D., Govindaraju, D. R. & Omholt, S. Phenomics: the next challenge. Nat Rev Genet. 11, 855–66 (2010).
Southgate, P. C. & Lucas, J. S. The pearl oyster. (ed. Elsevier) (Oxford, 2008).
Clark, M. S. et al. Insights into shell deposition in the Antarctic bivalve Laternula elliptica: gene discovery in the mantle transcriptome using 454 pyrosequencing. BMC Genomics 11, 362 (2010).
Kawakami, I. K. Studies on pearl sac formation. 1. On the regeneration and transplantation of the mantle piece in the pearl oyster. Mem Fac Science Kyushu University (Ser E) 1, 83–8 (1952).
Farn, A. E. 1986, Pearls Naturals, cultured and imitation (ed. Butterworth Gem Books) 150 pp (1986).
Ky, C. L., Okura, R., Nakasai, S. & Devaux, D. Quality Trait Signature at Archipelago Scale of the Cultured Pearls Produced by the Black-Lipped Pearl Oyster (Pinctada margaritifera Var. cumingi) in French Polynesia. Journal of Shellfish Research 35(4), 827–835 (2016).
Jerry, D. R. et al. Donor oyster derived heritability estimates and the effect of genotype x environment interaction on the production of pearl quality traits in the silver-lip pearl oyster. Pinctada maxima. Aquaculture 338, 66–71 (2012).
McGinty, E. L., Evans, B. S., Taylor, J. U. U. & Jerry, D. R. Xenografts and pearl production in two pearl oyster species, P. maxima and P. margaritifera: effect on pearl quality and a key to understanding genetic contribution. Aquaculture 302, 175–181 (2010).
Ky, C. L. et al. Is pearl colour produced from Pinctada margaritifera predictable through shell phenotypes and rearing environments selections? Aqua. Research 48(3), 1041–1057 (2017).
Wada, K. T. & Komaru, A. Color and weight of pearls produced by grafting the mantle tissue from a selected population for white shell color of the Japanese pearl oyster Pinctada fucata martensii (Dunker). Aquaculture 142, 25–32 (1996).
Blay, C., Planes S. & Ky, C.L. Donor and recipient contribution to phenotypic traits and the expression of biomineralisation genes in the pearl oyster model Pinctada margaritifera. Scientific Reports, 7(1), 2696 (1–12) (2017).
McDougall, C., Moase, P. & Degnan, B. M. Host and donor influence on pearls produced by the silver-lip pearl oyster. Pinctada maxima. Aquaculture 450, 313–320 (2016).
Le Pabic, L. et al. Culture site dependence on pearl size realization in Pinctada margaritifera in relation to recipient oyster growth and mantle graft biomineralization gene expression using the same donor phenotype. Estuarine, Coastal and Shelf Science 182, 294–303 (2016).
Ky, C. L. et al. Macro-geographical differences influenced by family-based expression on cultured pearl grade, shape and colour in the black-lip pearl oyster Pinctada margaritifera: a preliminary case study in French Polynesia. Aquaculture Research 48, 270–282 (2017).
Ellis, S. & Haws, M. Producing pearls using the black-lip pearl oyster (Pinctada margaritifera). Aquafarmer Information Sheet, 141, 8 pp (1999).
Inoue, N. et al. Gene expression patterns and pearl formation in the Japanese pearl oyster (Pinctada fucata): a comparison of gene expression patterns between the pearl sac and mantle tissues. Aquaculture 308, 268–274 (2010).
Kishore, P. & Southgate, P. C. A detailed description of pearl‐sac development in the black‐lip pearl oyster, Pinctada margaritifera (Linnaeus 1758). Aquaculture Research 47, 2215–2226 (2014).
Liu, X. et al. The role of matrix proteins in the control of nacreous layer deposition during pearl formation. Proceedings of the Royal Society of London B: Biological Sciences 279(1730), 1000–1007 (2012).
Zhan, X. et al. Expressed sequence tags 454 sequencing and biomineralization gene expression for pearl sac of the pearl oyster. Pinctada fucata martensii. Aquaculture Research 46(3), 745–758 (2015).
Sudo, S. et al. Structures of mollusc shell framework proteins. Nature (London) 387, 563–564 (1997).
Marin, F. & Luquet, G. Molluscan shell proteins. Comptes Rendus Palevol 3, 469–492 (2004).
Zhang, C. & Zhang, R. Matrix proteins in the outer shells of molluscs. Mar Biotechnol (NY) 8, 572–586 (2006).
Marie, B. et al. Different secretory repertoires control the biomineralization processes of prism and nacre deposition of the pearl oyster shell. PNAS 109, 20986–20991 (2012).
Suzuki, M. et al. An Acidic Matrix Protein, Pif, Is a Key Macromolecule for Nacre Formation. Science 325, 1388–1390 (2009).
Miyamoto, H., Miyoshi, F. & Kohno, J. The Carbonic anhydrase domain protein nacrein is expressed in the epithelial cells of the mantle and acts as a negative regulator in calcification in the mollusk Pinctada fucata. Zoological Science 22, 311–315 (2005).
Tayale, A. et al. Evidence of donor effect on cultured pearl quality from a duplicated grafting experiment on Pinctada margaritifera using wild donors. Aquatic Living Resources 25(3), 269–280 (2012).
Ky, C. L. et al. Family effect on cultured pearl quality in black-lipped pearl oyster Pinctada margaritifera and insights for genetic improvement. Aquatic Living Resources 26, 133–145 (2013).
Ky, C. L., Lo, C. & Planes, S. Mono- and polychromatic inner shell phenotype diversity in Pinctada margaritifera donor pearl oysters and its relation with cultured pearl colour. Aquaculture 468, 199–205 (2017).
Ky, C. L. et al. Is pearl colour produced from Pinctada margaritifera predictable through shell phenotypes and rearing environments selections? Aquaculture Research 48(3), 1041–1057 (2017).
Cartwright, J. H. E. & Checa, A. G. The dynamics of nacre self assembly. Journal of the royal society 4, 491–504 (2007).
Cartwright, J. H. E., Checa, A. G., Escribano, B. & Sainz-Diaz, C. I. Spriral and target patterns in bivalve nacre manifest a natural excitable medium from layer growth of a biological liquid crystal. PNAS 106(26), 10449–10504 (2009).
Cartwright, J. H. E., Checa, A. G. & Rousseau, M. Pearls are self-organised natural ratchets. Langmuir american chemical society publications 29, 8370–8376 (2013).
Snow, M. R., Pring, A., Self, P., Losic, D. & Shapter, J. The origin of the color of pearl in iridescence from nanocomposite structures of the nacre. American mineralogist 89, 1353–1358 (2004).
Xiang, L. et al. Patterns of expression in the matrix proteins responsible for nucleation and growth of aragonite crystals in flat pearls of Pinctada fucata. PloS one 8(6), e66564 (2013).
Joubert, C. et al. Temperature and Food Influence Shell Growth and Mantle Gene Expression of Shell Matrix Proteins in the Pearl Oyster Pinctada margaritifera. PLoS ONE 9(8), e103944, https://doi.org/10.1371/journal.pone.0103944 (2014).
Bédouet, L. et al. Proteomics analysis of the nacre soluble and insoluble proteins from the oyster Pinctada margaritifera. Marine biotechnology 9(5), 638–649 (2007).
Kono, M., Hayashi, N. & Samata, T. Molecular mechanism of the nacreous layer formation in Pinctada maxima. Biochemical and biophysical research communications 269(1), 213–218 (2000).
Cuif, J. P. et al. Structural, mineralogical and biochemical diversity in the lower part of the pearl layer of cultivated seawater pearls from Polynesia. Microsc. Microanal. 14, 405–417 (2008).
Cuif, J. P. et al. Evidence of a biological control over origins, growth and end of the calcite prisms in the shells of Pinctada margaritifera (Pelecypod, Pterioidea). Minerals 4, 815–834 (2014).
Ky, C. L., Lau, C., Sham Koua, M. & Lo, C. Growth Performance Comparison of Pinctada margaritifera Juveniles Produced by Thermal Shock or Gonad Scarification Spawning Procedures. Journal of Shellfish Research 34(3), 811–817 (2015).
Ky, C. L., Molinari, N., Moe, E. & Pommier, S. Impact of season and grafter skill on nucleus retention and pearl oyster mortality rate in Pinctada margaritifera aquaculture. Aquaculture International 22(5), 1689–1701 (2014).
Ky, C. L., Demmer, J., Sham Koua, M. & Cabral, P. Development of cultured pearl circles and shape after initial graft and second nucleus insertion in the black-lipped pearl oyster Pinctada margaritifera. Journal Of Shellfish Research 34(2), 319–328 (2015).
Gervis, M. H. & Sims, N. A. The biology and culture of pearl oysters (Bivalvia: Pteriidae). International Centre for Living Aquatic Resources Management 21, 1–49 (1992).
McGinty, E. L., Zenger, K. R., Jones, D. B. & Jerry, D. R. Transcriptome analysis of biomineralisation-related genes within the pearl sac: host and donor oyster contribution. Marine Genomics 5, 27–33 (2012).
Lemer, S., Saulnier, D., Gueguen, Y. & Planes, S. Identification of genes associated with shell color in the black lipped pearl oyster, Pinctada margaritifera. BMC Genomics 16, 568 (2015).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta C(T)) Method. Methods 25, 402–8 (2001).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Research 64, 5245–5250 (2004).
Acknowledgements
This study was supported by grants from the “Direction des Ressources Marines et Minières”, through the TripaGEN project (2015–2017). The authors especially would like to thank the host site, SCA Regahiga Pearls (Mangareva island, Gambier archipelago, French Polynesia) for their generous support.
Author information
Authors and Affiliations
Contributions
C.L.K. designed the experiment and wrote the manuscript. V.Q. realized the molecular gene expression analysis. C.S. performed the electron microscopy observations. F.B. analyzed the data. D.D. supported the logistical and technical organization for the realization of the experimental grafts. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ky, CL., Quillien, V., Broustal, F. et al. Phenome of pearl quality traits in the mollusc transplant model Pinctada margaritifera. Sci Rep 8, 2122 (2018). https://doi.org/10.1038/s41598-018-20564-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20564-1
This article is cited by
-
Pearl shape classification using deep convolutional neural networks from Tahitian pearl rotation in Pinctada margaritifera
Scientific Reports (2023)
-
Colour plasticity in the shells and pearls of animal graft model Pinctada margaritifera assessed by HSV colour quantification
Scientific Reports (2019)
-
Expressions of Shell Matrix Protein Genes in the Pearl Sac and Its Correlation with Pearl Weight in the First 6 Months of Pearl Formation in Hyriopsis cumingii
Marine Biotechnology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.